

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135464.5 - phase: 0 /pseudo
(1087 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
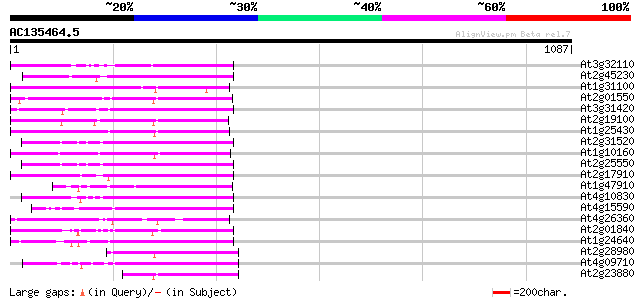
Score E
Sequences producing significant alignments: (bits) Value
At3g32110 non-LTR reverse transcriptase, putative 190 4e-48
At2g45230 putative non-LTR retroelement reverse transcriptase 171 2e-42
At1g31100 hypothetical protein 169 6e-42
At2g01550 putative non-LTR retroelement reverse transcriptase 165 1e-40
At3g31420 hypothetical protein 163 4e-40
At2g19100 putative non-LTR retroelement reverse transcriptase 160 3e-39
At1g25430 hypothetical protein 159 6e-39
At2g31520 putative non-LTR retroelement reverse transcriptase 157 2e-38
At1g10160 putative reverse transcriptase 157 3e-38
At2g25550 putative non-LTR retroelement reverse transcriptase 157 4e-38
At2g17910 putative non-LTR retroelement reverse transcriptase 156 5e-38
At1g47910 reverse transcriptase, putative 154 2e-37
At4g10830 putative protein 152 1e-36
At4g15590 reverse transcriptase like protein 150 4e-36
At4g26360 putative protein 149 6e-36
At2g01840 putative non-LTR retroelement reverse transcriptase 146 7e-35
At1g24640 hypothetical protein 145 1e-34
At2g28980 putative non-LTR retroelement reverse transcriptase 144 4e-34
At4g09710 RNA-directed DNA polymerase -like protein 143 5e-34
At2g23880 putative non-LTR retroelement reverse transcriptase 143 6e-34
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 190 bits (482), Expect = 4e-48
Identities = 138/444 (31%), Positives = 206/444 (46%), Gaps = 52/444 (11%)
Query: 1 DFNAVRYAEERKSSRGISGVIDYHHFSDFIVDNSLIDLSLCGRRFTWFKGDGNAM---SR 57
DFN + +ER G D F ++I D+SLIDL G +FTW +G R
Sbjct: 702 DFNTIVRLDERSGGNGRLSS-DSLAFGEWINDHSLIDLGFKGNKFTWKRGREERFFVAKR 760
Query: 58 IDRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEE---NWGPRPSRMLKCWQDLPG 114
+DR L L+W + + L SDH PL + + E N G RP R W PG
Sbjct: 761 LDRVLCCAHARLKWQEASVLHLPFLASDHAPLYVQLTPEVSGNRGRRPFRFEAAWLSHPG 820
Query: 115 YNQFVKDKLKSFQIEGWGGFVLREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDD 174
+ + + + W + + ++ L H S+ D LKK + D
Sbjct: 821 FKELL--------LTSWNKDISTPEALKVQELLDL----HQSD-----DLLKKEEELLKD 863
Query: 175 KGDVGDLSADEILELRGITHDLHSLSRVNSSILW-QQSRLLWLKDGDANSKYFHSVLSGR 233
D +LE ++W Q+SR W GD N+K+FH+ R
Sbjct: 864 --------FDVVLE--------------QEEVVWMQKSREKWFVHGDRNTKFFHTSTIIR 901
Query: 234 RRRNSIISLLA-DSNLVEGVQPIRNAVLSHFRDHFAARHTSRVGIDNLP---FKCLSHTD 289
RRRN I L D + Q + + +++ ++ V ++ LP F LS D
Sbjct: 902 RRRNQIEMLQDNDGRWLSNAQELETHAIDYYKRLYSLDDLDAV-VEQLPQEGFTALSEAD 960
Query: 290 GSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKL 349
S LTKPFS EV+ A+ YK+PGPDG F ++ W+ + E V +F+ DF +G
Sbjct: 961 FSSLTKPFSPLEVEGAIRSMGKYKAPGPDGFQPVFYQQGWEVVGESVTKFVMDFFSSGSF 1020
Query: 350 TRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAF 409
+ N + LI+K+ P++I FRPISL L+K ++KV+ RLK V+ K+I +QT+F
Sbjct: 1021 PQETNDVLVVLIAKVLKPEKITQFRPISLCNVLFKTITKVMVGRLKGVINKLIGPAQTSF 1080
Query: 410 VKDRQILDGILIANEVVDEARKNK 433
+ R D I++ EVV R+ K
Sbjct: 1081 IPGRLSTDNIVVVQEVVHSMRRKK 1104
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 171 bits (434), Expect = 2e-42
Identities = 120/424 (28%), Positives = 208/424 (48%), Gaps = 33/424 (7%)
Query: 26 FSDFIVDNSLIDLSLCGRRFTWF--KGDGNAMSRIDRFLLSEEWCLQWPNSFQVALLRGL 83
F + L +++ G +F+W+ + D R+DR + ++ W +P + L +
Sbjct: 160 FRQMLNSCGLWEVNHSGYQFSWYGNRNDELVQCRLDRTVANQAWMELFPQAKATYLQKIC 219
Query: 84 SDHCPLQLSVDEENWGPRPS-RMLKCWQDLPGYNQFVKDKLKSF--QIEGWGGFVLREKL 140
SDH PL ++ +NW + K W G+ KD L +F Q ++ EK+
Sbjct: 220 SDHSPLINNLVGDNWRKWAGFKYDKRWVQREGF----KDLLCNFWSQQSTKTNALMMEKI 275
Query: 141 KFIKTALKEWHIGHASNIPGKIDSLK------KRQADFDDKGDVGDLSADEILELRGITH 194
+ + +W + +I L+ +Q FD + EL +
Sbjct: 276 ASCRREISKWKRVSKPSSAVRIQELQFKLDAATKQIPFDRR------------ELARLKK 323
Query: 195 DLHSLSRVNSSILWQQ-SRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLADSNLV-EGV 252
+L S N WQ+ SR++W+++GD N+KYFH+ RR +N I L+ +
Sbjct: 324 EL-SQEYNNEEQFWQEKSRIMWMRNGDRNTKYFHAATKNRRAQNRIQKLIDEEGREWTSD 382
Query: 253 QPIRNAVLSHFRDHFAARHTSRV--GIDNLPFKCLSHTDGSGLTKPFSADEVKSAVWDCD 310
+ + ++F+ FA+ ++NL +S + L P + +EV+ A + +
Sbjct: 383 EDLGRVAEAYFKKLFASEDVGYTVEELENLT-PLVSDQMNNNLLAPITKEEVQRATFSIN 441
