

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135463.1 - phase: 0 /pseudo
(488 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
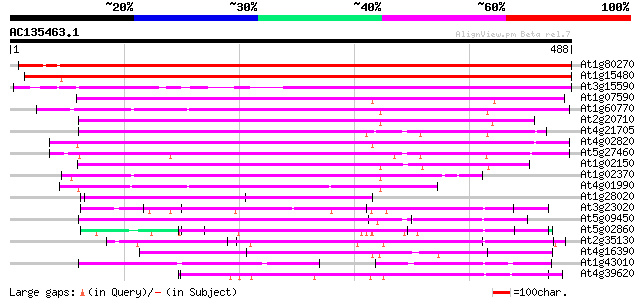
Score E
Sequences producing significant alignments: (bits) Value
At1g80270 unknown protein (At1g80270) 561 e-160
At1g15480 hypothetical protein 550 e-157
At3g15590 DNA-binding protein, putative 343 9e-95
At1g07590 DNA-binding protein like protein 179 2e-45
At1g60770 unknown protein (At1g60770) 170 1e-42
At2g20710 unknown protein 166 3e-41
At4g21705 unknown protein 161 7e-40
At4g02820 Unknown protein (At4g02820) 158 6e-39
At5g27460 putative protein 149 3e-36
At1g02150 hypothetical protein 139 3e-33
At1g02370 unknown protein 124 9e-29
At4g01990 unknown protein 124 2e-28
At1g28020 hypothetical protein 108 7e-24
At3g23020 hypothetical protein 103 3e-22
At5g09450 unknown protein 102 5e-22
At5g02860 putative protein 97 2e-20
At2g35130 hypothetical protein 94 2e-19
At4g16390 salt-inducible protein homolog 84 2e-16
At1g43010 hypothetical protein 83 3e-16
At4g39620 putative protein 82 7e-16
>At1g80270 unknown protein (At1g80270)
Length = 596
Score = 561 bits (1445), Expect = e-160
Identities = 280/482 (58%), Positives = 367/482 (76%), Gaps = 5/482 (1%)
Query: 8 TEPDADGDSRDTAEIDINELELSDTETDSSDKKSFSSRRRSELFKAIVSVSGLSVDSALD 67
T D D E + EL+L ETD S K +++SELFK IVS GLS+ SALD
Sbjct: 94 TSSDEDEGKLSADEEEEEELDL--IETDVSRKTV--EKKQSELFKTIVSAPGLSIGSALD 149
Query: 68 KWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASKLDLIAKL 127
KWVE+G E++R EI A+ LRRR+MYGRALQ +WLE+NKK+E TE++YAS+LDL K+
Sbjct: 150 KWVEEGNEITRVEIAKAMLQLRRRRMYGRALQMSEWLEANKKIEMTERDYASRLDLTVKI 209
Query: 128 RGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQ 187
RGL K E ++ +P SF+GE+LYRTLLANC + N++K+E FNKM++LGFP++ F C+Q
Sbjct: 210 RGLEKGEACMQKIPKSFKGEVLYRTLLANCVAAGNVKKSELVFNKMKDLGFPLSGFTCDQ 269
Query: 188 LLLIYKKIDKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGC 247
+LL++K+ID+KKIADVLL+MEKEN+KPS TYKILIDVKG +NDI GM QI+ETMK EG
Sbjct: 270 MLLLHKRIDRKKIADVLLLMEKENIKPSLLTYKILIDVKGATNDISGMEQILETMKDEGV 329
Query: 248 ELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVER 307
ELD T+A ARHY+ AGL +K E +LKE+EGE+L+ N LL +YA LGR DEV+R
Sbjct: 330 ELDFQTQALTARHYSGAGLKDKAEKVLKEMEGESLEANRRAFKDLLSIYASLGREDEVKR 389
Query: 308 IWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFE-MMSNKWKLTARNYESLLKIYIRHK 366
IWK+CESKP E+ LAAI+A+G+L K++EAEA+FE ++ + ++ Y LL++Y+ HK
Sbjct: 390 IWKICESKPYFEESLAAIQAFGKLNKVQEAEAIFEKIVKMDRRASSSTYSVLLRVYVDHK 449
Query: 367 MLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTF 426
ML+KGKDL+K M +SGC I TTWDAL+ LYV+AGEVEKAD++L KA +Q+ K M +F
Sbjct: 450 MLSKGKDLVKRMAESGCRIEATTWDALIKLYVEAGEVEKADSLLDKASKQSHTKLMMNSF 509
Query: 427 MTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADN 486
M IM++Y+KRGDVHN EKIF ++R+A Y SR+ F AL QAY NAK PAYG+R+R+KADN
Sbjct: 510 MYIMDEYSKRGDVHNTEKIFLKMREAGYTSRLRQFQALMQAYINAKSPAYGMRDRLKADN 569
Query: 487 LF 488
+F
Sbjct: 570 IF 571
>At1g15480 hypothetical protein
Length = 623
Score = 550 bits (1417), Expect = e-157
Identities = 269/493 (54%), Positives = 364/493 (73%), Gaps = 18/493 (3%)
Query: 14 GDSRDTAEIDINELELSDTETDSSDKKSFS-----------------SRRRSELFKAIVS 56
GD D + +++ +T +DS D + FS S+R SE+FKAIVS
Sbjct: 106 GDDDDLEDKNVDLATPDETSSDSEDGEEFSGDEGDIEGAELELHVPESKRPSEMFKAIVS 165
Query: 57 VSGLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKE 116
VSGLSV SALDKWVE+GK+ +R+E A+ LR+R+M+GRALQ +WL+ NK+ E E++
Sbjct: 166 VSGLSVGSALDKWVEQGKDTNRKEFESAMLQLRKRRMFGRALQMTEWLDENKQFEMEERD 225
Query: 117 YASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMREL 176
YA +LDLI+K+RG K E Y++ +P SFRGEL+YRTLLAN + N+R E FNKM++L
Sbjct: 226 YACRLDLISKVRGWYKGEAYIKTIPESFRGELVYRTLLANHVATSNVRTAEAVFNKMKDL 285
Query: 177 GFPVTAFACNQLLLIYKKIDKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMS 236
GFP++ F CNQ+L++YK++DKKKIADVLL++EKEN+KP+ TYKILID KG SNDI GM
Sbjct: 286 GFPLSTFTCNQMLILYKRVDKKKIADVLLLLEKENLKPNLNTYKILIDTKGSSNDITGME 345
Query: 237 QIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLY 296
QIVETMK+EG ELD RA +ARHYA+AGL EK E +LKE+EGE+L+EN +C LL +Y
Sbjct: 346 QIVETMKSEGVELDLRARALIARHYASAGLKEKAEKVLKEMEGESLEENRHMCKDLLSVY 405
Query: 297 AILGRADEVERIWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFE-MMSNKWKLTARNY 355
L R DEV R+WK+CE PR + LAAI A+G++ K+++AEAVFE ++ ++++ Y
Sbjct: 406 GYLQREDEVRRVWKICEENPRYNEVLAAILAFGKIDKVKDAEAVFEKVLKMSHRVSSNVY 465
Query: 356 ESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQ 415
LL++Y+ HKM+++GKDL+K M DSGC IG TWDA++ LYV+AGEVEKA++ L KA+Q
Sbjct: 466 SVLLRVYVDHKMVSEGKDLVKQMSDSGCNIGALTWDAVIKLYVEAGEVEKAESSLSKAIQ 525
Query: 416 QNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPA 475
++KP+ ++FM +M +Y +RGDVHN EKIF R++QA Y SR + L QAY NAK PA
Sbjct: 526 SKQIKPLMSSFMYLMHEYVRRGDVHNTEKIFQRMKQAGYQSRFWAYQTLIQAYVNAKAPA 585
Query: 476 YGIRERMKADNLF 488
YG++ERMKADN+F
Sbjct: 586 YGMKERMKADNIF 598
>At3g15590 DNA-binding protein, putative
Length = 526
Score = 343 bits (881), Expect = 9e-95
Identities = 195/486 (40%), Positives = 292/486 (59%), Gaps = 73/486 (15%)
Query: 4 EDRLTEPDADGDSRDTAEIDINELELSDTETDSSDKKSFSSRRRSELFKAIVSVSGLSVD 63
+D L EP+ D+ D LE+ + + K + R +SEL+++IV+ SV
Sbjct: 87 DDSLFEPELGSDNDD--------LEIEEKHSKDGGKPT-KKRGQSELYESIVAYK--SVK 135
Query: 64 SALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASKLDL 123
L+KWV++GK+LS+ E+ LA+++LR+RK Y LQ +WL +N + EFTE YAS+LDL
Sbjct: 136 HVLEKWVKEGKDLSQAEVTLAIHNLRKRKSYAMCLQLWEWLGANTQFEFTEANYASQLDL 195
Query: 124 IAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGFPVTAF 183
+AKL L Y H LL EN++ + T++ +
Sbjct: 196 VAKLLLL-----YSMHDRKKISDVLL-------LMERENIKPSRATYHFL---------- 233
Query: 184 ACNQLLLIYKKIDKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMK 243
I+ K +A + MEK IVET+K
Sbjct: 234 -----------INSKGLAGDITGMEK----------------------------IVETIK 254
