

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135463.4 - phase: 2 /pseudo
(1134 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
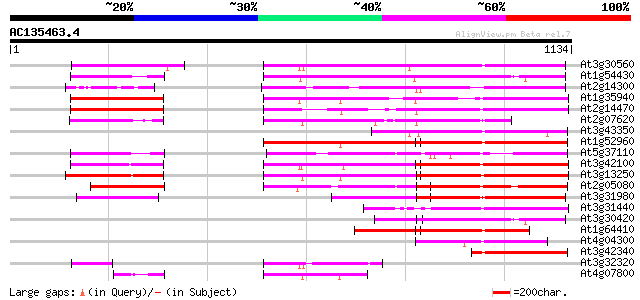
Score E
Sequences producing significant alignments: (bits) Value
At3g30560 hypothetical protein 451 e-127
At1g54430 hypothetical protein 445 e-125
At2g14300 pseudogene; similar to MURA transposase of maize Muta... 409 e-114
At1g35940 hypothetical protein 398 e-110
At2g14470 pseudogene 355 8e-98
At2g07620 putative helicase 327 2e-89
At3g43350 putative protein 305 7e-83
At1g52960 hypothetical protein 286 4e-77
At5g37110 putative helicase 286 6e-77
At3g42100 putative protein 280 4e-75
At3g13250 hypothetical protein 271 1e-72
At2g05080 putative helicase 260 3e-69
At3g31980 hypothetical protein 258 1e-68
At3g31440 hypothetical protein 248 2e-65
At3g30420 hypothetical protein 231 2e-60
At1g64410 unknown protein 223 6e-58
At4g04300 hypothetical protein 199 6e-51
At3g42340 putative protein 163 6e-40
At3g32320 hypothetical protein 156 7e-38
At4g07800 hypothetical protein 135 1e-31
>At3g30560 hypothetical protein
Length = 1473
Score = 451 bits (1161), Expect = e-127
Identities = 262/651 (40%), Positives = 373/651 (57%), Gaps = 45/651 (6%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P++ RRDTG + KN DNR VVPYN L+ KY AHIN+E+CN+S S+KYLFKY+ K
Sbjct: 648 YPIYRRRDTGRFIKKNKYPCDNRYVVPYNDFLLRKYRAHINVEWCNQSVSVKYLFKYVNK 707
Query: 574 GVDRVTATLE------TSEEPSVDE----------IQ*YYDCRYLSPSESIWRIFGFDVH 617
G DRVT +LE SEE +V E ++ Y+DCRY+S E++WRI G+ +H
Sbjct: 708 GPDRVTVSLEPHRKEVVSEENNVGETNNDPQEQNQVEDYFDCRYVSACEAMWRIKGYPIH 767
Query: 618 CRYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYV 677
R V +LTFH G+Q I ++ ++VL R T F AW + N+ ++ L Y
Sbjct: 768 YRQTLVTKLTFHEKGKQPIYVKEGETAESVLYRVNDDETQFTAWFELNKRDPEAAKLLYE 827
Query: 678 QFPRKFTYNVENRSWHL*K-SGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTV 736
Q P +T+N +++++ K G VGR+ VPP + Y +R+L+N G F+DI+T+
Sbjct: 828 QIPNFYTWNGKDKNFRRRKMPGFVVGRINHVPPKIDDAYHLRILINNIRGPKGFDDIKTI 887
Query: 737 DGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVW 796
+G V+ +YR+AC ALGLL +D+ +I+ I E +RK+F +L+ +S P VW
Sbjct: 888 EGVVHKTYRDACYALGLLDDDKDYINGIEEANFWCFHKYVRKLFVIMLIFESLSSPAVVW 947
Query: 797 NQTWHILSEGI-----------------------LYERRRTLNSPG*LLNSDQLSAYEKI 833
TW ILSE L E+ PG +QL E+
Sbjct: 948 EHTWKILSEDFQRKVRDKLKCPAAKNIETTWNRSLEEKMLPQLKPGDEPAFNQLILDERN 1007
Query: 834 VNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEG-KIVLNVASSSIASLLLPRGRTA 892
N + H ++ K + ++G I LNVASS IASLLL GRTA
Sbjct: 1008 YNRETLKTIHDDWLKMLTTEQKKVYDKIMDAVLNNKGGDICLNVASSGIASLLLEGGRTA 1067
Query: 893 HSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMRDIMSN 952
HS+F IPL E S C +E+ + A+L+T A LIIWDEAPM+S+ FE+ D++++DI+S
Sbjct: 1068 HSRFGIPLTPHETSTCNMERGSDLAELVTAAKLIIWDEAPMMSKYCFESLDKSLKDILST 1127
Query: 953 VVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRVLKLTKNIR 1012
D +PFGGK ++ GGDFRQILPV+ GR IV + +NSS LW+ C+V KLTKN+R
Sbjct: 1128 PED----MPFGGKLIIFGGDFRQILPVILAAGRELIVKSSLNSSHLWQYCKVFKLTKNMR 1183
Query: 1013 LQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKNSGNHVGDIVEST 1072
L + + + F++WIL VG+GKL NDG I+I DDI + N + I+++
Sbjct: 1184 LLQDIDINEAREIEDFSKWILAVGEGKLNQPNDGVTQIQIRDDILIPEGDNPIESIIKAV 1243
Query: 1073 YPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSCDTV 1123
Y ++ FFQDRAIL PT + V +ND++LS + GEEK Y S D++
Sbjct: 1244 YGTSFDEERDPKFFQDRAILCPTNDDVNSINDHMLSKLTGEEKIYRSSDSI 1294
Score = 165 bits (418), Expect = 1e-40
Identities = 97/234 (41%), Positives = 136/234 (57%), Gaps = 7/234 (2%)
Query: 126 RSLVQDLMKLMDECNLLVKKFRMVRD-FREANVDVPVKLRLFRNRNFDSRVYNVPEISEV 184
+ V L+K++DE N V FR+ RD F D +R+ NR+ D RVYN+P + EV
Sbjct: 252 KETVSALLKMLDEINPHVANFRIARDRFNIEKEDANFHMRIISNRDTDGRVYNLPSVGEV 311
Query: 185 VALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDRYEEKIKLNH 244
ALI GDFD D RDIV++ K LRRIHE H Y+ LQYP+LFP GED Y IK
Sbjct: 312 -ALIPGDFDDNLDKRDIVLQIKSEKLRRIHECHVSYLSLQYPLLFPKGEDGYRLGIKKTE 370
Query: 245 RTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLSYVR 304
T+ KK+ VS+R++ YRLQ+R +E ++ + RL QQFIVD ++M+E+ RL +++
Sbjct: 371 TNTSKRKKKQKDVSMRQWFDYRLQERKDEKHILLRSKRLLQQFIVDAFTMIESNRLRFIK 430
Query: 305 KNQSQIRSGLLWL-----DSRDKFDSPKAINSIICAELPDSQMYPELFWFDALG 353
KNQ+++RS D+ D S K + II Y + + DA+G
Sbjct: 431 KNQTKLRSTNKQAVQDASDAGDNDLSNKGKSVIIPPSFTGGPAYMQQNYLDAMG 484
Score = 32.7 bits (73), Expect = 1.2
Identities = 13/34 (38%), Positives = 22/34 (64%)
Query: 314 LLWLDSRDKFDSPKAINSIICAELPDSQMYPELF 347
++W+D R KF ++ II AE+PD + P+L+
Sbjct: 569 IVWMDPRYKFPIADHVDKIIFAEIPDKENDPKLY 602
>At1g54430 hypothetical protein
Length = 1639
Score = 445 bits (1145), Expect = e-125
Identities = 258/665 (38%), Positives = 376/665 (55%), Gaps = 63/665 (9%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
F L+ RR+ VLK +LDNR VVP+N +++ KY AHIN+E+CNKS++IKYLFKYITK
Sbjct: 794 FILYRRRNDQRYVLKGQTRLDNRFVVPHNLEILKKYKAHINVEWCNKSSAIKYLFKYITK 853
Query: 574 GVDRVTATLETS------------EEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHCRYP 621
GVD+ T ++ EE +EI Y DCRYLS E++WRIF F++H P
Sbjct: 854 GVDKATFIIQKGNSVNGQGSGNGFEEKPRNEINEYLDCRYLSACEAMWRIFMFNIHHHNP 913
Query: 622 AVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPR 681
V+RL HL GEQ +F + +NV R H+RTM + + N+ + L YVQ P
Sbjct: 914 PVQRLPLHLPGEQSTIFEEEENLENVEYRYGHERTMLTEYFELNKICEDARKLKYVQVPT 973
Query: 682 KFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVY 741
F ++ N+ + K +++GR+ + P +LY +R+LLN+ G TSF+ ++TV G V+
Sbjct: 974 MFVWDSTNKMYTRRKQRENIGRIVNILPTAGDLYYLRILLNKVKGATSFDYLKTVGGVVH 1033
Query: 742 NSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWH 801
S++ AC GLL D+++ D + E + +R +F +L+ +S+PL +W+ W
Sbjct: 1034 ESFKAACHTRGLLDGDKEWHDAMDEAAQWSTSYLLRSLFVLILIYCEVSEPLKLWSHCWE 1093
Query: 802 ILSEGILYERRRTLN-------------------------------------SPG*LLNS 824
+++ + ++++ LN P LN
Sbjct: 1094 SMADDVFRKQQKVLNFPQLELKAEELEKYTLIEIETLLRQHEKSLSDYPEMPQPENKLNE 1153
Query: 825 DQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSIASL 884
Q Y+ ++ V N+ +FF+ G GG GKTFL+ + RS GK V+ VASS+IA+L
Sbjct: 1154 QQRIIYDDVLKSVINKEGKLFFLYGAGGTGKTFLYKTIISALRSNGKNVMPVASSAIAAL 1213
Query: 885 LLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDR 944
LLP GRTAHS+F IP+ + E+S C I+ A +L+ LIIWDEAPM R FEA DR
Sbjct: 1214 LLPGGRTAHSRFKIPINVHEDSICDIKIGSMLANVLSKVDLIIWDEAPMAHRHTFEAVDR 1273
Query: 945 TMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRV 1004
T+RDI+S + A FGGKTV+LGGDFRQILPV+P+ R + V A IN S LW C
