

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135463.3 + phase: 0 /pseudo
(924 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
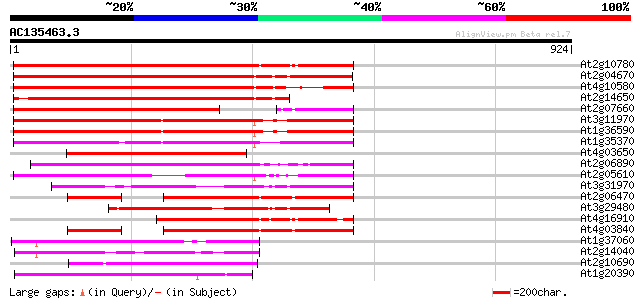
Score E
Sequences producing significant alignments: (bits) Value
At2g10780 pseudogene 625 e-179
At2g04670 putative retroelement pol polyprotein 624 e-179
At4g10580 putative reverse-transcriptase -like protein 576 e-164
At2g14650 putative retroelement pol polyprotein 525 e-149
At2g07660 putative retroelement pol polyprotein 452 e-127
At3g11970 hypothetical protein 442 e-124
At1g36590 hypothetical protein 436 e-122
At1g35370 hypothetical protein 406 e-113
At4g03650 putative reverse transcriptase 390 e-108
At2g06890 putative retroelement integrase 388 e-108
At2g05610 putative retroelement pol polyprotein 382 e-106
At3g31970 hypothetical protein 350 2e-96
At2g06470 putative retroelement pol polyprotein 289 4e-78
At3g29480 hypothetical protein 286 4e-77
At4g16910 retrotransposon like protein 285 1e-76
At4g03840 putative transposon protein 284 1e-76
At1g37060 Athila retroelment ORF 1, putative 241 1e-63
At2g14040 putative retroelement pol polyprotein 204 1e-52
At2g10690 putative retroelement pol polyprotein 200 4e-51
At1g20390 hypothetical protein 196 4e-50
>At2g10780 pseudogene
Length = 1611
Score = 625 bits (1612), Expect = e-179
Identities = 312/562 (55%), Positives = 407/562 (71%), Gaps = 7/562 (1%)
Query: 6 AELKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRI 65
A+LKKQLE+LLDK F+RPS+SPWGAPVL VKKKDG+ RLCIDYR LNKVT+KN+YPLPRI
Sbjct: 673 AKLKKQLEELLDKGFIRPSSSPWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPLPRI 732
Query: 66 DDLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVF 125
D+LMDQL GA FSKIDL SGYHQI ++ D++KTA RTRY H+E+ VMPFG+TNAP F
Sbjct: 733 DELMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAFRTRYDHFEFVVMPFGLTNAPAAF 792
Query: 126 MEYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWL 185
M+ MN +F +LD+FV++FI+DIL+YS++ E H HL+ VL L+E +L+AKLSKC FW
Sbjct: 793 MKMMNGVFRDFLDEFVIIFINDILVYSKSWEAHQEHLRAVLERLREHELFAKLSKCSFWQ 852
Query: 186 SEVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALP 245
V F GH+IS G++VDP K+ ++ +W P++ TEIRSFLGLAGYYRRF+ F+ +A P
Sbjct: 853 RSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQP 912
Query: 246 LTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQ 305
LT+LT K +F W +CE SF ELK L P+L+LP+ EP+ VY DAS +GLG VLMQ
Sbjct: 913 LTRLTGKDTAFNWSDECEKSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGCVLMQ 972
Query: 306 DGKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQ 365
G V+AYASR LR HEKNYPTHDLE+AAVV LKIWR YLYG++ ++++DHKSLKY+F Q
Sbjct: 973 KGSVIAYASRQLRKHEKNYPTHDLEMAAVVFFLKIWRSYLYGAKVQIYTDHKSLKYIFTQ 1032
Query: 366 KELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSRKTLHMSALMVKEFDLLEQFRDLS 425
ELN+RQRRW+EL+ DY+ + YHPGKAN VADALSR+ + A + DL+ L
Sbjct: 1033 PELNLRQRRWMELVADYNLDIAYHPGKANQVADALSRRRSEVEAER-SQVDLVNMMGTLH 1091
Query: 426 LVCELTPQSVQLGMLKINN-DFLNSIREAQEVDVKLVDLMVAGNGTKDSDFKVDDQGVLR 484
V L+ + LG+ + D L+ IR AQE D ++ N T +++ + G +
Sbjct: 1092 -VNALSKEVEPLGLGAADQADLLSRIRLAQERDEEIKG-WAQNNKT---EYQTSNNGTIV 1146
Query: 485 FRGRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVYACL 544
GR+C+P++ LK+ IL E+H+S SI P + KMY DLK+ + W G+K+DVA++V C
Sbjct: 1147 VNGRVCVPNDRALKEEILREAHQSKFSIHPGSNKMYRDLKRYYHWVGMKKDVARWVAKCP 1206
Query: 545 TCLKSKVEHQKPAELLTPLDVP 566
TC K EHQ P+ LL L +P
Sbjct: 1207 TCQLVKAEHQVPSGLLQNLPIP 1228
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 624 bits (1609), Expect = e-179
Identities = 306/559 (54%), Positives = 399/559 (70%), Gaps = 9/559 (1%)
Query: 7 ELKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRID 66
ELKKQLEDLL K F+RPS SPWGAPVL VKKKDG+ RLCIDYR LN VT+KN+YPLPRID
Sbjct: 532 ELKKQLEDLLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCIDYRGLNWVTVKNKYPLPRID 591
Query: 67 DLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFM 126
+L+DQL GA FSKIDL SGYHQI + + D++KTA RTRYGH+E+ VMPF +TNAP FM
Sbjct: 592 ELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAFRTRYGHFEFVVMPFALTNAPAAFM 651
Query: 127 EYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLS 186
MN +F +LD+FV++FIDDIL+YS++ EEH HL+ V+ L+E+KL+AKLSKC FW
Sbjct: 652 RLMNSVFQEFLDEFVIIFIDDILVYSKSPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQR 711
Query: 187 EVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPL 246
E+ F GHI+S G++VDP K++A+ W P + TEIRSFL L GYYRRF++GF+ +A P+
Sbjct: 712 EIGFLGHIVSAEGVSVDPEKIEAIRDWPRPTNATEIRSFLRLTGYYRRFVKGFASMAQPM 771
Query: 247 TQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQD 306
T+LT K FVW +CE F LK+ L + P+L LP+ +P++VY DAS++GLG VLMQ
Sbjct: 772 TKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQR 831
Query: 307 GKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQK 366