Query: 244 AEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRAD 303
EG ELD ++ LA++Y AGL E+ + ++KEIEG+ L++ WVC +LL LYA +G +D
Sbjct: 255 EEGIELDPELQSILAKYYIRAGLKERAQDLMKEIEGKGLQQTPWVCRSLLPLYADIGDSD 314
Query: 304 EVERIWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMSNKWKL-TARNYESLLKIY 362
V R+ + + PR ++C++AI+AWG+LK++EEAEAVFE + K+K+ Y +L++IY
Sbjct: 315 NVRRLSRFVDQNPRYDNCISAIKAWGKLKEVEEAEAVFERLVEKYKIFPMMPYFALMEIY 374
Query: 363 IRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPM 422
+KML KG+DL+K MG++G IGP+TW ALV LY++AGEV KA+ +L +A + NKM+PM
Sbjct: 375 TENKMLAKGRDLVKRMGNAGIAIGPSTWHALVKLYIKAGEVGKAELILNRATKDNKMRPM 434
Query: 423 FTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERM 482
FTT+M I+E+YAKRGDVHN EK+F ++++A+Y +++ + + AY NAK PAYG+ ERM
Sbjct: 435 FTTYMAILEEYAKRGDVHNTEKVFMKMKRASYAAQLMQYETVLLAYINAKTPAYGMIERM 494
Query: 483 KADNLF 488
KADN+F
Sbjct: 495 KADNVF 500
>At1g07590 DNA-binding protein like protein
Length = 534
Score = 179 bits (455), Expect = 2e-45
Identities = 126/434 (29%), Positives = 212/434 (48%), Gaps = 10/434 (2%)
Query: 59 GLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYA 118
G++V SAL W+ G + ++ A+N LR+ RAL+ ++W+ + E EY+
Sbjct: 78 GVTVGSALQSWMGDGFPVHGGDVYHAINRLRKLGRNKRALELMEWIIRERPYRLGELEYS 137
Query: 119 SKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGF 178
L+ KL G+ + EK VP F+ ELLY L+ C +R E KMRELG+
Sbjct: 138 YLLEFTVKLHGVSQGEKLFTRVPQEFQNELLYNNLVIACLDQGVIRLALEYMKKMRELGY 197
Query: 179 PVTAFACNQLLLIYKKIDKKK-IADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQ 237
+ N+L++ ++K IA L +M+ + P TY IL+ ++ ++IDG+ +
Sbjct: 198 RTSHLVYNRLIIRNSAPGRRKLIAKDLALMKADKATPHVSTYHILMKLEANEHNIDGVLK 257
Query: 238 IVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYA 297
+ MK G E + ++ LA +A A L EA +EIE +N L+ LY
Sbjct: 258 AFDGMKKAGVEPNEVSYCILAMAHAVARLYTVAEAYTEEIEKSITGDNWSTLDILMILYG 317
Query: 298 ILGRADEVERIWKVCES--KPRVEDCLAAIEAWGRLKKIEEAEAVF-EMMSNKWKLTARN 354
LG+ E+ R W V R + L A EA+ R+ ++ AE ++ EM + K
Sbjct: 318 RLGKEKELARTWNVIRGFHHVRSKSYLLATEAFARVGNLDRAEELWLEMKNVKGLKETEQ 377
Query: 355 YESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKAL 414
+ SLL +Y + ++ K + + M +G T+ L +A +++A ++ L
Sbjct: 378 FNSLLSVYCKDGLIEKAIGVFREMTGNGFKPNSITYRHLALGCAKAKLMKEALKNIEMGL 437
Query: 415 QQNKMK------PMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAY 468
K P T ++I+E +A++GDV N+EK+F ++ A Y ++AL +AY
Sbjct: 438 NLKTSKSIGSSTPWLETTLSIIECFAEKGDVENSEKLFEEVKNAKYNRYAFVYNALFKAY 497
Query: 469 KNAKLPAYGIRERM 482
AK+ + +RM
Sbjct: 498 VKAKVYDPNLFKRM 511
>At1g60770 unknown protein (At1g60770)
Length = 491
Score = 170 bits (431), Expect = 1e-42
Identities = 131/476 (27%), Positives = 227/476 (47%), Gaps = 16/476 (3%)
Query: 24 INELELSDTETDSSDKKSFSSRRRSELFKAIVSVSGLSVDSALDKWVEKGKELSRQEIGL 83
+ L S T S KK + LFK + + V L+++++ K + + E+G
Sbjct: 3 MRHLSRSRDVTKRSTKKYIEEPLYNRLFKD--GGTEVKVRQQLNQFLKGTKHVFKWEVGD 60
Query: 84 ALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNS 143
+ LR R +Y AL+ + +E + + T + A LDL+AK R + E Y +P +
Sbjct: 61 TIKKLRNRGLYYPALKLSEVMEE-RGMNKTVSDQAIHLDLVAKAREITAGENYFVDLPET 119
Query: 144 FRGELLYRTLLANCASLENL-RKTEETFNKMRELGFPVTAFACNQLLLIYKKI-DKKKIA 201
+ EL Y +LL NC E L K E NKM+EL ++ + N L+ +Y K + +K+
Sbjct: 120 SKTELTYGSLL-NCYCKELLTEKAEGLLNKMKELNITPSSMSYNSLMTLYTKTGETEKVP 178
Query: 202 DVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEG-CELDHLTRASLARH 260
++ ++ ENV P SYTY + + +NDI G+ +++E M +G D T +++A
Sbjct: 179 AMIQELKAENVMPDSYTYNVWMRALAATNDISGVERVIEEMNRDGRVAPDWTTYSNMASI 238
Query: 261 YAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCE-SKPRVE 319
Y AGL++K E L+E+E +N + + L+ LY LG+ EV RIW+ + P+
Sbjct: 239 YVDAGLSQKAEKALQELEMKNTQRDFTAYQFLITLYGRLGKLTEVYRIWRSLRLAIPKTS 298
Query: 320 DC--LAAIEAWGRLKKIEEAEAVF-EMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIK 376
+ L I+ +L + AE +F E +N R L+ Y + ++ K +L +
Sbjct: 299 NVAYLNMIQVLVKLNDLPGAETLFKEWQANCSTYDIRIVNVLIGAYAQEGLIQKANELKE 358
Query: 377 TMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKAL-----QQNKMKPMFTTFMTIME 431
G + TW+ + YV++G++ +A + KA+ K P T +M
Sbjct: 359 KAPRRGGKLNAKTWEIFMDYYVKSGDMARALECMSKAVSIGKGDGGKWLPSPETVRALMS 418
Query: 432 QYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNL 487
+ ++ DV+ AE + L+ F L + Y A +R R+K +N+
Sbjct: 419 YFEQKKDVNGAENLLEILKNGTDNIGAEIFEPLIRTYAAAGKSHPAMRRRLKMENV 474
>At2g20710 unknown protein
Length = 490
Score = 166 bits (420), Expect = 3e-41
Identities = 117/404 (28%), Positives = 196/404 (47%), Gaps = 8/404 (1%)
Query: 61 SVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASK 120
S+ LD W+++G + E+ + LR+ + ALQ DW+ ++ E +E + A +
Sbjct: 53 SIIKVLDGWLDQGNLVKTSELHSIIKMLRKFSRFSHALQISDWMSEHRVHEISEGDVAIR 112
Query: 121 LDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGFPV 180
LDLIAK+ GL +AEK+ E +P R LY LL AS + L K E+ F +M+ELGF
Sbjct: 113 LDLIAKVGGLGEAEKFFETIPMERRNYHLYGALLNCYASKKVLHKAEQVFQEMKELGFLK 172
Query: 181 TAFACNQLLLIYKKIDKKKIADVLLM-MEKENVKPSSYTYKILIDVKGLSNDIDGMSQIV 239
N +L +Y + K + + LL ME E VKP +T + + +D++GM + +
Sbjct: 173 GCLPYNVMLNLYVRTGKYTMVEKLLREMEDETVKPDIFTVNTRLHAYSVVSDVEGMEKFL 232
Query: 240 ETMKA-EGCELDHLTRASLARHYAAAGLTEKT-EAILKEIEGENLKENMWVCPTLLRLYA 297
+A +G LD T A A Y AGLTEK E + K + N ++ L+ Y
Sbjct: 233 MRCEADQGLHLDWRTYADTANGYIKAGLTEKALEMLRKSEQMVNAQKRKHAYEVLMSFYG 292
Query: 298 ILGRADEVERIWKVCESKPRVEDC--LAAIEAWGRLKKIEEAEAVFEMMSNKWKL-TARN 354
G+ +EV R+W + + + ++ I A ++ IEE E + E L R
Sbjct: 293 AAGKKEEVYRLWSLYKELDGFYNTGYISVISALLKMDDIEEVEKIMEEWEAGHSLFDIRI 352
Query: 355 YESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKAL 414
L+ Y + M+ K ++++ + +TW+ L Y AG++EKA ++A+
Sbjct: 353 PHLLITGYCKKGMMEKAEEVVNILVQKWRVEDTSTWERLALGYKMAGKMEKAVEKWKRAI 412
Query: 415 QQNK--MKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYIS 456
+ +K +P M+ ++ + D+ KI L + +IS
