

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135396.1 - phase: 0 /pseudo
(1436 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
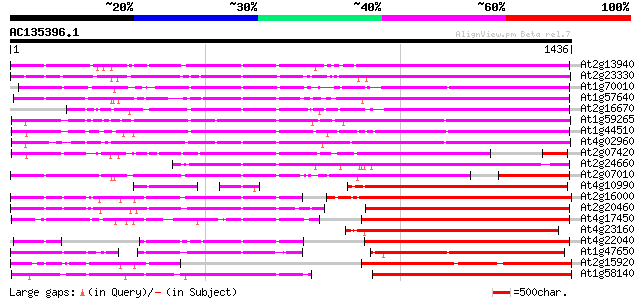
At2g13940 putative retroelement pol polyprotein 862 0.0
At2g23330 putative retroelement pol polyprotein 847 0.0
At1g70010 hypothetical protein 827 0.0
At1g57640 785 0.0
At2g16670 putative retroelement pol polyprotein 722 0.0
At1g59265 polyprotein, putative 702 0.0
At1g44510 polyprotein, putative 702 0.0
At4g02960 putative polyprotein of LTR transposon 693 0.0
At2g07420 putative retroelement pol polyprotein 676 0.0
At2g24660 putative retroelement pol polyprotein 645 0.0
At2g07010 putative retroelement pol polyprotein 595 e-170
At4g10990 putative retrotransposon polyprotein 520 e-147
At2g16000 putative retroelement pol polyprotein 517 e-146
At2g20460 putative retroelement pol polyprotein 513 e-145
At4g17450 retrotransposon like protein 504 e-142
At4g23160 putative protein 485 e-137
At4g22040 LTR retrotransposon like protein 474 e-133
At1g47650 hypothetical protein 469 e-132
At2g15920 putative retroelement pol polyprotein 446 e-125
At1g58140 hypothetical protein 440 e-123
>At2g13940 putative retroelement pol polyprotein
Length = 1501
Score = 862 bits (2226), Expect = 0.0
Identities = 528/1495 (35%), Positives = 808/1495 (53%), Gaps = 115/1495 (7%)
Query: 1 ITTIRLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGADYDTWDAENSMVMTWLV 60
I+++ LNG NY +W+ + ++ + K G+I G +P +Y+ W A NSM++ W+
Sbjct: 44 ISSVELNGDNYNQWATEMLNALQAKRKTGFINGTIPRPPPNDPNYENWTAVNSMIVGWIR 103
Query: 61 NSLTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGC 120
S+ ++ A + A LW+++ Q +S +GN+ +I+++ QL RQ +V +Y+G
Sbjct: 104 TSIEPKVKATVTFISDAHLLWKDLKQRFS-VGNKVRIHQIRAQLSSCRQDGQAVIEYYGR 162
Query: 121 LKRIWQDLDLFNEYEWKSPEDCK-----HYQKMVDVSRVFKFLAGLNVE-FDEVRGRILG 174
L +W++ +++ + C+ K + ++ +F+ GL+ F + ++
Sbjct: 163 LSNLWEEYNIYKPVTVCTCGLCRCGATSEPTKEREEEKIHQFVLGLDESRFGGLCATLIN 222
Query: 175 RNPIPPIGEVFAEVRREESRRQVM--LGKKKAAVPPPVEGSALAVPQVNH-------KSF 225
+P+P +GE+++ V REE R + +K+ AV G Q++H S
Sbjct: 223 MDPLPSLGEIYSRVIREEQRLASVHVREQKEEAV-----GFLARREQLDHHSRVDASSSR 277
Query: 226 RNPRGGDKTHLV------CDYCGRNRHTQETCFKLHGRPN-------------------N 260
GG +++ + C CGR H ++ C+++ G P+
Sbjct: 278 SEHTGGSRSNSIIKGRVTCSNCGRTGHEKKECWQIVGFPDWWSERNGGRGSNGRGRGGRG 337
Query: 261 SKAGKFGDRPMP---TTSNAASSP-FTKEQMDHLLKLLKFNSSPNTPIGTVAQTGKDSWA 316
S G+ + M T+SN++ P FT+E M L +L+K S+ G+ + D
Sbjct: 338 SNGGRGQGQVMAAHATSSNSSVFPEFTEEHMRVLSQLVKEKSNS----GSTSNNNSDR-- 391
Query: 317 LSVQNHSNPWIIDSGASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKGNIKISEQ 376
LS + I+DSGAS HMT ++ P V ADGS + G + +S
Sbjct: 392 LSGKTKLGDIILDSGASHHMTGTLSSLTNVVPVPPCP-VGFADGSKAFALSVGVLTLSNT 450
Query: 377 ITLQSVLHVPKFACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYYF 436
++L +VL VP C L+SV KL + + C F + C QD++S IG+ E G+YY
Sbjct: 451 VSLTNVLFVPSLNCTLISVSKLLKQTQCLATFTDTLCFLQDRSSKTLIGSGEERGGVYYL 510
Query: 437 -DETPLGNTAMFGSSRTSPPLVSNKIMLWHKRLGHPSFSYLKHLFPEFSKEISSSQFH-C 494
D TP ++ V + LWH+RLGHPSFS L L P FSK S+ H C
Sbjct: 511 TDVTP---------AKIHTANVDSDQALWHQRLGHPSFSVLSSL-PLFSKTSSTVTSHSC 560
Query: 495 DACHLAKDHRVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDHTRVCWVY 554
D C AK R F + F LIH DVWGP ++ +F+T +DD++R W Y
Sbjct: 561 DVCFRAKQTREVFPESINKTEECFSLIHCDVWGPYRVPASCGAVYFLTIVDDYSRAVWTY 620
Query: 555 LMEKKSEVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLVTNGIIHQSTCRD 614
L+ +KSEV Q +F + QF T+ ++RSDNGTE+ LS++ NGIIHQ++C
Sbjct: 621 LLLEKSEVRQVLTNFLKYAEKQFGKTVKMVRSDNGTEFM--CLSSYFRENGIIHQTSCVG 678
Query: 615 TPQQNGIAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACHLINRMPSRVLQYETPVQV 674
TPQQNG ERK+RH+L V RA++F ++P WG ++LTA +LINR PS +L TP +V
Sbjct: 679 TPQQNGRVERKHRHILNVARALLFQASLPIKFWGESILTAAYLINRTPSSILSGRTPYEV 738
Query: 675 LQNNFPTSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYKCFNPV 734
L + P L+VFG CYV+ + K ++ C+F+GY KKG+K ++
Sbjct: 739 LHGSKPVYS-----QLRVFGSACYVHRVTRDKDKFGQRSRSCIFVGYPFGKKGWKVYDIE 793
Query: 735 TKKFFESMDVHFVEDQ-PFFRENS--LQGESQSSNEEDNFWEILPV-----LDDI----- 781
+F S DV F E+ P+ NS L S + ED+ W I P+ +D +
Sbjct: 794 RNEFLVSRDVIFREEVFPYAGVNSSTLASTSLPTVSEDDDWAIPPLEVRGSIDSVETERV 853
Query: 782 -VTNDH--LDAKITEPRNPNLELLEHMVFETGGEEGETLIKNPELRVYVRKKFHKDGTNP 838
T D LD +++ PN E + L +P T P
Sbjct: 854 VCTTDEVVLDTSVSDSEIPNQEFVPDDT-----PPSSPLSVSPS-------GSPNTPTTP 901
Query: 839 IVFPAEVQSDSPSEGPTDNPSSSSSGNSSHSSNDLPDLSFPDINLPIAVRKNIPDLNIPI 898
IV P S P P S ++H L D + + +I L
Sbjct: 902 IVVPVA------SPIPVSPPKQRKSKRATHPPPKLNDYVLYNA---MYTPSSIHALPADP 952
Query: 899 AERKRTRTCTKHPMSNYLSYDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEM 958
++ + P+++Y+S S SH+AY++ I++ P+ +EA+ W A+ E+
Sbjct: 953 SQSSTVPGKSLFPLTDYVSDAAFSSSHRAYLAAITDNVEPKHFKEAVQIKVWNDAMFTEV 1012
Query: 959 NALNKNNTWCITDLPYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQET 1018
+AL N TW I DLP K A+G +WVF K +DG+VERYKARLV +G Q G DY+ET
Sbjct: 1013 DALEINKTWDIVDLPPGKVAIGSQWVFKTKYNSDGTVERYKARLVVQGNKQVEGEDYKET 1072
Query: 1019 FAPVAKINSIRILLSLAVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLGRGKVC 1078
FAPV ++ ++R LL W ++Q DV NAFL+G+L EEVYM+LPPGF KVC
Sbjct: 1073 FAPVVRMTTVRTLLRNVAANQWEVYQMDVHNAFLHGDLEEEVYMKLPPGFRHS-HPDKVC 1131
Query: 1079 RLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIM 1138
RL+KSLYGLKQ+PR WF++ + GF QS D+++F ++R +++YVDD+++
Sbjct: 1132 RLRKSLYGLKQAPRCWFKKLSDSLLRFGFVQSYEDYSLF-SYTRNNIELRVLIYVDDLLI 1190
Query: 1139 TGDDIGEISDLKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMT 1198
G+D + K L F +KDLGKLKYFLG+E +R EGIFL+QRKY LD++ ++G
Sbjct: 1191 CGNDGYMLQKFKDYLSRCFSMKDLGKLKYFLGIEVSRGPEGIFLSQRKYALDVIADSGNL 1250
Query: 1199 GCKAAETPMDPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHAP 1258
G + A TP++ N L S + D + Y+RL GRL+YL HTRP+++++V V++QFM P
Sbjct: 1251 GSRPAHTPLEQNHHLASDDGPLLSDPKPYRRLVGRLLYLLHTRPELSYSVHVLAQFMQNP 1310
Query: 1259 GPAHFEAVFRILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGG 1318
AHF+A R++RYLKG+PG+G++ + +E Y D+DW RRS S Y +GG
Sbjct: 1311 REAHFDAALRVVRYLKGSPGQGILLNADPDLTLEVYCDSDWQSCPLTRRSISAYVVLLGG 1370
Query: 1319 NLVTWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAI 1378
+ ++W++KKQ+ V+ SSAEAE+R++++ E+ W++K ++EL I TP ++YCD+KAAI
Sbjct: 1371 SPISWKTKKQDTVSHSSAEAEYRAMSYALKEIKWLRKLLKELGIEQSTPARLYCDSKAAI 1430
Query: 1379 SIAHNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVADVLTKSLPKRQF 1433
IA NPV H+RTKH+E D H +++ + G I ++ T Q+ADV TK+L + QF
Sbjct: 1431 HIAANPVFHERTKHIESDCHSVRDAVRDGIITTQHVRTTEQLADVFTKALGRNQF 1485
>At2g23330 putative retroelement pol polyprotein
Length = 1496
Score = 847 bits (2187), Expect = 0.0
Identities = 531/1513 (35%), Positives = 808/1513 (53%), Gaps = 144/1513 (9%)
Query: 1 ITTIRLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGADYDTWDAENSMVMTWLV 60
I+++ L NY WS+ +Q ++R + K+G+I G +P + W A NSM++ W+
Sbjct: 34 ISSVVLKENNYAEWSEELQNFLRAKQKLGFIDGSIPKP-AADPELSLWIAINSMIVGWIR 92
Query: 61 NSLTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGC 120
S+ I + + A LWEN+ + +S +GN + L ++ Q V Y+G
Sbjct: 93 TSIDPTIRSTVGFVSEASQLWENLRRRFS-VGNGVRKTLLKDEIAACTQDGQPVLAYYGR 151
Query: 121 LKRIWQDLDLFNEYEWKSPEDCK-----HYQKMVDVSRVFKFLAGLNVEFDEVRGRILGR 175
L ++W++L +KS +CK +K + RV KFL GL+ F +R I
Sbjct: 152 LIKLWEELQ-----NYKSGRECKCEAASDIEKEREDDRVHKFLLGLDSRFSSIRSSITDI 206
Query: 176 NPIPPIGEVFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVPQVNHKSFRNPRGGDKTH 235
P+P + +V++ V REE + + + K V G ++ +S PR DK+
Sbjct: 207 EPLPDLYQVYSRVVREE--QNLNASRTKDVVKTEAIGFSV-------QSSTTPRFRDKST 257
Query: 236 LVCDYCGRNRHTQETCFKLHGRPN----------------------NSKAGKFGDRPM-P 272
L C +C R H CF +HG P+ S +G+ G+R P
Sbjct: 258 LFCTHCNRKGHEVTQCFLVHGYPDWWLEQNPQENQPSTRGRGSNGRGSSSGRGGNRSSAP 317
Query: 273 TT--------SNAASSPFTKEQMDHLLKLLKFNSSPNTPIGTVAQTGKDSWALSVQNHSN 324
TT + AA+ + + D + +L+ + + S LS
Sbjct: 318 TTRGRGRANNAQAAAPTVSGDGNDQIAQLISLLQAQ--------RPSSSSERLSGNTCLT 369
Query: 325 PWIIDSGASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKGNIKISEQITLQSVLH 384
+ID+GAS HMT + F + S K DG S G + + + L VL
Sbjct: 370 DGVIDTGASHHMTGDCSILVDVFDITPSPVTK-PDGKASQATKCGTLLLHDSYKLHDVLF 428
Query: 385 VPKFACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYYFDETPLGNT 444
VP F C L+SV KL + ++ +F + C QD+ IG E G+YYF G
Sbjct: 429 VPDFDCTLISVSKLLKQTSSIAIFTDTFCFLQDRFLRTLIGAGEEREGVYYFT----GVL 484
Query: 445 AMFGSSRTSPPLVSNKIMLWHKRLGHPSFSYLKHLFPEFSKEISSSQFH----CDACHLA 500
A +S +S LWH+RLGHPS S L L PE ++ SS F CD C +