Query: 311 SYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRI 370
+K PGPDG+N ++FW+ + + + + F R+G + G+N T I LI KI +++
Sbjct: 442 PHKCPGPDGMNGFLYQQFWETMGDQITEMVQAFFRSGSIEEGMNKTNICLIPKILKAEKM 501
Query: 371 NDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQILDGILIANEVVDEAR 430
DFRPISL +YK++ K++ANRLK +L +IS++Q AFVK R I D ILIA+E++
Sbjct: 502 TDFRPISLCNVIYKVIGKLMANRLKKILPSLISETQAAFVKGRLISDNILIAHELLHALS 561
Query: 431 KNKR 434
N +
Sbjct: 562 SNNK 565
>At1g31100 hypothetical protein
Length = 1090
Score = 169 bits (429), Expect = 6e-42
Identities = 121/442 (27%), Positives = 207/442 (46%), Gaps = 21/442 (4%)
Query: 1 DFNAVRYAEERKSSRGISGVIDYHHFSDFIVDNSLIDLSLCGRRFTWFKGDGN--AMSRI 58
DFN V E + ++ F D + + L DL G FTW+ ++
Sbjct: 55 DFNQVLCPAEHSQATSLNVNRRMKVFRDCLFEAELCDLVFKGNTFTWWNKSATRPVAKKL 114
Query: 59 DRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDE-ENWGPRPSRMLKCWQDLPGYNQ 117
DR L++E WC ++P+++ V SDH + ++ + RP R P +
Sbjct: 115 DRILVNESWCSRFPSAYAVFGEPDFSDHASCGVIINPLMHREKRPFRFYNFLLQNPDFIS 174
Query: 118 FVKDKLKSFQIEGWGGFVLREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDDKGD 177
V + S + G F + +KLK +K ++ + + + SN+ ++ +K
Sbjct: 175 LVGELWYSINVVGSSMFKMSKKLKALKNPIRTFSMENFSNLEKRVKEAHNLVLYRQNKTL 234
Query: 178 VGDLSADEILELRGITHDLHSLSRVNSSILWQQSRLLWLKDGDANSKYFHSVLSGRRRRN 237
+ LE+ L L + S Q+SR+ W+ +GD+N+ YFH + R+ N
Sbjct: 235 SDPTIPNAALEMEAQRKWL-ILVKAEESFFCQRSRVTWMGEGDSNTSYFHRMADSRKAVN 293
Query: 238 SIISLLADSNLVEGVQPIRNAVLSHFRDHFAARHTSRVGIDNL---------PFKCLSHT 288
+I ++ D+ + Q + H ++F+ VG L PF+C SH
Sbjct: 294 TIHIIIDDNGVKIDTQL---GIKEHCIEYFSNLLGGEVGPPMLIQEDFDLLLPFRC-SHD 349
Query: 289 DGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGK 348
L FS ++KSA + S K+ GPDG F KE W + +V +S+F +
Sbjct: 350 QKKELAMSFSRQDIKSAFFSFPSNKTSGPDGFPVEFFKETWSVIGTEVTDAVSEFFTSSV 409
Query: 349 LTRGINTTFIALISKIDSPQRINDFRPISLVG----SLYKILSKVLANRLKLVLGKIISD 404
L + N T + LI KI + ++NDFRPIS +LYK+++++L NRL+ +L ++IS
Sbjct: 410 LLKQWNATTLVLIPKITNASKMNDFRPISCNDFGPITLYKVIARLLTNRLQCLLSQVISP 469
Query: 405 SQTAFVKDRQILDGILIANEVV 426
Q+AF+ R + + +L+A E+V
Sbjct: 470 FQSAFLPGRFLAENVLLATELV 491
>At2g01550 putative non-LTR retroelement reverse transcriptase
Length = 1449
Score = 165 bits (418), Expect = 1e-40
Identities = 130/454 (28%), Positives = 213/454 (46%), Gaps = 29/454 (6%)
Query: 1 DFNAVRYAEERKSSRG----ISGVIDYHHFSDFIVDNSLIDLSLCGRRFTWFKGDGN--A 54
DFN + +E SG+ D+ ++ S DL+ G FTW N
Sbjct: 557 DFNEILDMDEHSRMEDHPAVTSGMRDFQSLVNYC---SFSDLASHGPLFTWCNKRDNDPI 613
Query: 55 MSRIDRFLLSEEWCLQWPNSFQVALLRGLSDH--CPLQLSVDE--ENWGPRPSRMLKCWQ 110
++DR +++E W + +P S+ V G SDH C + L+++ + G +P + +
Sbjct: 614 WKKLDRVMVNEAWKMVYPQSYNVFEAGGCSDHLRCRINLNMNSGAQVRGNKPFKFVNAVA 673
Query: 111 DLPGYNQFVKD---KLKSFQIEGWGGFVLREKLKFIKTALKEWHIGHASNIPGKIDS--L 165
D+ + V++ + + + F +KLK +K L+ N+ + L
Sbjct: 674 DMEEFKPLVENFWRETEPIHMSTSSLFRFTKKLKALKPKLRGLAKEKMGNLVKRTREAYL 733
Query: 166 KKRQADFDDKGDVGDLSADEILELRGITHDLHSLSRVNSSILWQQSRLLWLKDGDANSKY 225
QA + + A EI + D ++ + L Q S+L WLK GD N+K
Sbjct: 734 SLCQAQQSNSQNPSQ-RAMEIESEAYVRWD--RIASIEEKYLKQVSKLHWLKVGDKNNKT 790
Query: 226 FHSVLSGRRRRNSIISLLA-DSNLVEGVQPIRNAVLSHFRDHFAARHTSRVGI------D 278
FH + R +NSI + D + I+N F++ GI
Sbjct: 791 FHRAATARAAQNSIREIQKEDGSTATTKDDIKNETERFFQEFLQLIPNDYEGITVEKLTS 850
Query: 279 NLPFKCLSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMR 338
LP+ C S + LT SA E++ A++ + KSPGPDG F K WD + + +
Sbjct: 851 LLPYHC-SPAEKDMLTASVSAKEIRGALFSMPNDKSPGPDGYTSEFYKRAWDIIGAEFVL 909
Query: 339 FISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVL 398
+ F G L +G+NTT +ALI K + + D+RPIS +YK++SK++ANRLK VL
Sbjct: 910 AVKSFFEKGFLPKGVNTTILALIPKKLEAKEMKDYRPISCCNVIYKVISKIIANRLKHVL 969
Query: 399 GKIISDSQTAFVKDRQILDGILIANEVVDEARKN 432
I+ +Q+AFVKDR +++ +L+A E+V + K+
Sbjct: 970 PNFIAGNQSAFVKDRLLIENLLLATELVKDYHKD 1003
>At3g31420 hypothetical protein
Length = 1491
Score = 163 bits (413), Expect = 4e-40
Identities = 128/446 (28%), Positives = 211/446 (46%), Gaps = 24/446 (5%)
Query: 1 DFNAVRYAEERKSSRGISGVIDYHHFSDFIVDNSLIDLSLCGRRFTWF--KGDGNAMSRI 58
DFN + E++ R + + F + + + DL G RF+W +GD + +
Sbjct: 317 DFNEILSNREKRGGR-LRPERTFQDFRNMVRGCNFTDLKSVGDRFSWAGKRGDHHVTCSL 375