Sbjct: 1274 TLRDILSVGDEKALTKTFGGKTVLLGGDFRQILPVIPQGTRQETVSAAINRSYLWESCHK 1333
Query: 1005 LKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKL-----GIDNDGEA-VIEIPDDICL 1058
L++N+R+Q P + FA WIL VGDG+ GID+D E I I ++ L
Sbjct: 1334 YLLSQNMRVQ-PEEIK-------FAEWILQVGDGEAPRKTHGIDDDQEEDNIIIDKNLLL 1385
Query: 1059 KNSGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYL 1118
+ N + + S +P+ + ++ + A+L P E V+++NDY+LS VPG KEY
Sbjct: 1386 PETENPLEVLCRSVFPDFTNTFQDLENLKGTAVLTPRNETVDEINDYLLSKVPGLAKEYF 1445
Query: 1119 SCDTV 1123
S D++
Sbjct: 1446 SADSI 1450
Score = 97.1 bits (240), Expect = 5e-20
Identities = 59/189 (31%), Positives = 91/189 (47%), Gaps = 29/189 (15%)
Query: 124 LDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNFDSRVYNVPEISE 183
L+ +V+DL+ +D N L K FR RD EA ++L + + Y++P E
Sbjct: 424 LNEKIVKDLITTVDTFNCLAKVFRKARDRYEAGDCPEFSIKLIGQKK-KGKQYDMPTTDE 482
Query: 184 VVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDRYEEKIKLN 243
+ LIVGDF RD++V K GL++I + H ++ LQYP+LFP GE + E I +
Sbjct: 483 IAGLIVGDFSKNIGERDVIVHHKSSGLQQISDLHPLFMTLQYPLLFPYGEIGFHEGIPVV 542
Query: 244 HRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLSYV 303
+ T I + RL Q+IVD Y+ +E +RL +
Sbjct: 543 EKGMT----------------------------IVRSKRLLHQYIVDAYTSIEQERLRWY 574
Query: 304 RKNQSQIRS 312
R NQ ++R+
Sbjct: 575 RLNQKKLRA 583
>At2g14300 pseudogene; similar to MURA transposase of maize Mutator
transposon
Length = 1230
Score = 409 bits (1052), Expect = e-114
Identities = 243/658 (36%), Positives = 352/658 (52%), Gaps = 102/658 (15%)
Query: 509 VEFKCFPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLF 568
+++K +P++ RRD+G + KN Q DN VVPYN L+ KY AHIN+E+CN+S SIKYLF
Sbjct: 495 LDYKGYPIYRRRDSGRFIEKNKYQCDNWYVVPYNDVLLRKYRAHINVEWCNQSVSIKYLF 554
Query: 569 KYITKGVDRVTATL--ETSEEPSV-DEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAVKR 625
KY+ KG DRVT E + +P +++Q Y+DCR G+ +H R +V +
Sbjct: 555 KYVNKGPDRVTQNNVGEINNDPQERNQVQDYFDCR------------GYPIHYRQTSVTK 602
Query: 626 LTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPRKFTY 685
LTFH G+Q + ++ ++VL R + T F+AW + N+ ++ L Y Q P +T
Sbjct: 603 LTFHEKGKQSVYVKEGETAESVLYRVNNDETQFIAWFELNKRDPEAAKLLYEQIPNFYTI 662
Query: 686 NVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYR 745
N VPP + Y +R+L+N F+DI+TV+G V+ +YR
Sbjct: 663 N-------------------HVPPKIDDAYHLRILINNIRAPKGFDDIKTVEGVVHKTYR 703
Query: 746 EACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSE 805
+AC ALGLL +D+++I I E S +RK F +L++ +S P+ VW TW IL E
Sbjct: 704 DACYALGLLDDDKEYIHGIEEANFWCSPKYVRKSFVIMLISESLSSPVVVWEHTWKILFE 763
Query: 806 GILYERRRTLNSPG*L-----------------------LNSDQLSA------------- 829
+ R L P N + L
Sbjct: 764 DFQRKVRDKLERPDLWRYKMLLEPGDEPAFNPLIIDERNYNRESLKKKHDNWLKTLTPEH 823
Query: 830 ----YEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSIASLL 885
+++I++ V N+ +FF+ +GG GKTFLW LS R +G LNVASSSIASLL
Sbjct: 824 KKVYHDEIMDDVLNDKGGVFFLYAFGGTGKTFLWKVLSAAIRCKGDTCLNVASSSIASLL 883
Query: 886 LPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRT 945
L GRTAHS+F IPL E S C +E+ + A+L+T A LIIWDE
Sbjct: 884 LEGGRTAHSRFGIPLTPHETSTCNMERGSDLAELVTAAKLIIWDE--------------- 928
Query: 946 MRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRVL 1005
++PFG K ++ GGDFRQIL V+P GR IV + +NSS LW+ C+VL
Sbjct: 929 -------------DMPFGRKVILFGGDFRQILHVIPAAGRELIVKSSLNSSYLWQHCKVL 975
Query: 1006 KLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKNSGNHV 1065
KLTKN+RL + + + F +WIL VG+GKL +DG I+IPDDI + N +
Sbjct: 976 KLTKNMRLLQDIDINEAREIEDFLKWILTVGEGKLNEPSDGVTHIQIPDDILIPEGDNPI 1035
Query: 1066 GDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSCDTV 1123
I+++ Y + ++ FFQ +AIL PT + V +ND++LS + GEE+ Y S +++
Sbjct: 1036 ESIIKAVYGTTFAQKRDPKFFQHKAILCPTNDDVNSINDHMLSKLTGEERIYRSSNSI 1093
Score = 106 bits (264), Expect = 9e-23
Identities = 70/180 (38%), Positives = 100/180 (54%), Gaps = 22/180 (12%)
Query: 114 YHSELSDGNGLDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNFDS 173
+HS L +D+ L +D++ L C + +F + ++ EAN +R+ R D
Sbjct: 138 FHSHLLVARWIDQ-LKRDVVHL---CLIAHDRFNIEKE--EANFH----MRIISKRETDG 187
Query: 174 RVYNVPEISEVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGE 233
RVYN+P ++EV ALI GDFD D +DIV++ K G LRRIHE H YP+LFP GE
Sbjct: 188 RVYNLPSVAEVAALIPGDFDDNLDKKDIVLQMKSGKLRRIHECH-------YPLLFPKGE 240
Query: 234 DRYEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYS 293
D Y IK T+ K S+R++ YRLQ+R +E ++ + RL QQF+ L S
Sbjct: 241 DGYRLGIKKTPTKTSKGKK-----SMRQWFDYRLQERKDEKHILLRSKRLLQQFMTKLRS 295
Score = 37.7 bits (86), Expect = 0.038
Identities = 14/34 (41%), Positives = 24/34 (70%)
Query: 314 LLWLDSRDKFDSPKAINSIICAELPDSQMYPELF 347
++W+D R KF + ++ II AE+PD + +PEL+
Sbjct: 421 IVWMDPRYKFHTADHVDKIIFAEIPDKEKHPELY 454
>At1g35940 hypothetical protein
Length = 1678
Score = 398 bits (1022), Expect = e-110
Identities = 248/684 (36%), Positives = 355/684 (51%), Gaps = 130/684 (19%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P++ RR T V K GI+ DNR VVPYN KL ++Y+AHIN+E+CN++ SIKYLFKYI K
Sbjct: 883 YPIYRRRKTDDYVEKGGIKCDNRYVVPYNKKLSLRYNAHINVEWCNQNGSIKYLFKYINK 942
Query: 574 GVDRVTATLET-----------------SEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDV 616
G D+V +E S E +EI+ ++DCRY+S SE++WRIF + +
Sbjct: 943 GPDKVVFIVEPTQQTTAGDSETPQQEQGSAEKKKNEIKDWFDCRYVSASEAVWRIFKYPI 1002
Query: 617 HCRYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANR------NYSQ 670
V++L+FH+ G+Q F S ++VL R + + F+AW+ NR N +
Sbjct: 1003 QHISTPVQKLSFHVEGKQPAYFDPKSNIEDVLERVANVDSQFMAWLTLNRRNAVGKNGKR 1062
Query: 671 SWNLTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSF 730
+ Y + P FT++ EN+S+ G S+GR+ +V ++ Y +R+LLN G TS+
Sbjct: 1063 ARECLYAEIPAYFTWDGENKSFKKRTRGFSIGRIHYVSRKMEDEYFLRVLLNIVRGPTSY 1122
Query: 731 EDIRTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMS 790
+I+T DG VY +++EAC A G+L +D+ FID + E
Sbjct: 1123 AEIKTYDGVVYKTFKEACFARGILDDDQVFIDGLVEAT---------------------- 1160
Query: 791 DPLNVWNQTWHILSEGILYERRRTLNSPG------------------------------- 819
+V +QTWH+L+E IL +R +P
Sbjct: 1161 ---HVRSQTWHLLAEDILKTKRDEFKNPDLTLTETEIKNYTLQEIEKIMLSNGATLEDID 1217
Query: 820 -----------*LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRS 868
+L +Q Y I V N L +FFV G+GG GKTF+W LS R
Sbjct: 1218 EFPKPTRDEWKQMLTPEQRGVYNAITEAVFNNLGGVFFVYGFGGTGKTFIWKTLSAAIRC 1277
Query: 869 EGKIVLNVASSSIASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIW 928
G+IVLNVASS IASLLL GRTAHS+F IPL E S
Sbjct: 1278 RGQIVLNVASSGIASLLLEGGRTAHSRFGIPLNHDEFS---------------------- 1315