Sbjct: 485 APRVHKASSDFAISGD--LWHRRLGHPSTSVLLSL-PECNR--SSQGFDKIDSCDTCFRS 539
Query: 501 KDHRVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDHTRVCWVYLMEKKS 560
K R F + F LIH DVWGP + + + +F+T +DD++R W YLM K+
Sbjct: 540 KQTREVFPISNNKTMECFSLIHGDVWGPYRTPSTTGAVYFLTLVDDYSRSVWTYLMSSKT 599
Query: 561 EVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLVTNGIIHQSTCRDTPQQNG 620
EV Q ++F M + QF + R+DNGTE+ L+ + T+GI+HQ++C DTPQQNG
Sbjct: 600 EVSQLIKNFCAMSERQFGKQVKAFRTDNGTEFM--CLTPYFQTHGILHQTSCVDTPQQNG 657
Query: 621 IAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACHLINRMPSRVLQYETPVQVLQNNFP 680
ERK+RH+L V RA +F N+P WG ++LTA HLINR PS VL+ +TP ++L P
Sbjct: 658 RVERKHRHILNVARACLFQGNLPVKFWGESILTATHLINRTPSAVLKGKTPYELLFGERP 717
Query: 681 TSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYKCFNPVTKKFFE 740
+ + L+ FGCLCY +I + K ++ KCVF+GY KK ++ ++ T K F
Sbjct: 718 SYDM-----LRSFGCLCYAHIRPRNKDKFTSRSRKCVFIGYPHGKKAWRVYDLETGKIFA 772
Query: 741 SMDVHFVED-QPFFRENSLQGESQSSNEEDNFWEILPVLDDIVTNDHLDAKITEPRNPNL 799
S DV F ED P+ + SN + P +V +D T+ + N+
Sbjct: 773 SRDVRFHEDIYPY-------ATATQSNVP-----LPPPTPPMVNDDWFLPISTQVDSTNV 820
Query: 800 ELLEHMVFETGGEEGETLIKNPELRVYVRKKFHKDGTNPIVFPAEVQSDSPSEGPTDNPS 859
+ G I P + + TNP+ P E+ S PS P+ +
Sbjct: 821 DSSSSSSPAQSGS-----IDQPPRSI---DQSPSTSTNPV--PEEIGSIVPSSSPSRSID 870
Query: 860 SSSSGNSSHSSNDL---PDLSFPDI-NLPIAVRKN---------IPDLNIPIAERKRTRT 906
S+S S+ + +L + S P LP + K + D +K+T +
Sbjct: 871 RSTSDLSASDTTELLSTGESSTPSSPGLPELLGKGCREKKKSVLLKDFVTNTTSKKKTAS 930
Query: 907 CTKH-------------------------PMSNYLSYDKLSHSHKAYVSRISNLFVPRTI 941
H P+S++L+ S +H A+++ I + P+
Sbjct: 931 HNIHSPSQVLPSGLPTSLSADSVSGKTLYPLSDFLTNSGYSANHIAFMAAILDSNEPKHF 990
Query: 942 QEALGDPNWKLAVKEEMNALNKNNTWCITDLPYDKKAVGCKWVFTVKCKADGSVERYKAR 1001
++A+ W A+ +E++AL N+TW ITDLP+ KKA+ KWV+ +K +DG++ER+KAR
Sbjct: 991 KDAILIKEWCEAMSKEIDALEANHTWDITDLPHGKKAISSKWVYKLKYNSDGTLERHKAR 1050
Query: 1002 LVAKGFTQTHGIDYQETFAPVAKINSIRILLSLAVNFNWTLHQYDVKNAFLNGELHEEVY 1061
LV G Q G+D++ETFAPVAK+ ++R +L++A +W +HQ DV NAFL+G+L EEVY
Sbjct: 1051 LVVMGNHQKEGVDFKETFAPVAKLTTVRTILAVAAAKDWEVHQMDVHNAFLHGDLEEEVY 1110
Query: 1062 MRLPPGFEDKLGRGKVCRLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHS 1121
MRLPPGF+ KVCRL+KSLYGLKQ+PR WF + + ++ GFTQS D+++F +
Sbjct: 1111 MRLPPGFKCS-DPSKVCRLRKSLYGLKQAPRCWFSKLSTALRNIGFTQSYEDYSLFSLKN 1169
Query: 1122 REGKIAILIVYVDDIIMTGDDIGEISDLKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIF 1181
+ I +L VYVDD+I+ G+++ I K +L F +KDLGKLKYFLG+E +R +G
Sbjct: 1170 GDTIIHVL-VYVDDLIVAGNNLDAIDRFKSQLHKCFHMKDLGKLKYFLGLEVSRGPDGFC 1228
Query: 1182 LNQRKYILDLLTETGMTGCKAAETPMDPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTR 1241
L+QRKY LD++ ETG+ GCK + P+ N KL S+ + E+Y+RL GR IYL+ TR
Sbjct: 1229 LSQRKYALDIVKETGLLGCKPSAVPIALNHKLASITGPVFTNPEQYRRLVGRFIYLTITR 1288
Query: 1242 PDIAFAVSVISQFMHAPGPAHFEAVFRILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAG 1301
PD+++AV ++SQFM AP AH+EA R++RYLKG+P +G+ R+ + + AY D+D+
Sbjct: 1289 PDLSYAVHILSQFMQAPLVAHWEAALRLVRYLKGSPAQGIFLRSDSSLIINAYCDSDYNA 1348
Query: 1302 NINDRRSTSGYCTFVGGNLVTWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELK 1361
RRS S Y ++G + ++W++KKQ+ V+ SSAEAE+R++A+ E+ W+K +++L
Sbjct: 1349 CPLTRRSLSAYVVYLGDSPISWKTKKQDTVSYSSAEAEYRAMAYTLKELKWLKALLKDLG 1408
Query: 1362 IAGPTPMKVYCDNKAAISIAHNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVA 1421
+ +PMK++CD++AAI IA NPV H+RTKH+E D H +++ + I +I T+ QVA
Sbjct: 1409 VHHSSPMKLHCDSEAAIHIAANPVFHERTKHIESDCHKVRDAVLDKLITTEHIYTEDQVA 1468
Query: 1422 DVLTKSLPKRQFD 1434
D+LTKSLP+ F+
Sbjct: 1469 DLLTKSLPRPTFE 1481
>At1g70010 hypothetical protein
Length = 1315
Score = 827 bits (2137), Expect = 0.0
Identities = 503/1443 (34%), Positives = 755/1443 (51%), Gaps = 179/1443 (12%)
Query: 22 IRGRGKIGYITGDKKQPDKVGADYDTWDAENSMVMTWLVNSLTEEIGANYLCYATAKDLW 81
I + K+G++ G +PD W NSMV +WL+NS+++EI + L + TA +W
Sbjct: 5 IEAKNKLGFVDGSIPKPDDDDPYCKIWRRCNSMVKSWLLNSVSKEIYTSILYFPTAAAIW 64
Query: 82 ENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGCLKRIWQDLDLFNEYEWKSPED 141
+++ + + ++Y+L Q+ +RQ ++ Y + +W++L P
Sbjct: 65 KDLYTRFHK-SSLPRLYKLRQQIHSLRQGNLDLSSYHTRTQTLWEELTSLQAV----PRT 119
Query: 142 CKHYQKMVDVSRVFKFLAGLNVEFDEVRGRILGRNPIPPIGEVFAEVRREESRRQVMLGK 201
+ + +RV FL GLN +D VR +IL + +P + EVF + ++E++R +
Sbjct: 120 VEDLLIERETNRVIDFLMGLNDCYDTVRSQILMKKTLPSLSEVFNMIDQDETQRSARIST 179
Query: 202 KKAAVPPPVEGSALAVPQVNHKSFRNPRGGD----KTHLVCDYCGRNRHTQETCFKLHGR 257
G +V V+++S ++ GD K VC YC R H ++TC+K HG
Sbjct: 180 --------TPGMTSSVFPVSNQSSQSALNGDTYQKKERPVCSYCSRPGHVEDTCYKKHGY 231
Query: 258 PNNSKAGK------------FGDRPMPTTSNAASSPFTKEQMDHLLKLLKFN-SSPNTPI 304
P + K+ + G + ++ ++ T Q+ L+ L P+TP
Sbjct: 232 PTSFKSKQKFVKPSISANAAIGSEEVVNNTSVSTGDLTTSQIQQLVSFLSSKLQPPSTP- 290
Query: 305 GTVAQTGKDSWALSVQNHSNPWIIDSGASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSS 364
VQ + + S S T C P SGS
Sbjct: 291 --------------VQPEVHSISVSSDPSSSSTVC--------PISGS------------ 316
Query: 365 IAGKGNIKISEQITLQSVLHVPKFACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTI 424
+ + + L VL +P+F NLLSV L++ C + F ++CV QD +
Sbjct: 317 ------VHLGRHLILNDVLFIPQFKFNLLSVSSLTKSMGCRIWFDETSCVLQDATRELMV 370
Query: 425 GTAREINGLYYFDETPLGNTAMFGSSRTSPPLVSNKIMLWHKRLGHPSFSYLKHLFP--E 482
G +++ LY D L + SS T + S+ LWHKRLGHPS L+ +
Sbjct: 371 GMGKQVANLYIVDLDSLSHPGT-DSSITVASVTSHD--LWHKRLGHPSVQKLQPMSSLLS 427
Query: 483 FSKEISSSQFHCDACHLAKDHRVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSRKKWFVT 542
F K+ +++ FHC CH++K + F S +S+PF LIH D WGP ++T ++F+T
Sbjct: 428 FPKQKNNTDFHCRVCHISKQKHLPFVSHNNKSSRPFDLIHIDTWGPFSVQTHDGYRYFLT 487
Query: 543 FIDDHTRVCWVYLMEKKSEVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLV 602
+DD++R WVYL+ KS+V F M++NQF TTI +RSDN E + F
Sbjct: 488 IVDDYSRATWVYLLRNKSDVLTVIPTFVTMVENQFETTIKGVRSDNAPEL---NFTQFYH 544
Query: 603 TNGIIHQSTCRDTPQQNGIAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACHLINRMP 662
+ GI+ +C +TPQQN + ERK++H+L V R++ F ++P WG+ +LTA +LINR+P
Sbjct: 545 SKGIVPYHSCPETPQQNSVVERKHQHILNVARSLFFQSHIPISYWGDCILTAVYLINRLP 604
Query: 663 SRVLQYETPVQVLQNNFPTSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYA 722
+ +L+ + P +VL PT I KVFGCLCY R K P+A+ C F+GY
Sbjct: 605 APILEDKCPFEVLTKTVPTYDHI-----KVFGCLCYASTSPKDRHKFSPRAKACAFIGYP 659
Query: 723 SNKKGYKCFNPVTKKFFESMDVHFVEDQ-PFFRENSLQGESQSSNEEDNFWEILPVLDDI 781
S KGYK + T S V F E+ PF S S EE NF+
Sbjct: 660 SGFKGYKLLDLETHSIIVSRHVVFHEELFPFL-------GSDLSQEEQNFFP-------- 704
Query: 782 VTNDHLDAKITEPRNPNLELLEHMVFETGGEEGETLIKNPELRVYVRKKFHKDGTNPIVF 841
D T P ++ D NP
Sbjct: 705 ------DLNPTPP---------------------------------MQRQSSDHVNPSDS 725
Query: 842 PAEVQSDSPSEGPTDN---PSSSSSGNSSHSSNDLPDLSFPDI--NLPIAVRK--NIPDL 894
+ V+ PS PT+N PS +S + L D + + P +RK + +
Sbjct: 726 SSSVEI-LPSANPTNNVPEPSVQTSHRKAKKPAYLQDYYCHSVVSSTPHEIRKFLSYDRI 784
Query: 895 NIPIAERKRTRTCTKHPMSNYLSYDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAV 954
N P TK P SNY +KL W+ A+
Sbjct: 785 NDPYLTFLACLDKTKEP-SNYTEAEKLQ--------------------------VWRDAM 817
Query: 955 KEEMNALNKNNTWCITDLPYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGID 1014
E + L +TW + LP DK+ +GC+W+F +K +DGSVERYKARLVA+G+TQ GID
Sbjct: 818 GAEFDFLEGTHTWEVCSLPADKRCIGCRWIFKIKYNSDGSVERYKARLVAQGYTQKEGID 877
Query: 1015 YQETFAPVAKINSIRILLSLAVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFE----D 1070
Y ETF+PVAK+NS+++LL +A F +L Q D+ NAFLNG+L EE+YMRLP G+ D
Sbjct: 878 YNETFSPVAKLNSVKLLLGVAARFKLSLTQLDISNAFLNGDLDEEIYMRLPQGYASRQGD 937
Query: 1071 KLGRGKVCRLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILI 1130
L VCRLKKSLYGLKQ+ R W+ +F S + G GF QS DHT F K S +G ++
Sbjct: 938 SLPPNAVCRLKKSLYGLKQASRQWYLKFSSTLLGLGFIQSYCDHTCFLKIS-DGIFLCVL 996
Query: 1131 VYVDDIIMTGDDIGEISDLKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILD 1190
VY+DDII+ ++ + LK ++++ F ++DLG+LKYFLG+E RS +GI ++QRKY LD
Sbjct: 997 VYIDDIIIASNNDAAVDILKSQMKSFFKLRDLGELKYFLGLEIVRSDKGIHISQRKYALD 1056
Query: 1191 LLTETGMTGCKAAETPMDPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSV 1250
LL ETG GCK + PMDP++ + + ++ Y+RL GRL+YL+ TRPDI FAV+
Sbjct: 1057 LLDETGQLGCKPSSIPMDPSMVFAHDSGGDFVEVGPYRRLIGRLMYLNITRPDITFAVNK 1116
Query: 1251 ISQFMHAPGPAHFEAVFRILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTS 1310
++QF AP AH +AV++IL+Y+KGT G+GL + +Q++ Y +AD+ + RRSTS
Sbjct: 1117 LAQFSMAPRKAHLQAVYKILQYIKGTIGQGLFYSATSELQLKVYANADYNSCRDSRRSTS 1176
Query: 1311 GYCTFVGGNLVTWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKV 1370