Query: 59 DRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENWGPR-----PSRMLKCWQDLP 113
DR + + EW +P S V + G SDH PL ++ + R SR+
Sbjct: 376 DRTMANNEWHTLFPESETVFMEYGESDHRPLVTNISAQKEERRGFFSYDSRLTH----KE 431
Query: 114 GYNQFVKDKLKSFQIEGWGGFVLREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFD 173
G+ Q V ++ G L KL + A+ W + N +I ++ + D
Sbjct: 432 GFKQVVLNQWHRRNGSFEGDSSLNRKLVECRQAISRWKKNNRVNAEERIKIIRHK---LD 488
Query: 174 DKGDVGDLSADEILELRGITHDLHSLSRVNSSILWQ-QSRLLWLKDGDANSKYFHSVLSG 232
G + E ++R DL+ + + I WQ +SR WL GD N++YFHS
Sbjct: 489 RAIATGTATPRERRQMR---QDLNQ-AYADEEIYWQTKSRSRWLNAGDRNTRYFHSTTKT 544
Query: 233 RRRRNSIISLL-ADSNLVEGVQPIRNAVLSHFRDHFAAR-HTSRVGIDNLPFKCLSHTD- 289
RR RN ++S+ +D ++ G + I +++F D + + +TS D TD
Sbjct: 545 RRCRNRLLSVQDSDGDICRGDENIAKVAINYFDDLYKSTPNTSLRYADVFQGFQQKITDE 604
Query: 290 -GSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGK 348
L +P + E++ +V+ ++P PDG F ++FW ++K+ V+ ++ F +
Sbjct: 605 INEDLIRPVTELEIEESVFSVAPSRTPDPDGFTADFYQQFWPDIKQKVIDEVTRFFERSE 664
Query: 349 LTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTA 408
L N T + LI K+++P I FRPI+L YKI+SK+L NRLK LG I+++Q A
Sbjct: 665 LDERHNHTNLCLIPKVETPTTIAKFRPIALCNVSYKIISKILVNRLKKHLGGAITENQAA 724
Query: 409 FVKDRQILDGILIANEVVDEARKNKR 434
FV R I + +IA+EV + KR
Sbjct: 725 FVPGRLITNNAIIAHEVYYALKARKR 750
>At2g19100 putative non-LTR retroelement reverse transcriptase
Length = 1447
Score = 160 bits (406), Expect = 3e-39
Identities = 123/445 (27%), Positives = 209/445 (46%), Gaps = 27/445 (6%)
Query: 1 DFNAVRYAEERKSSRGISGV-IDYHHFSDFIVDNSLIDLSLCGRRFTWFKGDGNAM--SR 57
DFN EE V + F + SL D++ G +TW + + +
Sbjct: 550 DFNETLELEEHSKVEDNPVVSMGMRDFRSMVNYCSLTDMAHHGPLYTWSNKREHDLIAKK 609
Query: 58 IDRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENW----GPRPSRMLKCWQDLP 113
+DR ++++ W +P S+ V G DH ++++++ G RP + + ++
Sbjct: 610 LDRVMVNDVWTQSFPQSYSVFEAGGCLDHLRGRINLNDGPGSIVRGKRPFKFVNVLTEME 669
Query: 114 GYNQFVKDKLKSFQ---IEGWGGFVLREKLKFIKTALKEWHIGHASNIPGKI----DSLK 166
+ V K + + F +KLK +K L+ N+ K D+L
Sbjct: 670 DFKPTVDSYWKETEPIFLSTSSLFRFSKKLKSLKPLLRNLAKERLGNLVKKTREAYDTLC 729
Query: 167 KRQADFDDKGDVGDLSADEILELRGITHDLHSLSRVNSSILWQQSRLLWLKDGDANSKYF 226
K+Q + + + + + + E ++ + L ++S+L WL GD N+K F
Sbjct: 730 KKQ-----ESTLNNPTPNAMKEEVEAHDRWEHVAGLEEKFLKKKSKLHWLDGGDKNNKAF 784
Query: 227 HSVLSGRRRRNSIISLLA-DSNLVEGVQPIRNAVLSHFRDHFAARHTSRVGI------DN 279
H + R +NSI + D ++ I+ FR+ G+ D
Sbjct: 785 HRAVVTREAQNSISEIQCQDGSVTAKGDEIKAYAERFFREFLQLIPNEYEGVTMADLQDL 844
Query: 280 LPFKCLSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRF 339
LPF+C S T+ LT+ +A+E+K ++ + KSPGPDG F K W+ L + +
Sbjct: 845 LPFRC-SETEHELLTRVVTAEEIKKVLFSMPNDKSPGPDGFTSEFFKATWEILGNEFILA 903
Query: 340 ISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLG 399
I F G L +GINTT +ALI K + + D+RPIS +YK++SK++ANRLKLV
Sbjct: 904 IQSFFAKGFLPKGINTTILALIPKKKEAKEMKDYRPISCCNVIYKVISKIIANRLKLVPP 963
Query: 400 KIISDSQTAFVKDRQILDGILIANE 424
K I+ +Q+AFVKDR +++ +L+A +
Sbjct: 964 KFIAGNQSAFVKDRLLIENVLLATD 988
>At1g25430 hypothetical protein
Length = 1213
Score = 159 bits (403), Expect = 6e-39
Identities = 119/436 (27%), Positives = 202/436 (46%), Gaps = 13/436 (2%)
Query: 1 DFNAVRYAEERKSSRGISGVIDYHHFSDFIVDNSLIDLSLCGRRFTWFKGDGNA--MSRI 58
DFN V +E + ++ I+ F D ++ L DL G FTW+ +I
Sbjct: 144 DFNQVLNPQEHSNPVSLNVDINMRDFRDCLLAAELSDLRYKGNTFTWWNKSHTTPVAKKI 203
Query: 59 DRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENW-GPRPSRMLKCWQDLPGYNQ 117
DR L+++ W +P+S + SDH + ++E + RP + +
Sbjct: 204 DRILVNDSWNALFPSSLGIFGSLDFSDHVSCGVVLEETSIKAKRPFKFFNYLLKNLDFLN 263
Query: 118 FVKDKLKSFQIEGWGGFVLREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDDKGD 177
V+D + + G F + +KLK +K +K++ + S + + D+
Sbjct: 264 LVRDNWFTLNVVGSSMFRVSKKLKALKKPIKDFSRLNYSELEKRTKEAHDFLIGCQDRTL 323
Query: 178 VGDLSADEILELRGITHDLHSLSRVNSSILWQQSRLLWLKDGDANSKYFHSVLSGRRRRN 237
+ EL H L+ S Q+SR+ W +GD N+KYFH + R N
Sbjct: 324 ADPTPINASFELEA-ERKWHILTAAEESFFRQKSRISWFAEGDGNTKYFHRMADARNSSN 382
Query: 238 SIISLLADSN--LVEGVQPIRNAVLSHFRDHFAARHTSRVGIDN-----LPFKCLSHTDG 290
SI S L D N LV+ + I + S+F + N L ++C S
Sbjct: 383 SI-SALYDGNGKLVDSQEGILDLCASYFGSLLGDEVDPYLMEQNDMNLLLSYRC-SPAQV 440
Query: 291 SGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLT 350