Sbjct: 413 EVSKPGWRPHQVVLMSCVDYLEGQRDMEGLRKILRLLSERGHIS 456
>At4g21705 unknown protein
Length = 492
Score = 161 bits (408), Expect = 7e-40
Identities = 113/422 (26%), Positives = 206/422 (48%), Gaps = 21/422 (4%)
Query: 61 SVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASK 120
SV L WV+ GK++S E+ ++ LRRRK + AL+ W+ F+ E+A
Sbjct: 40 SVYPELQNWVQCGKKVSVAELIRIVHDLRRRKRFLHALEVSKWMNETGVCVFSPTEHAVH 99
Query: 121 LDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGFPV 180
LDLI ++ G AE+Y E++ ++ + Y LL +N+ K+ F KM+E+GF
Sbjct: 100 LDLIGRVYGFVTAEEYFENLKEQYKNDKTYGALLNCYVRQQNVEKSLLHFEKMKEMGFVT 159
Query: 181 TAFACNQLLLIYKKIDK-KKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIV 239
++ N ++ +Y I + +K+ VL M++ENV P +Y+Y+I I+ G D++ + +
Sbjct: 160 SSLTYNNIMCLYTNIGQHEKVPKVLEEMKEENVAPDNYSYRICINAFGAMYDLERIGGTL 219
Query: 240 ETM-KAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAI 298
M + + +D T A A+ Y G ++ +LK E K++ L+ LYA
Sbjct: 220 RDMERRQDITMDWNTYAVAAKFYIDGGDCDRAVELLKMSENRLEKKDGEGYNHLITLYAR 279
Query: 299 LGRADEVERIW----KVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMSNKWKLTARN 354
LG+ EV R+W VC+ + +D L +++ ++ + EAE V +WK +
Sbjct: 280 LGKKIEVLRLWDLEKDVCKRRIN-QDYLTVLQSLVKIDALVEAEEVL----TEWKSSGNC 334
Query: 355 YE-----SLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTV 409
Y+ ++++ YI M K + +++ + G P +W+ + + Y + G +E A
Sbjct: 335 YDFRVPNTVIRGYIGKSMEEKAEAMLEDLARRGKATTPESWELVATAYAEKGTLENAFKC 394
Query: 410 LQKAL----QQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALA 465
++ AL K +P T +++ G + E LR +++ +HAL
Sbjct: 395 MKTALGVEVGSRKWRPGLTLVTSVLSWVGDEGSLKEVESFVASLRNCIGVNK-QMYHALV 453
Query: 466 QA 467
+A
Sbjct: 454 KA 455
>At4g02820 Unknown protein (At4g02820)
Length = 532
Score = 158 bits (400), Expect = 6e-39
Identities = 124/460 (26%), Positives = 213/460 (45%), Gaps = 8/460 (1%)
Query: 35 DSSDKKSFSSRRRSELFKAIVSV--SGLSVDSALDKWVEKGKELSRQEIGLALNSLRRRK 92
+S++KK R L ++S+ + S + KW E+G + + E+ + LR+ K
Sbjct: 48 ESANKKETVVGGRDTLGGRLLSLVYTKRSAVVTIRKWKEEGHSVRKYELNRIVRELRKIK 107
Query: 93 MYGRALQALDWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRT 152
Y AL+ +W+ + ++ +YA LDLI+K+RGL AEK+ E +P+ RG +
Sbjct: 108 RYKHALEICEWMVVQEDIKLQAGDYAVHLDLISKIRGLNSAEKFFEDMPDQMRGHAACTS 167
Query: 153 LLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKKIADVLLMMEKENV 212
LL + + K E F KM E GF + N +L +Y + + VL+ K
Sbjct: 168 LLHSYVQNKLSDKAEALFEKMGECGFLKSCLPYNHMLSMYISRGQFEKVPVLIKELKIRT 227
Query: 213 KPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEA 272
P TY + + ND++G ++ K E D +T + L YA EK
Sbjct: 228 SPDIVTYNLWLTAFASGNDVEGAEKVYLKAKEEKLNPDWVTYSVLTNLYAKTDNVEKARL 287
Query: 273 ILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCES---KPRVEDCLAAIEAWG 329
LKE+E K+N +L+ L+A LG D V WK +S K + L+ I A
Sbjct: 288 ALKEMEKLVSKKNRVAYASLISLHANLGDKDGVNLTWKKVKSSFKKMNDAEYLSMISAVV 347
Query: 330 RLKKIEEAEAVF-EMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPT 388
+L + E+A+ ++ E S AR +L Y+ + G+ + + + G +
Sbjct: 348 KLGEFEQAKGLYDEWESVSGTGDARIPNLILAEYMNRDEVLLGEKFYERIVEKGINPSYS 407
Query: 389 TWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMT-IMEQYAKRGDVHNAEKIFY 447
TW+ L Y++ ++EK KA+ K + + ++ ++G+V AEK+
Sbjct: 408 TWEILTWAYLKRKDMEKVLDCFGKAIDSVKKWTVNVRLVKGACKELEEQGNVKGAEKLMT 467
Query: 448 RLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNL 487
L++A Y++ +++L + Y A A + ERM DN+
Sbjct: 468 LLQKAGYVN-TQLYNSLLRTYAKAGEMALIVEERMAKDNV 506
>At5g27460 putative protein
Length = 491
Score = 149 bits (376), Expect = 3e-36
Identities = 127/472 (26%), Positives = 221/472 (45%), Gaps = 27/472 (5%)
Query: 35 DSSDKKSFSSRRRSELFKAIVSVSG--LSVDSALDKWVEKGKELSRQEIGLALNSLRRRK 92
D SD S ++R K I+ +G SV S L + ++ G +S E+ L L R
Sbjct: 28 DGSDTSSVANRNS---LKEILRKNGPRRSVTSLLQERIDSGHAVSLSELRLISKRLIRSN 84
Query: 93 MYGRALQALDWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLE---HVPNSFR-GEL 148
Y ALQ ++W+E+ K +EF+ + A +LDLI K GL + E+Y E H S R +
Sbjct: 85 RYDLALQMMEWMENQKDIEFSVYDIALRLDLIIKTHGLKQGEEYFEKLLHSSVSMRVAKS 144
Query: 149 LYRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDK-KKIADVLLMM 207
Y LL + +++ E K+ LGF VT N+++ +Y+ + +K+ V+ MM
Sbjct: 145 AYLPLLRAYVKNKMVKEAEALMEKLNGLGFLVTPHPFNEMMKLYEASGQYEKVVMVVSMM 204
Query: 208 EKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAE-GCELDHLTRASLARHYAAAGL 266
+ + + +Y + ++ + + + + + M + E+ + +LA Y +G
Sbjct: 205 KGNKIPRNVLSYNLWMNACCEVSGVAAVETVYKEMVGDKSVEVGWSSLCTLANVYIKSGF 264
Query: 267 TEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCESKPRVEDCLAAIE 326
EK +L++ E + N L+ LYA LG + V R+W+V +S C+ I
Sbjct: 265 DEKARLVLEDAEKMLNRSNRLGYFFLITLYASLGNKEGVVRLWEVSKSVCGRISCVNYIC 324
Query: 327 AWGRLKK---IEEAEAVFEMMSNKWKLTARNYE-----SLLKIYIRHKMLNKGKDLIKTM 378
L K +EEAE VF ++W+ NY+ LL Y+R+ + K + L +
Sbjct: 325 VLSSLVKTGDLEEAERVF----SEWEAQCFNYDVRVSNVLLGAYVRNGEIRKAESLHGCV 380
Query: 379 GDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKA---LQQNKMKPMFTTFMTIMEQYAK 435
+ G T TW+ L+ +V+ +EKA + + +++ +P M I E + K
Sbjct: 381 LERGGTPNYKTWEILMEGWVKCENMEKAIDAMHQVFVLMRRCHWRPSHNIVMAIAEYFEK 440
Query: 436 RGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNL 487
+ A L + ++ + + L + +++AK PAY I E MK D L
Sbjct: 441 EEKIEEATAYVRDLHRLG-LASLPLYRLLLRMHEHAKRPAYDIYEMMKLDKL 491
>At1g02150 hypothetical protein
Length = 524
Score = 139 bits (351), Expect = 3e-33
Identities = 100/407 (24%), Positives = 194/407 (47%), Gaps = 18/407 (4%)
Query: 60 LSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESN-KKLEFTEKEYA 118
L S L++W + G++L++ E+ + LR+ K +AL+ DW+ + ++ + + A
Sbjct: 81 LGAASVLNQWEKAGRKLTKWELCRVVKELRKYKRANQALEVYDWMNNRGERFRLSASDAA 140
Query: 119 SKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMRELGF 178
+LDLI K+RG+P AE++ +P +F+ +Y +LL ++ K E N MR+ G+
Sbjct: 141 IQLDLIGKVRGIPDAEEFFLQLPENFKDRRVYGSLLNAYVRAKSREKAEALLNTMRDKGY 200
Query: 179 PVTAFACNQLLLIYKKI-DKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQ 237