GYC F+G +L+ W+S+KQ+VV++SSAEAE+RS++ E++W+ F++EL++ P +
Sbjct: 1177 GYCMFLGDSLICWKSRKQDVVSKSSAEAEYRSLSVATDELVWLTNFLKELQVPLSKPTLL 1236
Query: 1371 YCDNKAAISIAHNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVADVLTKSLPK 1430
+CDN+AAI IA+N V H+RTKH+E D H ++E++ G + +I T+ Q+AD TK L
Sbjct: 1237 FCDNEAAIHIANNHVFHERTKHIESDCHSVRERLLKGLFELYHINTELQIADPFTKPLYP 1296
Query: 1431 RQF 1433
F
Sbjct: 1297 SHF 1299
>At1g57640
Length = 1444
Score = 785 bits (2026), Expect = 0.0
Identities = 493/1493 (33%), Positives = 776/1493 (51%), Gaps = 177/1493 (11%)
Query: 10 NYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGADYDTWDAENSMVMTWLVNSLTEEIGA 69
NY W+ + +R R K G++ G QP D + W N+++++W+ ++ E+
Sbjct: 42 NYEEWACGFKTALRSRKKFGFLDGTIPQPLDGSPDLEDWLTINALLVSWMKMTIDSELLT 101
Query: 70 NYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGCLKRIWQDLD 129
N A+DLWE + + +S + N + ++ L +Q +V Y+G L +IW +++
Sbjct: 102 NISHRDVARDLWEQIRKRFS-VSNGPKNQKMKADLATCKQEGMTVEGYYGKLNKIWDNIN 160
Query: 130 LFNEYEWKSPEDC-----KHYQKMVDVSRVFKFLAGLN-VEFDEVRGRILGRNPIPPIGE 183
+ C +K + V ++L GLN +F +R + R P+P + E
Sbjct: 161 SYRPLRICKCGRCICNLGTDQEKYREDDMVHQYLYGLNETKFHTIRSSLTSRVPLPGLEE 220
Query: 184 VFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVP-----QVNHKSFRNPRGGDKTHLVC 238
V+ VR+EE + +++ + +A AV +V + F N L C
Sbjct: 221 VYNIVRQEED-----MVNNRSSNEERTDVTAFAVQMRPRSEVISEKFANSEKLQNKKL-C 274
Query: 239 DYCGRNRHTQETCFKLHGRP----------NNSKA------GKFGD-----RPMPTTSNA 277
+C R H+ E CF L G P +NS G+FG +P PT N
Sbjct: 275 THCNRGGHSPENCFVLIGYPEWWGDRPRGKSNSNGSTSRGRGRFGPGFNGGQPRPTYVNV 334
Query: 278 ----------------------ASSPFTKEQMDHLLKLLKFNSSPNTPIGTVAQTGKDSW 315
A S T EQ ++KLL S N Q+G S
Sbjct: 335 VMTGPFPSSEHVNRVITDSDRDAVSGLTDEQWRGVVKLLNAGRSDNKSNAHETQSGTCSL 394
Query: 316 ALSVQNHSNPWIIDSGASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKGNIKISE 375
S WI+D+GAS HMT L S S + +ADG+ +G +++
Sbjct: 395 FTS-------WILDTGASHHMTGNLELLSD-MRSMSPVLIILADGNKRVAVSEGTVRLGS 446
Query: 376 QITLQSVLHVPKFACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYY 435
+ L+SV +V + +L+SV ++ ++++C
Sbjct: 447 HLILKSVFYVKELESDLISVGQMMDENHCV------------------------------ 476
Query: 436 FDETPLGNTAMFGSSRTSPPLVSNKIMLWHKRLGHPSFSYLKHLFPEF---SKEISSSQF 492
N A +S +P LWH+RLGH S + L E KEI +
Sbjct: 477 -------NAAAVHTSVKAP------FDLWHRRLGHASDKIVNLLPRELLSSGKEILENV- 522
Query: 493 HCDACHLAKDHRVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDHTRVCW 552
CD C AK R +F + F LIH DVWGP + + S ++F+T +DD++R W
Sbjct: 523 -CDTCMRAKQTRDTFPLSDNRSMDSFQLIHCDVWGPYRAPSYSGARYFLTIVDDYSRGVW 581
Query: 553 VYLMEKKSEVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLVTNGIIHQSTC 612
VYLM KSE ++ +DF +++ QF T I I+RSDNGTE+ + + + GI H+++C
Sbjct: 582 VYLMTDKSETQKHLKDFIALVERQFDTEIKIVRSDNGTEFL--CMREYFLHKGIAHETSC 639
Query: 613 RDTPQQNGIAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACHLINRMPSRVLQYETPV 672
TP QNG ERK+RH+L + RA+ F +P WG +L+A +LINR PS +LQ ++P
Sbjct: 640 VGTPHQNGRVERKHRHILNIARALRFQSYLPIQFWGECILSAAYLINRTPSMLLQGKSPY 699
Query: 673 QVLQNNFPTSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYKCFN 732
++L P L+VFG LCY + N K ++ +CVF+GY +KG++ F+
Sbjct: 700 EMLYKTAPKYS-----HLRVFGSLCYAHNQNHKGDKFAARSRRCVFVGYPHGQKGWRLFD 754
Query: 733 PVTKKFFESMDVHFVEDQPFFRENSLQGESQSSNEEDNFWEILPVLDDIVTNDHLDAKIT 792
+KFF S DV F+E S NEED VL D V ++ I
Sbjct: 755 LEEQKFFVSRDV-------IFQETEFPYSKMSCNEEDE-----RVLVDCVGPPFIEEAIG 802
Query: 793 EPRNPNLELLEHMVFETGGEEGETLIKNPELRVYVRKKFHKDGTNPIVFPAEVQSDSPSE 852
PR G GE + P + T PI+ +S SPSE
Sbjct: 803 -PRTI-----------IGRNIGEATV-GPNV-----------ATGPIIPEINQESSSPSE 838
Query: 853 GPTDNPSSSSSGNSSHSSNDLPDLSFPDINLPIAVRKNIPDLNIPIAERK---------- 902
+ + +S+ + DLP S PI +R++ P+ +
Sbjct: 839 FVSLSSLDPFLASSTVQTADLPLSSTTPA--PIQLRRSSRQTQKPMKLKNFVTNTVSVES 896
Query: 903 ---RTRTCTKHPMSNYLSYDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEMN 959
+ + +P+ Y+ + + SHKA+++ ++ P T EA+ D W+ A+ E+
Sbjct: 897 ISPEASSSSLYPIEKYVDCHRFTSSHKAFLAAVTAGMEPTTYNEAMVDKAWREAMSAEIE 956
Query: 960 ALNKNNTWCITDLPYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETF 1019
+L N T+ I +LP K+A+G KWV+ +K ++DG++ERYKARLV G Q G+DY ETF
Sbjct: 957 SLRVNQTFSIVNLPPGKRALGNKWVYKIKYRSDGAIERYKARLVVLGNCQKEGVDYDETF 1016
Query: 1020 APVAKINSIRILLSLAVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLGRGKVCR 1079
APVAK++++R+ L +A +W +HQ DV NAFL+G+L EEVYM+LP GF+ KVCR
Sbjct: 1017 APVAKMSTVRLFLGVAAARDWHVHQMDVHNAFLHGDLKEEVYMKLPQGFQCD-DPSKVCR 1075
Query: 1080 LKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIMT 1139
L KSLYGLKQ+PR WF + S +K +GFTQS +D+++F ++ +G ++VYVDD+I++
Sbjct: 1076 LHKSLYGLKQAPRCWFSKLSSALKQYGFTQSLSDYSLF-SYNNDGIFVHVLVYVDDLIIS 1134
Query: 1140 GDDIGEISDLKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMTG 1199
G ++ K LE+ F +KDLG LKYFLG+E +R+ +G +L+QRKY+LD+++E G+ G
Sbjct: 1135 GSCPDAVAQFKSYLESCFHMKDLGLLKYFLGIEVSRNAQGFYLSQRKYVLDIISEMGLLG 1194
Query: 1200 CKAAETPMDPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHAPG 1259
+ + P++ N KL + D RY+RL GRLIYL TRP+++++V ++QFM P
Sbjct: 1195 ARPSAFPLEQNHKLSLSTSPLLSDSSRYRRLVGRLIYLVVTRPELSYSVHTLAQFMQNPR 1254
Query: 1260 PAHFEAVFRILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGGN 1319
H+ A R++RYLK PG+G++ + +Q+ + D+D+A RRS +GY +G
Sbjct: 1255 QDHWNAAIRVVRYLKSNPGQGILLSSTSTLQINGWCDSDYAACPLTRRSLTGYFVQLGDT 1314
Query: 1320 LVTWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAIS 1379
++W++KKQ V+RSSAEAE+R++A E++W+K+ + +L ++ M+++ D+K+AI+
Sbjct: 1315 PISWKTKKQPTVSRSSAEAEYRAMAFLTQELMWLKRVLYDLGVSHVQAMRIFSDSKSAIA 1374
Query: 1380 IAHNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVADVLTKSLPKRQ 1432
++ NPV H+RTKHVEVD HFI++ I G I S++P+ Q+AD+LTK+L +++
Sbjct: 1375 LSVNPVQHERTKHVEVDCHFIRDAILDGIIATSFVPSHKQLADILTKALGEKE 1427
>At2g16670 putative retroelement pol polyprotein
Length = 1333
Score = 722 bits (1864), Expect = 0.0
Identities = 457/1317 (34%), Positives = 692/1317 (51%), Gaps = 117/1317 (8%)
Query: 146 QKMVDVSRVFKFLAGLN-VEFDEVRGRILGRNPIPPIGEVFAEVRREESRRQVMLGKKKA 204
+K + +V +FL+GL+ F VR ++ R P+ P+ EV+ VR+EE +L
Sbjct: 89 EKDREEDKVHEFLSGLDDALFRTVRSSLVSRIPVQPLEEVYNIVRQEED----LLRNGAN 144
Query: 205 AVPPPVEGSALAVPQVNHKSFRNPRGGDKTH-LVCDYCGRNRHTQETCFKLHGRPNNSKA 263
+ E +A A Q+ K ++ RG +K +VC +C R+ H E+C+ + G P
Sbjct: 145 VLDDQREVNAFAA-QMRPKLYQG-RGDEKDKSMVCKHCNRSGHASESCYAVIGYPE---- 198
Query: 264 GKFGDRPMPTTSNAASSPFTKEQMDHLLKLLKFNSSPNTPI------GTVAQTGKDSWAL 317
+GDRP + T + + P A T +D +
Sbjct: 199 -WWGDRPRSRSLQTRGRGGTNSSGGRGRGAAAYANRVTVPNHDTYEQANYALTDEDRDGV 257
Query: 318 SVQNHSNPW-----IIDSG---ASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKG 369
+ S W I++SG A+E +T+ + + +ADG +G
Sbjct: 258 NGLTDSQ-WRTIKSILNSGKDAATEKLTDMDPIL-----------IVLADGRERISVKEG 305
Query: 370 NIKISEQITLQSVLHVPKFACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTARE 429
+++ + + SV +V +F +L+S+ +L +++ C + V QD+ S +G R
Sbjct: 306 TVRLGSNLVMISVFYVEEFQSDLISIGQLMDENRCVLQMSDRFLVVQDRTSRMVMGAGRR 365
Query: 430 INGLYYFDETPLGNTAMFGSSRTSPPLVSNKIMLWHKRLGHPSFSYLKHLFPEFSKEISS 489
+ G ++F T + + + LWH R+GHP+ + L PE S +SS
Sbjct: 366 VGGTFHFRSTEIAASVTVKEEKNYE--------LWHSRMGHPAARVVS-LIPESSVSVSS 416
Query: 490 SQFH--CDACHLAKDHRVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDH 547
+ + CD CH AK R SF + F LI+ D+WGP + + + ++F+T IDD+
Sbjct: 417 THLNKACDVCHRAKQTRNSFPLSINKTLRIFELIYCDLWGPYRTPSHTGARYFLTIIDDY 476
Query: 548 TRVCWVYLMEKKSEVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLVTNGII 607
+R W+YL+ KSE ++FF M QF+ I +RSDNGTE+ L+ F G+I
Sbjct: 477 SRGVWLYLLNDKSEAPCHLKNFFAMTDRQFNVKIKTVRSDNGTEFL--CLTKFFQEQGVI 534
Query: 608 HQSTCRDTPQQNGIAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACHLINRMPSRVLQ 667
H+ +C TP++N ERK+RHLL V RA+ F N+P WG +LTA +LINR PS VL
Sbjct: 535 HERSCVATPERNDRVERKHRHLLNVARALRFQANLPIQFWGECVLTAAYLINRTPSSVLN 594
Query: 668 YETPVQVLQNNFPTSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKG 727
TP + L P L+VFG LCY + N K ++ +CVF+GY +KG
Sbjct: 595 DSTPYERLHKKQPRFD-----HLRVFGSLCYAHNRNRGGDKFAERSRRCVFVGYPHGQKG 649
Query: 728 YKCFNPVTKKFFESMDVHFVEDQPFFRENSLQGESQSSNEEDNFWEILPVLDDIVTND-H 786
++ F+ +FF S DV F E + FR + Q + EE+ W P++D ++ + H
Sbjct: 650 WRLFDLEQNEFFVSRDVVFSELEFPFRISHEQNVIEE--EEEALWA--PIVDGLIEEEVH 705
Query: 787 LDAK--------ITEPRNPNL--ELLEHMVFETGGEEGETLIKNPELRVYVRKKFHKDGT 836
L ++ P +P+ EH T ++ P T
Sbjct: 706 LGQNAGPTPPICVSSPISPSATSSRSEH---STSSPLDTEVVPTPATST-TSASSPSSPT 761
Query: 837 NPIVFPAEVQSDSPSEGPTDNPSSSSSGNSSHSSNDLPDLSFPDINLPIAVRKNIPDLNI 896
N P + P+ P + + N P ++ D + V + P