L FS +++++A++ KS GPDG F + W + +V I +F +G L
Sbjct: 441 CELESTFSNEDIRAALFSLPRNKSCGPDGFTAEFFIDSWSIVGAEVTDAIKEFFSSGCLL 500
Query: 351 RGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFV 410
+ N T I LI KI +P +DFRPIS + +LYK+++++L +RL+ +L +IS +Q+AF+
Sbjct: 501 KQWNATTIVLIPKIVNPTCTSDFRPISCLNTLYKVIARLLTDRLQRLLSGVISSAQSAFL 560
Query: 411 KDRQILDGILIANEVV 426
R + + +L+A ++V
Sbjct: 561 PGRSLAENVLLATDLV 576
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 157 bits (398), Expect = 2e-38
Identities = 121/422 (28%), Positives = 205/422 (47%), Gaps = 22/422 (5%)
Query: 23 YHHFSDFIVDNSLIDLSLCGRRFTWF--KGDGNAMSRIDRFLLSEEWCLQWPNSFQVALL 80
+ +F + + + D+ G RF+W + +DR ++ W +P + L
Sbjct: 312 FRNFRNMVSHCDIEDMRSKGDRFSWVGERHTHTVKCCLDRVFINSAWTATFPYAETEFLD 371
Query: 81 RGLSDHCPLQLSVDEENWGPRPSRMLKCWQ---DLPGYNQFVKDKLKSFQIEGWGGFVLR 137
SDH P+ + +E PR S++ + D+P + + V+ ++ + +
Sbjct: 372 FTGSDHKPVLVHFNESF--PRRSKLFRFDNRLIDIPTFKRIVQTSWRTNRNSR--STPIT 427
Query: 138 EKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDDKGDVGDLSADEILELRGITHDLH 197
E++ + A+ + HASN+ + +KK Q+ + + + ++ + I
Sbjct: 428 ERISSCRQAMAR--LKHASNLNSE-QRIKKLQSSLNRA-----MESTRRVDRQLIPQLQE 479
Query: 198 SLSRVNSS--ILWQQ-SRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLADSN-LVEGVQ 253
SL++ S I W+Q SR W+K+GD N+ YFH+ R +N + +++ D + G +
Sbjct: 480 SLAKAFSDEEIYWKQKSRNQWMKEGDQNTGYFHACTKTRYSQNRVNTIMDDQGRMFTGDK 539
Query: 254 PIRNAVLSHFRDHFAARHTSRVGIDNLPFKC-LSHTDGSGLTKPFSADEVKSAVWDCDSY 312
I N F + F+ ID FK +++T LTK FS E+ A+
Sbjct: 540 EIGNHAQDFFTNIFSTNGIKVSPIDFADFKSTVTNTVNLDLTKEFSDTEIYDAICQIGDD 599
Query: 313 KSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRIND 372
K+PGPDG+ F K WD + DV+ + F + IN T I +I KI +P ++D
Sbjct: 600 KAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPSINHTNICMIPKITNPTTLSD 659
Query: 373 FRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQILDGILIANEVVDEARKN 432
+RPI+L LYK++SK L NRLK L I+SDSQ AF+ R I D ++IA+EV+ +
Sbjct: 660 YRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVMIAHEVMHSLKVR 719
Query: 433 KR 434
KR
Sbjct: 720 KR 721
>At1g10160 putative reverse transcriptase
Length = 1118
Score = 157 bits (397), Expect = 3e-38
Identities = 119/441 (26%), Positives = 205/441 (45%), Gaps = 19/441 (4%)
Query: 1 DFNAVRYAEERKS-SRGISGVIDYHHFSDFIVDNSLIDLSLCGRRFTW--FKGDGNAMSR 57
DFN + A E S ++ + + + D+ L DL G FTW + D + +
Sbjct: 142 DFNQIAAASEHYSINQSLLNLRGMEDLQCCLRDSQLSDLPSRGVFFTWSNHQQDNPILRK 201
Query: 58 IDRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENWGPRPSRMLKCWQDL---PG 114
+DR L + EW +P++ V G SDH P + +D N P + K + L P
Sbjct: 202 LDRALANGEWFAVFPSALAVFDPPGDSDHAPCIILID--NQPPPSKKSFKYFSFLSSHPS 259
Query: 115 YNQFVKDKLKSFQIEGWGGFVLREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDD 174
Y + + + G F LR+ LK K + + SNI + R D
Sbjct: 260 YLAALSTAWEENTLVGSHMFSLRQHLKVAKLCCRTLNRLRFSNIQQRTAQSLTRLEDI-- 317
Query: 175 KGDVGDLSADEILELRGITHDLHSL-SRVNSSILWQQSRLLWLKDGDANSKYFHSVLSGR 233
+ ++ +D + + + S Q+SR+ WL +GDAN+++FH +
Sbjct: 318 QVELLTSPSDTLFRREHVARKQWIFFAAALESFFRQKSRIRWLHEGDANTRFFHRAVIAH 377
Query: 234 RRRNSIISLLADSNL-VEGVQPIRNAVLSHFRDHFA--ARHTSRVGIDN----LPFKCLS 286
+ N I L D VE V I+ +++++ + + + ++ LPF+C S
Sbjct: 378 QATNLIKFLRGDDGFRVENVDQIKGMLIAYYSHLLGIPSENVTPFSVEKIKGLLPFRCDS 437
Query: 287 HTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRN 346
S LT S +E+ ++ K+PGPDG F E W +K V+ I +F +
Sbjct: 438 FL-ASQLTTIPSEEEITQVLFSMPRNKAPGPDGFPVEFFIEAWAIVKSSVVAAIREFFIS 496
Query: 347 GKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQ 406
G L RG N T I LI K+ R+ FRP++ ++YK+++++++ RLKL + + + +Q
Sbjct: 497 GNLPRGFNATAITLIPKVTGADRLTQFRPVACCTTIYKVITRIISRRLKLFIDQAVQANQ 556
Query: 407 TAFVKDRQILDGILIANEVVD 427
F+K R + + +L+A+E+VD
Sbjct: 557 VGFIKGRLLCENVLLASELVD 577
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 157 bits (396), Expect = 4e-38
Identities = 121/422 (28%), Positives = 205/422 (47%), Gaps = 22/422 (5%)
Query: 23 YHHFSDFIVDNSLIDLSLCGRRFTWF--KGDGNAMSRIDRFLLSEEWCLQWPNSFQVALL 80
+ +F + + + D+ G RF+W + +DR ++ W +P + L
Sbjct: 538 FRNFRNMVSHCDIEDMRSKGDRFSWVGERHTHTVKCCLDRVFINSAWTATFPYAEIEFLD 597
Query: 81 RGLSDHCPLQLSVDEENWGPRPSRMLKCWQ---DLPGYNQFVKDKLKSFQIEGWGGFVLR 137
SDH P+ + +E PR S++ + D+P + + V+ ++ + +