Query: 929 DEAPMISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADI 988
+ D++ DI+ N +N FGGK VV GGDFRQ+LPV+ GRA+I
Sbjct: 1316 -----------VSLDKSFSDIIKN----TNNKVFGGKVVVFGGDFRQVLPVINGAGRAEI 1360
Query: 989 VDACINSSMLWRRCRVLKLTKNIRLQFPTNGQDDA-SLRLFARWILDVGDGKLGIDNDGE 1047
V + +N+S LW C+VLKLTKN+RL +A ++ F+ W+L V DG++ NDG
Sbjct: 1361 VMSSLNASYLWDHCKVLKLTKNMRLLANNLSATEAKEIQEFSDWLLAVSDGRINEPNDGV 1420
Query: 1048 AVIEIPDDICLKNSGNHVGDIVESTY--PNLLSNMKESSFFQDRAILAPTLELVEKVNDY 1105
A I+IP+D+ + N+ + I Y P +L + + FFQ RAILAP E V +N+Y
Sbjct: 1421 ATIDIPEDLLITNADKPIETITNEIYGDPKILHEITDPKFFQGRAILAPKNEDVNTINEY 1480
Query: 1106 VLSLVPGEEKEYLSCDTVLKCDEE 1129
+L + EE+ YLS D++ D +
Sbjct: 1481 LLEQLDAEERIYLSADSIDPTDSD 1504
Score = 136 bits (343), Expect = 6e-32
Identities = 69/190 (36%), Positives = 122/190 (63%), Gaps = 3/190 (1%)
Query: 124 LDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRN-FDSRVYNVPEIS 182
LD++L++ ++K+++ CN V++FR R+ + N + P +R+ +R + R Y++P S
Sbjct: 484 LDKNLIEAIIKMLNRCNPYVRRFRTARERIQTNDEEPFHMRIIADRQGVEGRTYSMPTTS 543
Query: 183 EVVALIVGDFDSFEDGRDIVVREKDGG-LRRIHETHSKYIPLQYPVLFPLGEDRYEEKIK 241
EV ALI GDF RDIV+ +K G L+RI++ H Y+ LQYP++F GED + I+
Sbjct: 544 EVAALIPGDFRHGMPDRDIVIGKKSNGHLKRINQIHISYLALQYPLIFCYGEDGFRPGIE 603
Query: 242 LNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLS 301
++ + K+ +S+R++ A+R+Q+R+ E + + RLFQQ + D Y+ +E+ RL+
Sbjct: 604 KCSKSKSKKKNKKC-ISMRQWFAFRIQEREVECQTLLRSKRLFQQCLCDAYTTIESNRLN 662
Query: 302 YVRKNQSQIR 311
Y++ NQS++R
Sbjct: 663 YIKFNQSKLR 672
>At2g14470 pseudogene
Length = 1265
Score = 355 bits (911), Expect = 8e-98
Identities = 228/664 (34%), Positives = 322/664 (48%), Gaps = 145/664 (21%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P++ RR V K GI+ DNR V+PYN K ++Y+AHIN+E+CN+++SIKYLFKYI K
Sbjct: 542 YPIYRRRKIDDYVEKGGIKCDNRYVMPYNKKFSLRYNAHINVEWCNQNDSIKYLFKYINK 601
Query: 574 GVDRVTATLETSEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAVKRLTFHLHGE 633
G D+V +E +++ +
Sbjct: 602 GPDKVIFIVEPTQQATAG------------------------------------------ 619
Query: 634 QRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNY-----SQSWNLTYVQFPRKFTYNVE 688
DS P K+ W D RN ++ Y + P FT++ E
Sbjct: 620 ------DSETPQQEQRSAEKKKNEIKDWFDCRRNAVGKNGKRARECLYAEIPAYFTWDGE 673
Query: 689 NRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREAC 748
N+++ G S+GR+ +V ++ Y +R+LLN +
Sbjct: 674 NKAFKKRTRGFSIGRIHYVSRKMEDDYFLRVLLNISV----------------------- 710
Query: 749 AALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGIL 808
F L E F LLL+ +S P +VW+QTWHIL+E IL
Sbjct: 711 ----------LFWRLSQEF------------FAMLLLSDSLSRPAHVWSQTWHILAEDIL 748
Query: 809 YERRRTLNSPG*L----------------------------------------LNSDQLS 828
++R +P + L +Q
Sbjct: 749 KKKRDEFKNPEDIDEFPKPTIDGIDNSNRLIVEELRYNRESNLKEKHEEWKQMLTPEQRG 808
Query: 829 AYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSIASLLLPR 888
Y +I V N L +FFV G+GG GKTF+W LS R +IVLNVASS IASLLL
Sbjct: 809 VYNEITEAVFNNLGGVFFVYGFGGTGKTFIWKTLSATIRYRDQIVLNVASSGIASLLLEG 868
Query: 889 GRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMRD 948
GRTAHS+F IPL E S C I+ + + A L+ ASL+IWDEAPM+SR FEA D++ D
Sbjct: 869 GRTAHSRFGIPLNPDEFSVCKIKPKSDLANLVKKASLVIWDEAPMMSRFCFEALDKSFSD 928
Query: 949 IMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRVLKLT 1008
I+ N N FGGK VV GGDFRQ+ PV+ GRA+IV + +N+S LW C+VLKLT
Sbjct: 929 IIKN----TDNTVFGGKVVVFGGDFRQVFPVINGAGRAEIVMSSLNASYLWDNCKVLKLT 984
Query: 1009 KNIRLQFPTNGQDDA-SLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKNSGNHVGD 1067
KN RL + +A ++ F+ W+L VGDG++ NDG A+I+IP+D+ + N+ +
Sbjct: 985 KNTRLLANNLSETEAKEIQEFSDWLLAVGDGRINESNDGVAIIDIPEDLLITNADKPIES 1044
Query: 1068 IVESTY--PNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSCDTVLK 1125
I Y P +L + + FFQ RAILA E V +N+Y+L + EE+ YLS D++
Sbjct: 1045 ITNEIYGDPKILHEITDPKFFQGRAILASKNEDVNTINEYLLDQLHAEERIYLSADSIDP 1104
Query: 1126 CDEE 1129
D +
Sbjct: 1105 TDSD 1108
Score = 139 bits (350), Expect = 9e-33
Identities = 71/190 (37%), Positives = 122/190 (63%), Gaps = 3/190 (1%)
Query: 124 LDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRN-FDSRVYNVPEIS 182
LD++L++ ++K+++ CN V+KFR R+ + N + P +R+ +R D R Y++ S
Sbjct: 143 LDKNLIEVIIKMLNRCNPYVRKFRTARERIQTNDEEPFHMRIIADRQGVDGRTYSMHTTS 202
Query: 183 EVVALIVGDFDSFEDGRDIVVREKDGG-LRRIHETHSKYIPLQYPVLFPLGEDRYEEKIK 241
EV ALI GDF RDIV+ +K G L+RI++ H Y+ LQYP++F GED + I+
Sbjct: 203 EVAALIPGDFRHGMPDRDIVIEKKSNGHLKRINQIHISYLALQYPLIFCYGEDGFRPGIE 262
Query: 242 LNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLS 301
++ + K+ +S+R++ A+R+Q+R+ E + + RLFQQF+ D Y+ +E+ RL+
Sbjct: 263 KCFKSKSKKKNKKC-ISMRQWFAFRIQEREVECQTLLRSKRLFQQFLCDAYTTIESNRLN 321
Query: 302 YVRKNQSQIR 311
Y++ NQS++R
Sbjct: 322 YIKFNQSKLR 331
Score = 30.0 bits (66), Expect = 7.8
Identities = 12/34 (35%), Positives = 22/34 (64%)
Query: 314 LLWLDSRDKFDSPKAINSIICAELPDSQMYPELF 347
LL++ ++ K + I+ +I AE+PD + PEL+
Sbjct: 463 LLFMHAKSKLPTSDDIDKLISAEIPDKEKEPELY 496
>At2g07620 putative helicase
Length = 1241
Score = 327 bits (838), Expect = 2e-89
Identities = 203/546 (37%), Positives = 294/546 (53%), Gaps = 56/546 (10%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P++ RR++ V K GI+ DN VVPYN L ++Y AHIN+E+C +S SIKYLFKYI K
Sbjct: 499 YPIYRRRESEHFVEKGGIKCDNTYVVPYNRMLSLRYRAHINVEWCKQSGSIKYLFKYINK 558
Query: 574 GVDRVTATLETSEEPS---------------VD---EIQ*YYDCRYLSPSESIWRIFGFD 615
G DRV +E ++ S +D EI+ Y+DCRY+S SE++WRIF F
Sbjct: 559 GQDRVAIVVEPKDKTSNMVLFSGSQKLLVAVIDDDKEIKDYFDCRYVSASEAVWRIFKFP 618
Query: 616 VHCRYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLT 675
+ R V +L++HL G+Q + F D+ D + + ++ MF+ ++ N+ +
Sbjct: 619 IQYRTTPVMKLSYHLPGKQPLCFEDTQNIDELSEKKANEDFMFIGFLKLNQECEFARQFI 678
Query: 676 YVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRT 735
Y + P FT++ +N+ W L + G +GR+ + + Y M++LL G S EDIRT
Sbjct: 679 YTEIPPYFTWDGQNKQWKLRERGFYIGRMNYASIKMEPEYYMKILLGIVCGPKSDEDIRT 738
Query: 736 VD----------GHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLL- 784
G ++ Y C + +L + + ++++V S + K+ +LL
Sbjct: 739 YKDVVRRKFFFLGIIFRIYFLCCFWINVLLDQ----SMCGKMLLVLSDAE--KINYALLE 792
Query: 785 ---LTSCMSDPLNVWNQTWHILSEGILYERRRTLNSPG*--------------LLNSDQL 827
+ C L + EG + R + S+Q
Sbjct: 793 IEDMLLCNGSTLEDFKHMPKPTKEGTDHSNRFITEEKNYNVEKLKEDHDDWFNKMTSEQK 852
Query: 828 SAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSIASLLLP 887
Y++I+ V +FFV G+GG GKTF+W LS R +G I +NVASS IA LLL
Sbjct: 853 EIYDEIMKAVLENSGGIFFVYGFGGTGKTFMWKTLSAAVRMKGLISVNVASSGIAFLLLQ 912
Query: 888 RGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMR 947