Sbjct: 762 NLQFLP--LSRAKPTTAQAVAPPAVPPPRRQSTRNKAPPVTLKDFVVNTTVCQESPSKLN 819
Query: 897 PIAERKRTRTCTKHPMSNYLSYDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKE 956
I + + R T+ + S SH YV+
Sbjct: 820 SILYQLQKRDDTR----------RFSASHTTYVA-------------------------- 843
Query: 957 EMNALNKNNTWCITDLPYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQ 1016
++A +N+TW I DLP K+A+G +WV+ VK +DGSVERYKARLVA G Q G DY
Sbjct: 844 -IDAQEENHTWTIEDLPPGKRAIGSQWVYKVKHNSDGSVERYKARLVALGNKQKEGEDYG 902
Query: 1017 ETFAPVAKINSIRILLSLAVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLGRGK 1076
ETFAPVAK+ ++R+ L +AV NW +HQ DV NAFL+G+L EEVYM+LPPGFE K
Sbjct: 903 ETFAPVAKMATVRLFLDVAVKRNWEIHQMDVHNAFLHGDLREEVYMKLPPGFEAS-HPNK 961
Query: 1077 VCRLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDI 1136
VCRL+K+LYGLKQ+PR WFE+ + +K +GF QS AD+++F +I ILI YVDD+
Sbjct: 962 VCRLRKALYGLKQAPRCWFEKLTTALKRYGFQQSLADYSLFTLVKGSVRIKILI-YVDDL 1020
Query: 1137 IMTGDDIGEISDLKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETG 1196
I+TG+ K L + F +KDLG LKYFLG+E ARS GI++ QRKY LD+++ETG
Sbjct: 1021 IITGNSQRATQQFKEYLASCFHMKDLGPLKYFLGIEVARSTTGIYICQRKYALDIISETG 1080
Query: 1197 MTGCKAAETPMDPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMH 1256
+ G K A P++ N KL + D +RY+RL GRLIYL+ TR D+AF+V ++++FM
Sbjct: 1081 LLGVKPANFPLEQNHKLGLSTSPLLTDPQRYRRLVGRLIYLAVTRLDLAFSVHILARFMQ 1140
Query: 1257 APGPAHFEAVFRILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFV 1316
P H+ A R++RYLK PG+G+ R G Q+ + D+DWAG+ RRS +GY
Sbjct: 1141 EPREDHWAAALRVVRYLKADPGQGVFLRRSGDFQITGWCDSDWAGDPMSRRSVTGYFVQF 1200
Query: 1317 GGNLVTWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKA 1376
G + ++W++KKQ+ V++SSAEAE+R+++ E+LW+K+ + L ++ PM + CD+K+
Sbjct: 1201 GDSPISWKTKKQDTVSKSSAEAEYRAMSFLASELLWLKQLLFSLGVSHVQPMIMCCDSKS 1260
Query: 1377 AISIAHNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVADVLTKSLPKRQF 1433
AI IA NPV H+RTKH+E+D HF++++ G I ++ T SQ+AD+ TK L + F
Sbjct: 1261 AIYIATNPVFHERTKHIEIDYHFVRDEFVKGVITPRHVGTTSQLADIFTKPLGRDCF 1317
>At1g59265 polyprotein, putative
Length = 1466
Score = 702 bits (1813), Expect = 0.0
Identities = 471/1480 (31%), Positives = 735/1480 (48%), Gaps = 100/1480 (6%)
Query: 5 RLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPD---------KVGADYDTWDAENSMV 55
+L NY WS+ V G G++ G P +V DY W ++ ++
Sbjct: 25 KLTSTNYLMWSRQVHALFDGYELAGFLDGSTTMPPATIGTDAAPRVNPDYTRWKRQDKLI 84
Query: 56 MTWLVNSLTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVT 115
+ ++ +++ + TA +WE + ++Y++ + + +L QL + + ++
Sbjct: 85 YSAVLGAISMSVQPAVSRATTAAQIWETLRKIYAN-PSYGHVTQLRTQLKQWTKGTKTID 143
Query: 116 KYFGCLKRIWQDLDLFNEYEWKSPEDCKHYQKMVDVSRVFKFLAGLNVEFDEVRGRILGR 175
Y L + L L + M +V + L L E+ V +I +
Sbjct: 144 DYMQGLVTRFDQLALLGK-------------PMDHDEQVERVLENLPEEYKPVIDQIAAK 190
Query: 176 NPIPPIGEVFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVPQVNHKSFRNPRGGDKTH 235
+ P + E+ + ES+ +L A V P +A AV N + N G++ +
Sbjct: 191 DTPPTLTEIHERLLNHESK---ILAVSSATVIPI---TANAVSHRNTTTTNNNNNGNRNN 244
Query: 236 LVCDYCGRNRHT-----QETCFKLHGRPNNSKAGKFGDRPMPTTSNAASSPFTKEQMDHL 290
Y RN + Q++ H PNN+++ + + S+ Q+ H
Sbjct: 245 R---YDNRNNNNNSKPWQQSSTNFH--PNNNQSKPYLGKCQICGVQGHSAKRCS-QLQHF 298
Query: 291 LKLLKFNS--SPNTPIGTVAQTGKDSWALSVQNHSNPWIIDSGASEHMTNCSHLFSSYFP 348
L + SP TP A AL SN W++DSGA+ H+T+ + S + P
Sbjct: 299 LSSVNSQQPPSPFTPWQPRANL-----ALGSPYSSNNWLLDSGATHHITSDFNNLSLHQP 353
Query: 349 SSGSEKVKIADGSYSSIAGKGNIKISEQ---ITLQSVLHVPKFACNLLSVHKLSEDSNCS 405
+G + V +ADGS I+ G+ +S + + L ++L+VP NL+SV++L + S
Sbjct: 354 YTGGDDVMVADGSTIPISHTGSTSLSTKSRPLNLHNILYVPNIHKNLISVYRLCNANGVS 413
Query: 406 VLFCRSTCVFQDQNSGKTIGTAREINGLYYFDETPLGNTAMFGSSRTSPPLVSNKIMLWH 465
V F ++ +D N+G + + + LY + ++F S + S WH
Sbjct: 414 VEFFPASFQVKDLNTGVPLLQGKTKDELYEWPIASSQPVSLFASPSSKATHSS-----WH 468
Query: 466 KRLGHPSFSYLKHLFPEFSKEI---SSSQFHCDACHLAKDHRVSFNSRGYSASKPFYLIH 522
RLGHP+ S L + +S + S C C + K ++V F+ ++++P I+
Sbjct: 469 ARLGHPAPSILNSVISNYSLSVLNPSHKFLSCSDCLINKSNKVPFSQSTINSTRPLEYIY 528
Query: 523 SDVWGPSKIKTLSRKKWFVTFIDDHTRVCWVYLMEKKSEVEQRFQDFFNMIKNQFHTTIG 582
SDVW S I + +++V F+D TR W+Y +++KS+V++ F F N+++N+F T IG
Sbjct: 529 SDVWS-SPILSHDNYRYYVIFVDHFTRYTWLYPLKQKSQVKETFITFKNLLENRFQTRIG 587
Query: 583 ILRSDNGTEYFNKYLSTFLVTNGIIHQSTCRDTPQQNGIAERKNRHLLEVTRAIMFSMNV 642
SDNG E+ L + +GI H ++ TP+ NG++ERK+RH++E ++ ++
Sbjct: 588 TFYSDNGGEFVA--LWEYFSQHGISHLTSPPHTPEHNGLSERKHRHIVETGLTLLSHASI 645
Query: 643 PKYLWGNALLTACHLINRMPSRVLQYETPVQVLQNNFPTSRIITNIPLKVFGCLCYVYIP 702
PK W A A +LINR+P+ +LQ E+P Q L P L+VFGC CY ++
Sbjct: 646 PKTYWPYAFAVAVYLINRLPTPLLQLESPFQKLFGTSPNYD-----KLRVFGCACYPWLR 700
Query: 703 NIFRSKLDPKAEKCVFLGYASNKKGYKCFNPVTKKFFESMDVHFVEDQ-PFFRENSLQGE 761
+ KLD K+ +CVFLGY+ + Y C + T + + S V F E+ PF +
Sbjct: 701 PYNQHKLDDKSRQCVFLGYSLTQSAYLCLHLQTSRLYISRHVRFDENCFPFSNYLATLSP 760
Query: 762 SQSSNEEDN-FWEI-------LPVLDDIVTNDHLDAKITEPRNPNLELLEHMVFETGGEE 813
Q E + W PVL +D A T P +P+ V + +
Sbjct: 761 VQEQRRESSCVWSPHTTLPTRTPVLPAPSCSDPHHAA-TPPSSPSAPFRNSQVSSSNLDS 819
Query: 814 G--ETLIKNPELRVYVRKKFHKDGTNPIVFPAEVQSDSPS-----------EGPTDNPSS 860
+ +PE ++G P P + Q+ + S E P+ S
Sbjct: 820 SFSSSFPSSPEPTAP-----RQNGPQPTTQPTQTQTQTHSSQNTSQNNPTNESPSQLAQS 874
Query: 861 SSSGNSSHSSNDLPDLSFPDINL---PIAVRKNIPDLNIPIAERKRTRTCTKHPMSNYLS 917
S+ S SS+ P S + P ++ + P I H M
Sbjct: 875 LSTPAQSSSSSPSPTTSASSSSTSPTPPSILIHPPPPLAQIVNNNNQAPLNTHSMGTRAK 934
Query: 918 YDKLSHSHKAYVS-RISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTW-CITDLPYD 975
+ + K ++ ++ PRT +AL D W+ A+ E+NA N+TW + P
Sbjct: 935 AGIIKPNPKYSLAVSLAAESEPRTAIQALKDERWRNAMGSEINAQIGNHTWDLVPPPPSH 994
Query: 976 KKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILLSLA 1035
VGC+W+FT K +DGS+ RYKARLVAKG+ Q G+DY ETF+PV K SIRI+L +A
Sbjct: 995 VTIVGCRWIFTKKYNSDGSLNRYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVLGVA 1054
Query: 1036 VNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLGRGKVCRLKKSLYGLKQSPRAWF 1095
V+ +W + Q DV NAFL G L ++VYM PPGF DK VC+L+K+LYGLKQ+PRAW+
Sbjct: 1055 VDRSWPIRQLDVNNAFLQGTLTDDVYMSQPPGFIDKDRPNYVCKLRKALYGLKQAPRAWY 1114
Query: 1096 ERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIMTGDDIGEISDLKRRLEA 1155
+ + GF S +D T F R I ++VYVDDI++TG+D + + L
Sbjct: 1115 VELRNYLLTIGFVNSVSD-TSLFVLQRGKSIVYMLVYVDDILITGNDPTLLHNTLDNLSQ 1173
Query: 1156 EFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMTGCKAAETPMDPNVKLKS 1215
F +KD +L YFLG+E R G+ L+QR+YILDLL T M K TPM P+ KL
Sbjct: 1174 RFSVKDHEELHYFLGIEAKRVPTGLHLSQRRYILDLLARTNMITAKPVTTPMAPSPKLSL 1233
Query: 1216 VAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHAPGPAHFEAVFRILRYLKG 1275
+ ++ D Y+ + G L YL+ TRPDI++AV+ +SQFMH P H +A+ RILRYL G
Sbjct: 1234 YSGTKLTDPTEYRGIVGSLQYLAFTRPDISYAVNRLSQFMHMPTEEHLQALKRILRYLAG 1293
Query: 1276 TPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNVVARSS 1335
TP G+ + + + AY+DADWAG+ +D ST+GY ++G + ++W SKKQ V RSS
Sbjct: 1294 TPNHGIFLKKGNTLSLHAYSDADWAGDKDDYVSTNGYIVYLGHHPISWSSKKQKGVVRSS 1353
Query: 1336 AEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAISIAHNPVLHDRTKHVEV 1395
EAE+RSVA+ E+ WI + EL I P +YCDN A + NPV H R KH+ +
Sbjct: 1354 TEAEYRSVANTSSEMQWICSLLTELGIRLTRPPVIYCDNVGATYLCANPVFHSRMKHIAI 1413
Query: 1396 DKHFIKEKIDSGEICMSYIPTKSQVADVLTKSLPKRQFDD 1435
D HFI+ ++ SG + + ++ T Q+AD LTK L + F +
Sbjct: 1414 DYHFIRNQVQSGALRVVHVSTHDQLADTLTKPLSRTAFQN 1453
>At1g44510 polyprotein, putative
Length = 1459
Score = 702 bits (1812), Expect = 0.0
Identities = 470/1481 (31%), Positives = 726/1481 (48%), Gaps = 120/1481 (8%)
Query: 5 RLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKV---------GADYDTWDAENSMV 55
+L NY WS + + G GY+ P + + W ++ ++
Sbjct: 32 KLTSTNYLMWSIQIHALLDGYDLAGYLDNSVVIPPETTTINSVVSANPSFTLWKRQDKLI 91
Query: 56 MTWLVNSLTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVT 115
+ L+ +++ + + + +W ++ Y+ + I +L Q+ + + ++
Sbjct: 92 FSALIGAISPAVQSLVSRATNSSQIWSTLNNTYAK-PSYGHIKQLRQQIQRLTKGTKTID 150
Query: 116 KYFGCLKRIWQDLDLFNEYEWKSPEDCKHYQKMVDVSRVFKFLAGLNVEFDEVRGRILGR 175
+Y L + + M +V L GL E+ V +I G+
Sbjct: 151 EYVQSHTTRLDQLAILGK-------------PMEHEEQVEHILKGLPEEYKTVVDQIEGK 197
Query: 176 NPIPPIGEVFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVPQVNHKSFRNPRGGDKTH 235
+ P I E+ + ES K + PP ++ V ++F N
Sbjct: 198 DNTPTITEIHERLINHES-------KLLSDEVPPSSSFPMSANAVQQRNFNNN------- 243
Query: 236 LVCDYCGRNRHTQETCFKLHGRPNNSKAGKFGDRPMPTTSNAASSPFTKEQMD------- 288
C +N+H H N+ + P+T N + K +
Sbjct: 244 -----CNQNQHKNRYQGNTHNNNTNTNS-------QPSTYNKSGQRTFKPYLGKCQICSV 291