GKV+AYASR LR HE NYPTHDLE+A V+ LKIWR YLYG + +VF+DHKSLKY+F+Q
Sbjct: 832 GKVIAYASRQLRKHEGNYPTHDLEMAVVIFALKIWRSYLYGGKVQVFTDHKSLKYIFNQP 891
Query: 367 ELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSRKTLHMSALMVKEFDLLEQFRDLSL 426
ELN+RQ RW+EL+ DYD + YHPGKANVVADALSRK + A + + L
Sbjct: 892 ELNLRQMRWMELVADYDLEIAYHPGKANVVADALSRK--RVGAAPGQSVEALVSEIGALR 949
Query: 427 VCELTPQSVQLGMLKINN-DFLNSIREAQEVDVKLVDLMVAGNGTKDSDFKVDDQGVLRF 485
+C + + LG+ ++ D L R AQE D + ++A + + S+++ G +
Sbjct: 950 LCVVARE--PLGLEAVDRADLLTRARLAQEKD----EGLIAASKAEGSEYQFAANGTIFV 1003
Query: 486 RGRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVYACLT 545
GR+C+P ++EL++ IL E+H S SI P ATKMY DLK+ + W G+KRDVA +V C
Sbjct: 1004 YGRVCVPKDEELRREILSEAHASMFSIHPGATKMYRDLKRYYQWVGMKRDVANWVAECDV 1063
Query: 546 CLKSKVEHQKPAELLTPLD 564
C K EHQ P + +D
Sbjct: 1064 CQLVKAEHQVPDAIWVIMD 1082
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 576 bits (1484), Expect = e-164
Identities = 292/562 (51%), Positives = 376/562 (65%), Gaps = 59/562 (10%)
Query: 6 AELKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRI 65
AELKKQL+DLL K F+RPS SPWGAPVL VKKKDG+ RLCIDYR+LN+VT+KNRYPLPRI
Sbjct: 505 AELKKQLKDLLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCIDYRELNRVTVKNRYPLPRI 564
Query: 66 DDLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVF 125
D+L+DQL GA FSKIDL SGYHQI + + D++KTA RTRYGH+E+ VMPFG+TNAP VF
Sbjct: 565 DELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAFRTRYGHFEFVVMPFGLTNAPAVF 624
Query: 126 MEYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWL 185
M MN +F +LD+FV++FIDDIL+YS++ EE HL+ V+ L+E+KL+AKLSKC FW
Sbjct: 625 MRLMNSVFQEFLDEFVIIFIDDILVYSKSPEEQEVHLRRVMEKLREQKLFAKLSKCSFWQ 684
Query: 186 SEVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALP 245
E+ F GHI+S G++VDP K++A+ W P + TEIRSFLG AGYYRRF++GF+ +A P
Sbjct: 685 REMGFLGHIVSAEGVSVDPEKIEAIRDWPRPTNATEIRSFLGWAGYYRRFVKGFASMAQP 744
Query: 246 LTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQ 305
+T+LT K FVW +CE F LK+ L + P+L LP+ +P++VY DAS++GLG VLMQ
Sbjct: 745 MTKLTGKDVPFVWSQECEEGFVSLKEMLTSTPVLALPEHGQPYMVYTDASRVGLGCVLMQ 804
Query: 306 DGKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQ 365
GKV+AYASR L HE NYPTHDLE+AAV+ LKIWR YLYG + +VF+DHKSLKY+F Q
Sbjct: 805 HGKVIAYASRQLMKHEGNYPTHDLEMAAVIFALKIWRSYLYGGKVQVFTDHKSLKYIFTQ 864
Query: 366 KELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSRKTLHMSALMVKEFDLLEQFRDLS 425
ELN+RQRRW+EL+ DYD + YHPGKANVV DALSRK + A + + ++L
Sbjct: 865 PELNLRQRRWMELVADYDLEIAYHPGKANVVVDALSRK--RVGAALGQSVEVLVSEIGAL 922
Query: 426 LVCELTPQSVQLGMLKINN-DFLNSIREAQEVDVKLVDLMVAGNGTKDSDFKVDDQGVLR 484
+C + + LG+ ++ D L +R AQ+ D+G+
Sbjct: 923 RLCAVAREP--LGLEAVDRADLLTRVRLAQK----------------------KDEGL-- 956
Query: 485 FRGRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVYACL 544
ATKMY DLK+ + W G+K DVA +V C
Sbjct: 957 ------------------------------RATKMYRDLKRYYQWVGMKMDVANWVAECD 986
Query: 545 TCLKSKVEHQKPAELLTPLDVP 566
C K EHQ +L L +P
Sbjct: 987 VCQLVKAEHQVLGGMLQSLPIP 1008
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 525 bits (1351), Expect = e-149
Identities = 263/457 (57%), Positives = 331/457 (71%), Gaps = 17/457 (3%)
Query: 6 AELKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRI 65
AELKKQLEDLL GAPVL VKKKDG+ RLCIDYR LN VT+KN+YPLPRI
Sbjct: 492 AELKKQLEDLL------------GAPVLFVKKKDGSFRLCIDYRGLNWVTVKNKYPLPRI 539
Query: 66 DDLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVF 125
D+L+DQL GA FSKIDL SGYH I + + D++KTA RTRYGH+E+ VMPFG+TNAP F
Sbjct: 540 DELLDQLRGATCFSKIDLTSGYHLIPIAEADVRKTAFRTRYGHFEFVVMPFGLTNAPAAF 599
Query: 126 MEYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWL 185
M MN +F LD+FV++FIDDIL+YS++ EEH HL+ V+ L+E+KL+AKLSKC FW
Sbjct: 600 MRLMNSVFQEVLDEFVIIFIDDILVYSKSLEEHEVHLRRVMEKLREQKLFAKLSKCSFWQ 659
Query: 186 SEVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALP 245
E+ F GHI+S G++VDP K++A+ W TP + TEIRSFLGLAGYYRRF++GF+ +A P
Sbjct: 660 REMGFLGHIVSAEGVSVDPEKIEAIRDWHTPTNATEIRSFLGLAGYYRRFVKGFASMAQP 719
Query: 246 LTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQ 305
+T+LT K FVW +CE F LK+ L + P+L LP+ EP++VY DAS +GLG VLMQ
Sbjct: 720 MTKLTGKDVPFVWSPECEEGFVSLKEMLTSTPVLALPEHGEPYMVYTDASGVGLGCVLMQ 779
Query: 306 DGKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQ 365
GKV+AYASR LR HE NYPTHDLE+AAV+ LKIWR YLYG +VF+DHKSLKY+F Q
Sbjct: 780 RGKVIAYASRQLRKHEGNYPTHDLEMAAVIFALKIWRSYLYGGNVQVFTDHKSLKYIFTQ 839
Query: 366 KELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSRKTLHMSALMVKEFDLLEQFRDLS 425
ELN+RQR+W+EL+ DYD + YHPGKANVVADALS K + A + + L
Sbjct: 840 PELNLRQRQWMELVADYDLEIAYHPGKANVVADALSHK--RVGAAPGQSVEALVSEIGAL 897
Query: 426 LVCELTPQSVQLGMLKINN-DFLNSIREAQEVDVKLV 461
+C + + LG+ ++ D L +R AQE D L+
Sbjct: 898 RLCAVARE--PLGLEAVDRADLLTRVRLAQEKDEGLI 932
Score = 30.4 bits (67), Expect = 4.8
Identities = 14/32 (43%), Positives = 17/32 (52%)