GRTAHS+F IP+ + + C I + A +L ASLIIWDEAPM+SR FE+ DR++
Sbjct: 913 GGRTAHSRFGIPINPDDFTTCHIVPNSDLANMLKEASLIIWDEAPMMSRYCFESLDRSLN 972
Query: 948 DIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRVLKL 1007
D++ N VDG PFGGK VV GGDFRQ+L V+ GRA+IV A +NSS LW C VL L
Sbjct: 973 DVIGN-VDGK---PFGGKVVVFGGDFRQVLHVIHGAGRAEIVLAALNSSYLWEHCNVLTL 1028
Query: 1008 TKNIRL 1013
TKN+ L
Sbjct: 1029 TKNMSL 1034
Score = 90.5 bits (223), Expect = 5e-18
Identities = 57/192 (29%), Positives = 97/192 (49%), Gaps = 35/192 (18%)
Query: 122 NGLDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNR-NFDSRVYNVPE 180
N + + L++ L++++D CN ++ FR+ + ++N P +++ +R D R Y P
Sbjct: 136 NQVRKDLIEKLIRMLDACNPYIENFRLAKYKLDSNNGEPFYMQIVSDRVGKDGRTYCNPR 195
Query: 181 ISEVVALIVGDFDSFEDGRDIVVREKDGG-LRRIHETHSKYIPLQYPVLFPLGEDRYEEK 239
SEV ALI GDF RDI+V++K G L RI + H Y+P+QYP++F GED +
Sbjct: 196 TSEVAALIPGDFRPKMHTRDIIVQDKKTGQLSRISKVHPSYVPMQYPLIFNYGEDDFRPG 255
Query: 240 IKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQR 299
I+ + TG K+ N+ ++ Y+ +E+ R
Sbjct: 256 IQKGYTGRTG-------------------KQANK--------------CINGYTTIESNR 282
Query: 300 LSYVRKNQSQIR 311
L+Y++ NQS +R
Sbjct: 283 LAYIKFNQSNLR 294
>At3g43350 putative protein
Length = 830
Score = 305 bits (782), Expect = 7e-83
Identities = 174/434 (40%), Positives = 255/434 (58%), Gaps = 41/434 (9%)
Query: 731 EDIRTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMS 790
+D++TV G V+ S+R+A ALGLL +D+++I+ I + S +R++F +LL+ ++
Sbjct: 53 KDLKTVKGVVHKSFRDAVFALGLLDDDKEYINAIKDANFWCSAKYVRRLFVIMLLSESLT 112
Query: 791 DPLNVWNQTWHILSEGI--------------LYERRRTLNSPG*LLNS------------ 824
P VW++TW ILS+ I L E R L G L+
Sbjct: 113 KPEMVWDETWRILSKDIEHLQLSDEERQQYCLQEIARLLTKNGVSLSKWNRCHKFQMNTM 172
Query: 825 ------DQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVAS 878
Y++I++VV ++ +FFV G+GG GKTFLW LS RS+G I LNVAS
Sbjct: 173 VDYGDFRAKKIYDEIMDVVLHDRGGVFFVYGFGGTGKTFLWKLLSAAVRSKGDISLNVAS 232
Query: 879 SSIASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLA 938
S IA+L L GRTAHS+F IP+ E S C I + + +L+ A LIIWDEAPM+S+
Sbjct: 233 SGIAALRLDGGRTAHSRFDIPINPNESSTCNISRGSDLGELVKEAKLIIWDEAPMMSKHC 292
Query: 939 FEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSML 998
FE+ DRT++DI++N D P GGK +V GGDFRQ+LPV+ GR +IV A +NSS +
Sbjct: 293 FESLDRTLKDIVNNPGD----KPLGGKVIVFGGDFRQVLPVINGAGREEIVFAALNSSYI 348
Query: 999 WRRCRVLKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICL 1058
W +VL+LTKN+RL + + + F++WILDVGDGK+ NDG A+I+IP++ +
Sbjct: 349 WEHSKVLELTKNMRLLADISEHEKRDIEDFSKWILDVGDGKISQPNDGIALIDIPEEFLI 408
Query: 1059 KNSGNHVGDIVESTYPNLLSNMKESS-----FFQDRAILAPTLELVEKVNDYVLSLVPGE 1113
+ V I+E+ Y N K+ +Q RAIL PT E V +N++++ ++ GE
Sbjct: 409 NGDNDPVESIIEAVYGNTFMEEKDPKKTDYPQYQGRAILCPTNEDVNSINEHMMRMLDGE 468
Query: 1114 EKEYLSCDTVLKCD 1127
E+ YLS D++ D
Sbjct: 469 ERIYLSSDSIDPAD 482
>At1g52960 hypothetical protein
Length = 924
Score = 286 bits (733), Expect = 4e-77
Identities = 145/307 (47%), Positives = 204/307 (66%), Gaps = 4/307 (1%)
Query: 821 LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSS 880
++ S+Q Y++I++ V ++ +FFV G+GG GKTFLW LS RS+G I LNVASS
Sbjct: 471 MVTSEQKKIYDEIMDAVLHDRGGVFFVYGFGGTGKTFLWKLLSAAIRSKGDISLNVASSG 530
Query: 881 IASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFE 940
IA+LLL GRT HS+F IP+ E S C I + + +L+ A+LIIWDE PM+S+ FE
Sbjct: 531 IAALLLDGGRTTHSRFGIPINPNESSTCNISRGSDLGELVKEANLIIWDETPMMSKHCFE 590
Query: 941 AFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWR 1000
+ DRT+RDIM+N D PFGGK +V GGDFRQ+LPV+ GR +IV A +NSS +W
Sbjct: 591 SLDRTLRDIMNNPGDK----PFGGKGIVFGGDFRQVLPVINGAGREEIVFAALNSSYIWE 646
Query: 1001 RCRVLKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKN 1060
C+VL+LTKN+RL + + + F++WILDVGDGK+ NDG A+I+IP++ +
Sbjct: 647 HCKVLELTKNMRLLANISEHEKRDIEYFSKWILDVGDGKISQPNDGIALIDIPEEFLING 706
Query: 1061 SGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSC 1120
+ V I+E+ Y N K+ FFQ RAIL PT E V +N++++S++ GEE+ YLS
Sbjct: 707 DNDPVESIIEAVYGNTFMEEKDPKFFQGRAILCPTNEDVNSINEHMMSMLDGEERIYLSS 766
Query: 1121 DTVLKCD 1127
D++ D
Sbjct: 767 DSIDPAD 773
Score = 257 bits (656), Expect = 3e-68
Identities = 129/337 (38%), Positives = 204/337 (60%), Gaps = 20/337 (5%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
FP++ RRDTG+ V KNG Q DNR V+PYN K+ ++Y AHIN+E CN+S SIKYLFKY+ K
Sbjct: 77 FPVYRRRDTGIYVEKNGFQCDNRYVIPYNEKVSLRYQAHINVELCNQSGSIKYLFKYVHK 136
Query: 574 GVDRVTATLETSEEPSV----DEIQ*YYDCRYLSPSESIWRIFGFDVHCRYPAVKRLTFH 629
G DRVT T+E +++ + DE++ Y+DCRY+S E++WRIF F +H R V +L FH
Sbjct: 137 GHDRVTVTVEPNDQDTAKKEKDEVKDYFDCRYVSACEAMWRIFKFPIHYRTTPVVKLFFH 196
Query: 630 LHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRN----------------YSQSWN 673
G+Q + ++ ++V+ R + T FLAW N+
Sbjct: 197 EEGKQPVYYKPGETTESVMDRLSSEATQFLAWFQLNKKPPSRTIRANAKKLPKAAPDPTK 256
Query: 674 LTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDI 733
L + + P FT+N + + + + + G ++GR+ FVP ++ Y +R+LLN + G TS++D+
Sbjct: 257 LLFEEIPNHFTWNSKEKKFMIRERGFAIGRINFVPRTIEDAYYLRILLNIKRGVTSYKDL 316
Query: 734 RTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPL 793
+TV G V+ S+R+A ALGLL +D+++I+ I + S +R++F +LL+ ++ P
Sbjct: 317 KTVKGVVHKSFRDAVFALGLLDDDKEYINGIKDAKFWCSAKYVRRLFVIMLLSESLTKPE 376
Query: 794 NVWNQTWHILSEGILYERRRTLNSPG*LLNSDQLSAY 830
VW++TW ILSE I +R+ P L+ ++ Y
Sbjct: 377 MVWDETWRILSEDIERRKRKEWKRPDLQLSDEERQQY 413
>At5g37110 putative helicase
Length = 1307
Score = 286 bits (731), Expect = 6e-77
Identities = 202/652 (30%), Positives = 308/652 (46%), Gaps = 123/652 (18%)
Query: 519 RRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITKGVDRV 578
+R T V K + DNR V+PYN L ++Y AHIN+E+CN+S S+KY+FKYI KG DRV
Sbjct: 571 KRRTDDFVEKKDFKCDNRYVIPYNRSLSLRYRAHINVEWCNQSGSVKYIFKYIHKGPDRV 630
Query: 579 TATLETSEEPSVDEIQ*YYDCRYLSPSESIWRIFGFDV-HCRYPAVKRLTFHLHGEQRIM 637
T +E+S E D SE+ + D +CR
Sbjct: 631 TVVVESSLNSKNKENGKQKDNADTDGSETKKKNEVEDYFNCR------------------ 672
Query: 638 FRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYVQFPRKFTYNVENRSWHL*KS 697
R + VL R+ +MFLAW + N+ + LTY P +FTY+ + + ++L K
Sbjct: 673 -----RIETVLNRSDLDGSMFLAWFELNKVSKIARKLTYADIPTRFTYDSKEKKFNLRKK 727
Query: 698 GQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREACAALGLLAND 757
G ++GR+ +VP ++ Y +R+LLN Q G FE++RTV+ +Y +++AC ALGLL ND
Sbjct: 728 GFAIGRINYVPRDIEDGYYLRILLNVQPGPRCFEELRTVNDVLYKEWKDACEALGLLDND 787
Query: 758 RQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGILYERRRTLNS 817
+++ID + SG +R++F +++ + P NVW TW LSE I +R+ N
Sbjct: 788 QEYIDDLKRTSFWSSGGYLRQLF--VIMLDALISPENVWAATWQHLSEDIQNNKRKYFNR 845