Query: 289 --HLLKLLKFNSSPNTPIGTVAQTGKDSW------ALSVQNHSNPWIIDSGASEHMTNCS 340
H + + P + A + W A+ +NPW++DSGA+ H+T+
Sbjct: 292 QGHSARRCPQLQAMQLPASSSAHSPFTPWQPRANLAIGSPYAANPWLLDSGATHHITSDL 351
Query: 341 HLFSSYFPSSGSEKVKIADGSYSSIAGKGNIKISEQ---ITLQSVLHVPKFACNLLSVHK 397
+ S + P +G E V IADG+ +I G+ + Q + L VL+VP NL+SV++
Sbjct: 352 NALSLHQPYNGGEYVMIADGTGLTIKQTGSTFLPSQNRDLALHKVLYVPDIRKNLISVYR 411
Query: 398 LSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYYFDETPLGNTAMFGSSRTSPPLV 457
L + SV F ++ +D N+G + R + LY + T TA+F S L
Sbjct: 412 LCNTNQVSVEFFPASFQVKDLNTGTLLLQGRTKDDLYEWPVTNPPATALFTSPSPKTTLS 471
Query: 458 SNKIMLWHKRLGHPSFSYLKHLFPEFSKEIS---SSQFHCDACHLAKDHRVSFNSRGYSA 514
S WH RLGHPS S L L +FS +S S++ C C + K H++ F + +
Sbjct: 472 S-----WHSRLGHPSASILNTLLSKFSLPVSVASSNKTSCSDCLINKSHKLPFATSSIHS 526
Query: 515 SKPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDHTRVCWVYLMEKKSEVEQRFQDFFNMIK 574
S P I +DVW S I + K+++ +D +TR W+Y +++KS+V+ F F +++
Sbjct: 527 SSPLEYIFTDVW-TSPIISHDNYKYYLVLVDHYTRYTWLYPLQQKSQVKATFIAFKALVE 585
Query: 575 NQFHTTIGILRSDNGTEYFNKYLSTFLVTNGIIHQSTCRDTPQQNGIAERKNRHLLEVTR 634
N+F I L SDNG E+ L FLV+NGI H ++ TP+ NG++ERK+RH++E
Sbjct: 586 NRFQAKIRTLYSDNGGEFIA--LRDFLVSNGISHLTSPPHTPEHNGLSERKHRHIVETGL 643
Query: 635 AIMFSMNVPKYLWGNALLTACHLINRMPSRVLQYETPVQVLQNNFPTSRIITNIPLKVFG 694
++ +VP+ W A TA +LINRMP+ VL ++P Q L + P + L+VFG
Sbjct: 644 TLLTQASVPREYWTYAFATAVYLINRMPTPVLCLQSPFQKLFGSSPNYQ-----RLRVFG 698
Query: 695 CLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYKCFNPVTKKFFESMDVHFVEDQ-PF- 752
CLC+ ++ R+KL+ ++++CVFLGY+ + Y C + + + S V F E PF
Sbjct: 699 CLCFPWLRPYTRNKLEERSKRCVFLGYSLTQTAYLCLDVDNNRLYTSRHVMFDESTYPFA 758
Query: 753 --FRENSLQG-----ESQSSNEEDNFWEILPVLDDIVTNDHLDAKITEPRNPNLELLEHM 805
RE S ES SS+ N VL A E +P + +
Sbjct: 759 ASIREQSQSSLVTPPESSSSSSPANSGFPCSVL----RLQSPPASSPETPSPPQQQNDSP 814
Query: 806 VF--ETGGE--------EGETLIKNPELRVYVRKKFHKDGTNPIVFPAEVQSDSPSEGPT 855
V +TG TL +P + H++G P A+ +SP GP
Sbjct: 815 VSPRQTGSPTPSHHSQVRDSTLSPSPSVSNSEPTAPHENGPEP---EAQSNPNSPFIGPL 871
Query: 856 DNPSSSSSGNSSHSSNDLPD---LSFPDINLPIAVRKNIPDLNIPIAERKRTRTCTKHPM 912
NP+ ++ +SS + + P IA N + P + +T +K
Sbjct: 872 PNPNPETNPSSSIEQRPVDKSTTTALPPNQTTIAATSN--SRSQPPKNNHQMKTRSK--- 926
Query: 913 SNYLSYDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTWCITDL 972
N ++ K S +++ +L P T+ +AL D W+ A+ +E +A +N+TW +
Sbjct: 927 -NNITKPKTKTSLTVALTQ-PHLSEPNTVTQALKDKKWRFAMSDEFDAQQRNHTWDLVPP 984
Query: 973 PYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILL 1032
+ VGC+WVF +K +G +++YKARLVAKGF Q +G+DY ETF+PV K +IR++L
Sbjct: 985 NPTQHLVGCRWVFKLKYLPNGLIDKYKARLVAKGFNQQYGVDYAETFSPVIKATTIRVVL 1044
Query: 1033 SLAVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLGRGKVCRLKKSLYGLKQSPR 1092
+AV NW L Q DV NAFL G L EEVYM PPGF DK VCRL+K++YGLKQ+PR
Sbjct: 1045 DVAVKKNWPLKQLDVNNAFLQGTLTEEVYMAQPPGFVDKDRPSHVCRLRKAIYGLKQAPR 1104
Query: 1093 AWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIMTGDDIGEISDLKRR 1152
AW+ + GF S AD ++F +S + L+VYVDDII+TG D +S +
Sbjct: 1105 AWYMELKQHLLNIGFVNSLADTSLFI-YSHGTTLLYLLVYVDDIIVTGSDHKSVSAVLSS 1163
Query: 1153 LEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMTGCKAAETPMDPNVK 1212
L F IKD L YFLG+E R+ G+ L QRKY+ DLL + M K TP+ + K
Sbjct: 1164 LAERFSIKDPTDLHYFLGIEATRTNTGLHLMQRKYMTDLLAKHNMLDAKPVATPLPTSPK 1223
Query: 1213 LKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHAPGPAHFEAVFRILRY 1272
L ++ D Y+ + G L YL+ TRPDIAFAV+ +SQFMH P H++A R+LRY
Sbjct: 1224 LTLHGGTKLNDASEYRSVVGSLQYLAFTRPDIAFAVNRLSQFMHQPTSDHWQAAKRVLRY 1283
Query: 1273 LKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNVVA 1332
L GT G+ + I + A++DADWAG+ D ST+ Y ++G N ++W SKKQ V+
Sbjct: 1284 LAGTTTHGIFLNSSSPIHLHAFSDADWAGDSADYVSTNAYVIYLGRNPISWSSKKQRGVS 1343
Query: 1333 RSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAISIAHNPVLHDRTKH 1392
RSS E+E+R+VA+ E+ W+ + EL I P ++CDN A I NPV H R KH
Sbjct: 1344 RSSTESEYRAVANAASEIRWLCSLLTELHIRLPHGPTIFCDNIGATYICANPVFHSRMKH 1403
Query: 1393 VEVDKHFIKEKIDSGEICMSYIPTKSQVADVLTKSLPKRQF 1433
+ +D HF++ I S + +S++ T Q+AD LTKSL + F
Sbjct: 1404 IALDYHFVRGMIQSRALRVSHVSTNDQLADALTKSLSRPHF 1444
>At4g02960 putative polyprotein of LTR transposon
Length = 1456
Score = 693 bits (1789), Expect = 0.0
Identities = 460/1478 (31%), Positives = 727/1478 (49%), Gaps = 113/1478 (7%)
Query: 5 RLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPD---------KVGADYDTWDAENSMV 55
+L NY WS+ V G G++ G P +V DY W ++ ++
Sbjct: 25 KLTSTNYLMWSRQVHALFDGYELAGFLDGSTPMPPATIGTDAVPRVNPDYTRWRRQDKLI 84
Query: 56 MTWLVNSLTEEIGANYLCYATAKDLWENVSQMYSD--LGNQSQIYELTLQLGEIRQAEDS 113
+ ++ +++ + TA +WE + ++Y++ G+ +Q+ +T
Sbjct: 85 YSAILGAISMSVQPAVSRATTAAQIWETLRKIYANPSYGHVTQLRFITR----------- 133
Query: 114 VTKYFGCLKRIWQDLDLFNEYEWKSPEDCKHYQKMVDVSRVFKFLAGLNVEFDEVRGRIL 173
F L + + +D + E + L L ++ V +I
Sbjct: 134 ----FDQLALLGKPMDHDEQVE--------------------RVLENLPDDYKPVIDQIA 169
Query: 174 GRNPIPPIGEVFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVPQVNHKSFRNPRGGDK 233
++ P + E+ + ES+ +L A V P + + + N +N RG ++
Sbjct: 170 AKDTPPSLTEIHERLINRESK---LLALNSAEVVP-ITANVVTHRNTNTNRNQNNRGDNR 225
Query: 234 THLVCDYCGRNRHTQETCFKLHGRPNNSKAGKFGDRPMPTTSNAASSPFTKEQMDHLLKL 293
+ N + + ++ P++S + +P P +L
Sbjct: 226 NY-------NNNNNRSNSWQ----PSSSGSRSDNRQPKPYLGRCQICSVQGHSAKRCPQL 274
Query: 294 LKFNSSPNTPIGTVAQTG---KDSWALSVQNHSNPWIIDSGASEHMTNCSHLFSSYFPSS 350
+F S+ N T T + + A++ ++N W++DSGA+ H+T+ + S + P +
Sbjct: 275 HQFQSTTNQQQSTSPFTPWQPRANLAVNSPYNANNWLLDSGATHHITSDFNNLSFHQPYT 334
Query: 351 GSEKVKIADGSYSSIAGKGNIKI---SEQITLQSVLHVPKFACNLLSVHKLSEDSNCSVL 407
G + V IADGS I G+ + S + L VL+VP NL+SV++L + SV
Sbjct: 335 GGDDVMIADGSTIPITHTGSASLPTSSRSLDLNKVLYVPNIHKNLISVYRLCNTNRVSVE 394
Query: 408 FCRSTCVFQDQNSGKTIGTAREINGLYYFDETPLGNTAMFGSSRTSPPLVSNKIMLWHKR 467
F ++ +D N+G + + + LY + +MF S P WH R
Sbjct: 395 FFPASFQVKDLNTGVPLLQGKTKDELYEWPIASSQAVSMFAS-----PCSKATHSSWHSR 449
Query: 468 LGHPSFSYLKHLFPEFSKEI---SSSQFHCDACHLAKDHRVSFNSRGYSASKPFYLIHSD 524
LGHPS + L + S + S C C + K H+V F++ ++SKP I+SD
Sbjct: 450 LGHPSLAILNSVISNHSLPVLNPSHKLLSCSDCFINKSHKVPFSNSTITSSKPLEYIYSD 509
Query: 525 VWGPSKIKTLSRKKWFVTFIDDHTRVCWVYLMEKKSEVEQRFQDFFNMIKNQFHTTIGIL 584
VW S I ++ +++V F+D TR W+Y +++KS+V+ F F ++++N+F T IG L
Sbjct: 510 VWS-SPILSIDNYRYYVIFVDHFTRYTWLYPLKQKSQVKDTFIIFKSLVENRFQTRIGTL 568
Query: 585 RSDNGTEYFNKYLSTFLVTNGIIHQSTCRDTPQQNGIAERKNRHLLEVTRAIMFSMNVPK 644
SDNG E+ L +L +GI H ++ TP+ NG++ERK+RH++E+ ++ +VPK
Sbjct: 569 YSDNGGEFV--VLRDYLSQHGISHFTSPPHTPEHNGLSERKHRHIVEMGLTLLSHASVPK 626
Query: 645 YLWGNALLTACHLINRMPSRVLQYETPVQVLQNNFPTSRIITNIPLKVFGCLCYVYIPNI 704
W A A +LINR+P+ +LQ ++P Q L P LKVFGC CY ++
Sbjct: 627 TYWPYAFSVAVYLINRLPTPLLQLQSPFQKLFGQPPNYE-----KLKVFGCACYPWLRPY 681
Query: 705 FRSKLDPKAEKCVFLGYASNKKGYKCFNPVTKKFFESMDVHFVEDQ-PFFRENS--LQGE 761
R KL+ K+++C F+GY+ + Y C + T + + S V F E PF N +
Sbjct: 682 NRHKLEDKSKQCAFMGYSLTQSAYLCLHIPTGRLYTSRHVQFDERCFPFSTTNFGVSTSQ 741
Query: 762 SQSSNEEDNF--WEILPVLDDIVT-----NDHLDAKITEPRNPN---------LELLEHM 805
Q S+ N+ LP ++ HLD P +P+ L
Sbjct: 742 EQRSDSAPNWPSHTTLPTTPLVLPAPPCLGPHLDTSPRPPSSPSPLCTTQVSSSNLPSSS 801
Query: 806 VFETGGEEGETLIKN-PELRVYVRKKFHKDGTNPIVFPAEVQSDSPSEGPTDNPSSSSSG 864
+ E N P+ + + + +PI+ S SP+ ++P S
Sbjct: 802 ISSPSSSEPTAPSHNGPQPTAQPHQTQNSNSNSPILNNPNPNSPSPNSPNQNSPLPQSPI 861
Query: 865 NSSHSSNDLPDLSFPDINLPIAVRKNIPDLNI-----PIAERKRTRTCTKHPMSNYLSYD 919
+S H P S + N P + + P L PI + H M+
Sbjct: 862 SSPHIPT--PSTSISEPNSPSSSSTSTPPLPPVLPAPPIIQVNAQAPVNTHSMATRAKDG 919
Query: 920 KLSHSHK-AYVSRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTW-CITDLPYDKK 977
+ K +Y + ++ PRT +A+ D W+ A+ E+NA N+TW + P
Sbjct: 920 IRKPNQKYSYATSLAANSEPRTAIQAMKDDRWRQAMGSEINAQIGNHTWDLVPPPPPSVT 979
Query: 978 AVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILLSLAVN 1037
VGC+W+FT K +DGS+ RYKARLVAKG+ Q G+DY ETF+PV K SIRI+L +AV+
Sbjct: 980 IVGCRWIFTKKFNSDGSLNRYKARLVAKGYNQRPGLDYAETFSPVIKSTSIRIVLGVAVD 1039
Query: 1038 FNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLGRGKVCRLKKSLYGLKQSPRAWFER 1097
+W + Q DV NAFL G L +EVYM PPGF DK VCRL+K++YGLKQ+PRAW+
Sbjct: 1040 RSWPIRQLDVNNAFLQGTLTDEVYMSQPPGFVDKDRPDYVCRLRKAIYGLKQAPRAWYVE 1099
Query: 1098 FGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIMTGDDIGEISDLKRRLEAEF 1157
+ + GF S +D T F R I ++VYVDDI++TG+D + L F
Sbjct: 1100 LRTYLLTVGFVNSISD-TSLFVLQRGRSIIYMLVYVDDILITGNDTVLLKHTLDALSQRF 1158
Query: 1158 DIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMTGCKAAETPMDPNVKLKSVA 1217
+K+ L YFLG+E R +G+ L+QR+Y LDLL T M K TPM + KL +