+ N ++ +Y + + K+ ++ M++++++ Y+Y I + G ++ M
Sbjct: 201 ALHPLPFNVMMTLYMNLREYDKVDAMVFEMKQKDIRLDIYSYNIWLSSCGSLGSVEKMEL 260
Query: 238 IVETMKAE-GCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLY 296
+ + MK++ + T +++A Y G TEK E L+++E N LL LY
Sbjct: 261 VYQQMKSDVSIYPNWTTFSTMATMYIKMGETEKAEDALRKVEARITGRNRIPYHYLLSLY 320
Query: 297 AILGRADEVERIWKVCES-KPRVEDC--LAAIEAWGRLKKIEEAEAVFEMMSNKWKLTAR 353
LG E+ R+W V +S P + + A + + R+ IE AE V+E +W
Sbjct: 321 GSLGNKKELYRVWHVYKSVVPSIPNLGYHALVSSLVRMGDIEGAEKVYE----EWLPVKS 376
Query: 354 NYES-----LLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADT 408
+Y+ L+ Y+++ L + L M + G +TW+ L + + + +A T
Sbjct: 377 SYDPRIPNLLMNAYVKNDQLETAEGLFDHMVEMGGKPSSSTWEILAVGHTRKRCISEALT 436
Query: 409 VLQKALQ---QNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQA 452
L+ A + +P + + DV + E + LRQ+
Sbjct: 437 CLRNAFSAEGSSNWRPKVLMLSGFFKLCEEESDVTSKEAVLELLRQS 483
>At1g02370 unknown protein
Length = 537
Score = 124 bits (312), Expect = 9e-29
Identities = 99/375 (26%), Positives = 186/375 (49%), Gaps = 13/375 (3%)
Query: 46 RRSELFK--AIVSVSGLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDW 103
R+ EL+K +++SV+G +V L++++ +G + + ++ +LR+ + A + DW
Sbjct: 69 RQRELYKKLSMLSVTGGTVAETLNQFIMEGITVRKDDLFRCAKTLRKFRRPQHAFEIFDW 128
Query: 104 LESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLE-N 162
+E +K+ F+ ++A LDLI K +GL AE Y ++ S + L NC +E
Sbjct: 129 MEK-RKMTFSVSDHAICLDLIGKTKGLEAAENYFNNLDPSAKNHQSTYGALMNCYCVELE 187
Query: 163 LRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKKIADVLL-MMEKENVKPSSYTYKI 221
K + F M EL F + N ++ +Y ++ + + VL+ M++ + P TY I
Sbjct: 188 EEKAKAHFEIMDELNFVNNSLPFNNMMSMYMRLSQPEKVPVLVDAMKQRGISPCGVTYSI 247
Query: 222 LIDVKGLSNDIDGMSQIVETM-KAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGE 280
+ G ND+DG+ +I++ M K + T ++LA Y AGL EK ++ LK +E +
Sbjct: 248 WMQSCGSLNDLDGLEKIIDEMGKDSEAKTTWNTFSNLAAIYTKAGLYEKADSALKSMEEK 307
Query: 281 NLKENMWVCPTLLRLYAILGRADEVERIWK-VCESKPRVEDC--LAAIEAWGRLKKIEEA 337
N L+ LYA + + EV R+W+ + +++P V + L ++A +L ++
Sbjct: 308 MNPNNRDSHHFLMSLYAGISKGPEVYRVWESLKKARPEVNNLSYLVMLQAMSKLGDLDGI 367
Query: 338 EAVF-EMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSL 396
+ +F E S W R + Y++ M + + ++ G + GP + A L
Sbjct: 368 KKIFTEWESKCWAYDMRLANIAINTYLKGNMYEEAEKILD--GAMKKSKGPFS-KARQLL 424
Query: 397 YVQAGEVEKADTVLQ 411
+ E +KAD ++
Sbjct: 425 MIHLLENDKADLAMK 439
Score = 29.6 bits (65), Expect = 3.9
Identities = 23/135 (17%), Positives = 54/135 (39%), Gaps = 1/135 (0%)
Query: 335 EEAEAVFEMMSN-KWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDAL 393
E+A+A FE+M + + + +++ +Y+R K L+ M G + T+
Sbjct: 189 EKAKAHFEIMDELNFVNNSLPFNNMMSMYMRLSQPEKVPVLVDAMKQRGISPCGVTYSIW 248
Query: 394 VSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQAN 453
+ +++ + ++ + + ++ K + TF + Y K G A+ + +
Sbjct: 249 MQSCGSLNDLDGLEKIIDEMGKDSEAKTTWNTFSNLAAIYTKAGLYEKADSALKSMEEKM 308
Query: 454 YISRISPFHALAQAY 468
+ H L Y
Sbjct: 309 NPNNRDSHHFLMSLY 323
>At4g01990 unknown protein
Length = 502
Score = 124 bits (310), Expect = 2e-28
Identities = 92/340 (27%), Positives = 173/340 (50%), Gaps = 14/340 (4%)
Query: 44 SRRRSELFKAIVSVS---GLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQA 100
+++ ++K + S+ G ++ L+++V +G + + ++ LR+ + RAL+
Sbjct: 35 AKKHRSIYKKLSSLGTRGGGKMEETLNQFVMEGVPVKKHDLIRYAKDLRKFRQPQRALEI 94
Query: 101 LDWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASL 160
+W+E K++ FT ++A +L+LIAK +GL AE Y + +S + + Y +LL NC +
Sbjct: 95 FEWME-RKEIAFTGSDHAIRLNLIAKSKGLEAAETYFNSLDDSIKNQSTYGSLL-NCYCV 152
Query: 161 ENLR-KTEETFNKMRELGFPVTAFACNQLLLIYKKIDK-KKIADVLLMMEKENVKPSSYT 218
E K + F M +L + N L+ +Y + + +K+ +++ M+++++ P T
Sbjct: 153 EKEEVKAKAHFENMVDLNHVSNSLPFNNLMAMYMGLGQPEKVPALVVAMKEKSITPCDIT 212
Query: 219 YKILIDVKGLSNDIDGMSQIVETMKAEGCEL-DHLTRASLARHYAAAGLTEKTEAILKEI 277
Y + I G D+DG+ ++++ MKAEG + T A+LA Y GL K E LK +
Sbjct: 213 YSMWIQSCGSLKDLDGVEKVLDEMKAEGEGIFSWNTFANLAAIYIKVGLYGKAEEALKSL 272
Query: 278 EGENLKENMWVC-PTLLRLYAILGRADEVERIWKVCESK-PRVEDC--LAAIEAWGRLKK 333
E N+ ++ C L+ LY + A EV R+W + + + P V + L + A +L
Sbjct: 273 E-NNMNPDVRDCYHFLINLYTGIANASEVYRVWDLLKKRYPNVNNSSYLTMLRALSKLDD 331
Query: 334 IEEAEAVF-EMMSNKWKLTARNYESLLKIYIRHKMLNKGK 372
I+ + VF E S W R + Y++ M + +
Sbjct: 332 IDGVKKVFAEWESTCWTYDMRMANVAISSYLKQNMYEEAE 371
>At1g28020 hypothetical protein
Length = 612
Score = 108 bits (270), Expect = 7e-24
Identities = 69/254 (27%), Positives = 134/254 (52%), Gaps = 4/254 (1%)
Query: 66 LDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASKLDLIA 125
L++W ++G +++ + + + LR +ALQ +W+ K +++A++L LI
Sbjct: 56 LEQWRQQGNQVNPSHVRVIIKKLRDSDQSLQALQVSEWMSKEKICNLIPEDFAARLHLIE 115
Query: 126 KLRGLPKAEKYLEHVPNSFRGELLYRTLLANCA-SLENLRKTEETFNKMRELGFPVTAFA 184
+ GL +AEK+ E +P + RG+ +Y +LL + A S + L K E TF KMR+LG +
Sbjct: 116 NVVGLEEAEKFFESIPKNARGDSVYTSLLNSYARSDKTLCKAEATFQKMRDLGLLLRPVP 175
Query: 185 CNQLLLIYKKI-DKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMK 243
N ++ +Y + +++K+ ++LL M+ +V+ + T ++ + D+ M + + +
Sbjct: 176 YNAMMSLYSALKNREKVEELLLEMKDNDVEADNVTVNNVLKLYSAVCDVTEMEKFLNKWE 235
Query: 244 A-EGCELDHLTRASLARHYAAAGLTEKTEAILKEIEG-ENLKENMWVCPTLLRLYAILGR 301
G +L+ T +A+ Y A + K +L+ E + K L++LY G
Sbjct: 236 GIHGIKLEWHTTLDMAKAYLRARSSGKAMKMLRLTEQLVDQKSLKSAYDHLMKLYGEAGN 295
Query: 302 ADEVERIWKVCESK 315
+EV R+WK+ +SK
Sbjct: 296 REEVLRVWKLYKSK 309
Score = 59.3 bits (142), Expect = 5e-09
Identities = 37/146 (25%), Positives = 72/146 (48%), Gaps = 5/146 (3%)
Query: 62 VDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASKL 121
V L++W G ++ ++ + +LR K + +ALQ +W+ + ++YA++L
Sbjct: 387 VTPLLEQW---GDQMKPSDLKCLIKNLRDSKQFSKALQVSEWMGEKQVCNLYLEDYAARL 443
Query: 122 DLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCA-SLENL-RKTEETFNKMRELGFP 179
L + GL +AEKY E++P + + +Y LL++ A S +NL +E +M E