Sbjct: 598 FTGSDHKPVLVHFNESF--PRRSKLFRFDNRLIDIPTFKRIVQTSWRTNRNSR--STPIT 653
Query: 138 EKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDDKGDVGDLSADEILELRGITHDLH 197
E++ + A+ + HASN+ + +KK Q+ + + + ++ + I
Sbjct: 654 ERISSCRQAMAR--LKHASNLNSE-QRIKKLQSSLNRA-----MESTRRVDRQLIPQLQE 705
Query: 198 SLSRVNSS--ILWQQ-SRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLADSN-LVEGVQ 253
SL++ S I W+Q SR W+K+GD N+ YFH+ R +N + +++ D + G +
Sbjct: 706 SLAKAFSDEEIYWKQKSRNQWMKEGDQNTGYFHACTKTRYSQNRVNTIMDDQGRMFTGDK 765
Query: 254 PIRNAVLSHFRDHFAARHTSRVGIDNLPFKC-LSHTDGSGLTKPFSADEVKSAVWDCDSY 312
I N F + F+ ID FK +++T LTK FS E+ A+
Sbjct: 766 EIGNHAQDFFTNIFSTNGIKVSPIDFADFKSTVTNTVNLDLTKEFSDTEIYDAICQIGDD 825
Query: 313 KSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRIND 372
K+PGPDG+ F K WD + DV+ + F + IN T I +I KI +P ++D
Sbjct: 826 KAPGPDGLTARFYKNCWDIVGYDVILEVKKFFETSFMKPSINHTNICMIPKITNPTTLSD 885
Query: 373 FRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQILDGILIANEVVDEARKN 432
+RPI+L LYK++SK L NRLK L I+SDSQ AF+ R I D ++IA+EV+ +
Sbjct: 886 YRPIALCNVLYKVISKCLVNRLKSHLNSIVSDSQAAFIPGRIINDNVMIAHEVMHSLKVR 945
Query: 433 KR 434
KR
Sbjct: 946 KR 947
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 156 bits (395), Expect = 5e-38
Identities = 133/447 (29%), Positives = 207/447 (45%), Gaps = 30/447 (6%)
Query: 1 DFNAVRYAEERKSSRGISGVIDYHHFSDFIVDNSLIDLSLCGRRFTWF--KGDGNAMSRI 58
DFN +R +E+ S + F +++ S+ +L G FTW + D ++
Sbjct: 114 DFNDIRSNDEKLGGPRRSPS-SFQCFEHMLLNCSMHELGSTGNSFTWGGNRNDQWVQCKL 172
Query: 59 DRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENWGPRPS-RMLKCWQDLPGYNQ 117
DR + W +PN+ Q L + SDH P+ + +N R R K D P +
Sbjct: 173 DRCFGNPAWFSIFPNAHQWFLEKFGSDHRPVLVKFTNDNELFRGQFRYDKRLDDDPYCIE 232
Query: 118 FVKDKLKSFQIEGWGGFVLREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDDKGD 177
+ S +G L + A+ W +N +I L+K
Sbjct: 233 VIHRSWNSAMSQGTHSSFF--SLIECRRAISVWKHSSDTNAQSRIKRLRK---------- 280
Query: 178 VGDLSADEILELR-----GITHDLHSLSRVNSSILWQQ-SRLLWLKDGDANSKYFHSVLS 231
DL A++ +++ D SL+ + + W+Q SR WL GD N+ +FH+ +
Sbjct: 281 --DLDAEKSIQIPCWPRIEYIKDQLSLAYGDEELFWRQKSRQKWLAGGDKNTGFFHATVH 338
Query: 232 GRRRRNSIISLLADSNLVEGVQPIRNAVL--SHFRDHFAARH--TSRVGIDNLPFKCLSH 287
R +N + S L D N E + + S F + F + + T ++ L K S
Sbjct: 339 SERLKNEL-SFLLDENDQEFTRNSDKGKIASSFFENLFTSTYILTHNNHLEGLQAKVTSE 397
Query: 288 TDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNG 347
+ + L + + EV +AV+ + +PGPDG F ++ WD +K ++ I F G
Sbjct: 398 MNHN-LIQEVTELEVYNAVFSINKESAPGPDGFTALFFQQHWDLVKHQILTEIFGFFETG 456
Query: 348 KLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQT 407
L + N T I LI KI SPQR++D RPISL LYKI+SK+L RLK L I+S +Q+
Sbjct: 457 VLPQDWNHTHICLIPKITSPQRMSDLRPISLCSVLYKIISKILTQRLKKHLPAIVSTTQS 516
Query: 408 AFVKDRQILDGILIANEVVDEARKNKR 434
AFV R I D IL+A+E++ R N R
Sbjct: 517 AFVPQRLISDNILVAHEMIHSLRTNDR 543
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 154 bits (390), Expect = 2e-37
Identities = 123/362 (33%), Positives = 184/362 (49%), Gaps = 31/362 (8%)
Query: 84 SDHCPLQLSV-DEENWGPRPSRMLKCWQDLPGYNQFVKDKLKSFQIEGW--------GGF 134
SDH P+ ++ D+ G + R K W KD L +GW G F
Sbjct: 4 SDHSPVIATIADKIPRGKQNFRFDKRW--------IGKDGLLEAISQGWNLDSGFREGQF 55
Query: 135 VLREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDDKGDVGDLSADEILELRGITH 194
V EKL + A+ +W S IP +++ +A+ D D S +EI EL T
Sbjct: 56 V--EKLTNCRRAISKWR---KSLIPFGRQTIEDLKAELDVAQRDDDRSREEITEL---TL 107
Query: 195 DLHSLSRVNSSILWQQSRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLADSNLVEGVQP 254
L R +Q+SR LW+K GD NSK+FH++ RR RN I L D N + ++
Sbjct: 108 RLKEAYRDEEQYWYQKSRSLWMKLGDNNSKFFHALTKQRRARNRITGL-HDENGIWSIED 166
Query: 255 --IRNAVLSHFRDHFAARHTSRVGIDNLPFKCLSHTDGSG--LTKPFSADEVKSAVWDCD 310
I+N +S+F++ F + +V + L + TD LT + EV++A++
Sbjct: 167 DDIQNIAVSYFQNLFTTANP-QVFDEALGEVQVLITDRINDLLTADATECEVRAALFMIH 225
Query: 311 SYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRI 370
K+PGPDG+ F ++ W +K D++ ++ F + G + +NTT I LI K + P R+
Sbjct: 226 PEKAPGPDGMTALFFQKSWAIIKSDLLSLVNSFLQEGVFDKRLNTTNICLIPKTERPTRM 285
Query: 371 NDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQILDGILIANEVVDEAR 430
+ RPISL YK++SK+L RLK VL +IS++Q+AFV R I D ILIA E+ R
Sbjct: 286 TELRPISLCNVGYKVISKILCQRLKTVLPNLISETQSAFVDGRLISDNILIAQEMFHGLR 345