Sbjct: 1159 SVKEHEDLHYFLGIEAKRVPQGLHLSQRRYTLDLLARTNMLTAKPVATPMATSPKLTLHS 1218
Query: 1218 EDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHAPGPAHFEAVFRILRYLKGTP 1277
++ D Y+ + G L YL+ TRPD+++AV+ +SQ+MH P H+ A+ R+LRYL GTP
Sbjct: 1219 GTKLPDPTEYRGIVGSLQYLAFTRPDLSYAVNRLSQYMHMPTDDHWNALKRVLRYLAGTP 1278
Query: 1278 GKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNVVARSSAE 1337
G+ + + + AY+DADWAG+ +D ST+GY ++G + ++W SKKQ V RSS E
Sbjct: 1279 DHGIFLKKGNTLSLHAYSDADWAGDTDDYVSTNGYIVYLGHHPISWSSKKQKGVVRSSTE 1338
Query: 1338 AEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAISIAHNPVLHDRTKHVEVDK 1397
AE+RSVA+ E+ WI + EL I P +YCDN A + NPV H R KH+ +D
Sbjct: 1339 AEYRSVANTSSELQWICSLLTELGIQLSHPPVIYCDNVGATYLCANPVFHSRMKHIALDY 1398
Query: 1398 HFIKEKIDSGEICMSYIPTKSQVADVLTKSLPKRQFDD 1435
HFI+ ++ SG + + ++ T Q+AD LTK L + F +
Sbjct: 1399 HFIRNQVQSGALRVVHVSTHDQLADTLTKPLSRVAFQN 1436
>At2g07420 putative retroelement pol polyprotein
Length = 1664
Score = 676 bits (1743), Expect = 0.0
Identities = 432/1265 (34%), Positives = 658/1265 (51%), Gaps = 127/1265 (10%)
Query: 4 IRLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDK--------VGADYDTWDAENSMV 55
+ L G NY W+++ + + RG +I + + V + W E+ V
Sbjct: 11 VTLKGVNYLLWARTTKTTLCSRGLWAHILTSEAPSEATIREGMEIVHVGEEKWFQEDQSV 70
Query: 56 MTWLVNSLTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVT 115
+ L NSL + Y TAK+LWE + ++ + N S+++E+ + ++ Q + T
Sbjct: 71 LALLQNSLEASLLEAYSYCETAKELWETLFNVFGNQSNLSRVFEVKKAINDLSQGDMEFT 130
Query: 116 KYFGCLKRIWQDLDLFNEYEWKSPEDCKHYQKMVDVSRVFKFLAGLNVEFDEVRGRILGR 175
++FG + +W +L++ + D K + + +VF L L+ ++++ +L
Sbjct: 131 QHFGKFRSLWAELEMLRP----NTLDPKVLIERREQDKVFGLLLTLSSTYNDLIKHLLRA 186
Query: 176 NPIPPIGEVFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVPQVNHKSFRNPRGGDKTH 235
+ +P + EV +++++E++ R N+K N K
Sbjct: 187 DKLPNLEEVCSQIQKEQANRG------------------------NYKYDNN-----KKA 217
Query: 236 LVCDYCGRNRHTQETCFKLHG---------RPNNSKAGKFGDRPMPTTSN---------- 276
L C++C R+ HT+E C+ LH R N FG + TSN
Sbjct: 218 LWCEHCKRSGHTKEKCWTLHPHLRPGRREPRANQVTGENFGTQEQSGTSNQHLGGNGAAM 277
Query: 277 -AASSPFTKEQMDHLLKLLKFNSSPNTPIGTVAQTGKDSWALSVQNHSNPWIIDSGASEH 335
A+S + + L+K LK +S GK ALS P IIDSGAS H
Sbjct: 278 AASSDLVRRSDLKALIKALKESS------------GKSYHALS---SLKPLIIDSGASHH 322
Query: 336 MTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKGNIKISEQITLQSVLHVPKFACNLLSV 395
M + S L S+ P+ G+ V IA+G + G G++ + ++ + ++P F NLLSV
Sbjct: 323 MISDSKLISNIEPALGN--VVIANGDRIPVKGVGDLDLFDKSS--KAFYMPTFTSNLLSV 378
Query: 396 HKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYYFDETPLGNTAMFGSSRTSPP 455
K + D NC +F + FQD + + +G +GLY ++T ++ SS S
Sbjct: 379 KKATTDLNCYAIFGPNEVHFQDIETSRVLGQGVTKDGLYVLEDT---KPSVPLSSHFSSI 435
Query: 456 LVSNKIMLWHKRLGHPSFSYLKHLFPEFSKEISSSQFHCDACHLAKDHRVSFNSRGYSAS 515
L + WH RLGHP LK L P S C+AC L K + F
Sbjct: 436 LGNANSESWHARLGHPHSRALKLLLPS----TSFKNDECEACILGKHCKSVFPKSSTIYE 491
Query: 516 KPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDHTRVCWVYLMEKKSEVEQRFQDFFNMIKN 575
K F LIHSDVW S + K+FVTFID+ ++ W L+ K V + F +F + N
Sbjct: 492 KCFDLIHSDVW-TSPCLSRENHKYFVTFIDEKSKFTWFTLLPSKDRVLEAFTNFQTYVTN 550
Query: 576 QFHTTIGILRSDNGTEYFNKYLSTFLVTNGIIHQSTCRDTPQQNGIAERKNRHLLEVTRA 635
+ I ILRSDN EY + L +GIIHQ++C TPQQNG+AERKNRHL+EV R
Sbjct: 551 HYDAKIKILRSDNRGEYTSHAFKQHLNKHGIIHQTSCPYTPQQNGVAERKNRHLMEVRRV 610
Query: 636 IMFSMNVPKYLWGNALLTACHLINRMPSRVLQYETPVQVLQNNFPTSRIITNIPLKVFGC 695
+MF NVPK+ W + +++AC+LIN+ P+++L +P +VL P L+VFGC
Sbjct: 611 MMFHTNVPKHFWIDGVVSACYLINQTPTKILLDSSPFEVLNKVKPFIN-----HLRVFGC 665
Query: 696 LCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYKCFNPVTKKFFESMDVHFVEDQPFFRE 755
+C+V I R+KL PK+ K +F+GY+ N+KGYKC+ T+K S DV F+E + ++ +
Sbjct: 666 VCFVLISGEQRNKLQPKSTKGMFIGYSINQKGYKCYVLETRKVLISRDVKFLESKSYYDK 725
Query: 756 NSLQG-ESQSSNEEDNFWEILPVLDDI-VTNDHLDAKITEPRNPNLELL---EHMVFETG 810
+ + + + + D + +L+ + V+N T PR N E + E+M E
Sbjct: 726 KNWEDIQDLTDSPSDRATNLRIILERLGVSNIQTQ---TTPRTSNPETITQPENMEEEEE 782
Query: 811 GEEGETLIKNPELRVYVRKKFHKDGTNPIVFPAEVQSDSPSEGPTD--NPSSSSSGNSSH 868
EE E + E ++ E +S E T N + + N
Sbjct: 783 EEEEEEEKQGKE--------------QELITLEETESSKVQEKDTSLLNDDNGHTNNQEE 828
Query: 869 SSNDLPDLSFPDINLPIAVRKNIPDLNIPIAERKRTRTCTKHPMSNYLSYDKLSHSHKAY 928
SN + P + +++ D + + +HP+ + L H+ +
Sbjct: 829 DSNSREEPRIPRRS------EHLKDKRVYYNNQVYFDNVVEHPIQVVCTLAHLPEEHQVF 882
Query: 929 VSRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTWCITDLPYDKKAVGCKWVFTVK 988
++ ++P+T +EA+ W+ A+ E A+ N+TW +LP KK V KWVF +K
Sbjct: 883 FGKVDQHWIPQTYEEAITHQVWRDAIAAEKQAMENNHTWDEDELPRGKKVVTSKWVFAIK 942
Query: 989 CKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILLSLAVNFNWTLHQYDVK 1048
K+DG +ERYKARLVA+GFTQT+G DY +TFAPVAK++++R++LSL N W L Q DVK
Sbjct: 943 YKSDGEIERYKARLVARGFTQTYGEDYLDTFAPVAKLHTVRVVLSLTTNLEWDLWQMDVK 1002
Query: 1049 NAFLNGELHEEVYMRLPPGFEDKLGRGKVCRLKKSLYGLKQSPRAWFERFGSVVKGHGFT 1108
NAFL GEL E+VYM+ PPG ED KV +LKK++YGLKQSPRAW+ + + + G GF
Sbjct: 1003 NAFLQGELEEKVYMKPPPGLEDINAPNKVFKLKKAIYGLKQSPRAWYHKLSTTLMGRGFK 1062
Query: 1109 QSQADHTMFFKHSREGKIAILIVYVDDIIMTGDDIGEISDLKRRLEAEFDIKDLGKLKYF 1168
+S+AD+T+F S++G I +++VYVDDII++G+D I D K L++ FDIKDLG+LKYF
Sbjct: 1063 RSEADNTLFTLPSQKG-IVVILVYVDDIIISGNDKVGIQDTKTFLKSVFDIKDLGELKYF 1121
Query: 1169 LGMEFARSKEGIFLNQRKYILDLLTETGMTGCKAAETPMDPNVKLKSVAEDE---IIDRE 1225
LG+E RSKEG+FL+QRKY LDLL + G G K A+TP++ + K K E + D
Sbjct: 1122 LGIEVCRSKEGLFLSQRKYTLDLLAQVGKLGVKPAKTPLEDDYKAKRKGEHDNKPFEDAT 1181
Query: 1226 RYQRL 1230
RY+RL
Sbjct: 1182 RYRRL 1186
Score = 69.7 bits (169), Expect = 1e-11
Identities = 31/62 (50%), Positives = 43/62 (69%)
Query: 1365 PTPMKVYCDNKAAISIAHNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVADVL 1424
P P+ ++CDN+AAI IA N V H+RTKH+EVD H ++E++ G I Y + Q+ADV
Sbjct: 1194 PNPIPMHCDNQAAIHIASNSVFHERTKHIEVDCHKVREQVQLGVILPHYTEGEEQLADVF 1253
Query: 1425 TK 1426
TK
Sbjct: 1254 TK 1255
>At2g24660 putative retroelement pol polyprotein
Length = 1156
Score = 645 bits (1663), Expect = 0.0
Identities = 399/1098 (36%), Positives = 588/1098 (53%), Gaps = 119/1098 (10%)
Query: 417 DQNSGKTIGTAREINGLYYFDETPLGNTAMFGSSRTSPPLVSNKIMLWHKRLGHPSFSYL 476
D+ S IG+ E +G+YY + TA +++ VS+ LWH+RLGHPSFS L
Sbjct: 52 DRFSRTLIGSGEERDGVYYLTDVA---TAKIHTAK-----VSSDQALWHQRLGHPSFSVL 103
Query: 477 KHLFPEFSKEISSSQFHCDACHLAKDHRVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSR 536
L S +S CD C AK R F + + F LIH DVWGP ++ +
Sbjct: 104 SSLPVLTSSSLSVGSRSCDVCFRAKQTREVFPVSTNKSIECFSLIHCDVWGPYRVPSSCG 163
Query: 537 KKWFVTFIDDHTRVCWVYLMEKKSEVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKY 596
+F+T +DD +R W YL+ KSEV +F + QF ++ +LRSDNGTE+
Sbjct: 164 AVYFLTIVDDFSRAVWTYLLLAKSEVRTVLTNFLVYTEKQFGKSVKVLRSDNGTEFM--C 221
Query: 597 LSTFLVTNGIIHQSTCRDTPQQNGIAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACH 656
L+++ +GI+HQ++C TPQQNG ERK+RH+L V RAI+F ++P WG A+LTA +
Sbjct: 222 LASYFREHGIVHQTSCVGTPQQNGRVERKHRHILNVARAILFQASLPIQFWGEAVLTAAY 281
Query: 657 LINRMPSRVLQYETPVQVLQNNFPTSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKC 716
LINR P+ + +P ++L N+ P L+VFG CYV+ + + K ++ C
Sbjct: 282 LINRTPTSLHNGLSPYEILHNSKPNYE-----HLRVFGSACYVHRASRDKDKFGERSRLC 336
Query: 717 VFLGYASNKKGYKCFNPVTKKFFESMDVHFVEDQ-PFFRENSLQ-GESQSSNEEDNFWEI 774
VF+GY +KG+K F+ K+F S DV F ED P+ N+ S + D W +
Sbjct: 337 VFIGYPFAQKGWKVFDMEKKEFLVSRDVVFREDVFPYAATNTDHVSASPAPLLVDEDWMV 396
Query: 775 LPVLDD------IVTNDHLDAKITEPRNP-NLELLEHMVFETGGEEGETLIKNPELRVYV 827
D + N + D K P + E + E E++ K + V
Sbjct: 397 STTTLDRGSSAVLSANKNTDPKPNSVTKPDDFAEPESVTKPDDFAEPESVTKPDDFAEPV 456
Query: 828 RKKFHKDGTNP--IVFPAEV-------QSDSPSEGPTDN--PSSSSSGNSSHSSNDLPDL 876
D P ++ P V Q+DS E +D+ +S + + +S D D
Sbjct: 457 SVTKPDDFAEPDSVIKPVVVTESNSALQTDSVFEPDSDSDVAPASIASDLQNSRVDSSDS 516
Query: 877 SFPDINL----PIAVRKNIPDL-------NIPI-----------AERKRTR--------- 905
S PD NL +A + PD+ N P+ + R+R R
Sbjct: 517 STPDKNLASGDTLAQIDDSPDIVSTPNRNNQPLFVVDSPFVEATSPRQRKRQIRQSVRLQ 576
Query: 906 -----TCTKHPMSNYLSYDKLSHS-------------------------HKAYVSRISNL 935
T P++ + D S S HKA+++ I+
Sbjct: 577 DYVLYNATVSPINPHALPDSSSQSSSMVQGTSSLYPLSDYVSDDCFSAGHKAFLAAITAN 636
Query: 936 FVPRTIQEALGDPNWKLAVKEEMNALNKNNTWCITDLPYDKKAVGCKWVFTVKCKADGSV 995
P+ +EA+ W A+ +E++AL N TW I DLP K A+G +WV+ K ADGS+
Sbjct: 637 DEPKHFKEAVRIKVWNDAMFKEVDALEINKTWDIVDLPPGKVAIGSQWVYKTKYNADGSI 696
Query: 996 ERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILLSLAVNFNWTLHQYDVKNAFLNGE 1055
ERYKARLV +G Q G DY ETFAPV K+ ++R LL L W ++Q DV NAFL+G+