Query: 818 PG*LLNSDQLSAYEKIVNVVDNELDHMFFVDG-----YGGIGKT-----FLWNALSYHFR 867
P +L+ ++ Y E+DH+ +G Y + + F N L +
Sbjct: 846 PDLILSDEEKKLYAL------QEIDHILRRNGTSLTYYKTMPQVPRDPRFDTNVLILDEK 899
Query: 868 SEGKIVLNVASSSIASLLLPR------------------------------GRTAHSQFS 897
+ L + +L P GRTAHS+F
Sbjct: 900 GYDRDNLTEKHAKWIKMLTPEQKSIYDDIIGAVNENVGVVVFVYGFGGTEGGRTAHSRFG 959
Query: 898 IPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMRDIMSNVVDGA 957
IPL E + C ++ ++A L+ ASLIIWDEAPM+SR FE+ DR++ DI N
Sbjct: 960 IPLNPNEFTTCNMKVGSDRANLVKEASLIIWDEAPMMSRYCFESLDRSLSDICGN----G 1015
Query: 958 SNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRVLKLTKNIRLQFPT 1017
N PFGGK VV GG
Sbjct: 1016 DNKPFGGKVVVFGG---------------------------------------------L 1030
Query: 1018 NGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKNSGNHVGDIVESTYPNL- 1076
+ + ++ F+ WIL VGDG++ NDGEA+I IP + + + + + I Y ++
Sbjct: 1031 SVSEAKDIKEFSEWILAVGDGRIVEPNDGEALIVIPSEFLITKAKDPIEAICTEIYGDIT 1090
Query: 1077 -LSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSCDTVLKCD 1127
+ + FFQ++AIL PT E V ++N+ +L + GEE +LS D++ D
Sbjct: 1091 KIHEQNDPIFFQEKAILCPTNEDVNQINETMLDNLQGEEFTFLSSDSLDPAD 1142
Score = 107 bits (268), Expect = 3e-23
Identities = 67/189 (35%), Positives = 95/189 (49%), Gaps = 36/189 (19%)
Query: 124 LDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNFDSRVYNVPEISE 183
L + ++ ++KL++ N V R RD AN + + +R+ R D RVYNVP SE
Sbjct: 253 LRKEIIDAVIKLLNSINPYVSHLRTARDRFNANPEETLHMRIVSKREKDGRVYNVPTTSE 312
Query: 184 VVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPLGEDRYEEKIKLN 243
V LI GDF RDIVV+EK G L+RI E Y+PLQYP+LFP GED + I+
Sbjct: 313 VAMLIPGDFTIDMPSRDIVVQEKSGKLQRISEILPCYLPLQYPLLFPYGEDGFRIGIE-K 371
Query: 244 HRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLSYV 303
H+T L+QQF+VD Y+ +E+ L Y+
Sbjct: 372 HQT-----------------------------------GLWQQFLVDSYTAIESNWLGYI 396
Query: 304 RKNQSQIRS 312
+ NQ+ +R+
Sbjct: 397 KLNQTSLRA 405
>At3g42100 putative protein
Length = 1752
Score = 280 bits (715), Expect = 4e-75
Identities = 149/312 (47%), Positives = 208/312 (65%), Gaps = 7/312 (2%)
Query: 821 LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSS 880
+LN++Q Y++I V N+L +FF+ G+GG GKTF+W L+ RS G+IVLNVASS
Sbjct: 1271 MLNTEQRGIYDEITGAVFNDLGGVFFIYGFGGTGKTFIWKTLAAAVRSRGQIVLNVASSG 1330
Query: 881 IASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFE 940
IASLLL GRTAHS+F+IPL E S C I + + A L+ ASLIIWDEAPM+S+ FE
Sbjct: 1331 IASLLLEGGRTAHSRFAIPLNPDEFSVCKITPKSDLANLIKEASLIIWDEAPMMSKFCFE 1390
Query: 941 AFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWR 1000
+ D++ DI++N N FGGK VV GGDFRQ+LPV+ GR +IV + +N+S LW
Sbjct: 1391 SLDKSFYDILNN----KDNKVFGGKVVVFGGDFRQVLPVINGAGRVEIVMSSLNASYLWD 1446
Query: 1001 RCRVLKLTKNIRLQFPTNGQDDA-SLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLK 1059
C+VLKLTKN+RL ++A ++ F+ W+L VGDG++ NDGEA+I+IP+++ +K
Sbjct: 1447 HCKVLKLTKNMRLLSGGLSSEEAKEIQQFSDWLLAVGDGRINEPNDGEALIDIPEELLIK 1506
Query: 1060 NSGNHVGDIVESTY--PNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEY 1117
+GN + I + Y P+ L + + FFQ RAILAPT E V +N Y+L + EE+ Y
Sbjct: 1507 EAGNPIEAISKEIYGDPSELHMINDPKFFQRRAILAPTNEDVNTINQYMLEHLKSEERIY 1566
Query: 1118 LSCDTVLKCDEE 1129
LS D++ D +
Sbjct: 1567 LSADSIDPTDSD 1578
Score = 236 bits (602), Expect = 6e-62
Identities = 132/340 (38%), Positives = 200/340 (58%), Gaps = 23/340 (6%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P++ RR T + K G + DN VVPYN KL ++Y AHIN+E+CN+S SIKYLFKYI K
Sbjct: 873 YPIYRRRMTEDYIEKGGFKCDNGYVVPYNKKLSLRYQAHINVEWCNQSGSIKYLFKYINK 932
Query: 574 GVDRVTATLE-----------TSEEP------SVDEIQ*YYDCRYLSPSESIWRIFGFDV 616
G DRV +E TS EP DEI+ ++DCRY+S SE++WRI+ F +
Sbjct: 933 GADRVVFIVEPVNQDKTTENATSGEPPNSTEKKKDEIKDWFDCRYVSASEAVWRIYKFPL 992
Query: 617 HCRYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWN--- 673
R AV+RL+FH G+Q + + + ++VL R ++ +MF+AW+ N+N N
Sbjct: 993 QDRSTAVQRLSFHDEGKQPVYAKPDADIEDVLERISNEDSMFMAWLTLNKNNDVGKNGKR 1052
Query: 674 ---LTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSF 730
L Y Q P FT++ +N+ W G S+GR+ +V + Y +R+LLN G S+
Sbjct: 1053 ARELLYSQIPAYFTWDGKNKQWVKRIRGFSLGRINYVCRKMEVEYYLRVLLNIVKGPMSY 1112
Query: 731 EDIRTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMS 790
+DI+T +G VY S++EAC A G+L +D+ +ID + E G +R F LLL+ ++
Sbjct: 1113 DDIKTFNGVVYPSFKEACFARGILDDDQVYIDGLHEASQFCFGDYLRNFFAMLLLSDSLA 1172
Query: 791 DPLNVWNQTWHILSEGILYERRRTLNSPG*LLNSDQLSAY 830
P +VW++TWH+L+E I ++R +P L ++ Y
Sbjct: 1173 RPEHVWSETWHLLAEDIENKKREDFKNPDLKLTLAEIRNY 1212
Score = 126 bits (317), Expect = 6e-29
Identities = 72/192 (37%), Positives = 116/192 (59%), Gaps = 7/192 (3%)
Query: 124 LDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRN-FDSRVYNVPEIS 182
L + +++ L++++++ N V KFR R+ + + D P +R+ +R D R Y++P S
Sbjct: 474 LKKEVIEALIEMLNKVNPYVDKFRQARERIQDDNDEPFHMRIVADRKGVDRRTYSMPTSS 533
Query: 183 EVVALIVGDFDSFEDGRDIVVREKDGG-LRRIHETHSKYIPLQYPVLFPLGEDRYEEKIK 241
EV ALI G F RDIV+ EK G L RI + H Y+ LQYP++ GED Y I+
Sbjct: 534 EVAALIPGGFQPSMFDRDIVLEEKTTGHLTRISQIHISYLALQYPLILCYGEDGYTPGIE 593
Query: 242 LNHRTTTGSVKKRVR--VSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQR 299
+ S KK+ + +S+R++ A+R+Q+R NE + + RLFQQF+ D Y+ +E+ R
Sbjct: 594 ---KCLPNSAKKKKKKCISMRQWFAFRIQERPNECKTLTRSKRLFQQFLCDAYTTIESNR 650
Query: 300 LSYVRKNQSQIR 311
LSY++ QS++R
Sbjct: 651 LSYIKFKQSKLR 662
Score = 32.7 bits (73), Expect = 1.2
Identities = 14/34 (41%), Positives = 22/34 (64%)
Query: 314 LLWLDSRDKFDSPKAINSIICAELPDSQMYPELF 347
LL++D++ K + I+ II AE+PD PEL+
Sbjct: 794 LLFMDAKSKLPTADDIDKIISAEIPDKDKEPELY 827
>At3g13250 hypothetical protein
Length = 1419
Score = 271 bits (694), Expect = 1e-72
Identities = 147/311 (47%), Positives = 202/311 (64%), Gaps = 7/311 (2%)
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
L +Q Y++I N V N+L +FFV G+GG GKTF+W L+ RS+G+I LNVASS I
Sbjct: 940 LTPEQRGIYDQITNAVFNDLGGVFFVYGFGGTGKTFIWKTLAAAVRSKGQICLNVASSGI 999
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
ASLLL GRTAHS+FSIPL E S C I+ + + A L+ ASLIIWDEAPM+S+ FEA
Sbjct: 1000 ASLLLEGGRTAHSRFSIPLNPDEFSVCKIKPKSDLADLIKEASLIIWDEAPMMSKFCFEA 1059
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
D++ DI+ V N FGGK +V GGDFRQ+LPV+ GRA+IV + +N+S LW
Sbjct: 1060 LDKSFSDIIKRV----DNKVFGGKVMVFGGDFRQVLPVINGAGRAEIVMSSLNASYLWDH 1115
Query: 1002 CRVLKLTKNIRLQFPTNGQDDA-SLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKN 1060
C+VL+LTKN+RL D+A ++ F+ W+L VGDG++ NDGE +I+IP+++ ++