Query: 431 KN 432
N
Sbjct: 346 TN 347
>At4g10830 putative protein
Length = 1294
Score = 152 bits (383), Expect = 1e-36
Identities = 122/426 (28%), Positives = 205/426 (47%), Gaps = 30/426 (7%)
Query: 23 YHHFSDFIVDNSLIDLSLCGRRFTWF--KGDGNAMSRIDRFLLSEEWCLQWPNSFQVALL 80
+ F + + L D+ G RF+W + +DR ++ E +P + L
Sbjct: 518 FRGFRNMVSTCDLKDIRSIGDRFSWVGERHSHTVKCCLDRAFINSEGAFLFPFAELEFLE 577
Query: 81 RGLSDHCPLQLSVDE-ENWGPRPSRMLKCWQDLPGYNQFVKDKLKSFQIEGWGGFV---- 135
SDH PL LS+++ E RP R K ++P + +VK GW +
Sbjct: 578 FTGSDHKPLFLSLEKTETRKMRPFRFDKRLLEVPHFKTYVK--------AGWNKAINGQR 629
Query: 136 --LREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDDKGDVGDLSADEILELRGIT 193
L ++++ + A+ + + H SN+ +I + + QA D +S+ E R I+
Sbjct: 630 KHLPDQVRTCRQAMAK--LKHKSNLNSRI-RINQLQAALDKA-----MSSVNRTERRTIS 681
Query: 194 HDLHSLSRV--NSSILWQQ-SRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLADSNLV- 249
H L+ + WQQ SR W+K+GD N+++FH+ R N ++++ + ++
Sbjct: 682 HIQRELTVAYRDEERYWQQKSRNQWMKEGDRNTEFFHACTKTRFSVNRLVTIKDEEGMIY 741
Query: 250 EGVQPIRNAVLSHFRDHFAARHTSRVGIDNLPFK-CLSHTDGSGLTKPFSADEVKSAVWD 308
G + I F + + ID FK ++ LTK S E+ +A+
Sbjct: 742 RGDKEIGVHAQEFFTKVYESNGRPVSIIDFAGFKPIVTEQINDDLTKDLSDLEIYNAICH 801
Query: 309 CDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQ 368
K+PGPDG+ F K W+ + DV++ + F R + + IN T I +I KI +P+
Sbjct: 802 IGDDKAPGPDGLTARFYKSCWEIVGPDVIKEVKIFFRTSYMKQSINHTNICMIPKITNPE 861
Query: 369 RINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQILDGILIANEVVDE 428
++D+RPI+L LYKI+SK L RLK L I+SDSQ AF+ R + D ++IA+E++
Sbjct: 862 TLSDYRPIALCNVLYKIISKCLVERLKGHLDAIVSDSQAAFIPGRLVNDNVMIAHEMMHS 921
Query: 429 ARKNKR 434
+ KR
Sbjct: 922 LKTRKR 927
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 150 bits (379), Expect = 4e-36
Identities = 121/403 (30%), Positives = 186/403 (46%), Gaps = 38/403 (9%)
Query: 42 GRRFTWFKG---DGNAMSRIDRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENW 98
G RFTW +G R+DR L L+W Q ALL CP Q +VD
Sbjct: 5 GNRFTWRRGLVESTFVAKRLDRVLFCAHARLKW----QEALL------CPAQ-NVDARR- 52
Query: 99 GPRPSRMLKCWQDLPGYNQFVKDKLKSFQIEGWGGFVLREKLKFIKTALKEWHIGHASNI 158
RP R W G+ + + + G L ++ LK+W+ NI
Sbjct: 53 --RPFRFEAAWLSHEGFKELLTASWDT-------GLSTPVALNRLRWQLKKWNKEVFGNI 103
Query: 159 ---PGKIDSLKKRQADFDDKGDVGDLSADEILELRGITHDLHSLSRVNSSILWQQSRLLW 215
K+ S K D + DL E L+ LH ++ +Q+SR
Sbjct: 104 HVRKEKVVSDLKAVQDLLEVVQTDDLLMKEDTLLKEFDVLLHQ----EETLWFQKSREKL 159
Query: 216 LKDGDANSKYFHSVLSGRRRRNSIISLLADSN--LVEGVQPIRNAVLSHFRDHFAARHTS 273
L GD N+ +FH+ RRRRN I +L DS V + + + ++R ++ S
Sbjct: 160 LALGDRNTTFFHTSTVIRRRRNRI-EMLKDSEDRWVTEKEALEKLAMDYYRKLYSLEDVS 218
Query: 274 RVGIDNLP---FKCLSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWD 330
V LP F L+ + + L +PF+ DEV AV +K+PGPDG F ++ W+
Sbjct: 219 VVR-GTLPTEGFPRLTREEKNNLNRPFTRDEVVVAVRSMGRFKAPGPDGYQPVFYQQCWE 277
Query: 331 ELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVL 390
+ E V +F+ +F +G L + N + L++K+ P+RI FRP+SL L+KI++K++
Sbjct: 278 TVGESVSKFVMEFFESGVLPKSTNDVLLVLLAKVAKPERITQFRPVSLCNVLFKIITKMM 337
Query: 391 ANRLKLVLGKIISDSQTAFVKDRQILDGILIANEVVDEARKNK 433
RLK V+ K+I +Q +F+ R D I++ E V R+ K
Sbjct: 338 VIRLKNVISKLIGPAQASFIPGRLSFDNIVVVQEAVHSMRRKK 380
>At4g26360 putative protein
Length = 1141
Score = 149 bits (377), Expect = 6e-36
Identities = 125/451 (27%), Positives = 207/451 (45%), Gaps = 57/451 (12%)
Query: 1 DFNAVRYAEERKSSRGISGVID--YHHFSDFIVDNSLIDLSLCGRRFTWFKGDGN--AMS 56
DFN V + E SR +S +D F + ++D L DL G FTW+
Sbjct: 136 DFNQVLHPHEH--SRHVSLNVDRRIRDFRECLLDAELSDLVYKGSSFTWWNKSKTRPVAK 193
Query: 57 RIDRFLLSEEWCLQWPNSFQVALLRGLSDH--CPLQLSVDEENWGPRPSRMLKCWQDLPG 114
+IDR L++E W +P+SF + SDH C + L +D RP + P
Sbjct: 194 KIDRILVNESWSNLFPSSFGLFGPPDFSDHASCGVVLELDPIK-AKRPFKFFNFLLKNPE 252
Query: 115 YNQFVKDKLKSFQIEGWGGFVLREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDD 174
+ V D S + G F + +KLK +K +K++ + SN+ + + + F +
Sbjct: 253 FLNLVWDVWYSTNVVGSSMFRVSKKLKALKKPIKDFSRLNYSNLEKRTEEAHETLLSFQN 312
Query: 175 KGDVGDLSADEILELRGITHDLHS------LSRVNSSILWQQSRLLWLKDGDANSKYFHS 228
L+ D L H+L + L+ S Q+SR+ W +GD N++YFH
Sbjct: 313 ------LTLDNP-SLENAAHELEAQRKWQILATAEESFFRQRSRVTWFAEGDGNTRYFHR 365
Query: 229 VLSGRRRRNSIISLLADSNL-VEGVQPIRNAVLSHFRDHFAARHTSRVGIDNLPFKC--- 284