Sbjct: 697 ERYKARLVVQGNKQVEGEDYNETFAPVVKMTTVRTLLRLVAANQWEVYQMDVNNAFLHGD 756
Query: 1056 LHEEVYMRLPPGFEDKLGRGKVCRLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADHT 1115
L EEVYM+LPPGF KVCRL+KSLYGLKQ+PR WF++ + GF Q D++
Sbjct: 757 LDEEVYMKLPPGFRHS-HPDKVCRLRKSLYGLKQAPRCWFKKLSDALLRFGFVQGHEDYS 815
Query: 1116 MFFKHSREGKIAILIVYVDDIIMTGDDIGEISDLKRRLEAEFDIKDLGKLKYFLGMEFAR 1175
FF ++R G ++VYVDD+++ G+D + K L F +KDLGKLKYFLG+E +R
Sbjct: 816 -FFSYTRNGIELRVLVYVDDLLICGNDGYMLQKFKEYLGRCFSMKDLGKLKYFLGIEVSR 874
Query: 1176 SKEGIFLNQRKYILDLLTETGMTGCKAAETPMDPNVKLKSVAEDEIIDRERYQRLAGRLI 1235
EGIFL+QRKY LD++T++G GC+ A TP++ N L + + D + Y+RL GRL+
Sbjct: 875 GSEGIFLSQRKYALDIITDSGNLGCRPALTPLEQNHHLATDDGPLLADAKPYRRLVGRLL 934
Query: 1236 YLSHTRPDIAFAVSVISQFMHAPGPAHFEAVFRILRYLKGTPGKGLMFRNRGHIQVEAYT 1295
YL HTRP+++++V V+SQFM P AH A RI+R+LKG+PG+G++ +Q++ Y
Sbjct: 935 YLLHTRPELSYSVHVLSQFMQTPREAHLAAAMRIVRFLKGSPGQGILLNADTELQLDVYC 994
Query: 1296 DADWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKK 1355
D+D+ RRS S Y +GG+ ++W++KKQ+ V+ SSAEAE+R+++ E+ W++K
Sbjct: 995 DSDFQTCPKTRRSLSAYVVLLGGSPISWKTKKQDTVSHSSAEAEYRAMSVALREIKWLRK 1054
Query: 1356 FMQELKIAGPTPMKVYCDNKAAISIAHNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIP 1415
++EL NPV H+RTKH+E D H +++ + G I ++
Sbjct: 1055 LLKELA---------------------NPVFHERTKHIESDCHSVRDAVRDGIITTHHVR 1093
Query: 1416 TKSQVADVLTKSLPKRQF 1433
T Q+AD+ TK+L + QF
Sbjct: 1094 TNEQLADIFTKALGRSQF 1111
>At2g07010 putative retroelement pol polyprotein
Length = 1413
Score = 595 bits (1533), Expect = e-170
Identities = 401/1240 (32%), Positives = 617/1240 (49%), Gaps = 120/1240 (9%)
Query: 1 ITTIRLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGADYDTWDAENSMVMTWLV 60
I+++ L G NY WS + ++ + K G+I G +P DY+ W A NSM++ W+
Sbjct: 39 ISSVMLTGDNYNEWSTKMLNALQAKRKTGFINGSISKPPLDNPDYENWQAVNSMIVGWIR 98
Query: 61 NSLTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGC 120
S+ ++ + A LW + Q +S +GN+ ++++ QL RQ V Y+G
Sbjct: 99 ASIEPKVKSTVTFICDAHQLWSELKQRFS-VGNKVHVHQIKTQLAACRQDGQPVIDYYGR 157
Query: 121 LKRIWQDLDLFNEYEWKSPEDCK-----HYQKMVDVSRVFKFLAGLN-VEFDEVRGRILG 174
L ++W++ ++ C K + ++ +F+ GL+ F + ++
Sbjct: 158 LCKLWEEFQIYKPITVCKCGLCTCGATLEPSKEREEEKIHQFVLGLDDSRFGGLSATLIA 217
Query: 175 RNPIPPIGEVFAEVRREESRRQVML--GKKKAAVPPPVEGSALAVPQVNHKSFRNPRGGD 232
+P P +GE+++ V REE R + ++++A+ S + S R D
Sbjct: 218 MDPFPSLGEIYSRVVREEQRLASVQIREQQQSAIGFLTRQSEVTADGRTDSSIIKSR--D 275
Query: 233 KTHLVCDYCGRNRHTQETCFKLHGRPN------------NSKAGKFG----------DRP 270
++ ++C +CGR+ H ++ C+++ G P+ +S G+ G R
Sbjct: 276 RS-VLCSHCGRSGHEKKDCWQIVGFPDWWTERTNGGGRGSSSRGRGGRSSGSNNSGRGRG 334
Query: 271 MPTTSNAASS---PFTKEQMDHLLKLLKFNSSPNTPIGTVAQTGKDSWALSVQNHSNPWI 327
T ++A +S PF + D L + + + N GT S LS + I
Sbjct: 335 QVTAAHATTSNLSPFPEFTPDQLRVITQMIQNKNN--GT-------SDKLSGKMKLGDVI 385
Query: 328 IDSGASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKGNIKISEQITLQSVLHVPK 387
+D+GAS HMT L ++ + S V ADG + G K+SE ++L +VL+VP
Sbjct: 386 LDTGASHHMTGQLSLLTNIV-TIPSCSVGFADGRKTFAISMGTFKLSETVSLSNVLYVPA 444
Query: 388 FACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYYFDETPLGNTAMF 447
C+L+SV KL + C LF + CV QD+ S IGT E +G+YY +
Sbjct: 445 LNCSLISVSKLVKQIKCLALFTDTICVLQDRFSRTLIGTGEERDGVYYLTDA-------- 496
Query: 448 GSSRTSPPLVSNKIMLWHKRLGHPSFSYLKHLFPEFS-KEISSSQFHCDACHLAKDHRVS 506
++ ++ LWH+RLGHPSFS L L P FS S S CD C AK R
Sbjct: 497 ATTTVHKVDITTDHALWHQRLGHPSFSVLSSL-PLFSGSSCSVSSRSCDVCFRAKQTREV 555
Query: 507 FNSRGYSASKPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDHTRVCWVYLMEKKSEVEQRF 566
F ++ F LIH DVWGP ++ + +F+T +DD +R W YL+ KSEV
Sbjct: 556 FPDSSNKSTDCFSLIHCDVWGPYRVPSSCGAVYFLTIVDDFSRSVWTYLLLAKSEVRSVL 615
Query: 567 QDFFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLVTNGIIHQSTCRDTPQQNGIAERKN 626
+F + QF ++ I+RSDNGTE+ LS++ GI+HQ++C TPQQNG ERK+
Sbjct: 616 TNFLAYTEKQFGKSVKIIRSDNGTEFM--CLSSYFKEQGIVHQTSCVGTPQQNGRVERKH 673
Query: 627 RHLLEVTRAIMFSMNVPKYLWGNALLTACHLINRMPSRVLQYETPVQVLQNNFPTSRIIT 686
RH+L V+RA++F ++P WG A++TA +LINR PS + +P ++L P
Sbjct: 674 RHILNVSRALLFQASLPIKFWGEAVMTAAYLINRTPSSIHNGLSPYELLHGCKPDYD--- 730
Query: 687 NIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYKCFNPVTKKFFESMDVHF 746
L+VFG CY + + K ++ C+F+GY +KG+K ++ T +F S DV
Sbjct: 731 --QLRVFGSACYAHRVTRDKDKFGERSRLCIFVGYPFGQKGWKVYDLSTNEFIVSRDV-- 786
Query: 747 VEDQPFFRENSLQGESQSSNEEDNFWEILPVLDDIVTNDHLDAKITEPRNPNLELLEHMV 806
FREN ++NE D + +T P + + L
Sbjct: 787 -----VFRENVFP---YATNEGDTIYT---------------PPVTCPITYDEDWLPFTT 823
Query: 807 FETGGEEGETLIKNPELRVYVRKKFHKDGTNPIVFPAEVQ---SDSPSEGPTDNPSSSSS 863
E G + E + +P + V + + P P V S S S PT P++SSS
Sbjct: 824 LEDRGSD-ENYLSDPPVCVTNVSESDTEHDTPQSLPTPVDDPLSPSTSVTPTQTPTNSSS 882
Query: 864 GNSSHSSNDLPDLSFPDI--NLPIAVRK---------------------NIPDLNIPIAE 900
S ++ P I N P K N P + P
Sbjct: 883 STSPSTNVSPPQQDTTPIIENTPPRQGKRQVQQPARLKDYILYNASCTPNTPHVLSPSTS 942
Query: 901 RKRT--RTCTKHPMSNYLSYDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEM 958
+ + + ++P+++Y+S + S HK +++ I+ P+ +E + W A+ +E+
Sbjct: 943 QSSSSIQGNLQYPLTDYISDECFSAGHKVFLAAITANDEPKHFKEDVKVKVWNDAMYKEV 1002
Query: 959 NALNKNNTWCITDLPYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQET 1018
+AL N TW I DLP K A+G +WV+ K ADG+VERYKARLV +G Q G DY ET
Sbjct: 1003 DALEVNKTWDIVDLPTGKVAIGSQWVYKTKFNADGTVERYKARLVVQGNNQIEGEDYTET 1062
Query: 1019 FAPVAKINSIRILLSLAVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLGRGKVC 1078
FAPV K+ ++R LL L W ++Q DV NAFL+G+L EEVYM+LPPGF KVC
Sbjct: 1063 FAPVVKMTTVRTLLRLVAANQWEVYQMDVHNAFLHGDLEEEVYMKLPPGFRHS-HPDKVC 1121
Query: 1079 RLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIM 1138
RL+KSLYGLKQ+PR WF++ +K GF Q D++ FF +S +G ++VYVDD+I+
Sbjct: 1122 RLRKSLYGLKQAPRCWFKKLSDALKRFGFIQGYEDYS-FFSYSCKGIELRVLVYVDDLII 1180
Query: 1139 TGDDIGEISDLKRRLEAEFDIKDLGKLKYFLGMEFARSKE 1178
G+D + K L F +KDLGKLKYFLG+E +R +
Sbjct: 1181 CGNDEYMVQKFKEYLGRCFSMKDLGKLKYFLGIEVSRGPD 1220
Score = 167 bits (423), Expect = 4e-41
Identities = 78/183 (42%), Positives = 121/183 (65%)
Query: 1251 ISQFMHAPGPAHFEAVFRILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTS 1310
+S+ AP AH EA RI+RYLKG+PG+G++ + +E Y D+D+ RRS S
Sbjct: 1215 VSRGPDAPREAHLEAAMRIVRYLKGSPGQGILLSANKDLTLEVYCDSDFQSCPLTRRSLS 1274
Query: 1311 GYCTFVGGNLVTWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKV 1370
Y +GG+ ++W++KKQ+ V+ SSAEAE+R+++ E+ W+ K ++EL I P ++
Sbjct: 1275 AYVVLLGGSPISWKTKKQDTVSHSSAEAEYRAMSVALKEIKWLNKLLKELGITLAAPTRL 1334
Query: 1371 YCDNKAAISIAHNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVADVLTKSLPK 1430
+CD+KAAISIA NPV H+RTKH+E D H +++ + G I ++ T Q+AD+ TK+L +
Sbjct: 1335 FCDSKAAISIAANPVFHERTKHIERDCHSVRDAVRDGIITTHHVRTSEQLADIFTKALGR 1394
Query: 1431 RQF 1433
QF
Sbjct: 1395 NQF 1397
>At4g10990 putative retrotransposon polyprotein
Length = 1203
Score = 520 bits (1340), Expect = e-147
Identities = 259/567 (45%), Positives = 370/567 (64%), Gaps = 12/567 (2%)
Query: 866 SSHSSNDLPDLSFPDINLPIAVRKNIPDLNIPIAERKRTRTCTKHPMSNYLSYDKLSHSH 925
S + N +P LS +L +I + I +K T T +PMS +SYDKL+
Sbjct: 460 SEYHCNSVPFLS----SLSPTTSTSIETPSSSIPPKKIT---TPYPMSTAISYDKLTPLF 512
Query: 926 KAYVSRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTWCITDLPYDKKAVGCKWVF 985
+Y+ + P+ +A+ W A EE++AL +N TW + L K VGCKWVF
Sbjct: 513 HSYICAYNVETEPKAFTQAMKSEKWTRAANEELHALEQNKTWIVESLTEGKNVVGCKWVF 572
Query: 986 TVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILLSLAVNFNWTLHQY 1045
T+K DGS+ERYKARLVA+GFTQ GIDY ETF+PVAK S+++LL LA W+L Q
Sbjct: 573 TIKYNPDGSIERYKARLVAQGFTQQEGIDYMETFSPVAKFGSVKLLLGLAAATGWSLTQM 632
Query: 1046 DVKNAFLNGELHEEVYMRLPPGFEDKLGRG----KVCRLKKSLYGLKQSPRAWFERFGSV 1101
DV NAFL+GEL EE+YM LP G+ G VCRL KSLYGLKQ+ R W++R SV
Sbjct: 633 DVSNAFLHGELDEEIYMSLPQGYTPPTGISLPSKPVCRLLKSLYGLKQASRQWYKRLSSV 692
Query: 1102 VKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIMTGDDIGEISDLKRRLEAEFDIKD 1161
G F QS AD+TMF K S I +++VYVDD+++ +D + +LK L +EF IKD
Sbjct: 693 FLGANFIQSPADNTMFVKVSCTS-IIVVLVYVDDLMIASNDSSAVENLKELLRSEFKIKD 751
Query: 1162 LGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMTGCKAAETPMDPNVKLKSVAEDEI 1221
LG ++FLG+E ARS EGI + QRKY +LL + G++GCK + PMDPN+ L +
Sbjct: 752 LGPARFFLGLEIARSSEGISVCQRKYAQNLLEDVGLSGCKPSSIPMDPNLHLTKEMGTLL 811
Query: 1222 IDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHAPGPAHFEAVFRILRYLKGTPGKGL 1281
+ Y+ L GRL+YL TRPDI FAV +SQF+ AP H +A ++LRYLKG PG+GL
Sbjct: 812 PNATSYRELVGRLLYLCITRPDITFAVHTLSQFLSAPTDIHMQAAHKVLRYLKGNPGQGL 871
Query: 1282 MFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNVVARSSAEAEFR 1341