Query: 535 DVAQFVYACLTCLKSKVEHQKPAELLTPLDVP 566
DVA +V C C K EHQ P +L L +P
Sbjct: 942 DVANWVAECDVCQLVKAEHQVPGGMLQSLPIP 973
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 452 bits (1164), Expect = e-127
Identities = 217/340 (63%), Positives = 267/340 (77%)
Query: 6 AELKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRI 65
AELKKQLE+LL K F+RPS+SPWGAPVL VKKKDG+ RLCIDYR LNKVT+KN+YPLPRI
Sbjct: 168 AELKKQLEELLAKGFIRPSSSPWGAPVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPLPRI 227
Query: 66 DDLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVF 125
D+LMDQL GA FSKIDL SGYHQI ++ D++KTA RTRYGH+E+ VMPFG+TNAP F
Sbjct: 228 DELMDQLGGAQWFSKIDLASGYHQIPIEPTDVRKTAFRTRYGHFEFVVMPFGLTNAPAAF 287
Query: 126 MEYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWL 185
M+ MN +F +LD+FV++FIDDIL++S++ E H HL+ VL L+E +L+AKLSK FW
Sbjct: 288 MKMMNGVFRDFLDEFVIIFIDDILVHSKSWEAHQEHLRAVLERLREHELFAKLSKFSFWQ 347
Query: 186 SEVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALP 245
V F GH+IS G++VDP K+ ++ +W P++ TEIRSFLGLAGYYRRF+ F+ +A P
Sbjct: 348 RSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQP 407
Query: 246 LTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQ 305
LT+LT K +F W +CE SF ELK L P+L+LP+ EP+ VY DAS +GLG VLMQ
Sbjct: 408 LTRLTGKDTAFNWSDECEKSFLELKAMLINAPVLVLPEEGEPYTVYTDASIVGLGCVLMQ 467
Query: 306 DGKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYL 345
G V+AYASR LR HEKNYPTHDLE+AAVV LKIWR YL
Sbjct: 468 KGSVIAYASRQLRKHEKNYPTHDLEMAAVVFALKIWRSYL 507
Score = 90.1 bits (222), Expect = 5e-18
Identities = 48/126 (38%), Positives = 75/126 (59%), Gaps = 6/126 (4%)
Query: 440 LKINNDFLNSIREAQEVDVKLVDLMVAGNGTKDSDFKVDDQGVLRFRGRICIPDNDELKK 499
LKI +L IR AQE D ++ + ++++ + G + GR+C+P++ LK+
Sbjct: 500 LKIWRSYL--IRLAQERDEEIKGWTL----NNKTEYQTSNNGTIVVNGRVCVPNDRALKE 553
Query: 500 LILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVYACLTCLKSKVEHQKPAEL 559
IL E+H+S SI P + KMY DLK+ + W G+++DVA++V C TC K EHQ P+ L
Sbjct: 554 EILREAHQSKFSIHPGSNKMYRDLKRYYHWVGMRKDVARWVAKCPTCQLVKAEHQVPSGL 613
Query: 560 LTPLDV 565
L L +
Sbjct: 614 LQNLPI 619
>At3g11970 hypothetical protein
Length = 1499
Score = 442 bits (1136), Expect = e-124
Identities = 244/565 (43%), Positives = 344/565 (60%), Gaps = 36/565 (6%)
Query: 7 ELKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRID 66
E+ K +EDLL V+ S+SP+ +PV+LVKKKDGT RLC+DYR+LN +T+K+ +P+P I+
Sbjct: 616 EIDKLVEDLLTNGTVQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDSFPIPLIE 675
Query: 67 DLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFM 126
DLMD+L GA +FSKIDLR+GYHQ+++ +D+QKTA +T GH+EY VMPFG+TNAP F
Sbjct: 676 DLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKTAFKTHSGHFEYLVMPFGLTNAPATFQ 735
Query: 127 EYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLS 186
MN IF +L KFV+VF DDIL+YS + EEH HLK V V++ KL+AKLSKC F +
Sbjct: 736 GLMNFIFKPFLRKFVLVFFDDILVYSSSLEEHRQHLKQVFEVMRANKLFAKLSKCAFAVP 795
Query: 187 EVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPL 246
+V + GH IS GI DP+K+ AV +W P ++ ++R FLGLAGYYRRF+ F +A PL
Sbjct: 796 KVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLKQLRGFLGLAGYYRRFVRSFGVIAGPL 855
Query: 247 TQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQD 306
LT K +F W A + +F +LK L P+L LP ++ FVV DA G+G VLMQ+
Sbjct: 856 HALT-KTDAFEWTAVAQQAFEDLKAALCQAPVLSLPLFDKQFVVETDACGQGIGAVLMQE 914
Query: 307 GKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQK 366
G +AY SR L+ + + ++ EL AV+ ++ WRHYL S F + +D +SLKYL +Q+
Sbjct: 915 GHPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLLQSHFIIKTDQRSLKYLLEQR 974
Query: 367 ELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSR----KTLHMSALMVKEFDLLEQFR 422
Q++WL L ++D+ + Y GK NVVADALSR + LHM A+ V E DLL
Sbjct: 975 LNTPIQQQWLPKLLEFDYEIQYRQGKENVVADALSRVEGSEVLHM-AMTVVECDLL---- 1029
Query: 423 DLSLVCELTPQSVQLGMLKINNDFLNSIREAQEVDVKLVDLMVAGNGTKDS-DFKVDDQG 481
+ +Q G D +L D++ A DS + Q
Sbjct: 1030 ----------KDIQAGYAN---------------DSQLQDIITALQRDPDSKKYFSWSQN 1064
Query: 482 VLRFRGRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVY 541
+LR + +I +P ND +K IL H S + + +K LF+W G+ +D+ ++
Sbjct: 1065 ILRRKSKIVVPANDNIKNTILLWLHGSGVGGHSGRDVTHQRVKGLFYWKGMIKDIQAYIR 1124
Query: 542 ACLTCLKSKVEHQKPAELLTPLDVP 566
+C TC + K + LL PL +P
Sbjct: 1125 SCGTCQQCKSDPAASPGLLQPLPIP 1149
>At1g36590 hypothetical protein
Length = 1499
Score = 436 bits (1122), Expect = e-122
Identities = 243/565 (43%), Positives = 343/565 (60%), Gaps = 36/565 (6%)
Query: 7 ELKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRID 66
E+ K +EDLL V+ S+SP+ +PV+LVKKKDGT RLC+DYR+LN +T+K+ +P+P I+
Sbjct: 616 EIDKLVEDLLTNGTVQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDSFPIPLIE 675
Query: 67 DLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFM 126
DLMD+L GA +FSKIDLR+GYHQ+++ +D+QKTA +T GH+EY VMPFG+TNAP F