Sbjct: 1116 CKVLRLTKNMRLLNNDLSVDEAKEIQEFSDWLLAVGDGRVNEPNDGEVIIDIPEELLIQE 1175
Query: 1061 SGNHVGDIVESTY--PNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYL 1118
+ N + I Y P L + + FFQ RAILAP E V +N Y+L + EE+ YL
Sbjct: 1176 ADNPIEAISREIYGDPTKLHEISDPKFFQRRAILAPKNEDVNTINQYMLEHLDSEERIYL 1235
Query: 1119 SCDTVLKCDEE 1129
S D++ D +
Sbjct: 1236 SADSIDPSDSD 1246
Score = 234 bits (596), Expect = 3e-61
Identities = 129/340 (37%), Positives = 201/340 (58%), Gaps = 23/340 (6%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P++ RR T + K G++ DNR VVPYN KL ++Y AHIN+E+CN++ SIKYLFKYI K
Sbjct: 541 YPIYRRRLTDDYIEKGGVKCDNRYVVPYNKKLSLRYQAHINVEWCNQNGSIKYLFKYINK 600
Query: 574 GVDRVTATLETSEEPSV-----------------DEIQ*YYDCRYLSPSESIWRIFGFDV 616
G DRV +E +E + DEI+ ++DCRY+S SE+IWRIF F +
Sbjct: 601 GPDRVVFIVEPIKEATSSDTTAPVVESDTTEKKKDEIKDWFDCRYVSASEAIWRIFKFPI 660
Query: 617 HCRYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANR------NYSQ 670
R V++L+FH G+Q F ++ +VL R ++ + FLAW+ NR N +
Sbjct: 661 QHRSTPVQKLSFHDKGKQPAYFDAKAKMADVLERVSNEDSQFLAWLTLNRKNAVGKNGKR 720
Query: 671 SWNLTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSF 730
+ + Y + P FT++ EN+ + G S+GR+ +V ++ Y +R+LLN G S+
Sbjct: 721 ARDCLYAEIPAYFTWDGENKQFKKRTRGFSLGRINYVSRKMEDEYYLRVLLNIVRGPQSY 780
Query: 731 EDIRTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMS 790
+DI+TV+G VY SY+ AC A G+L +D+ +I+ + E G +R F+ +LL+ ++
Sbjct: 781 DDIKTVNGVVYPSYKLACFARGILDDDQVYINGLIEASQFCFGDYLRNFFSMMLLSDSLA 840
Query: 791 DPLNVWNQTWHILSEGILYERRRTLNSPG*LLNSDQLSAY 830
P +VW++TWH+LSE IL ++R + L Q+ Y
Sbjct: 841 RPEHVWSETWHLLSEDILIKKRDEFKNQELTLTEAQIQNY 880
Score = 144 bits (364), Expect = 2e-34
Identities = 76/201 (37%), Positives = 127/201 (62%), Gaps = 4/201 (1%)
Query: 113 NYHSELSDGNGLDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRN-F 171
N S L++ L+ ++K++++ N V+KFR R+ ++ D P +R+ +R
Sbjct: 132 NNGSSTKGKKNLNKQLIDAIIKMLNQVNPYVEKFRSARERIDSTNDEPFHMRIVSDRKGT 191
Query: 172 DSRVYNVPEISEVVALIVGDFDSFEDGRDIVVREKDGG-LRRIHETHSKYIPLQYPVLFP 230
D R+YN+P EV ALI GDF S RDI++ +K G L+RI + H Y+ LQYP++F
Sbjct: 192 DGRLYNMPTAGEVAALIPGDFVSQMPVRDIILEKKSTGRLKRISQIHISYLALQYPLIFC 251
Query: 231 LGEDRYEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVD 290
GED Y I+ +++ G KK+ +S+R++ A+R+Q+R++E+ + Q RLFQQF+ D
Sbjct: 252 YGEDGYTPGIEKCYKS--GYTKKKKCISMRQWYAFRIQEREDESHTLLQSKRLFQQFLCD 309
Query: 291 LYSMVENQRLSYVRKNQSQIR 311
Y+ +E+ RL+Y++ NQS++R
Sbjct: 310 AYTTIESNRLAYIKFNQSKLR 330
>At2g05080 putative helicase
Length = 1219
Score = 260 bits (665), Expect = 3e-69
Identities = 145/308 (47%), Positives = 191/308 (61%), Gaps = 19/308 (6%)
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
L +Q S Y+ I+ V+ + +FFV G+GG GKTFLW LS RS+G IVLNVASS I
Sbjct: 747 LTLEQKSVYDNIIGAVNENVGGVFFVYGFGGTGKTFLWKTLSAALRSKGDIVLNVASSGI 806
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
ASLLL GRTAHS+ IPL E + C ++ ++A L+ ASLIIWDEAPM+SR FE+
Sbjct: 807 ASLLLEGGRTAHSRSGIPLNPNEFTTCNMKAGSDRANLVKEASLIIWDEAPMMSRHCFES 866
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
DR++ DI N N PFGGK VV GGDFRQ+LPV+P ADIV A +NSS LW
Sbjct: 867 LDRSLSDICGN----CDNKPFGGKVVVFGGDFRQVLPVIPGADTADIVMAALNSSYLWSH 922
Query: 1002 CRVLKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKNS 1061
C+VL LTKN+ L WIL VGDG++G NDGEA+I+IP + + +
Sbjct: 923 CKVLTLTKNMCL-------------FSEEWILAVGDGRIGEPNDGEALIDIPSEFLITKA 969
Query: 1062 GNHVGDIVESTYPNL--LSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLS 1119
+ + I Y ++ + K+ FFQ+RAIL PT E V ++N+ +L + GEE +LS
Sbjct: 970 KDPIQAICTEIYGDITKIHEQKDPVFFQERAILCPTNEDVNQINETMLDNLQGEELTFLS 1029
Query: 1120 CDTVLKCD 1127
D++ D
Sbjct: 1030 SDSLDTAD 1037
Score = 225 bits (573), Expect = 1e-58
Identities = 125/356 (35%), Positives = 199/356 (55%), Gaps = 33/356 (9%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
FP++ RR V K + DNR V+PYN L ++Y AHIN+E+CN+S S+KY+FKYI K
Sbjct: 359 FPIYRRRRIDDFVQKKDFKCDNRYVIPYNRSLSLRYRAHINVEWCNQSGSVKYIFKYIHK 418
Query: 574 GVDRVT--------------------ATLETSEEPSVDEIQ*YYDCRYLSPSESIWRIFG 613
G DRVT A + SE +E++ +++CRY+S E+ WRI
Sbjct: 419 GPDRVTVVVGSSLNSKNKEKGKQKVNADTDGSEPKKKNEVEDFFNCRYVSACEAAWRILK 478
Query: 614 FDVHCRYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWN 673
+ +H R +V +L+FHL GEQ I F+ + VL + D + + +
Sbjct: 479 YPIHYRSTSVMKLSFHLPGEQYIYFKGDEEVETVLNK-----------ADLDGSIQIARK 527
Query: 674 LTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDI 733
LTY P +FTY+ + + ++L K G ++GR+ +VP ++ Y +R+LLN G SFE++
Sbjct: 528 LTYPNIPTRFTYDPKEKKFNLRKKGFAIGRINYVPRDIEDGYYLRILLNVVPGPRSFEEL 587
Query: 734 RTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPL 793
+TV+G +Y +++AC ALGLL ND+++ID + SG +R++F +++ + P
Sbjct: 588 KTVNGVLYKEWKDACEALGLLDNDQEYIDDLKRTSFWSSGWYLRQLF--VIMLDALISPE 645
Query: 794 NVWNQTWHILSEGILYERRRTLNSPG*LLNSDQLSAYEKIVNVVDNELDHMFFVDG 849
NVW TW LSE I E+++ N P L +D + + E+ E+DH+ +G
Sbjct: 646 NVWAATWQHLSEDIQNEKKKYFNRPVTCLFTDLILSDEEKKVYALQEIDHILRRNG 701
Score = 134 bits (336), Expect = 4e-31
Identities = 68/150 (45%), Positives = 96/150 (63%), Gaps = 1/150 (0%)
Query: 163 LRLFRNRNFDSRVYNVPEISEVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIP 222
+R+ R D RVYNVP SEV LI GDF RDI+V EK G L+RI E Y+P
Sbjct: 1 MRIVSKRETDGRVYNVPTTSEVAMLIPGDFTIDIPCRDIIVEEKSGKLQRISEILPCYLP 60
Query: 223 LQYPVLFPLGEDRYEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSVIFQGMR 282
LQYP+LFP GED + I+ H+T G KK +S+R++ A+R+ +R +E ++ + R
Sbjct: 61 LQYPLLFPYGEDGFRTGIE-KHQTGAGKDKKNKFISIRQWFAFRIHERKHEKHILLRSKR 119
Query: 283 LFQQFIVDLYSMVENQRLSYVRKNQSQIRS 312
L+QQF+VD Y +E+ RL Y++ NQS +R+
Sbjct: 120 LWQQFLVDSYIAIESNRLGYIKLNQSSLRA 149
Score = 34.7 bits (78), Expect = 0.32
Identities = 16/34 (47%), Positives = 21/34 (61%)
Query: 314 LLWLDSRDKFDSPKAINSIICAELPDSQMYPELF 347
L+WLDS+ K + I+ I AE+PD PELF
Sbjct: 280 LIWLDSKCKLTRAEHIDKAISAEIPDKLKDPELF 313
>At3g31980 hypothetical protein
Length = 1099
Score = 258 bits (659), Expect = 1e-68
Identities = 140/305 (45%), Positives = 195/305 (63%), Gaps = 7/305 (2%)
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
+ S+Q Y++I+ V +FFV G+GG KTF+W LS R G I +NVASS I
Sbjct: 618 MTSEQKGIYDEIIKAVLENSGGIFFVYGFGGTSKTFMWKTLSAAVRMRGLISVNVASSGI 677
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
ASLLL GRTAHS+F IP+ + + C I + A +L ASLIIWDEAPM+SR FE+
Sbjct: 678 ASLLLQGGRTAHSRFGIPINPDDFTTCHIVPNSDLANMLKEASLIIWDEAPMMSRYCFES 737
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
DR++ D++ N +DG PFGGK VV GGDFRQ+LPV+ GRA+IV A +NSS LW
Sbjct: 738 LDRSLNDVIGN-IDGK---PFGGKVVVFGGDFRQVLPVIHGAGRAEIVLAALNSSYLWEH 793