+ R+ N+I +L+ DS ++ Q I DH A + + DN P+
Sbjct: 366 MADSRKSVNTITTLVDDSGTQIDSQQGIA--------DHCALYFENLLSDDNDPYSLEQD 417
Query: 285 ---------LSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKED 335
++ + L FS +++K+A + S K+ GPDG
Sbjct: 418 DMNLLLTYRCPYSQVADLEAMFSDEDIKAAFFGLPSNKACGPDGF--------------P 463
Query: 336 VMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLK 395
V + +F +G L + N T I LI K + +DFRPIS + +LYK+++++L +RL+
Sbjct: 464 VTAAVREFFISGNLLKQWNATTIVLIPKFPNASCTSDFRPISCMNTLYKVIARLLTDRLQ 523
Query: 396 LVLGKIISDSQTAFVKDRQILDGILIANEVV 426
+L +IS SQ+AF+ R + + +L+A E+V
Sbjct: 524 KLLSCVISPSQSAFLPGRLLAENVLLATEMV 554
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 146 bits (368), Expect = 7e-35
Identities = 134/459 (29%), Positives = 208/459 (45%), Gaps = 56/459 (12%)
Query: 1 DFNAVRYAEERKSSRGISGVIDYHHFSDFIVDNSLIDLSLCGRRFTWFKGDGNAM--SRI 58
DFN + E+K R S + +F++ I ++ DL G ++W N S +
Sbjct: 498 DFNEILNLNEKKGGRRRS-IGSLQNFTNMINCCNMKDLKSKGNPYSWVGKRQNETIESCL 556
Query: 59 DRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENWGPRPSRMLKCWQDLPGYNQF 118
DR ++ +W +P L SDH P+ + + EE R QF
Sbjct: 557 DRVFINSDWQASFPAFETEFLPIAGSDHAPVIIDIAEEVCTKR--------------GQF 602
Query: 119 VKDKLKSFQIE--------GW--------GGFVLREKLKFIKTALKEWHIGHASNIPGKI 162
D+ + FQ E GW GG+ EKL + L +W +N KI
Sbjct: 603 RYDR-RHFQFEDFVDSVQRGWNRGRSDSHGGYY--EKLHCCRQELAKWKRRTKTNTAEKI 659
Query: 163 DSLKKRQADFDDKGDVGDLSADEILELRGITHDLHSLSRVNSSILWQ-QSRLLWLKDGDA 221
++LK R D ++ L IL LR DL+ R + + W +SR W+ GD
Sbjct: 660 ETLKYR-VDAAERDHT--LPHQTILRLR---QDLNQAYR-DEELYWHLKSRNRWMLLGDR 712
Query: 222 NSKYFHSVLSGRRRRNSIISLLADSNLVEGVQP--IRNAVLSHFRDHFAARHTSR----- 274
N+ +F++ R+ RN I + D+ +E + I ++F D F TS
Sbjct: 713 NTMFFYASTKLRKSRNRI-KAITDAQGIENFRDDTIGKVAENYFADLFTTTQTSDWEEII 771
Query: 275 VGIDNLPFKCLSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKE 334
GI + ++H L + + EV+ AV+ + ++PG DG F WD +
Sbjct: 772 SGIAPKVTEQMNHE----LLQSVTDQEVRDAVFAIGADRAPGFDGFTAAFYHHLWDLIGN 827
Query: 335 DVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRL 394
DV + F + + IN T I LI KI P+ ++D+RPISL + YKI+SK+L RL
Sbjct: 828 DVCLMVRHFFESDVMDNQINQTQICLIPKIIDPKHMSDYRPISLCTASYKIISKILIKRL 887
Query: 395 KLVLGKIISDSQTAFVKDRQILDGILIANEVVDEARKNK 433
K LG +ISDSQ AFV + I D +L+A+E++ + +
Sbjct: 888 KQCLGDVISDSQAAFVPGQNISDNVLVAHELLHSLKSRR 926
>At1g24640 hypothetical protein
Length = 1270
Score = 145 bits (366), Expect = 1e-34
Identities = 121/450 (26%), Positives = 204/450 (44%), Gaps = 36/450 (8%)
Query: 1 DFNAVRYAEERKSSRGISGVIDYHHFSDFIVDNSLIDLSLCGRRFTWF--KGDGNAMSRI 58
DFN + + E+ S +D F++ I L+++ G FTW +GD R+
Sbjct: 99 DFNDILHNGEKNGGPRRSD-LDCKAFNEMIKGCDLVEMPAHGNGFTWAGRRGDHWIQCRL 157
Query: 59 DRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENWGPRPSRMLKCWQDLPGYNQF 118
DR ++EW +P S Q L SDH P+ + +++ G +F
Sbjct: 158 DRAFGNKEWFCFFPVSNQTFLDFRGSDHRPVLI------------KLMSSQDSYRGQFRF 205
Query: 119 -----VKDKLKSFQIEGWG------GFVLREKLKFIKTALKEWHIGHASNIPGKIDSLKK 167
K+ +K I W + ++L+ + +L W N +D + +
Sbjct: 206 DKRFLFKEDVKEAIIRTWSRGKHGTNISVADRLRACRKSLSSWK---KQNNLNSLDKINQ 262
Query: 168 RQADFDDKGDVGDLSADEILELRGITHDLHSLSRVNSSILWQQSRLLWLKDGDANSKYFH 227
+A + + + + L+ DL R + Q+SR WL+ G+ NSKYFH
Sbjct: 263 LEAALEKEQSLVWPIFQRVSVLK---KDLAKAYREEEAYWKQKSRQKWLRSGNRNSKYFH 319
Query: 228 SVLSGRRRRNSIISLLADSNLVEGVQPIRNAVLS-HFRDHFAARHTSRVG--IDNLPFKC 284
+ + R+R I L + ++ + + V + +F + F + + S L +
Sbjct: 320 AAVKQNRQRKRIEKLKDVNGNMQTSEAAKGEVAAAYFGNLFKSSNPSGFTDWFSGLVPR- 378
Query: 285 LSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFH 344
+S L SA E+K AV+ +PGPDG++ F + +W + V + F
Sbjct: 379 VSEVMNESLVGEVSAQEIKEAVFSIKPASAPGPDGMSALFFQHYWSTVGNQVTSEVKKFF 438
Query: 345 RNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISD 404
+G + N T + LI K P + D RPISL LYKI+SK++A RL+ L +I+SD
Sbjct: 439 ADGIMPAEWNYTHLCLIPKTQHPTEMVDLRPISLCSVLYKIISKIMAKRLQPWLPEIVSD 498
Query: 405 SQTAFVKDRQILDGILIANEVVDEARKNKR 434
+Q+AFV +R I D IL+A+E+V + + R
Sbjct: 499 TQSAFVSERLITDNILVAHELVHSLKVHPR 528
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 144 bits (362), Expect = 4e-34
Identities = 89/261 (34%), Positives = 135/261 (51%), Gaps = 7/261 (2%)
Query: 188 ELRGITHDLHSLSRVNSSILWQQSRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLA-DS 246