Sbjct: 444 YLTENVLGLEEAEKYFENIPENMKDYSVYVALLSSYAKSDKNLGNMVDEILREMEENNVD 503
Query: 180 VTAFACNQLLLIYKKIDKKKIADVLL 205
N +L +Y K + ++ +
Sbjct: 504 PDLITVNHVLKVYAAESKIQAMEMFM 529
Score = 33.5 bits (75), Expect = 0.27
Identities = 48/227 (21%), Positives = 90/227 (39%), Gaps = 32/227 (14%)
Query: 230 NDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVC 289
+DI G +I + ++ E DH LA Y G+TEK E ++ ++ + N V
Sbjct: 329 DDIVGAEEIYKVWESLPLEFDHRIPTMLASGYRDRGMTEKAEKLMNSKTIKDRRMNKPVT 388
Query: 290 PTLLRLYAILGRADEVERIWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMSNKWK 349
P L + W + KP CL I+ K+ +A V E M K
Sbjct: 389 PLLEQ--------------WG-DQMKPSDLKCL--IKNLRDSKQFSKALQVSEWMGEKQV 431
Query: 350 LTARNYESLLKIYIRHKMLNKG------KDLIKTMGDSGCTIGPTTWDALVSLYVQAGE- 402
+ ++Y+ +L +++ + M D + + AL+S Y ++ +
Sbjct: 432 CNLYLEDYAARLYLTENVLGLEEAEKYFENIPENMKDY------SVYVALLSSYAKSDKN 485
Query: 403 -VEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYR 448
D +L++ +++N + P T +++ YA + E R
Sbjct: 486 LGNMVDEILRE-MEENNVDPDLITVNHVLKVYAAESKIQAMEMFMRR 531
>At3g23020 hypothetical protein
Length = 842
Score = 103 bits (256), Expect = 3e-22
Identities = 95/425 (22%), Positives = 175/425 (40%), Gaps = 28/425 (6%)
Query: 62 VDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASK- 120
V S D+ + KG + G ++ + G + AL WL K+ E +
Sbjct: 206 VQSLWDEMIRKGIKPINSTYGTLIDVYSKG---GLKVHALCWLGKMSKIGMQPDEVTTGI 262
Query: 121 -LDLIAKLRGLPKAEKYLE-----------HVPNSFRGELLYRTLLANCASLENLRKTEE 168
L + K R KAE++ + HV S Y T++ +++ E
Sbjct: 263 VLQMYKKAREFQKAEEFFKKWSCDENKADSHVCLS---SYTYNTMIDTYGKSGQIKEASE 319
Query: 169 TFNKMRELGFPVTAFACNQLLLIYKKIDKKKIADVLLMMEKENVKPSSYTYKILIDVKGL 228
TF +M E G T N ++ IY + L+ K + P + TY ILI +
Sbjct: 320 TFKRMLEEGIVPTTVTFNTMIHIYGNNGQLGEVTSLMKTMKLHCAPDTRTYNILISLHTK 379
Query: 229 SNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWV 288
+NDI+ + MK +G + D ++ +L ++ + E+ E ++ E++ +N++ + +
Sbjct: 380 NNDIERAGAYFKEMKDDGLKPDPVSYRTLLYAFSIRHMVEEAEGLIAEMDDDNVEIDEYT 439
Query: 289 CPTLLRLYAILGRADEVERIWKVCE-----SKPRVEDCLAAIEAWGRLKKIEEAEAVFEM 343
L R+Y A+ +E+ W + E A I+A+G + EAE VF
Sbjct: 440 QSALTRMYV---EAEMLEKSWSWFKRFHVAGNMSSEGYSANIDAYGERGYLSEAERVFIC 496
Query: 344 MSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEV 403
K T Y ++K Y K K +L ++M G T T++ LV + A
Sbjct: 497 CQEVNKRTVIEYNVMIKAYGISKSCEKACELFESMMSYGVTPDKCTYNTLVQILASADMP 556
Query: 404 EKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHA 463
K L+K + + + ++ + K G ++ AE+++ + + N + +
Sbjct: 557 HKGRCYLEKMRETGYVSDCI-PYCAVISSFVKLGQLNMAEEVYKEMVEYNIEPDVVVYGV 615
Query: 464 LAQAY 468
L A+
Sbjct: 616 LINAF 620
Score = 55.8 bits (133), Expect = 5e-08
Identities = 58/296 (19%), Positives = 128/296 (42%), Gaps = 8/296 (2%)
Query: 150 YRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKKIAD-VLLMME 208
Y TL+ AS + K KMRE G+ ++ + K+ + +A+ V M
Sbjct: 543 YNTLVQILASADMPHKGRCYLEKMRETGYVSDCIPYCAVISSFVKLGQLNMAEEVYKEMV 602
Query: 209 KENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTE 268
+ N++P Y +LI+ + ++ VE MK G + + SL + Y G +
Sbjct: 603 EYNIEPDVVVYGVLINAFADTGNVQQAMSYVEAMKEAGIPGNSVIYNSLIKLYTKVGYLD 662
Query: 269 KTEAILKEIE---GENLKENMWVCPTLLRLYAILGRADEVERIWKVCESKPRVEDCLAAI 325
+ EAI +++ + +++ ++ LY+ + E I+ + + + A+
Sbjct: 663 EAEAIYRKLLQSCNKTQYPDVYTSNCMINLYSERSMVRKAEAIFDSMKQRGEANEFTFAM 722
Query: 326 E--AWGRLKKIEEAEAVFEMMSNKWKLT-ARNYESLLKIYIRHKMLNKGKDLIKTMGDSG 382
+ + + EEA + + M LT +Y S+L ++ + + K M SG
Sbjct: 723 MLCMYKKNGRFEEATQIAKQMREMKILTDPLSYNSVLGLFALDGRFKEAVETFKEMVSSG 782
Query: 383 CTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGD 438
+T+ +L ++ ++ G +KA +++ +++ ++K +++ + GD
Sbjct: 783 IQPDDSTFKSLGTILMKLGMSKKAVRKIEE-IRKKEIKRGLELWISTLSSLVGIGD 837
Score = 46.2 bits (108), Expect = 4e-05
Identities = 47/200 (23%), Positives = 85/200 (42%), Gaps = 7/200 (3%)
Query: 117 YASKLDLIAKLRGLPKAEK-YLEHVPNSFRGELLYRTLLANC-ASLENLRKTEETFNKMR 174
Y + + KL L AE+ Y E V + +++ +L N A N+++ M+
Sbjct: 578 YCAVISSFVKLGQLNMAEEVYKEMVEYNIEPDVVVYGVLINAFADTGNVQQAMSYVEAMK 637
Query: 175 ELGFPVTAFACNQLLLIYKKI----DKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSN 230
E G P + N L+ +Y K+ + + I LL + P YT +I++ +
Sbjct: 638 EAGIPGNSVIYNSLIKLYTKVGYLDEAEAIYRKLLQSCNKTQYPDVYTSNCMINLYSERS 697
Query: 231 DIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCP 290
+ I ++MK G E + T A + Y G E+ I K++ + +
Sbjct: 698 MVRKAEAIFDSMKQRG-EANEFTFAMMLCMYKKNGRFEEATQIAKQMREMKILTDPLSYN 756
Query: 291 TLLRLYAILGRADEVERIWK 310
++L L+A+ GR E +K
Sbjct: 757 SVLGLFALDGRFKEAVETFK 776
Score = 36.6 bits (83), Expect = 0.032
Identities = 28/134 (20%), Positives = 55/134 (40%), Gaps = 5/134 (3%)
Query: 342 EMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAG 401
EM+ K Y +L+ +Y + + + M G T ++ +Y +A
Sbjct: 212 EMIRKGIKPINSTYGTLIDVYSKGGLKVHALCWLGKMSKIGMQPDEVTTGIVLQMYKKAR 271
Query: 402 EVEKADTVLQK-ALQQNKMKPMFT----TFMTIMEQYAKRGDVHNAEKIFYRLRQANYIS 456
E +KA+ +K + +NK T+ T+++ Y K G + A + F R+ + +
Sbjct: 272 EFQKAEEFFKKWSCDENKADSHVCLSSYTYNTMIDTYGKSGQIKEASETFKRMLEEGIVP 331
Query: 457 RISPFHALAQAYKN 470
F+ + Y N
Sbjct: 332 TTVTFNTMIHIYGN 345
>At5g09450 unknown protein
Length = 409
Score = 102 bits (254), Expect = 5e-22
Identities = 65/296 (21%), Positives = 147/296 (48%), Gaps = 11/296 (3%)
Query: 61 SVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASK 120
S + L+KW+ +G +++ E+ LRR + Y AL+ +W+ +++ + ++ +YAS+
Sbjct: 70 SATTVLEKWIGEGNQMTINELREISKELRRTRRYKHALEVTEWMVQHEESKISDADYASR 129
Query: 121 LDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMREL-GFP 179
+DLI+K+ G+ AE+Y E + + Y +LL A+ + + E F ++ E
Sbjct: 130 IDLISKVFGIDAAERYFEGLDIDSKTAETYTSLLHAYAASKQTERAEALFKRIIESDSLT 189
Query: 180 VTAFACNQLLLIYKKIDK-KKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQI 238
A N+++ +Y + + +K+ +V+ +++++ V P +TY + + + +ID + +I
Sbjct: 190 FGAITYNEMMTLYMSVGQVEKVPEVIEVLKQKKVSPDIFTYNLWLSSCAATFNIDELRKI 249
Query: 239 VETMKAEGCELDHLTR-ASLARHYAAAGLTEKTEAILKEIEGENLKENMWVC-PTLLRLY 296