Sbjct: 676 DLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKTAFKTHSGHFEYLVMPFGLTNAPATFQ 735
Query: 127 EYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLS 186
MN IF +L KFV+VF DDIL+YS + EEH HLK V V++ KL+AKLSKC F +
Sbjct: 736 GLMNFIFKPFLRKFVLVFFDDILVYSSSLEEHRQHLKQVFEVMRANKLFAKLSKCAFAVP 795
Query: 187 EVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPL 246
+V + GH IS GI DP+K+ AV +W P ++ ++R FLGLAGYYRRF+ F +A PL
Sbjct: 796 KVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLKQLRGFLGLAGYYRRFVRSFGVIAGPL 855
Query: 247 TQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQD 306
LT K +F W A + +F +LK L P+L LP ++ FVV DA G+G VLMQ+
Sbjct: 856 HALT-KTDAFEWTAVAQQAFEDLKAALCQAPVLSLPLFDKQFVVETDACGQGIGAVLMQE 914
Query: 307 GKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQK 366
G +AY SR L+ + + ++ EL AV+ ++ WRHYL S F + +D +SLKYL +Q+
Sbjct: 915 GHPLAYISRQLKGKQLHLSIYEKELLAVIFAVRKWRHYLLQSHFIIKTDQRSLKYLLEQR 974
Query: 367 ELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSR----KTLHMSALMVKEFDLLEQFR 422
Q++WL L ++D+ + Y GK NVVADALSR + LHM A+ V E DLL
Sbjct: 975 LNTPIQQQWLPKLLEFDYEIQYRQGKENVVADALSRVEGSEVLHM-AMTVVECDLL---- 1029
Query: 423 DLSLVCELTPQSVQLGMLKINNDFLNSIREAQEVDVKLVDLMVAGNGTKDS-DFKVDDQG 481
+ +Q G D +L D++ A DS + Q
Sbjct: 1030 ----------KDIQAGYAN---------------DSQLQDIITALQRDPDSKKYFSWSQN 1064
Query: 482 VLRFRGRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVY 541
+LR + +I +P ND +K IL H S + + +K LF+ G+ +D+ ++
Sbjct: 1065 ILRRKSKIVVPANDNIKNTILLWLHGSGVGGHSGRDVTHQRVKGLFYSKGMIKDIQAYIR 1124
Query: 542 ACLTCLKSKVEHQKPAELLTPLDVP 566
+C TC + K + LL PL +P
Sbjct: 1125 SCGTCQQCKSDPAASPGLLQPLPIP 1149
>At1g35370 hypothetical protein
Length = 1447
Score = 406 bits (1043), Expect = e-113
Identities = 227/565 (40%), Positives = 334/565 (58%), Gaps = 44/565 (7%)
Query: 7 ELKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRID 66
E+ K ++D++ ++ S+SP+ +PV+LVKKKDGT RLC+DY +LN +T+K+R+ +P I+
Sbjct: 595 EIDKIVQDMIKSGTIQVSSSPFASPVVLVKKKDGTWRLCVDYTELNGMTVKDRFLIPLIE 654
Query: 67 DLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFM 126
DLMD+L G+ VFSKIDLR+GYHQ+++ +D+QKTA +T GH+EY VM FG+TNAP F
Sbjct: 655 DLMDELGGSVVFSKIDLRAGYHQVRMDPDDIQKTAFKTHNGHFEYLVMLFGLTNAPATFQ 714
Query: 127 EYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLS 186
MN +F +L KFV+VF DDILIYS + EEH HL++V V++ KL+AK SK
Sbjct: 715 SLMNSVFRDFLRKFVLVFFDDILIYSSSIEEHKEHLRLVFEVMRLHKLFAKGSK------ 768
Query: 187 EVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPL 246
GH IS I DP+K+ AV +W TP +V ++R FLG AGYYRRF+ F +A PL
Sbjct: 769 --EHLGHFISAREIETDPAKIQAVKEWPTPTTVKQVRGFLGFAGYYRRFVRNFGVIAGPL 826
Query: 247 TQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQD 306
LT K F W + +S+F+ LK L P+L LP ++ F+V DA G+ VLMQ
Sbjct: 827 HALT-KTDGFCWSLEAQSAFDTLKAVLCNAPVLALPVFDKQFMVETDACGQGIRAVLMQK 885
Query: 307 GKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQK 366
G +AY SR L+ + + ++ EL A + ++ WRHYL S F + +D +SLKYL +Q+
Sbjct: 886 GHPLAYISRQLKGKQLHLSIYEKELLAFIFAVRKWRHYLLPSHFIIKTDQRSLKYLLEQR 945
Query: 367 ELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSR----KTLHMSALMVKEFDLLEQFR 422
Q++WL L ++D+ + Y GK N+VADALSR + LHM+ +V+
Sbjct: 946 LNTPVQQQWLPKLLEFDYEIQYRQGKENLVADALSRVEGSEVLHMALSIVE--------- 996
Query: 423 DLSLVCELTPQSVQLGMLKINNDFLNSIREAQEVDVKLVDLMVAGNGTKDSDFKVD-DQG 481
C DFL I+ A E D L D++ A D+ Q
Sbjct: 997 -----C----------------DFLKEIQVAYESDGVLKDIISALQQHPDAKKHYSWSQD 1035
Query: 482 VLRFRGRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVY 541
+LR + +I +P++ E+ +L+ H S + R + +K LF+W G+ +D+ F+
Sbjct: 1036 ILRRKSKIVVPNDVEITNKLLQWLHCSGMGGRSGRDASHQRVKSLFYWKGMVKDIQAFIR 1095
Query: 542 ACLTCLKSKVEHQKPAELLTPLDVP 566
+C TC + K ++ LL PL +P
Sbjct: 1096 SCGTCQQCKSDNAAYPGLLQPLPIP 1120
>At4g03650 putative reverse transcriptase
Length = 839
Score = 390 bits (1002), Expect = e-108
Identities = 178/296 (60%), Positives = 229/296 (77%)
Query: 94 DEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSR 153
+ D++KTA RTRYGH+E+ VMPFG+TNAP FM MN +F +LD+FV++FIDDIL+YS+
Sbjct: 518 EADVRKTAFRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVYSK 577
Query: 154 TEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLSEVSFPGHIISGSGIAVDPSKVDAVSQW 213
+ EEH HL+ V+ L+E+KL+AKLSKC FW E+ F GHI+S G++VDP K++A+ W
Sbjct: 578 SPEEHEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDW 637
Query: 214 ETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPLTQLTCKGKSFVWDAQCESSFNELKQRL 273
P + TEIRSFLGLAGYYRRFI+GF+ +A P+T+LT K FVW +CE F LK+ L
Sbjct: 638 PRPTNATEIRSFLGLAGYYRRFIKGFASMAQPMTKLTGKDVPFVWSPECEEGFVSLKEML 697
Query: 274 PTNPILILPKPEEPFVVYFDASKLGLGVVLMQDGKVVAYASRLLRIHEKNYPTHDLELAA 333
+ P+L LP+ EP++VY DAS +GLG LMQ GKV+AYASR LR HE NYPTHDLE+AA
Sbjct: 698 TSTPVLALPEHGEPYMVYTDASGVGLGCALMQRGKVIAYASRQLRKHEGNYPTHDLEMAA 757
