

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
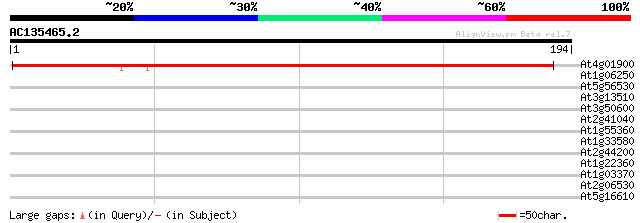
Query= AC135465.2 + phase: 0
(194 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g01900 P II nitrogen sensing protein GLB I 227 3e-60
At1g06250 lipase like protein 32 0.21
At5g56530 unknown protein (At5g56530) 32 0.28
At3g13510 unknown protein 31 0.36
At3g50600 unknown protein 30 0.61
At2g41040 hypothetical protein 29 1.4
At1g55360 unknown protein 28 2.3
At1g33580 En/Spm-like transposon protein, putative 28 2.3
At2g44200 putative protein 28 4.0
At1g22360 unknown protein 28 4.0
At1g03370 unknown protein 27 5.2
At2g06530 unknown protein 27 6.8
At5g16610 AT5g16610/MTG13_5 27 8.9
>At4g01900 P II nitrogen sensing protein GLB I
Length = 196
Score = 227 bits (579), Expect = 3e-60
Identities = 122/190 (64%), Positives = 145/190 (76%), Gaps = 3/190 (1%)
Query: 2 ALIAKPNVFNGLNFHINETQFPFSSFSVIRKRFGDS--SHRNVVLKS-NGNASILPKIRA 58
A + KP L F+ + FS I F S S ++V KS + N+ +LP + A
Sbjct: 3 ASMTKPISITSLGFYSDRKNIAFSDCISICSGFRHSRPSCLDLVTKSPSNNSRVLPVVSA 62
Query: 59 QNLPDYVPESKFYKVEAILRPWRIPQVSSGLLKMGIRGVTVSDVKGFGAQGGSKERQGGS 118
Q DY+P+SKFYKVEAI+RPWRI QVSS LLK+GIRGVTVSDV+GFGAQGGS ER GGS
Sbjct: 63 QISSDYIPDSKFYKVEAIVRPWRIQQVSSALLKIGIRGVTVSDVRGFGAQGGSTERHGGS 122
Query: 119 EFSEDNFVAKVKMEIVVRKDQVEAVINKIMETARTGEIGDGKIFLIPVSDVIRIRTGERG 178
EFSED FVAKVKMEIVV+KDQVE+VIN I+E ARTGEIGDGKIF++PVSDVIR+RTGERG
Sbjct: 123 EFSEDKFVAKVKMEIVVKKDQVESVINTIIEGARTGEIGDGKIFVLPVSDVIRVRTGERG 182
Query: 179 EQAERMAGGL 188
E+AE+M G +
Sbjct: 183 EKAEKMTGDM 192
>At1g06250 lipase like protein
Length = 423
Score = 32.0 bits (71), Expect = 0.21
Identities = 22/84 (26%), Positives = 36/84 (42%), Gaps = 4/84 (4%)
Query: 11 NGLNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQNLPDYVPESKF 70
N +N ++ + Q P + F+ R GD + +NVV + L +R N+PD P
Sbjct: 242 NNININLQKKQVPITVFAFGSPRIGDHNFKNVV----DSLQPLNILRIVNVPDVAPHYPL 297
Query: 71 YKVEAILRPWRIPQVSSGLLKMGI 94
I I ++S LK +
Sbjct: 298 LLYSEIGEVLEINTLNSTYLKRSL 321
>At5g56530 unknown protein (At5g56530)
Length = 420
Score = 31.6 bits (70), Expect = 0.28
Identities = 17/56 (30%), Positives = 28/56 (49%), Gaps = 1/56 (1%)
Query: 32 KRFGDSSHRNVVLKSNGNASILPKIRAQNLPDYVPESKFYKVEAILRPWRIPQVSS 87
KR+G H +V L + + ++ + Q+ YV KFY +A + W P+V S
Sbjct: 146 KRYGKKKHLSVPLPRSADPDLINQSGHQHAIAYVEGGKFYGAKATINVWE-PKVQS 200
>At3g13510 unknown protein
Length = 419
Score = 31.2 bits (69), Expect = 0.36
Identities = 14/49 (28%), Positives = 24/49 (48%)
Query: 32 KRFGDSSHRNVVLKSNGNASILPKIRAQNLPDYVPESKFYKVEAILRPW 80
KR+G HR+V + + ++ + Q+ YV K+Y +A L W
Sbjct: 145 KRYGKKKHRSVPIPKSAEPDLINQNGHQHAIAYVEGDKYYGAKATLNVW 193
>At3g50600 unknown protein
Length = 852
Score = 30.4 bits (67), Expect = 0.61
Identities = 24/91 (26%), Positives = 42/91 (45%), Gaps = 10/91 (10%)
Query: 78 RPWRIPQVSSGLLKMGIRGV---TVSDVKGFGAQGGSKERQG----GSEFSEDNFVAKVK 130
+P +P S + MG+ + V+D + A+ K G GS E+ K
Sbjct: 332 KPLALPAGESSSM-MGLESLGKQNVADEQAKAAEEFKKTMYGATGDGSSSDEEGVTKPKK 390
Query: 131 MEIVVRKDQVEAVI--NKIMETARTGEIGDG 159
++I +R+ + NK+ E A+T ++GDG
Sbjct: 391 LQIRIREKPTSTTVDVNKLKEAAKTFKLGDG 421
>At2g41040 hypothetical protein
Length = 262
Score = 29.3 bits (64), Expect = 1.4
Identities = 19/62 (30%), Positives = 34/62 (54%), Gaps = 3/62 (4%)