Query: 1002 CRVLKLTKNIRLQFPTNGQDDA-SLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKN 1060
C+VL LTKN+RL +D+A ++ F+ W+L VGDG++ NDGE +I+IP+++ +K+
Sbjct: 794 CKVLTLTKNMRLMSNDLDKDEAEEIKEFSNWLLAVGDGRVSEPNDGEVLIDIPEELLIKD 853
Query: 1061 SGNHVGDIVESTYP--NLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYL 1118
+ + + I ++ Y +LL + FFQ RAIL P V +ND +L + GE YL
Sbjct: 854 ANDPIEAITKAVYGDLDLLQPNNDPKFFQQRAILCPRNTDVNTINDIMLDKLNGELVTYL 913
Query: 1119 SCDTV 1123
S D++
Sbjct: 914 SADSI 918
Score = 112 bits (281), Expect = 9e-25
Identities = 63/169 (37%), Positives = 99/169 (58%), Gaps = 3/169 (1%)
Query: 135 LMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNR-NFDSRVYNVPEISEVVALIVGDFD 193
++D N V+KFR+ +D ++N P +++ +R D + Y P SEV ALI GDF
Sbjct: 1 MLDASNPYVEKFRLAKDKLDSNNGEPFYMQIVSDRVGKDGKTYCNPTTSEVTALIPGDFR 60
Query: 194 SFEDGRDIVVREKDGG-LRRIHETHSKYIPLQYPVLFPLGEDRYEEKIKLNHRTTTGSVK 252
RDI+V++K G L RI E H Y+P+QYP++F GED + I+ + TG
Sbjct: 61 PEMPTRDIIVQDKKTGHLSRISEVHPSYVPMQYPLIFNYGEDGFRPGIQKGYTGRTGKQA 120
Query: 253 KRVRVSLREFVAYRLQKRDNENSVIFQGMRLFQQFIVDLYSMVENQRLS 301
+ +S+R++ A+R+Q+R +E + + RLFQQF+VD Y+ N +S
Sbjct: 121 NKC-ISMRQWYAFRIQERSDEAQTLLRSKRLFQQFLVDGYTTGGNTNMS 168
Score = 111 bits (278), Expect = 2e-24
Identities = 63/200 (31%), Positives = 107/200 (53%), Gaps = 6/200 (3%)
Query: 650 RNRHKRTMFLAWMDANRNYSQSWNLTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPP 709
R ++ +MF+A++ N+ + TY + P+ FT++ +N+ W L + G +GR+ +
Sbjct: 379 RKANEDSMFMAFLKLNQECEFARQFTYTEIPQYFTWDGQNKQWKLRERGFCIGRMNYASI 438
Query: 710 GTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREACAALGLLANDRQFIDLISEIVV 769
Y MR+LL G TS EDIRT VY +Y+EAC A G+L +D+ +ID I E +
Sbjct: 439 KMDPEYYMRILLGIVCGPTSDEDIRTYKDVVYETYKEACLARGILTDDQAYIDTIVEGSL 498
Query: 770 VGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGILYERRRTLNSPG*LLNSDQLSA 829
G +R +F+ +LL C++ P VW + IL E I ++R+ ++P +L +
Sbjct: 499 YFFGDHLRNLFSMMLLDKCLARPEYVWEKCSRILIEDIETKKRKQYDNPDLVLTDAERRN 558
Query: 830 YEKIVNVVDNELDHMFFVDG 849
Y + E++ M +G
Sbjct: 559 YALL------EIEDMLLCNG 572
Score = 30.8 bits (68), Expect = 4.6
Identities = 14/34 (41%), Positives = 22/34 (64%)
Query: 314 LLWLDSRDKFDSPKAINSIICAELPDSQMYPELF 347
LL++D KF + I++II AE+PD P+L+
Sbjct: 281 LLFMDKSCKFPTSDDIDNIISAEIPDKSKDPKLY 314
>At3g31440 hypothetical protein
Length = 536
Score = 248 bits (632), Expect = 2e-65
Identities = 159/418 (38%), Positives = 229/418 (54%), Gaps = 39/418 (9%)
Query: 715 YCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGS 774
Y +R+LLN TS+ +I+T DG VY +++EAC A G+L +D+ FID + E +
Sbjct: 5 YFLRVLLNIVRRPTSYAEIKTYDGVVYKTFKEACFARGILDDDQVFIDGLVEATTLEDID 64
Query: 775 SIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGILYERRRTLNSPG*LLNSDQLSAYEKIV 834
K D ++ N+ ++ E + Y R L +E+
Sbjct: 65 EFPKP---------TRDGIDNSNR---LIVEELRYNRESNLKEK-----------HEEWK 101
Query: 835 NVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSIASLLLPRGRTAHS 894
++ +E G KT +W L R +IVLN+ASS IASLLL GRTAHS
Sbjct: 102 QMLTSE---------QRGTWKTIIWKTLFAAIRRRDQIVLNMASSGIASLLLEGGRTAHS 152
Query: 895 QFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMRDIMSNVV 954
+F I L E S C I+ + + A L+ ASL+I D+APM+SR FEA D++ DI+ N
Sbjct: 153 RFGIRLNPDEFSVCKIKPKSDLANLVKEASLVICDKAPMMSRFCFEALDKSFSDIIKNTY 212
Query: 955 DGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRRCRVLKLTKNIRLQ 1014
N FGGK VV GDFRQ+LPV+ GRA+IV + +N+S LW C+VLKLTKN+RL
Sbjct: 213 ----NKVFGGKVVVFSGDFRQVLPVINGAGRAEIVMSSLNASYLWDHCKVLKLTKNMRLL 268
Query: 1015 FPTNGQDDA-SLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKNSGNHVGDIVESTY 1073
+ +A + F+ W+L VGDG++ ND A+I+IP D+ + N+ + I Y
Sbjct: 269 ANNLSETEAKEIHEFSDWLLAVGDGRINEPNDDVAIIDIPKDLLITNADKPIEWITNEIY 328
Query: 1074 --PNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSCDTVLKCDEE 1129
P +L + + FFQ RAILAP E V +N+Y+L + EE+ YLS D++ D +
Sbjct: 329 GDPKILHEITDPKFFQGRAILAPKNEDVNTINEYLLEQLHAEERIYLSADSIDPTDSD 386
>At3g30420 hypothetical protein
Length = 837
Score = 231 bits (588), Expect = 2e-60
Identities = 135/308 (43%), Positives = 185/308 (59%), Gaps = 14/308 (4%)
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
LN Q Y+ ++ V N+ +FF+ G GG GKTFL+ + RS GK V+ VASS+I
Sbjct: 349 LNEQQRIIYDDVLKSVTNKEGKLFFLYGDGGTGKTFLYKTIISALRSNGKNVMPVASSAI 408
Query: 882 ASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFEA 941
A+LLLP GRTAHS F IP+ + E+ C I+ A +L+ LIIWDEAPM R FEA
Sbjct: 409 AALLLPGGRTAHSWFKIPINVHEDFICDIKIGSMLANVLSKVDLIIWDEAPMAHRHTFEA 468
Query: 942 FDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWRR 1001
DRT+RDI+S + A GGKTV+LGGDFRQILPV+P+R R + V A IN S LW
Sbjct: 469 VDRTLRDILSVGDEKALTKTLGGKTVLLGGDFRQILPVIPQRTRQETVSAAINRSYLWES 528
Query: 1002 CRVLKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKL-----GIDNDGEA-VIEIPDD 1055
C L++N+R+Q P + FA WIL +GDG+ GID+D E I I +
Sbjct: 529 CHKYLLSQNMRVQ-PEEIK-------FAEWILQIGDGEAPRKTHGIDDDQEEDNIIIDKN 580
Query: 1056 ICLKNSGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEK 1115
+ L + N + + +S P+ + ++ + A+L P E V+++NDY+LS VPG K
Sbjct: 581 LLLPETENPLEVLCQSVSPDFTNTFQDLENLKGTAVLTPRNETVDEINDYLLSKVPGLAK 640
Query: 1116 EYLSCDTV 1123
EY S D++
Sbjct: 641 EYFSADSI 648
Score = 60.1 bits (144), Expect = 7e-09
Identities = 29/96 (30%), Positives = 54/96 (56%)
Query: 738 GHVYNSYREACAALGLLANDRQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWN 797
G V+ S++ AC A GLL D+++ D + E + +R +F +L+ +S+PL +W+
Sbjct: 197 GVVHESFKAACHARGLLDGDKEWHDAMDEAAQWSTSYLLRSLFVLILIYCEVSEPLKLWS 256
Query: 798 QTWHILSEGILYERRRTLNSPG*LLNSDQLSAYEKI 833
W +++ +L +++R LN P L + +L Y I
Sbjct: 257 HCWESMADDVLRKQQRVLNFPQLELKAKELEKYTLI 292
>At1g64410 unknown protein
Length = 753
Score = 223 bits (567), Expect = 6e-58
Identities = 116/231 (50%), Positives = 158/231 (68%), Gaps = 4/231 (1%)
Query: 821 LLNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSS 880
++ +Q Y++I++VV ++ +FFV G+GG GKTFLW LS RS+G I LNVASS
Sbjct: 521 MVTLEQKKIYDEIMDVVLHDRGGVFFVYGFGGTGKTFLWKLLSAAIRSKGDISLNVASSG 580
Query: 881 IASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKAQLLTMASLIIWDEAPMISRLAFE 940
IA+LLL GRTAHS+F IP+ E S C I + + +L+ A+LIIWDEA M+S+ FE
Sbjct: 581 IAALLLDGGRTAHSRFGIPINPNESSTCNISRGLDFGELVKEANLIIWDEAHMMSKHCFE 640
Query: 941 AFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDACINSSMLWR 1000
+ DRT+RDIM+N D PFGGK +V GGDFRQ+L V+ GR +IV A +NSS +W
Sbjct: 641 SLDRTLRDIMNNPGD----KPFGGKVIVFGGDFRQVLSVINGAGREEIVFAALNSSYIWE 696
Query: 1001 RCRVLKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIE 1051