Query: 334 VVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQKELNMRQRRWLELLKDYDFGLNYH 389
V+ LKIWR YLYG + +VF+DHKSLKY+F Q ELN+RQRRW+EL+ DYD + YH
Sbjct: 758 VIFALKIWRSYLYGGKVQVFTDHKSLKYIFTQPELNLRQRRWMELVADYDLEIAYH 813
>At2g06890 putative retroelement integrase
Length = 1215
Score = 388 bits (997), Expect = e-108
Identities = 216/533 (40%), Positives = 317/533 (58%), Gaps = 33/533 (6%)
Query: 34 LVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGANVFSKIDLRSGYHQIKVK 93
L ++KDG+ R+C D R +N VT+K +P+PR+DD++D+L G+++FSKIDL+SGYHQI++
Sbjct: 461 LQRQKDGSWRMCFDCRAINNVTVKYCHPIPRLDDMLDELHGSSIFSKIDLKSGYHQIRMN 520
Query: 94 DEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSR 153
+ D KTA +T++G YE+ VMPFG+T+AP FM MN + A++ FV+V+ DDIL+YS
Sbjct: 521 EGDEWKTAFKTKHGLYEWLVMPFGLTHAPSTFMRLMNHVLRAFIGIFVIVYFDDILVYSE 580
Query: 154 TEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLSEVSFPGHIISGSGIAVDPSKVDAVSQW 213
+ EH HL VL VL++++LYA L KC F + F G ++S G+ VD KV A+ W
Sbjct: 581 SLREHIEHLDSVLNVLRKEELYANLKKCTFCTDNLVFLGFVVSADGVKVDEEKVKAIRDW 640
Query: 214 ETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPLTQLTCKGKSFVWDAQCESSFNELKQRL 273
+PK+V E+RSF GLAG+YRRF + FS + PLT++ K F W+ E +F LK +L
Sbjct: 641 PSPKTVGEVRSFHGLAGFYRRFFKDFSTIVAPLTEVMKKDVGFKWEKAQEEAFQSLKDKL 700
Query: 274 PTNPILILPKPEEPFVVYFDASKLGLGVVLMQDGKVVAYASRLLRIHEKNYPTHDLELAA 333
P+LIL + + F + DAS +G+G VLMQD K++A+ S L NYPT+D EL A
Sbjct: 701 TNAPVLILSEFLKTFEIECDASGIGIGAVLMQDQKLIAFFSEKLGGATLNYPTYDKELYA 760
Query: 334 VVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQKELNMRQRRWLELLKDYDFGLNYHPGKA 393
+V L+ W+HYL+ F + +DH+SLK+L Q++LN R RW+E ++ + + + Y GK
Sbjct: 761 LVRALQRWQHYLWPKVFVIHTDHESLKHLKGQQKLNKRHARWVEFIETFAYVIKYKKGKD 820
Query: 394 NVVADALSRKTLHMSALMVKEFDLLEQFRDLSLVCELTPQSVQLGMLKINNDFLNSIREA 453
NVVADALS++ +S L VK L F + V E ++DF
Sbjct: 821 NVVADALSQRYTLLSTLNVK----LMGFEQIKEVYE------------TDHDF------- 857
Query: 454 QEVDVKLVDLMVAGNGTKDSDFKVDDQGVLRFRGRICIPDNDELKKLILEESHKSSLSIR 513
QEV K + +G + F L + R+C+P N L+ L + E+H L
Sbjct: 858 QEV-YKACEKFASGRYFRQDKF-------LFYENRLCVP-NCSLRDLFVREAHGGGLMGH 908
Query: 514 PEATKMYHDLKKLFWWSGLKRDVAQFVYACLTCLKSKVEHQKPAELLTPLDVP 566
K + + F W +K DV + C TC ++K + Q P L TPL +P
Sbjct: 909 FGIAKTLEVMTEHFRWPHMKCDVKRICGRCNTCKQAKSKIQ-PNGLYTPLPIP 960
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 382 bits (982), Expect = e-106
Identities = 221/564 (39%), Positives = 316/564 (55%), Gaps = 85/564 (15%)
Query: 7 ELKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRID 66
E+ K ++D+L ++ S+SP+ +PV+LVKKKDGT RLC+DYR+LN +T+K+R+P+P I+
Sbjct: 58 EIDKIVDDMLASGTIQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDRFPIPLIE 117
Query: 67 DLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFM 126
DLMD+L G+NV+SKIDLR+GYHQ+++ D+ KTA +T GHYEY VMPFG+TNAP F
Sbjct: 118 DLMDELGGSNVYSKIDLRAGYHQVRMDPLDIHKTAFKTHNGHYEYLVMPFGLTNAPASFQ 177
Query: 127 EYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLS 186
MN F +L KFV+VF DDILIYS + EEH HL+ V V++ L+AK+SKC F +
Sbjct: 178 SLMNSFFKPFLRKFVLVFFDDILIYSTSMEEHKKHLEAVFEVMRVHHLFAKMSKCAFAVP 237
Query: 187 EVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPL 246
V + GH ISG GIA DP+K+ AV W P ++ ++ FLGL GYYRRF
Sbjct: 238 RVEYLGHFISGEGIATDPAKIKAVQDWPVPVNLKQLCGFLGLTGYYRRF----------- 286
Query: 247 TQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQD 306
FVV DA G+G VLMQ+
Sbjct: 287 -----------------------------------------FVVEMDACGHGIGAVLMQE 305
Query: 307 GKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQK 366
G +A+ SR L+ + + ++ EL AV+ V++ WRHYL S F + +D +SLKYL +Q+
Sbjct: 306 GHPLAFISRQLKGKQLHLSIYEKELLAVIFVVRKWRHYLISSHFIIKTDQRSLKYLLEQR 365
Query: 367 ELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSR----KTLHMSALMVKEFDLLEQFR 422
Q++WL L ++D+ + Y GK N+VADALSR + LHM AL V E DLL++ +
Sbjct: 366 LNTPIQQQWLPKLLEFDYEIQYKQGKENLVADALSRVEGSEVLHM-ALSVVECDLLKEIQ 424
Query: 423 DLSLVCELTPQSVQLGMLKINNDFLNSIREAQEVDVKLVDLMVAGNGTKDSDFKVDDQGV 482
+T +Q G++ I Q+ D K QGV
Sbjct: 425 ----AGYVTDGDIQ-GIITILQ---------QQADSK--------------KHYTWSQGV 456
Query: 483 LRFRGRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVYA 542
LR + +I +P+N +K IL H S + + +K LF+W + +D+ F+ +
Sbjct: 457 LRRKNKIVVPNNSGIKDTILRWLHCSGMGGHSGKEVTHQRVKGLFYWKSMVKDIQAFIRS 516
Query: 543 CLTCLKSKVEHQKPAELLTPLDVP 566
C TC + K ++ LL PL +P
Sbjct: 517 CGTCQQCKSDNAASPGLLQPLPIP 540
>At3g31970 hypothetical protein
Length = 1329
Score = 350 bits (899), Expect = 2e-96
Identities = 200/499 (40%), Positives = 282/499 (56%), Gaps = 82/499 (16%)
Query: 69 MDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFMEY 128
M + VG S I + + I + + +++KTA RTRYGH+E+ VMPFG+TN P FM
Sbjct: 552 MPESVGQVAVSDIRVVHEFEDIPIAEANVRKTAFRTRYGHFEFVVMPFGLTNIPTAFMRL 611
Query: 129 MNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLSEV 188
MN +F +LD+FV++FIDDIL+YS++ EEH ++ ++ S E +EV