+E M+ + + R L Y + E+ L +++ + W+ L+ L+
Sbjct: 250 LEEMRHDASSNEGWVRYIDLTSIYINSSRVTNAESTLPVEAEKSISQREWITYDFLMILH 309
Query: 297 AILGRADEVERIWKVCESKPRV---EDCLAAIEAWGRLKKIEEAEAVFEMMSNKWK 349
LG +++IWK + ++ + + ++ L + EAE + ++WK
Sbjct: 310 TGLGNKVMIDQIWKSLRNTNQILSSRSYICVLSSYLMLGHLREAEEII----HQWK 361
Score = 55.5 bits (132), Expect = 7e-08
Identities = 41/139 (29%), Positives = 67/139 (47%), Gaps = 3/139 (2%)
Query: 313 ESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMSNKWKLTARNYESLLKIYIRHKMLNKGK 372
ESK D + I+ ++ I+ AE FE + K TA Y SLL Y K + +
Sbjct: 118 ESKISDADYASRIDLISKVFGIDAAERYFEGLDIDSK-TAETYTSLLHAYAASKQTERAE 176
Query: 373 DLIKTMGDS-GCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIME 431
L K + +S T G T++ +++LY+ G+VEK V++ L+Q K+ P T+ +
Sbjct: 177 ALFKRIIESDSLTFGAITYNEMMTLYMSVGQVEKVPEVIE-VLKQKKVSPDIFTYNLWLS 235
Query: 432 QYAKRGDVHNAEKIFYRLR 450
A ++ KI +R
Sbjct: 236 SCAATFNIDELRKILEEMR 254
>At5g02860 putative protein
Length = 819
Score = 97.1 bits (240), Expect = 2e-20
Identities = 106/466 (22%), Positives = 184/466 (38%), Gaps = 63/466 (13%)
Query: 62 VDSALDKWVEKGK---ELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYA 118
+DS L + E K E + E+ L L K + AL+A DW +K+Y
Sbjct: 116 LDSVLSELFEPFKDKPESTSSELLAFLKGLGFHKKFDLALRAFDWF-------MKQKDYQ 168
Query: 119 SKLD--LIAKLRGLPKAEKYLEHVPNSFRG---------ELLYRTLLANCASLENLRKTE 167
S LD ++A + + E + N F G Y +L++ A+ R+
Sbjct: 169 SMLDNSVVAIIISMLGKEGRVSSAANMFNGLQEDGFSLDVYSYTSLISAFANSGRYREAV 228
Query: 168 ETFNKMRELGFPVTAFACNQLLLIYKKIDK--KKIADVLLMMEKENVKPSSYTYKILIDV 225
F KM E G T N +L ++ K+ KI ++ M+ + + P +YTY LI
Sbjct: 229 NVFKKMEEDGCKPTLITYNVILNVFGKMGTPWNKITSLVEKMKSDGIAPDAYTYNTLITC 288
Query: 226 KGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKEN 285
+ +Q+ E MKA G D +T +L Y + ++ +L E+ +
Sbjct: 289 CKRGSLHQEAAQVFEEMKAAGFSYDKVTYNALLDVYGKSHRPKEAMKVLNEMVLNGFSPS 348
Query: 286 MWVCPTLLRLYAILGRADEVERIWKVCE---SKPRVEDCLAAIEAWGRLKKIEEAEAVFE 342
+ +L+ YA G DE + +KP V + + R K+E A ++FE
Sbjct: 349 IVTYNSLISAYARDGMLDEAMELKNQMAEKGTKPDVFTYTTLLSGFERAGKVESAMSIFE 408
Query: 343 -------------------MMSNKWKLTAR-----------------NYESLLKIYIRHK 366
M N+ K T + +LL ++ ++
Sbjct: 409 EMRNAGCKPNICTFNAFIKMYGNRGKFTEMMKIFDEINVCGLSPDIVTWNTLLAVFGQNG 468
Query: 367 MLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTF 426
M ++ + K M +G T++ L+S Y + G E+A TV ++ L + P +T+
Sbjct: 469 MDSEVSGVFKEMKRAGFVPERETFNTLISAYSRCGSFEQAMTVYRRMLDAG-VTPDLSTY 527
Query: 427 MTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAK 472
T++ A+ G +EK+ + + +L AY N K
Sbjct: 528 NTVLAALARGGMWEQSEKVLAEMEDGRCKPNELTYCSLLHAYANGK 573
Score = 85.5 bits (210), Expect = 6e-17
Identities = 67/295 (22%), Positives = 126/295 (42%), Gaps = 6/295 (2%)
Query: 150 YRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKKIA-DVLLMME 208
Y TL+ C ++ + F +M+ GF N LL +Y K + K A VL M
Sbjct: 282 YNTLITCCKRGSLHQEAAQVFEEMKAAGFSYDKVTYNALLDVYGKSHRPKEAMKVLNEMV 341
Query: 209 KENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTE 268
PS TY LI +D ++ M +G + D T +L + AG E
Sbjct: 342 LNGFSPSIVTYNSLISAYARDGMLDEAMELKNQMAEKGTKPDVFTYTTLLSGFERAGKVE 401
Query: 269 KTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIW---KVCESKPRVEDCLAAI 325
+I +E+ K N+ +++Y G+ E+ +I+ VC P + +
Sbjct: 402 SAMSIFEEMRNAGCKPNICTFNAFIKMYGNRGKFTEMMKIFDEINVCGLSPDIVTWNTLL 461
Query: 326 EAWGRLKKIEEAEAVFEMMSNKWKLTAR-NYESLLKIYIRHKMLNKGKDLIKTMGDSGCT 384
+G+ E VF+ M + R + +L+ Y R + + + M D+G T
Sbjct: 462 AVFGQNGMDSEVSGVFKEMKRAGFVPERETFNTLISAYSRCGSFEQAMTVYRRMLDAGVT 521
Query: 385 IGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDV 439
+T++ +++ + G E+++ VL + ++ + KP T+ +++ YA ++
Sbjct: 522 PDLSTYNTVLAALARGGMWEQSEKVLAE-MEDGRCKPNELTYCSLLHAYANGKEI 575
Score = 70.5 bits (171), Expect = 2e-12
Identities = 60/307 (19%), Positives = 129/307 (41%), Gaps = 12/307 (3%)
Query: 170 FNKMRELGFPVTAFACNQLLLIYKKIDK-KKIADVLLMMEKENVKPSSYTYKILIDVKGL 228
F+++ G N LL ++ + +++ V M++ P T+ LI
Sbjct: 442 FDEINVCGLSPDIVTWNTLLAVFGQNGMDSEVSGVFKEMKRAGFVPERETFNTLISAYSR 501
Query: 229 SNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWV 288
+ + M G D T ++ A G+ E++E +L E+E K N
Sbjct: 502 CGSFEQAMTVYRRMLDAGVTPDLSTYNTVLAALARGGMWEQSEKVLAEMEDGRCKPNELT 561
Query: 289 CPTLLRLYAILGRADEVERIWKVCES------KPRVEDCLAAIEAWGRLKKIEEAEAVF- 341
+LL YA E+ + + E +PR + + + EAE F
Sbjct: 562 YCSLLHAYA---NGKEIGLMHSLAEEVYSGVIEPRAVLLKTLVLVCSKCDLLPEAERAFS 618
Query: 342 EMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAG 401
E+ + S++ IY R +M+ K ++ M + G T T+++L+ ++ ++
Sbjct: 619 ELKERGFSPDITTLNSMVSIYGRRQMVAKANGVLDYMKERGFTPSMATYNSLMYMHSRSA 678
Query: 402 EVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPF 461
+ K++ +L++ L + +KP ++ T++ Y + + +A +IF +R + + + +
Sbjct: 679 DFGKSEEILREILAKG-IKPDIISYNTVIYAYCRNTRMRDASRIFSEMRNSGIVPDVITY 737
Query: 462 HALAQAY 468
+ +Y
Sbjct: 738 NTFIGSY 744
Score = 48.1 bits (113), Expect = 1e-05
Identities = 43/203 (21%), Positives = 86/203 (42%), Gaps = 4/203 (1%)
Query: 148 LLYRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIY-KKIDKKKIADVLLM 206
+L +TL+ C+ + L + E F++++E GF N ++ IY ++ K VL
Sbjct: 595 VLLKTLVLVCSKCDLLPEAERAFSELKERGFSPDITTLNSMVSIYGRRQMVAKANGVLDY 654
Query: 207 MEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGL 266
M++ PS TY L+ + S D +I+ + A+G + D ++ ++ Y
Sbjct: 655 MKERGFTPSMATYNSLMYMHSRSADFGKSEEILREILAKGIKPDIISYNTVIYAYCRNTR 714
Query: 267 TEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRADE---VERIWKVCESKPRVEDCLA 323
I E+ + ++ T + YA +E V R +P +
Sbjct: 715 MRDASRIFSEMRNSGIVPDVITYNTFIGSYAADSMFEEAIGVVRYMIKHGCRPNQNTYNS 774
Query: 324 AIEAWGRLKKIEEAEAVFEMMSN 346
++ + +L + +EA+ E + N
Sbjct: 775 IVDGYCKLNRKDEAKLFVEDLRN 797
Score = 38.9 bits (89), Expect = 0.006
Identities = 54/326 (16%), Positives = 114/326 (34%), Gaps = 41/326 (12%)
Query: 150 YRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDK-KKIADVLLMME 208