EL+ T H LS + L Q+S+L W+ GD N+ YFH R+ RNSI + ++
Sbjct: 636 ELKAYTDWTH-LSELEEGFLKQKSKLHWMNVGDGNNSYFHKAAQVRKMRNSIREIRGPNA 694
Query: 247 NLVEGVQPIRNAVLSHFRDHFAARHTSRVGID-----NLPFKCLSHTDGSGLTKPFSADE 301
++ + I+ F + + GI NL S TD + LT+ + +E
Sbjct: 695 ETLQTSEEIKGEAERFFNEFLNRQSGDFHGISVEDLRNLMSYRCSVTDQNILTREVTGEE 754
Query: 302 VKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALI 361
++ ++ + KSPGPDG F K W D + I F G L +G+N T +ALI
Sbjct: 755 IQKVLFAMPNNKSPGPDGYTSEFFKATWSLTGPDFIAAIQSFFVKGFLPKGLNATILALI 814
Query: 362 SKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQILDGILI 421
K D + D+RPIS LYK++SK+LANRLKL+L I +Q+AFVK+R +++ +L+
Sbjct: 815 PKKDEAIEMKDYRPISCCNVLYKVISKILANRLKLLLPSFILQNQSAFVKERLLMENVLL 874
Query: 422 ANEVVDEARKNKRN*CCSKLI 442
A E+V + K C+ I
Sbjct: 875 ATELVKDYHKESVTPRCAMKI 895
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 143 bits (361), Expect = 5e-34
Identities = 124/431 (28%), Positives = 208/431 (47%), Gaps = 34/431 (7%)
Query: 26 FSDFIVDNSLIDLSLCGRRFTWFKGDGNA---MSRIDRFLLSEEWCLQWPNSFQVALLRG 82
F F+ N L D++ G +W +G + SR+DR L + W +P S L
Sbjct: 104 FRSFVSQNGLWDINHTGNSLSW-RGTRYSHFIKSRLDRALGNCSWSELFPMSKCEYLRFE 162
Query: 83 LSDHCPLQLSVDEENWGPRPSRMLKCWQDLPGYNQFVKDKLKSFQIEGWGGFVLRE---- 138
SDH PL +G P + K ++ + K+++++ E W + R+
Sbjct: 163 GSDHRPLVTY-----FGAPPLKRSKPFRFDRRLRE--KEEIRALVKEVWE--LARQDSVL 213
Query: 139 -KLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDD--KGDVGDLSADEILELRGITHD 195
K+ + ++ +W SN ++KK Q + D+ D S + IT +
Sbjct: 214 YKISRCRQSIIKWTKEQNSN---SAKAIKKAQQALESALSADIPDPSL-----IGSITQE 265
Query: 196 LHSLSRVNSSILWQQ-SRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLADSNLVE--GV 252
L + R + W+Q SR+ WL GD N YFH+ RR N++ S++ D + E
Sbjct: 266 LEAAYR-QEELFWKQWSRVQWLNSGDRNKGYFHATTRTRRMLNNL-SVIEDGSGQEFHEE 323
Query: 253 QPIRNAVLSHFRDHFAARHTSRVGIDNLPFK-CLSHTDGSGLTKPFSADEVKSAVWDCDS 311
+ I + + S+F++ F + S + + +S L K S E+K A++ +
Sbjct: 324 EQIASTISSYFQNIFTTSNNSDLQVVQEALSPIISSHCNEELIKISSLLEIKEALFSISA 383
Query: 312 YKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRIN 371
K+PGPDG + F +WD ++ DV R I F + L+ +N T + LI KI +P++++
Sbjct: 384 DKAPGPDGFSASFFHAYWDIIEADVSRDIRSFFVDSCLSPRLNETHVTLIPKISAPRKVS 443
Query: 372 DFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQILDGILIANEVVDEARK 431
D+RPI+L YKI++K+L RL+ L ++IS Q+AFV R I D +LI +E++ R
Sbjct: 444 DYRPIALCNVQYKIVAKILTRRLQPWLSELISLHQSAFVPGRAIADNVLITHEILHFLRV 503
Query: 432 NKRN*CCSKLI 442
+ CS I
Sbjct: 504 SGAKKYCSMAI 514
>At2g23880 putative non-LTR retroelement reverse transcriptase
Length = 1216
Score = 143 bits (360), Expect = 6e-34
Identities = 83/231 (35%), Positives = 129/231 (54%), Gaps = 8/231 (3%)
Query: 219 GDANSKYFHSVLSGRRRRNSIISLLA-DSNLVEGVQPIRNAVLSHFRDHFAARHTSRVGI 277
GD N+K FH ++ R NSI ++ D +V Q I+ +++F+D G+
Sbjct: 90 GDRNNKTFHRAITTREAVNSIREIVTRDGLVVTSQQDIQTEAVNYFQDFLQTIPADYEGM 149
Query: 278 ------DNLPFKCLSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDE 331
+ LPF+C S D LT+ + +E+K ++ KSPGPDG F K W+
Sbjct: 150 CVEELENLLPFRC-SEDDHRLLTRVVTGEEIKKVIFSMPKDKSPGPDGYTSEFYKASWEI 208
Query: 332 LKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLA 391
+ ++V+ I F G L +G+N+T +ALI K + I D+RPIS LYK +SK+LA
Sbjct: 209 IGDEVIIAIQSFFAKGFLPKGVNSTILALIPKKKEAREIKDYRPISCCNVLYKAISKILA 268
Query: 392 NRLKLVLGKIISDSQTAFVKDRQILDGILIANEVVDEARKNKRN*CCSKLI 442
NRLK +L K I +Q+AFVKDR +++ +L+A E+V + K+ + C+ I
Sbjct: 269 NRLKRILPKFIVGNQSAFVKDRLLIENVLLATELVKDYHKDSISTRCAMKI 319
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.344 0.155 0.538
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,369,083
Number of Sequences: 26719
Number of extensions: 997479
Number of successful extensions: 3833
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 53
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 3664
Number of HSP's gapped (non-prelim): 106
length of query: 1087
length of database: 11,318,596
effective HSP length: 110
effective length of query: 977
effective length of database: 8,379,506
effective search space: 8186777362
effective search space used: 8186777362
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 65 (29.6 bits)
Medicago: description of AC135464.5