Sbjct: 612 MNSVFQEFLDEFVIIFIDDILVYSKSPEEH--------------EVQSEESDGEAAGAEV 657
Query: 189 SFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPLTQ 248
G++VD K++A+ W P + TEIRSFLGLAGYY+RF++ F+ +A P+T+
Sbjct: 658 ----------GVSVDLEKIEAIRDWPRPTNATEIRSFLGLAGYYKRFVKVFASMAQPMTK 707
Query: 249 LTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQDGK 308
LT K FVW +C+ F LK+ L + P+L LP+ EP++VY DAS++GL VLMQ GK
Sbjct: 708 LTGKDVPFVWSPECDEGFMSLKEMLTSTPVLALPEHGEPYMVYTDASRVGLDCVLMQRGK 767
Query: 309 VVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQKEL 368
VF+DHKSLKY+F Q EL
Sbjct: 768 -------------------------------------------VFTDHKSLKYIFTQPEL 784
Query: 369 NMRQRRWLELLKDYDFGLNYHPGKANVVADALSRKTLHMSA-LMVKEFDLLEQFRDLSLV 427
N+RQRRW+EL+ DYD + YH GKANVVADALSRK + S +V E L +
Sbjct: 785 NLRQRRWMELVADYDLEIAYHSGKANVVADALSRKRVGGSVEALVSEIGALR-------L 837
Query: 428 CELTPQSVQLGMLKINN-DFLNSIREAQEVDVKLVDLMVAGNGTKDSDFKVDDQGVLRFR 486
C + + LG+ ++ D L +R AQE D + ++A + S+++ G +
Sbjct: 838 CVMAQE--PLGLEAVDRADLLTRVRLAQEKD----EGLIAASKADGSEYQFAANGTILVH 891
Query: 487 GRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVYACLTC 546
GR+C+P ++EL++ IL E+H S SI P ATKMY DLK+ + W G+KRDVA +V C C
Sbjct: 892 GRVCVPKDEELRREILSEAHASMFSIHPRATKMYRDLKRYYQWVGMKRDVANWVTECDVC 951
Query: 547 LKSKVEHQKPAELLTPLDV 565
K EHQ P LL L +
Sbjct: 952 QLVKAEHQVPGGLLQSLPI 970
>At2g06470 putative retroelement pol polyprotein
Length = 899
Score = 289 bits (740), Expect = 4e-78
Identities = 152/314 (48%), Positives = 204/314 (64%), Gaps = 7/314 (2%)
Query: 254 KSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQDGKVVAYA 313
K W CE SF ELK L P+L+LP+ EP+ VY DAS +GLG VLMQ G V+AYA
Sbjct: 371 KCSFWQRSCEKSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGCVLMQKGSVIAYA 430
Query: 314 SRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQKELNMRQR 373
SR L HEKNYPTHDLE+AAVV LKIWR YLYG++ ++++DHKSLKY+F Q ELN+RQR
Sbjct: 431 SRQLWKHEKNYPTHDLEMAAVVFALKIWRSYLYGAKVQIYTDHKSLKYIFIQPELNLRQR 490
Query: 374 RWLELLKDYDFGLNYHPGKANVVADALSRKTLHMSALMVKEFDLLEQFRDLSLVCELTPQ 433
RW+EL+ DYD + YHPGKAN VADALSR+ + A + DL+ L V L+ +
Sbjct: 491 RWMELVADYDLDIAYHPGKANQVADALSRRRSEVEAER-SQVDLVNMMGTLH-VNALSKE 548
Query: 434 SVQLGMLKINN-DFLNSIREAQEVDVKLVDLMVAGNGTKDSDFKVDDQGVLRFRGRICIP 492
LG+ + D L+ IR AQE D ++ + ++++ + G + GR+C+P
Sbjct: 549 VESLGLGAADQADLLSRIRLAQERDEEIKGWAL----NNKTEYQTSNNGTIVVNGRVCVP 604
Query: 493 DNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVYACLTCLKSKVE 552
++ LK+ IL E+H+S SI P + KMY DLK+ + W G+K+DVA++V C TC K E
Sbjct: 605 NDRALKEEILREAHQSKFSIHPGSNKMYRDLKRYYHWVGMKKDVARWVAKCPTCQLVKAE 664
Query: 553 HQKPAELLTPLDVP 566
HQ P+ LL L +P
Sbjct: 665 HQVPSGLLQNLPIP 678
Score = 121 bits (304), Expect = 2e-27
Identities = 54/89 (60%), Positives = 72/89 (80%)
Query: 96 DMQKTASRTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRTE 155
D++KTA RTRYGH+E+ VMPFG+TNAP FM+ MN +F +LD+FV++FIDDIL+YS++
Sbjct: 287 DVRKTAFRTRYGHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFIDDILVYSKSW 346
Query: 156 EEHA*HLKIVL*VLKEKKLYAKLSKCEFW 184
E H HL+ VL L+E +L+AKLSKC FW
Sbjct: 347 EAHQEHLRAVLERLREHELFAKLSKCSFW 375
>At3g29480 hypothetical protein
Length = 718
Score = 286 bits (732), Expect = 4e-77
Identities = 162/368 (44%), Positives = 232/368 (63%), Gaps = 32/368 (8%)
Query: 164 IVL*VLKEKKLYAKLSKCEFWLSEVSFPGH---IISGSGIAVDPSKVDAVSQWETPKSVT 220
+V+ ++ +K+ K CE +L +S + +G++VD K++A+ W P + T
Sbjct: 379 LVISAVQAEKMIKK--DCEAYLVTISMKSDGEATGAEAGVSVDLEKIEAIKDWPRPTNAT 436
Query: 221 EIRSFLGLAGYYRRFIEGFSKLALPLTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILI 280
EIRSFLGLAGYYRRF++GF+ +A PLT+LT K FVW +CE F LK+ L + P+L
Sbjct: 437 EIRSFLGLAGYYRRFVKGFASMAPPLTKLTGKNVPFVWSPECEEGFVSLKEILTSTPVLA 496
Query: 281 LPKPEEPFVVYFDASKLGLGVVLMQDGKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKI 340
LP+ E ++VY DAS++GLG VLMQ GKV+AYASR LR HE NYPTHDLE++A
Sbjct: 497 LPEHGESYMVYTDASRVGLGCVLMQRGKVIAYASRQLRKHEGNYPTHDLEMSA------- 549
Query: 341 WRHYLYGSRFEVFSDHKSLKYLFDQKELNMRQRRWLELLKDYDFGLNYHPGKANVVADAL 400
VF+DHKSLKY+ Q ELN+RQRRW+EL+ DYD + YH GKA+VVADAL
Sbjct: 550 -----------VFTDHKSLKYIITQPELNLRQRRWMELVADYDLEIAYHHGKASVVADAL 598
Query: 401 SRKTLHMSALMVKEFDLLEQFRDLSLVCELTPQSVQLGMLKINN-DFLNSIREAQEVDVK 459
SRK + ++ E L+ + R L L C + + LG+ ++ D L+ +R AQE D
Sbjct: 599 SRKRVGVAPGQSVE-ALVSEIRALRL-CAVARE--PLGLEAVDRVDLLSRVRLAQEKD-- 652
Query: 460 LVDLMVAGNGTKDSDFKVDDQGVLRFRGRICIPDNDELKKLILEESHKSSLSIRPEATKM 519
+ ++A + + S ++ G + GR+C+ +++EL++ IL E+H S SI P ATKM
Sbjct: 653 --EGLIAASKAEGSKYQFAANGTILVHGRVCVLNDEELRREILSEAHASMFSIHPGATKM 710
Query: 520 YHDLKKLF 527
Y DLK+ +
Sbjct: 711 YRDLKEYY 718
>At4g16910 retrotransposon like protein
Length = 687
Score = 285 bits (728), Expect = 1e-76
Identities = 155/326 (47%), Positives = 212/326 (64%), Gaps = 15/326 (4%)
Query: 242 LALPLTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGV 301