+ TLLA + F +M+ GF N L+ Y + ++ V M
Sbjct: 457 WNTLLAVFGQNGMDSEVSGVFKEMKRAGFVPERETFNTLISAYSRCGSFEQAMTVYRRML 516
Query: 209 KENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTE 268
V P TY ++ + +++ M+ C+ + LT SL YA
Sbjct: 517 DAGVTPDLSTYNTVLAALARGGMWEQSEKVLAEMEDGRCKPNELTYCSLLHAYANGKEIG 576
Query: 269 KTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCESK---PRVEDCLAAI 325
++ +E+ ++ + TL+ + + E ER + + + P + + +
Sbjct: 577 LMHSLAEEVYSGVIEPRAVLLKTLVLVCSKCDLLPEAERAFSELKERGFSPDITTLNSMV 636
Query: 326 EAWGRLKKIEEAEAVFEMMSNKW------------------------------------K 349
+GR + + +A V + M + K
Sbjct: 637 SIYGRRQMVAKANGVLDYMKERGFTPSMATYNSLMYMHSRSADFGKSEEILREILAKGIK 696
Query: 350 LTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTV 409
+Y +++ Y R+ + + M +SG T++ + Y E+A V
Sbjct: 697 PDIISYNTVIYAYCRNTRMRDASRIFSEMRNSGIVPDVITYNTFIGSYAADSMFEEAIGV 756
Query: 410 LQKALQQNKMKPMFTTFMTIMEQYAK 435
++ ++ +P T+ +I++ Y K
Sbjct: 757 VRYMIKHG-CRPNQNTYNSIVDGYCK 781
>At2g35130 hypothetical protein
Length = 591
Score = 94.0 bits (232), Expect = 2e-19
Identities = 65/288 (22%), Positives = 144/288 (49%), Gaps = 5/288 (1%)
Query: 190 LIYKKIDKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCEL 249
L+ +K + ++ DV M+++ KP++ TY ++I++ G ++ ++ M++ C+
Sbjct: 238 LMKRKGNTEEAIDVFQRMKRDRCKPTTETYNLMINLYGKASKSYMSWKLYCEMRSHQCKP 297
Query: 250 DHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGR---ADEVE 306
+ T +L +A GL EK E I ++++ + L+ +++V L+ Y+ G A E+
Sbjct: 298 NICTYTALVNAFAREGLCEKAEEIFEQLQEDGLEPDVYVYNALMESYSRAGYPYGAAEIF 357
Query: 307 RIWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMSNKWKL-TARNYESLLKIYIRH 365
+ + +P ++A+GR +AEAVFE M T +++ LL Y +
Sbjct: 358 SLMQHMGCEPDRASYNIMVDAYGRAGLHSDAEAVFEEMKRLGIAPTMKSHMLLLSAYSKA 417
Query: 366 KMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTT 425
+ + K + ++K M ++G +++++LY + G+ K + +L + ++ +T
Sbjct: 418 RDVTKCEAIVKEMSENGVEPDTFVLNSMLNLYGRLGQFTKMEKILAE-MENGPCTADIST 476
Query: 426 FMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKL 473
+ ++ Y K G + E++F L++ N+ + + + AY KL
Sbjct: 477 YNILINIYGKAGFLERIEELFVELKEKNFRPDVVTWTSRIGAYSRKKL 524
Score = 84.7 bits (208), Expect = 1e-16
Identities = 81/336 (24%), Positives = 146/336 (43%), Gaps = 11/336 (3%)
Query: 85 LNSLRRRKMYGRALQALDWLESNKK--LEFTEKEYASKLDLIAKL-RGLPKAEKYLEHVP 141
+ L +RK G +A+D + K+ + T + Y ++L K + + Y E
Sbjct: 235 IEGLMKRK--GNTEEAIDVFQRMKRDRCKPTTETYNLMINLYGKASKSYMSWKLYCEMRS 292
Query: 142 NSFRGELLYRTLLANCASLENL-RKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKK- 199
+ + + T L N + E L K EE F +++E G + N L+ Y +
Sbjct: 293 HQCKPNICTYTALVNAFAREGLCEKAEEIFEQLQEDGLEPDVYVYNALMESYSRAGYPYG 352
Query: 200 IADVLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLAR 259
A++ +M+ +P +Y I++D G + + E MK G + L
Sbjct: 353 AAEIFSLMQHMGCEPDRASYNIMVDAYGRAGLHSDAEAVFEEMKRLGIAPTMKSHMLLLS 412
Query: 260 HYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCESKPRVE 319
Y+ A K EAI+KE+ ++ + +V ++L LY LG+ ++E+I E+ P
Sbjct: 413 AYSKARDVTKCEAIVKEMSENGVEPDTFVLNSMLNLYGRLGQFTKMEKILAEMENGPCTA 472
Query: 320 DCLA---AIEAWGRLKKIEEAEAVF-EMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLI 375
D I +G+ +E E +F E+ ++ + S + Y R K+ K ++
Sbjct: 473 DISTYNILINIYGKAGFLERIEELFVELKEKNFRPDVVTWTSRIGAYSRKKLYVKCLEVF 532
Query: 376 KTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQ 411
+ M DSGC T L+S +VE+ +VL+
Sbjct: 533 EEMIDSGCAPDGGTAKVLLSACSSEEQVEQVTSVLR 568
Score = 59.7 bits (143), Expect = 4e-09
Identities = 67/294 (22%), Positives = 123/294 (41%), Gaps = 38/294 (12%)
Query: 198 KKIADVLLMME----KENVKPSSYTYKILIDVKGLSNDI-DGMSQIVETMKAEGCELDHL 252
KK ++L+ E K + +P + +LID G + S V+ +++ +
Sbjct: 133 KKWDSIILVCEWILRKSSFQPDVICFNLLIDAYGQKFQYKEAESLYVQLLESRYVPTED- 191
Query: 253 TRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVC 312
T A L + Y AGL E+ E +L E++ ++ + T+ Y +E + K
Sbjct: 192 TYALLIKAYCMAGLIERAEVVLVEMQNHHVSPKT-IGVTVYNAY--------IEGLMK-- 240
Query: 313 ESKPRVEDCLAAIEAWGRLKKIEEAEAVFEMMS-NKWKLTARNYESLLKIYIRHKMLNKG 371
R EEA VF+ M ++ K T Y ++ +Y +
Sbjct: 241 -----------------RKGNTEEAIDVFQRMKRDRCKPTTETYNLMINLYGKASKSYMS 283
Query: 372 KDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIME 431
L M C T+ ALV+ + + G EKA+ + ++ LQ++ ++P + +ME
Sbjct: 284 WKLYCEMRSHQCKPNICTYTALVNAFAREGLCEKAEEIFEQ-LQEDGLEPDVYVYNALME 342
Query: 432 QYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKL--PAYGIRERMK 483
Y++ G + A +IF ++ + ++ + AY A L A + E MK
Sbjct: 343 SYSRAGYPYGAAEIFSLMQHMGCEPDRASYNIMVDAYGRAGLHSDAEAVFEEMK 396
>At4g16390 salt-inducible protein homolog
Length = 777
Score = 83.6 bits (205), Expect = 2e-16
Identities = 75/365 (20%), Positives = 155/365 (41%), Gaps = 6/365 (1%)
Query: 114 EKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKM 173
+ Y+S + L L E + V F G+L + + ++ N N +
Sbjct: 181 DSRYSSLIKLAESLDACKPNEADVCDVITGFGGKLFEQDAVVTLNNMTNPETAPLVLNNL 240
Query: 174 RELGFPVT-AFACNQLLLIYKKI-DKKKIADVLLMMEKENVKPSSYTYKILIDVKGLSND 231
E P N + +++K D +K + M + +KP + T+ +I +
Sbjct: 241 LETMKPSREVILYNVTMKVFRKSKDLEKSEKLFDEMLERGIKPDNATFTTIISCARQNGV 300
Query: 232 IDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPT 291
+ E M + GCE D++T A++ Y AG + ++ E + + T
Sbjct: 301 PKRAVEWFEKMSSFGCEPDNVTMAAMIDAYGRAGNVDMALSLYDRARTEKWRIDAVTFST 360
Query: 292 LLRLYAILGRADEVERIWKVCES---KPRVEDCLAAIEAWGRLKKIEEAEAVF-EMMSNK 347
L+R+Y + G D I++ ++ KP + I++ GR K+ +A+ ++ ++++N
Sbjct: 361 LIRIYGVSGNYDGCLNIYEEMKALGVKPNLVIYNRLIDSMGRAKRPWQAKIIYKDLITNG 420
Query: 348 WKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKAD 407
+ Y +L++ Y R + + + + M + G ++ ++ L+S+ V++A
Sbjct: 421 FTPNWSTYAALVRAYGRARYGDDALAIYREMKEKGLSLTVILYNTLLSMCADNRHVDEAF 480
Query: 408 TVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANYISRISPFHALAQA 467
+ Q P TF +++ YA G V AE ++R+A + + ++ Q
Sbjct: 481 EIFQDMKNCETCDPDSWTFSSLITVYACSGRVSKAEAALLQIREAGFEPTLFVLTSVIQC 540