Query: 1 MALIAKPNVFNGLNFHINETQFP---FSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIR 57
+ L+ KP + N +F +++Q P FS+ + +R RF ++ V KS+ N + PKI
Sbjct: 11 ITLLNKPFLPNRSSFFSSDSQSPLLRFSASTSVRSRFPSAAISAVAPKSDINKNETPKIE 70
Query: 58 AQ 59
+
Sbjct: 71 IE 72
>At1g55360 unknown protein
Length = 422
Score = 28.5 bits (62), Expect = 2.3
Identities = 13/49 (26%), Positives = 23/49 (46%)
Query: 32 KRFGDSSHRNVVLKSNGNASILPKIRAQNLPDYVPESKFYKVEAILRPW 80
KR+G R+V L + ++ + Q+ YV K+Y +A + W
Sbjct: 148 KRYGKKKRRSVPLPKSAEPDLINQSGHQHAIAYVEGDKYYGAKATINVW 196
>At1g33580 En/Spm-like transposon protein, putative
Length = 428
Score = 28.5 bits (62), Expect = 2.3
Identities = 23/81 (28%), Positives = 36/81 (44%), Gaps = 11/81 (13%)
Query: 19 ETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILP----------KIRAQNLPDYVPES 68
E+ F ++ +IR + D + N N S P K++ P Y
Sbjct: 97 ESSSEFGAYELIRTAYFDGEDHSDSQNQNENDSKEPSTKEESDFREKLKDAETPLYYGCP 156
Query: 69 KFYKVEAILRPWRIPQVSSGL 89
K+ KV AI+R +RI +V SG+
Sbjct: 157 KYTKVSAIMRLYRI-KVKSGM 176
>At2g44200 putative protein
Length = 493
Score = 27.7 bits (60), Expect = 4.0
Identities = 19/60 (31%), Positives = 32/60 (52%), Gaps = 7/60 (11%)
Query: 111 SKERQGGSEFSEDNFVAKVK---MEIVVRKDQVEAVINKIMET----ARTGEIGDGKIFL 163
++ R+GGS+ SE+ A++K M+ V ++Q + K ET A ++ GK FL
Sbjct: 392 NRRRKGGSKLSEEERAARLKQMQMDAEVHEEQRWTRLKKADETDAVEADKNKVSTGKSFL 451
>At1g22360 unknown protein
Length = 481
Score = 27.7 bits (60), Expect = 4.0
Identities = 11/37 (29%), Positives = 25/37 (66%)
Query: 113 ERQGGSEFSEDNFVAKVKMEIVVRKDQVEAVINKIME 149
E+Q +FS D + +++ V++++VEAV+ ++M+
Sbjct: 401 EQQTNCKFSRDEWEVGIEIGGDVKREEVEAVVRELMD 437
>At1g03370 unknown protein
Length = 1859
Score = 27.3 bits (59), Expect = 5.2
Identities = 16/68 (23%), Positives = 32/68 (46%), Gaps = 4/68 (5%)
Query: 1 MALIAKPNVFNGLNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQN 60
M+++ K NG ++++ + +F+V + G + R + S NAS L +
Sbjct: 1 MSIVQKQEEMNGCGLNVDKVE----AFTVSPQEKGRKNKRKLADPSQPNASSLTEFPPYE 56
Query: 61 LPDYVPES 68
LP P++
Sbjct: 57 LPSLKPQN 64
>At2g06530 unknown protein
Length = 225
Score = 26.9 bits (58), Expect = 6.8
Identities = 15/51 (29%), Positives = 29/51 (56%), Gaps = 3/51 (5%)
Query: 123 DNFVAKVKMEIVVRKDQVEAVINKIMETARTGEIGDGKIFLIPVSDVIRIR 173
D + +++ E + Q + +IN+I +TA+ G++G K+ D+IR R
Sbjct: 24 DKSIREIERERQGLQTQEKKLINEIKKTAKQGQMGAVKVM---AKDLIRTR 71
>At5g16610 AT5g16610/MTG13_5
Length = 529
Score = 26.6 bits (57), Expect = 8.9
Identities = 15/60 (25%), Positives = 31/60 (51%), Gaps = 13/60 (21%)
Query: 64 YVPESKFYKVEAILRPWR------------IPQVSSGLLKMGIR-GVTVSDVKGFGAQGG 110
+ ++ K + RPW+ + + + GLL++ I+ GVTV +++ +G +GG
Sbjct: 240 FCTSDEYVKPSRVFRPWKRKKTATDSIETALEEDAPGLLQVLIQQGVTVDELRLYGNEGG 299
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.137 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,163,441
Number of Sequences: 26719
Number of extensions: 171962
Number of successful extensions: 449
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 442
Number of HSP's gapped (non-prelim): 14
length of query: 194
length of database: 11,318,596
effective HSP length: 94
effective length of query: 100
effective length of database: 8,807,010
effective search space: 880701000
effective search space used: 880701000
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC135465.2