+A P+TQLT K +F W +CE SF ELK L P+L+LP+ EP+ VY DAS +GLG
Sbjct: 1 MAQPMTQLTGKDTAFNWSEECERSFLELKAMLTNAPVLVLPEEGEPYTVYTDASIVGLGC 60
Query: 302 VLMQDGKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKY 361
VL+Q G V+AYASR LR HEKNYPT+DLE+AAVV LKIWR YLYG++ ++F+DHKSLKY
Sbjct: 61 VLIQKGSVIAYASRQLRKHEKNYPTNDLEMAAVVFALKIWRSYLYGAKVQIFTDHKSLKY 120
Query: 362 LFDQKELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSRKTLHMSALMVKEFDLLEQF 421
+F Q ELN+RQRRW++L+ DYD + YHPGKAN V DALSR + A + +L+
Sbjct: 121 IFTQPELNLRQRRWIKLVADYDLNIAYHPGKANQVVDALSRLRTVVEAER-NQVNLVNMM 179
Query: 422 RDLSLVCELTPQSVQLGMLKINN-DFLNSIREAQEVDVKLVDLMVAGNGTKDSDFKVDDQ 480
L L L+ + LG+ N D L+ IR AQE D ++ N T +++ +
Sbjct: 180 GTLHLNA-LSKEVEPLGLRAANQADLLSRIRSAQERDEEIKG-WAQNNKT---EYQSSNN 234
Query: 481 GVLRFRGRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFV 540
G + GR+C P++ LK+ IL+E+H+S SI P + KMY DLK+ + W G+K+DVA++V
Sbjct: 235 GTIVVNGRVCGPNDKALKEEILKEAHQSKFSIHPGSNKMYRDLKRYYHWVGMKKDVARWV 294
Query: 541 YACLTCLKSKVEHQKPAELLTPLDVP 566
+K EHQ P+ +L L +P
Sbjct: 295 --------AKEEHQVPSGMLQNLPIP 312
>At4g03840 putative transposon protein
Length = 973
Score = 284 bits (727), Expect = 1e-76
Identities = 149/314 (47%), Positives = 203/314 (64%), Gaps = 7/314 (2%)
Query: 254 KSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQDGKVVAYA 313
K W CE +F ELK L P+L+LP+ EP+ VY DAS +GL VLMQ G V+AYA
Sbjct: 371 KCSFWQRSCEKNFLELKAMLTNAPVLVLPEEGEPYTVYTDASVVGLECVLMQKGSVIAYA 430
Query: 314 SRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQKELNMRQR 373
SR LR HEKNYPTHDLE+AAVV LKIWR YLYG++ ++++DHKSLKY+F Q ELN+RQR
Sbjct: 431 SRQLRKHEKNYPTHDLEMAAVVFALKIWRSYLYGAKVQIYTDHKSLKYIFTQPELNLRQR 490
Query: 374 RWLELLKDYDFGLNYHPGKANVVADALSRKTLHMSALMVKEFDLLEQFRDLSLVCELTPQ 433
RW+EL+ DYD + YH GKAN VADALSR+ + A + DL+ L V L+ +
Sbjct: 491 RWMELVADYDLDIAYHAGKANQVADALSRRRSEVEAER-SQVDLVNMMGTLH-VNALSKE 548
Query: 434 SVQLGMLKINN-DFLNSIREAQEVDVKLVDLMVAGNGTKDSDFKVDDQGVLRFRGRICIP 492
LG+ + + L+ IR AQE D ++ + ++++ + G + GR+C+P
Sbjct: 549 VEPLGLGAADQANLLSRIRLAQERDEEIKGWAL----NNKTEYQTSNNGTIVVNGRVCVP 604
Query: 493 DNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVYACLTCLKSKVE 552
+N LK+ IL E+H+S SI P + K+Y DLK+ + W G+K+DVA++V C TC K E
Sbjct: 605 NNRALKEEILREAHQSKFSIHPGSNKIYRDLKRYYHWVGMKKDVARWVAKCPTCQLVKAE 664
Query: 553 HQKPAELLTPLDVP 566
HQ P+ LL L +P
Sbjct: 665 HQVPSGLLQNLPIP 678
Score = 118 bits (296), Expect = 1e-26
Identities = 53/89 (59%), Positives = 71/89 (79%)
Query: 96 DMQKTASRTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRTE 155
D++KTA TRYGH+E+ VMPFG+TNAP FM+ MN +F +LD+FV++FIDDIL+YS++
Sbjct: 287 DVRKTAFLTRYGHFEFVVMPFGLTNAPTAFMKMMNGVFRDFLDEFVIIFIDDILVYSKSW 346
Query: 156 EEHA*HLKIVL*VLKEKKLYAKLSKCEFW 184
E H HL+ VL L+E +L+AKLSKC FW
Sbjct: 347 EAHQEHLRAVLEQLREHELFAKLSKCSFW 375
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 241 bits (615), Expect = 1e-63
Identities = 150/428 (35%), Positives = 221/428 (51%), Gaps = 43/428 (10%)
Query: 3 NVCAELKKQLEDLLDKKFVRP-SASPWGAPVLLVKKKDGTM------------------R 43
N+ +KK++ LLD + P S S W +PV V KK G R
Sbjct: 910 NLKEVVKKEILKLLDAGVIYPISDSTWVSPVHCVPKKGGMTVVKNSKDELIPTRTITGHR 969
Query: 44 LCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASR 103
+CI+YR+LN + K +PLP ID ++++L + +D SG+ QI + D KT
Sbjct: 970 MCIEYRKLNVASRKEHFPLPFIDHMLERLANHPYYCFLDSYSGFFQIPIHPNDQGKTTFT 1029
Query: 104 TRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLK 163
YG + YK MPFG+ NAP F M IF +++ V VF+DD +Y + +L
Sbjct: 1030 CPYGTFAYKRMPFGLCNAPATFQRCMTSIFSDLIEEMVEVFMDDFSVYGSSFSSCLLNLC 1089
Query: 164 IVL*VLKEKKLYAKLSKCEFWLSEVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIR 223
VL +E L KC F + E G IS GI VD +K+D + Q + PK+V +IR
Sbjct: 1090 RVLKRCEETNLVLNWEKCHFMVREGIVLGRKISEEGIEVDKAKIDVMMQLQPPKTVKDIR 1149
Query: 224 SFLGLAGYYRRFIEGFSKLALPLTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPK 283
SFLG AG+YR FI+ FSKLA PLT+L CK F +D +C ++F +K+ L T PI+ P
Sbjct: 1150 SFLGHAGFYRIFIKDFSKLARPLTRLLCKETEFAFDDECLTAFKLIKEALITAPIVQAPN 1209
Query: 284 PEEPFVVYFDASKLGLGVVLMQDGKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRH 343
+ PF ++ M D +V Y T + EL AVV + +R
Sbjct: 1210 WDFPF-----------EIITMDDAQV-------------RYATTEKELLAVVFAFEKFRS 1245
Query: 344 YLYGSRFEVFSDHKSLKYLFDQKELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSRK 403
YL GS+ +++DH +L++++ +K+ R RW+ LL+++D + G N VAD LSR
Sbjct: 1246 YLVGSKVTIYTDHAALRHIYAKKDTKPRLLRWILLLQEFDMEIVDKKGIENGVADHLSRM 1305
Query: 404 TLHMSALM 411
+ L+
Sbjct: 1306 RIEDEVLI 1313
>At2g14040 putative retroelement pol polyprotein
Length = 841
Score = 204 bits (520), Expect = 1e-52
Identities = 141/423 (33%), Positives = 204/423 (47%), Gaps = 71/423 (16%)
Query: 8 LKKQLEDLLDKKFVRP-SASPWGAPVLLVKKKDGTM------------------RLCIDY 48
+KK++ LLD + P S S W +PV +V KK G + R+CIDY
Sbjct: 390 VKKEIMKLLDAGIIYPISDSTWVSPVHVVPKKGGVIVIKNEKNELIPTRTVTGHRMCIDY 449