C+VL+LTKN+RL + + + F++WILDVGDGK+ NDG A+I+
Sbjct: 697 HCKVLELTKNMRLLANISEHEKRDIEYFSKWILDVGDGKISQPNDGIALID 747
Score = 97.1 bits (240), Expect = 5e-20
Identities = 49/133 (36%), Positives = 83/133 (61%)
Query: 698 GQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTVDGHVYNSYREACAALGLLAND 757
G ++GR+ FVP ++ Y +R+LLN + G TS +D++TV VY S+R+A ALGLL +D
Sbjct: 331 GFAIGRINFVPRTIEDAYYLRILLNIKRGVTSSKDLKTVKAVVYKSFRDAVFALGLLDDD 390
Query: 758 RQFIDLISEIVVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGILYERRRTLNS 817
+++I+ I + S +R++F +LL+ ++ P VW++TW ILSE I ++R+
Sbjct: 391 KEYINEIKDANFWCSAKYVRRLFVIMLLSESLTKPEMVWDETWKILSEDIERKKRKEWKR 450
Query: 818 PG*LLNSDQLSAY 830
P L+ ++ Y
Sbjct: 451 PDLQLSDEERQQY 463
>At4g04300 hypothetical protein
Length = 286
Score = 199 bits (507), Expect = 6e-51
Identities = 119/286 (41%), Positives = 170/286 (58%), Gaps = 23/286 (8%)
Query: 821 LLNSDQLSAYEKIVNVVDNELDHMFF-VDGYGGIGKTFLWNALSYHFRSEGKIVLNVASS 879
+L +Q Y++I N V N++ +FF V G GGIGKTF+W L+ RS+G+ LN+ASS
Sbjct: 1 MLTPEQRGIYDQITNAVFNDMGGVFFFVYGSGGIGKTFIWKTLAAVGRSKGQTCLNIASS 60
Query: 880 SIASLLLPRGRTAHSQFSIPLMLLEESGCTIEKEGNKA---------------QLLTMAS 924
IASLLL GR AH +FSIPL E S C I+ + + A L+ AS
Sbjct: 61 GIASLLLEGGRIAHYRFSIPLNPDEFSVCKIKPKSDLADLIKEASLIIWDKLVDLIKKAS 120
Query: 925 LIIWDEAPMISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRG 984
LIIWD+APM S+ FEA D++ DI+ V N F GK +V GGDFRQ+LPV+ G
Sbjct: 121 LIIWDKAPMKSKFCFEALDKSFSDIIKRV----DNKVFCGKVMVFGGDFRQVLPVINGAG 176
Query: 985 RADIVDACINSSMLWRRCRVLKLTKNIRLQFPTNGQDDA-SLRLFARWILDVGDGKLGID 1043
RA+ V + +N+ +W C+VLKLTKN+RL D+A ++ F W+L VGDG++
Sbjct: 177 RAETVMSSLNAVYIWDHCKVLKLTKNMRLLNNDLSVDEAKEIQEFFDWLLVVGDGRVNEP 236
Query: 1044 NDGEAVIEIPDDICLKNSGNHVGDIVESTYPNL--LSNMKESSFFQ 1087
NDGEA+I+IP+++ ++ + + I Y + L + + FF+
Sbjct: 237 NDGEALIDIPEELLIQEADIPIEAISREIYGDATKLHEINDPKFFR 282
>At3g42340 putative protein
Length = 244
Score = 163 bits (412), Expect = 6e-40
Identities = 80/195 (41%), Positives = 124/195 (63%), Gaps = 4/195 (2%)
Query: 933 MISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGGDFRQILPVVPKRGRADIVDAC 992
M+S+ FE+ DRT+RD M+N+ D PFGGK +V GGDFRQ+L V+ GR +IV A
Sbjct: 1 MMSKHCFESLDRTLRDFMNNLEDK----PFGGKVIVFGGDFRQVLLVINGAGREEIVFAA 56
Query: 993 INSSMLWRRCRVLKLTKNIRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEI 1052
+NSS +W C+V +LTK ++L + + + F++WILDVGDGK+ N+G ++I+I
Sbjct: 57 LNSSYIWEHCKVFELTKKMKLLANISEHEKRDIEDFSKWILDVGDGKISQCNEGISLIDI 116
Query: 1053 PDDICLKNSGNHVGDIVESTYPNLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPG 1112
++ + + V I+E+ Y N K+ FFQ R IL PT E V +N++++S++ G
Sbjct: 117 REEFFINGDNDPVESIIEAVYGNTFMEEKDRKFFQGRVILCPTNEDVNSINEHMMSMLDG 176
Query: 1113 EEKEYLSCDTVLKCD 1127
EE+ YLS +++ D
Sbjct: 177 EERIYLSSNSIDPAD 191
>At3g32320 hypothetical protein
Length = 494
Score = 156 bits (394), Expect = 7e-38
Identities = 91/256 (35%), Positives = 139/256 (53%), Gaps = 30/256 (11%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P++ RRDTG + KN Q DN VVPYN L+ KY AHIN+E+CN+S S+KYLFKY+ K
Sbjct: 240 YPIYRRRDTGRFIKKNKYQCDNWYVVPYNDVLLRKYKAHINVEWCNQSVSVKYLFKYVNK 299
Query: 574 GVDRVTATLE------TSEEPSVDE----------IQ*YYDCRYLSPSESIWRIFGFDVH 617
G DRVT ++E SE+ +V E +Q Y+DC RI G+ +H
Sbjct: 300 GPDRVTVSVEPHRKEVVSEQNNVGETNNDQQERNQVQDYFDC----------RIRGYPIH 349
Query: 618 CRYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDANRNYSQSWNLTYV 677
R ++ +LTFH G+Q + ++ ++VL R + T F AW + N+ ++ L Y
Sbjct: 350 YRQTSITKLTFHEKGKQPVYVKEGETAESVLYRVNNDETQFTAWFELNKRDPEAAKLLYE 409
Query: 678 QFPRKFTYNVENRSWHL*K-SGQSVGRLTFVPPGTKELYCMRLLLNRQIGCTSFEDIRTV 736
Q P +T+N +++ + K G VGR+ VPP + Y +R+L+N +DI T
Sbjct: 410 QIPNFYTWNGKDKDFRGRKMPGFVVGRINHVPPKIDDAYHLRILINNM---RLLQDINTN 466
Query: 737 DGHVYNSYREACAALG 752
+ + + A+G
Sbjct: 467 EAREIEEFFKWILAVG 482
Score = 64.7 bits (156), Expect = 3e-10
Identities = 35/84 (41%), Positives = 49/84 (57%), Gaps = 1/84 (1%)
Query: 126 RSLVQDLMKLMDECNLLVKKFRMVRD-FREANVDVPVKLRLFRNRNFDSRVYNVPEISEV 184
+ V L+K++D+ N V FR+ RD F + +R+ R D RVYN+P ++EV
Sbjct: 31 KETVSALLKMLDKINPHVANFRIARDRFNIEKEEENFHMRIISRRETDGRVYNLPSVAEV 90
Query: 185 VALIVGDFDSFEDGRDIVVREKDG 208
ALI GDFD D RDIV++ K G
Sbjct: 91 AALIPGDFDDNLDKRDIVLQMKSG 114
Score = 34.7 bits (78), Expect = 0.32
Identities = 17/40 (42%), Positives = 22/40 (54%)
Query: 1007 LTKNIRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDG 1046
L N+RL N + + F +WIL VG+GKL NDG
Sbjct: 453 LINNMRLLQDINTNEAREIEEFFKWILAVGEGKLNEPNDG 492
>At4g07800 hypothetical protein
Length = 448
Score = 135 bits (340), Expect = 1e-31
Identities = 79/232 (34%), Positives = 128/232 (55%), Gaps = 24/232 (10%)
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITK 573
+P++ RR + K G++ DN VVPYN + +KY AHIN+E+CN++ SIKYLFKYI K
Sbjct: 213 YPVYRRRPIDGCIEKGGVKCDNMYVVPYN-RFSLKYQAHINVEWCNQNGSIKYLFKYINK 271
Query: 574 GVDRVTATLETSEEPS-----------------VDEIQ*YYDCRYLSPSESIWRIFGFDV 616
G DRV +E +E + D I+ ++D Y+S SE+IWRIF F +
Sbjct: 272 GPDRVVFIVELVKEETNSNTTTLGDETVTTKKKKDGIKDWFDYIYVSASEAIWRIFKFPI 331
Query: 617 HCRYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMFLAWMDAN------RNYSQ 670
R V++L+FH+ G+Q F ++ +VL R ++ + F+ W+ N +N +
Sbjct: 332 QHRSTPVQKLSFHVEGKQPAYFDAKAKMVDVLERVSNEDSQFMVWLTLNKKNVVGKNGKR 391
Query: 671 SWNLTYVQFPRKFTYNVENRSWHL*KSGQSVGRLTFVPPGTKELYCMRLLLN 722
+ N Y + P FT++ EN+ + G +GR+ +V ++ + +LLN
Sbjct: 392 ARNCLYAEIPTYFTWDGENKPFKKRTRGFFLGRINYVLRKMEDENYLIVLLN 443
Score = 62.8 bits (151), Expect = 1e-09
Identities = 35/103 (33%), Positives = 56/103 (53%), Gaps = 26/103 (25%)
Query: 210 LRRIHETHSKYIPLQYPVLFPLGEDRYEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQK 269
++RI + H Y+ LQYP++F GED Y I+ + + GS KK+
Sbjct: 3 VKRISQIHISYLTLQYPLMFCYGEDGYTPAIEKCYNS--GSTKKK--------------- 45
Query: 270 RDNENSVIFQGMRLFQQFIVDLYSMVENQRLSYVRKNQSQIRS 312
RLFQQF++D Y+++E+ RL+Y++ NQS++RS
Sbjct: 46 ---------SVSRLFQQFLIDAYTIIESNRLAYIKFNQSKLRS 79
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.337 0.148 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,691,683
Number of Sequences: 26719
Number of extensions: 1059251
Number of successful extensions: 3753
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 3535
Number of HSP's gapped (non-prelim): 126
length of query: 1134
length of database: 11,318,596
effective HSP length: 110
effective length of query: 1024
effective length of database: 8,379,506
effective search space: 8580614144
effective search space used: 8580614144
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC135463.4