Query: 468 YKNAK 472
Y AK
Sbjct: 541 YGKAK 545
Score = 45.4 bits (106), Expect = 7e-05
Identities = 46/216 (21%), Positives = 95/216 (43%), Gaps = 12/216 (5%)
Query: 207 MEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMK-AEGCELDHLTRASLARHYAAAG 265
M+++ + + Y L+ + + +D +I + MK E C+ D T +SL YA +G
Sbjct: 451 MKEKGLSLTVILYNTLLSMCADNRHVDEAFEIFQDMKNCETCDPDSWTFSSLITVYACSG 510
Query: 266 LTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIW-KVCESKPRVED--CL 322
K EA L +I + ++V ++++ Y + D+V R + +V E +D C
Sbjct: 511 RVSKAEAALLQIREAGFEPTLFVLTSVIQCYGKAKQVDDVVRTFDQVLELGITPDDRFCG 570
Query: 323 AAIEAWGRLKKIEEAEAVFEMMSNKWKLTARNYESLLKIYIRHKMLNKG---KDLIKTMG 379
+ + E + + + K KL ++K+ + + +G K+ + +
Sbjct: 571 CLLNVMTQTPSEEIGKLIGCVEKAKPKL-----GQVVKMLVEEQNCEEGVFKKEASELID 625
Query: 380 DSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQ 415
G + + L+ L V ++E+A +LQ L+
Sbjct: 626 SIGSDVKKAYLNCLIDLCVNLNKLERACEILQLGLE 661
>At1g43010 hypothetical protein
Length = 257
Score = 83.2 bits (204), Expect = 3e-16
Identities = 56/211 (26%), Positives = 109/211 (51%), Gaps = 8/211 (3%)
Query: 61 SVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLESNKKLEFTEKEYASK 120
S+ L++W ++G E++ + + +L K + AL+A W+ + + ++ A++
Sbjct: 51 SIIPLLEQWRKQGYEVNPSHLRGLIKNLSDCKNFTTALEASKWMFKHSVFDNFPEDCAAQ 110
Query: 121 LDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCAS-LENLRKTEETFNKMRELGFP 179
L L+ + GL +AEK +++P R Y LL++ + + K E TF KMRELGF
Sbjct: 111 LHLVNTVLGLEEAEKMFKNIPEKMRD---YSVLLSSYTKPVRTVDKAEATFKKMRELGFL 167
Query: 180 VTAFACNQLLLIYKKIDKKKIADVLL-MMEKENVKPSSYTYKILIDVKGLSNDIDGMSQI 238
+ + N ++ +Y ++ + + + LL ++K N++ S +V + +I+ M +
Sbjct: 168 LKPYLFNSMICLYGQLQRLDMVEKLLYKLKKNNMEVGSLKVN---NVSRVYANINAMEKF 224
Query: 239 VETMKAEGCELDHLTRASLARHYAAAGLTEK 269
+ EG EL+ T ++A+ Y AG EK
Sbjct: 225 KTWVSKEGIELERDTIVAMAKAYHRAGSIEK 255
Score = 42.4 bits (98), Expect = 6e-04
Identities = 37/154 (24%), Positives = 71/154 (46%), Gaps = 9/154 (5%)
Query: 319 EDCLAAIEAWGRLKKIEEAEAVFEMMSNKWKLTARNYESLLKIYIRH-KMLNKGKDLIKT 377
EDC A + + +EEAE +F+ + K R+Y LL Y + + ++K + K
Sbjct: 105 EDCAAQLHLVNTVLGLEEAEKMFKNIPEK----MRDYSVLLSSYTKPVRTVDKAEATFKK 160
Query: 378 MGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRG 437
M + G + P +++++ LY Q ++ + +L K L++N M+ + YA
Sbjct: 161 MRELGFLLKPYLFNSMICLYGQLQRLDMVEKLLYK-LKKNNMEVGSLKVNNVSRVYA--- 216
Query: 438 DVHNAEKIFYRLRQANYISRISPFHALAQAYKNA 471
+++ EK + + A+A+AY A
Sbjct: 217 NINAMEKFKTWVSKEGIELERDTIVAMAKAYHRA 250
>At4g39620 putative protein
Length = 563
Score = 82.0 bits (201), Expect = 7e-16
Identities = 80/335 (23%), Positives = 150/335 (43%), Gaps = 24/335 (7%)
Query: 148 LLYRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKK----KIADV 203
+L R+L++ + E L KT + + K+ C+ L+++++ K + +V
Sbjct: 69 VLVRSLMSRISDREPLVKTLDKYVKV---------VRCDHCFLLFEELGKSDKWLQCLEV 119
Query: 204 LLMMEKEN-VKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELD-HLTRASLARHY 261
M+K+ P + Y LI V G + MK GC D + A + H
Sbjct: 120 FRWMQKQRWYIPDNGVYSKLISVMGKKGQTRMAMWLFSEMKNSGCRPDASVYNALITAHL 179
Query: 262 AA---AGLTEKTEAILKEIEG-ENLKENMWVCPTLLRLYAILGRADEVERIWKVCESKPR 317
A EK L +++G E + N+ LLR +A G+ D+V ++K + P
Sbjct: 180 HTRDKAKALEKVRGYLDKMKGIERCQPNVVTYNILLRAFAQSGKVDQVNALFKDLDMSPV 239
Query: 318 VEDCLA---AIEAWGRLKKIEEAEAVF-EMMSNKWKLTARNYESLLKIYIRHKMLNKGKD 373
D ++A+G+ I+E EAV M SN+ K + L+ Y + + K +
Sbjct: 240 SPDVYTFNGVMDAYGKNGMIKEMEAVLTRMRSNECKPDIITFNVLIDSYGKKQEFEKMEQ 299
Query: 374 LIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQY 433
K++ S T+++++ Y +A ++KA+ V +K N + P F T+ ++ Y
Sbjct: 300 TFKSLMRSKEKPTLPTFNSMIINYGKARMIDKAEWVFKKMNDMNYI-PSFITYECMIMMY 358
Query: 434 AKRGDVHNAEKIFYRLRQANYISRISPFHALAQAY 468
G V A +IF + +++ + + S +A+ + Y
Sbjct: 359 GYCGSVSRAREIFEEVGESDRVLKASTLNAMLEVY 393
Score = 65.5 bits (158), Expect = 6e-11
Identities = 67/343 (19%), Positives = 150/343 (43%), Gaps = 12/343 (3%)
Query: 149 LYRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLL--IYKKIDKKKIADVLLM 206
+Y L++ R F++M+ G A N L+ ++ + K + V
Sbjct: 135 VYSKLISVMGKKGQTRMAMWLFSEMKNSGCRPDASVYNALITAHLHTRDKAKALEKVRGY 194
Query: 207 MEK----ENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYA 262
++K E +P+ TY IL+ S +D ++ + + + D T + Y
Sbjct: 195 LDKMKGIERCQPNVVTYNILLRAFAQSGKVDQVNALFKDLDMSPVSPDVYTFNGVMDAYG 254
Query: 263 AAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVC---ESKPRVE 319
G+ ++ EA+L + K ++ L+ Y +++E+ +K + KP +
Sbjct: 255 KNGMIKEMEAVLTRMRSNECKPDIITFNVLIDSYGKKQEFEKMEQTFKSLMRSKEKPTLP 314
Query: 320 DCLAAIEAWGRLKKIEEAEAVFEMMSNKWKLTAR-NYESLLKIYIRHKMLNKGKDLIKTM 378
+ I +G+ + I++AE VF+ M++ + + YE ++ +Y +++ +++ + +
Sbjct: 315 TFNSMIINYGKARMIDKAEWVFKKMNDMNYIPSFITYECMIMMYGYCGSVSRAREIFEEV 374
Query: 379 GDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGD 438
G+S + +T +A++ +Y + G +AD + A ++ P +T+ + + Y K D
Sbjct: 375 GESDRVLKASTLNAMLEVYCRNGLYIEADKLFHNA-SAFRVHPDASTYKFLYKAYTK-AD 432
Query: 439 VHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRER 481
+ +I + + + I F A ++LP G R
Sbjct: 433 MKEQVQILMKKMEKDGIVPNKRFFLEALEVFGSRLPGSGSENR 475
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.133 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,165,541
Number of Sequences: 26719
Number of extensions: 419373
Number of successful extensions: 4284
Number of sequences better than 10.0: 431
Number of HSP's better than 10.0 without gapping: 237
Number of HSP's successfully gapped in prelim test: 196
Number of HSP's that attempted gapping in prelim test: 1847
Number of HSP's gapped (non-prelim): 1577
length of query: 488
length of database: 11,318,596
effective HSP length: 103
effective length of query: 385
effective length of database: 8,566,539
effective search space: 3298117515
effective search space used: 3298117515
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135463.1