Query: 49 RQLNKVTIKNRYPLPRIDDLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGH 108
R+LN T K+ +PL ID ++++L + +D G+ QI + +D +KT YG
Sbjct: 450 RKLNSATRKDNFPLSFIDQMLERLSNQPYYCFLDGYLGFFQILIHPDDQEKTTFTCPYGT 509
Query: 109 YEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*V 168
+ Y+ MPFG+ NAP F M IF ++ F+ VFIDD L+ R E++H
Sbjct: 510 FAYRRMPFGLCNAPATFQHCMKYIFSDMIEDFMEVFIDDFLVLQRCEDKH---------- 559
Query: 169 LKEKKLYAKLSKCEFWLSEVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGL 228
L K F + + GH IS G+ VD +K+ EI FLG
Sbjct: 560 -----LVLNWEKSHFMVRDGIVLGHKISEKGVEVDRAKI-------------EIMRFLGH 601
Query: 229 AGYYRRFIEGFSKLALPLTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPF 288
AG+YRRFI+ FSK+A P TQL CK + +D C +FN +K+ L + I+ P+ E PF
Sbjct: 602 AGFYRRFIKDFSKIARPCTQLLCKEQKSEFDDTCLQAFNIVKESLVSALIVQPPEWELPF 661
Query: 289 VVYFDASKLGLGVVLMQDGKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGS 348
V DAS +G VL Q + ++H Y + L+ A V+
Sbjct: 662 EVICDASDYVVGAVLGQ--------RKDKKLHAIYYASRTLDGAQVI------------- 700
Query: 349 RFEVFSDHKSLKYLFDQKELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSRKTLHMS 408
V + H +L+YL +K+ R RW+ LL+D+D + G N VAD LSR +
Sbjct: 701 ---VHTVHAALRYLLSKKDAKPRLLRWILLLQDFDLEIKDKKGIENEVADNLSRLRVQEE 757
Query: 409 ALM 411
LM
Sbjct: 758 VLM 760
>At2g10690 putative retroelement pol polyprotein
Length = 622
Score = 200 bits (508), Expect = 4e-51
Identities = 123/315 (39%), Positives = 175/315 (55%), Gaps = 8/315 (2%)
Query: 98 QKTASRTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRTEEE 157
+KT G + YK M FG+ NAPG F M IF ++++ + VF+DD +S
Sbjct: 164 KKTTFTCPNGTFAYKRMSFGLCNAPGTFQRSMTSIFSDFIEEIMEVFMDDFSGFSSC--- 220
Query: 158 HA*HLKIVL*VLKEKKLYAKLSKCEFWLSEVSFPGHIISGSGIAVDPSKVDAVSQWETPK 217
+L VL +E L KC F + E GH ISG GI VD +K+D + Q + PK
Sbjct: 221 -LLNLCRVLERCEETNLVLNWEKCHFMVHEGIVLGHKISGKGIEVDKAKIDVMIQLQPPK 279
Query: 218 SVTEIRSFLGLAGYYRRFIEGFSKLALPLTQLTCKGKSFVWDAQCESSFNELKQRLPTNP 277
+V +IRSFLG A +YRRFI+ FSK+A PLT+L CK F +D C +F+ +K+ L + P
Sbjct: 280 TVKDIRSFLGHAEFYRRFIKDFSKIARPLTRLLCKETEFNFDEDCLKAFHLIKETLVSAP 339
Query: 278 ILILPKPEEPFVVYFDASKLGLGVVLMQ--DGK--VVAYASRLLRIHEKNYPTHDLELAA 333
I P + PF + DA +G VL Q D K V+ YASR + + Y T + EL A
Sbjct: 340 IFQAPNWDHPFEIMCDAFDYAVGAVLDQKIDDKLHVIYYASRTMDEAQTRYATIEKELLA 399
Query: 334 VVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQKELNMRQRRWLELLKDYDFGLNYHPGKA 393
VV + R YL G + +V+ DH +L++++ +KE R RW+ LL+++D + G
Sbjct: 400 VVFAFEKIRSYLVGFKVKVYIDHAALRHIYAKKETKSRLLRWILLLQEFDMEIIDKKGVE 459
Query: 394 NVVADALSRKTLHMS 408
N VAD LSR + S
Sbjct: 460 NGVADHLSRMRIEDS 474
>At1g20390 hypothetical protein
Length = 1791
Score = 196 bits (499), Expect = 4e-50
Identities = 122/399 (30%), Positives = 197/399 (48%), Gaps = 7/399 (1%)
Query: 8 LKKQLEDLLDK-KFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRID 66
+ +++E LL + + W A ++VKKK+G R+C+DY LNK K+ YPLP ID
Sbjct: 793 VNEEVEKLLKAGQIIEVKYPEWLANPVVVKKKNGKWRVCVDYTDLNKACPKDSYPLPHID 852
Query: 67 DLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFM 126
L++ G + S +D SGY+QI + +D +KT+ T G Y YKVM FG+ NA +
Sbjct: 853 RLVEATSGNGLLSFMDAFSGYNQILMHKDDQEKTSFVTDRGTYCYKVMSFGLKNAGATYQ 912
Query: 127 EYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLS 186
++N++ + + V V+IDD+L+ S E+H HL VL + +KC F ++
Sbjct: 913 RFVNKMLADQIGRTVEVYIDDMLVKSLKPEDHVEHLSKCFDVLNTYGMKLNPTKCTFGVT 972
Query: 187 EVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPL 246
F G++++ GI +P ++ A+ + +P++ E++ G RFI + LP
Sbjct: 973 SGEFLGYVVTKRGIEANPKQIRAILELPSPRNAREVQRLTGRIAALNRFISRSTDKCLPF 1032
Query: 247 TQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQD 306
L + F WD E +F +LK L T PIL+ P+ E +Y S + VL+++
Sbjct: 1033 YNLLKRRAQFDWDKDSEEAFEKLKDYLSTPPILVKPEVGETLYLYIAVSDHAVSSVLVRE 1092
Query: 307 G----KVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYL 362
+ + Y S+ L E YP + AVV + R Y V +D + L+
Sbjct: 1093 DRGEQRPIFYTSKSLVEAETRYPVIEKAALAVVTSARKLRPYFQSHTIAVLTD-QPLRVA 1151
Query: 363 FDQKELNMRQRRWLELLKDYDFGLNYHPG-KANVVADAL 400
+ R +W L +YD P K+ V+AD L
Sbjct: 1152 LHSPSQSGRMTKWAVELSEYDIDFRPRPAMKSQVLADFL 1190
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.341 0.151 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,329,342
Number of Sequences: 26719
Number of extensions: 802671
Number of successful extensions: 2698
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 50
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 2566
Number of HSP's gapped (non-prelim): 94
length of query: 924
length of database: 11,318,596
effective HSP length: 108
effective length of query: 816
effective length of database: 8,432,944
effective search space: 6881282304
effective search space used: 6881282304
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 65 (29.6 bits)
Medicago: description of AC135463.3