M+ + + ++DADW + RRS +G+C ++G +L+TW+SKKQ+VV+RSS E+E+R
Sbjct: 872 MYSASSELCLNGFSDADWGTCKDSRRSVTGFCIYLGTSLITWKSKKQSVVSRSSTESEYR 931
Query: 1342 SVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAISIAHNPVLHDRTKHVEVDKHFIK 1401
S+A CE++W+++ +++L + P K++CDNK+A+ +A NPV H+RTKH+E+D H ++
Sbjct: 932 SLAQATCEIIWLQQLLKDLHVTMTCPAKLFCDNKSALHLATNPVFHERTKHIEIDCHTVR 991
Query: 1402 EKIDSGEICMSYIPTKSQVADVLTKSL 1428
++I +G++ ++PT +Q+AD+LTK L
Sbjct: 992 DQIKAGKLKTLHVPTGNQLADILTKPL 1018
Score = 76.3 bits (186), Expect = 1e-13
Identities = 51/165 (30%), Positives = 81/165 (48%), Gaps = 3/165 (1%)
Query: 318 SVQN--HSNPWIIDSGASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKGNIKISE 375
S+QN S+ WIIDSGAS H+ + +F SG V + +G+ +I G I I+
Sbjct: 89 SLQNVLSSDAWIIDSGASSHVCSDLTMFRELIHVSGVT-VTLPNGTRVAITHTGTICITS 147
Query: 376 QITLQSVLHVPKFACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYY 435
+ L +VL VP F NL+SV L + + S F C Q+ G IG + N LY
Sbjct: 148 TLILHNVLLVPDFKFNLISVCCLVKTLSYSAHFFADCCYIQELTRGLMIGRGKTYNNLYI 207
Query: 436 FDETPLGNTAMFGSSRTSPPLVSNKIMLWHKRLGHPSFSYLKHLF 480
+ + ++ + V + +LWH+RLG + + ++L+
Sbjct: 208 LETQRTSFSPSLPAASSFTGTVQDDCLLWHQRLGIRHYLHYRNLY 252
Score = 75.9 bits (185), Expect = 2e-13
Identities = 42/115 (36%), Positives = 60/115 (51%), Gaps = 15/115 (13%)
Query: 536 RKKWFVTFIDDHTRVCWVYLMEKKSEVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNK 595
R +F+T +DD TR WVY+M+ KSEV F F +I Q++ I +RSDN E
Sbjct: 249 RNLYFLTLVDDCTRTTWVYMMKNKSEVSNIFPVFVKLIFTQYNAKIKAIRSDNVKEL--- 305
Query: 596 YLSTFLVTNGIIHQSTCRDTPQQNGIAER------------KNRHLLEVTRAIMF 638
+ F+ G+IHQ +C TPQQN + ER H + +TR ++F
Sbjct: 306 AFTKFVKEQGMIHQFSCAYTPQQNSVVERYPSGYKGYKVLDLESHSISITRNVVF 360
>At2g16000 putative retroelement pol polyprotein
Length = 1454
Score = 517 bits (1331), Expect = e-146
Identities = 273/639 (42%), Positives = 406/639 (62%), Gaps = 23/639 (3%)
Query: 811 GEEGETLIKNPELRVYVRK--KFHKDGTNPIVFPAEVQSDSPSEGPTDNP----SSSSSG 864
G +G L+ V++ + +FH++ VFP S S P SS
Sbjct: 812 GYKGYKLMDLESNTVFISRNVQFHEE-----VFPLAKNPGSESSLKLFTPMVPVSSGIIS 866
Query: 865 NSSHSSNDLPDLSFPDINLPIA---VRKNIPDLNIPIAERKRTRTCTKHPMSNYLSYDKL 921
+++HS + LP D+ I+ VRK P ++ ++ K+P+S+ +SY K+
Sbjct: 867 DTTHSPSSLPS-QISDLPPQISSQRVRK--PPAHLNDYHCNTMQSDHKYPISSTISYSKI 923
Query: 922 SHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTWCITDLPYDKKAVGC 981
S SH Y++ I+ + +P EA W AV E+ A+ K NTW IT LP KKAVGC
Sbjct: 924 SPSHMCYINNITKIPIPTNYAEAQDTKEWCEAVDAEIGAMEKTNTWEITTLPKGKKAVGC 983
Query: 982 KWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILLSLAVNFNWT 1041
KWVFT+K ADG++ERYKARLVAKG+TQ G+DY +TF+PVAK+ +I++LL ++ + W
Sbjct: 984 KWVFTLKFLADGNLERYKARLVAKGYTQKEGLDYTDTFSPVAKMTTIKLLLKVSASKKWF 1043
Query: 1042 LHQYDVKNAFLNGELHEEVYMRLPPGFEDKLG----RGKVCRLKKSLYGLKQSPRAWFER 1097
L Q DV NAFLNGEL EE++M++P G+ ++ G V RLK+S+YGLKQ+ R WF++
Sbjct: 1044 LKQLDVSNAFLNGELEEEIFMKIPEGYAERKGIVLPSNVVLRLKRSIYGLKQASRQWFKK 1103
Query: 1098 FGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIMTGDDIGEISDLKRRLEAEF 1157
F S + GF ++ DHT+F K +G+ I++VYVDDI++ + L L+ F
Sbjct: 1104 FSSSLLSLGFKKTHGDHTLFLK-MYDGEFVIVLVYVDDIVIASTSEAAAAQLTEELDQRF 1162
Query: 1158 DIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMTGCKAAETPMDPNVKLKSVA 1217
++DLG LKYFLG+E AR+ GI + QRKY L+LL TGM CK PM PN+K++
Sbjct: 1163 KLRDLGDLKYFLGLEVARTTAGISICQRKYALELLQSTGMLACKPVSVPMIPNLKMRKDD 1222
Query: 1218 EDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHAPGPAHFEAVFRILRYLKGTP 1277
D I D E+Y+R+ G+L+YL+ TRPDI FAV+ + QF AP H A +R+L+Y+KGT
Sbjct: 1223 GDLIEDIEQYRRIVGKLMYLTITRPDITFAVNKLCQFSSAPRTTHLTAAYRVLQYIKGTV 1282
Query: 1278 GKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNVVARSSAE 1337
G+GL + + ++ + D+DWA + RRST+ + FVG +L++WRSKKQ+ V+RSSAE
Sbjct: 1283 GQGLFYSASSDLTLKGFADSDWASCQDSRRSTTSFTMFVGDSLISWRSKKQHTVSRSSAE 1342
Query: 1338 AEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAISIAHNPVLHDRTKHVEVDK 1397
AE+R++A CE++W+ + L+ + P P+ +Y D+ AAI IA NPV H+RTKH+++D
Sbjct: 1343 AEYRALALATCEMVWLFTLLVSLQASPPVPI-LYSDSTAAIYIATNPVFHERTKHIKLDC 1401
Query: 1398 HFIKEKIDSGEICMSYIPTKSQVADVLTKSLPKRQFDDM 1436
H ++E++D+GE+ + ++ T+ QVAD+LTK L QF+ +
Sbjct: 1402 HTVRERLDNGELKLLHVRTEDQVADILTKPLFPYQFEHL 1440
Score = 401 bits (1030), Expect = e-111
Identities = 246/794 (30%), Positives = 401/794 (49%), Gaps = 75/794 (9%)
Query: 1 ITTIRLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGADYDTWDAENSMVMTWLV 60
I + RL+ NY WS ++ + + + K G+I G +P + ++ W NSMV +WL+
Sbjct: 74 IISHRLDETNYGDWSVAMLISLDAKNKTGFIDGTLSRPLESDLNFRLWSRCNSMVKSWLL 133
Query: 61 NSLTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGC 120
NS++ +I + L A D+W +++ ++ + N + Y LT ++ + RQ S+++Y+
Sbjct: 134 NSVSPQIYRSILRMNDASDIWRDLNSRFN-VTNLPRTYNLTQEIQDFRQGTLSLSEYYTR 192
Query: 121 LKRIWQDLDLFNEYEWKSPEDC---KHYQKMVDVSRVFKFLAGLNVEFDEVRGRILGRNP 177
LK +W LD + P C Q+ + +++ KFLAGLN + VR +I+ +
Sbjct: 193 LKTLWDQLDSTEALD--EPCTCGKAMRLQQKAEQAKIVKFLAGLNESYAIVRRQIIAKKA 250
Query: 178 IPPIGEVFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVPQVNHKSFRNP------RGG 231
+P +GEV+ + ++ S++ V PP +A V ++ +P G
Sbjct: 251 LPSLGEVYHILDQDNSQQSF-----SNVVAPP---AAFQVSEITQSPSMDPTVCYVQNGP 302
Query: 232 DKTHLVCDYCGRNRHTQETCFKLHGRPNN----SKAGKFGDRPMPTTSNAASSP------ 281
+K +C + R H E C+K HG P KAG+ +P P +N A S
Sbjct: 303 NKGRPICSFYNRVGHIAERCYKKHGFPPGFTPKGKAGEKLQKPKPLAANVAESSEVNTSL 362
Query: 282 ------FTKEQMDHLLKLL--KFNSSPNTPIGTVAQTGKDSWA-------------LSVQ 320
+KEQ+ + + + ++P + T + + D+ L+V
Sbjct: 363 ESMVGNLSKEQLQQFIAMFSSQLQNTPPSTYATASTSQSDNLGICFSPSTYSFIGILTVA 422
Query: 321 NH---SNPWIIDSGASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKGNIKISEQI 377
H S W+IDSGA+ H+++ LFSS +S V + G I+G G +K+++ I
Sbjct: 423 RHTLSSATWVIDSGATHHVSHDRSLFSS-LDTSVLSAVNLPTGPTVKISGVGTLKLNDDI 481
Query: 378 TLQSVLHVPKFACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYYFD 437
L++VL +P+F NL+S+ L++D V+F +++C QD G+ +G R + LY D
Sbjct: 482 LLKNVLFIPEFRLNLISISSLTDDIGSRVIFDKNSCEIQDLIKGRMLGQGRRVANLYLLD 541
Query: 438 ETPLGNTAMFGSSRTSPPLVSNKIMLWHKRLGHPSFSYLKHLFPEF--SKEISSSQFHCD 495
G S V + I +WH+RLGH S L + ++ + C
Sbjct: 542 ---------VGDQSISVNAVVD-ISMWHRRLGHASLQRLDAISDSLGTTRHKNKGSDFCH 591
Query: 496 ACHLAKDHRVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDHTRVCWVYL 555
CHLAK ++SF + + F L+H DVWGP ++T+ K+F+T +DDH+R W+YL
Sbjct: 592 VCHLAKQRKLSFPTSNKVCKEIFDLLHIDVWGPFSVETVEGYKYFLTIVDDHSRATWMYL 651
Query: 556 MEKKSEVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLVTNGIIHQSTCRDT 615
++ KSEV F F ++NQ+ + +RSDN E ++F GI+ +C +T
Sbjct: 652 LKTKSEVLTVFPAFIQQVENQYKVKVKAVRSDNAPEL---KFTSFYAEKGIVSFHSCPET 708
Query: 616 PQQNGIAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACHLINRMPSRVLQYETPVQVL 675
P+QN + ERK++H+L V RA+MF VP LWG+ +LTA LINR PS++L +TP ++L
Sbjct: 709 PEQNSVVERKHQHILNVARALMFQSQVPLSLWGDCVLTAVFLINRTPSQLLMNKTPYEIL 768
Query: 676 QNNFPTSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYKCFNPVT 735
P L+ FGCLCY R K P++ C+FLGY S KGYK + +
Sbjct: 769 TGTAPVYE-----QLRTFGCLCYSSTSPKQRHKFQPRSRACLFLGYPSGYKGYKLMDLES 823
Query: 736 KKFFESMDVHFVED 749
F S +V F E+
Sbjct: 824 NTVFISRNVQFHEE 837
>At2g20460 putative retroelement pol polyprotein
Length = 1461
Score = 513 bits (1322), Expect = e-145
Identities = 250/528 (47%), Positives = 360/528 (67%), Gaps = 6/528 (1%)
Query: 910 HPMSNYLSYDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTWCI 969
HP+S+ LSY ++S SH Y++ IS + +P++ EA W A+ +E+ A+ + +TW I
Sbjct: 919 HPISSSLSYSQISPSHMLYINNISKIPIPQSYHEAKDSKEWCGAIDQEIGAMERTDTWEI 978
Query: 970 TDLPYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINSIR 1029
T LP KKAVGCKWVFTVK ADGS+ER+KAR+VAKG+TQ G+DY ETF+PVAK+ +++
Sbjct: 979 TSLPPGKKAVGCKWVFTVKFHADGSLERFKARIVAKGYTQKEGLDYTETFSPVAKMATVK 1038
Query: 1030 ILLSLAVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLGRGK----VCRLKKSLY 1085
+LL ++ + W L+Q D+ NAFLNG+L E +YM+LP G+ D G VCRLKKS+Y
Sbjct: 1039 LLLKVSASKKWYLNQLDISNAFLNGDLEETIYMKLPDGYADIKGTSLPPNVVCRLKKSIY 1098
Query: 1086 GLKQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIMTGDDIGE 1145
GLKQ+ R WF +F + + GF + DHT+F + + +L+VYVDDI++
Sbjct: 1099 GLKQASRQWFLKFSNSLLALGFEKQHGDHTLFVR-CIGSEFIVLLVYVDDIVIASTTEQA 1157
Query: 1146 ISDLKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMTGCKAAET 1205
L L+A F +++LG LKYFLG+E AR+ EGI L+QRKY L+LLT M CK +
Sbjct: 1158 AQSLTEALKASFKLRELGPLKYFLGLEVARTSEGISLSQRKYALELLTSADMLDCKPSSI 1217
Query: 1206 PMDPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHAPGPAHFEA 1265
PM PN++L + D+E Y+RL G+L+YL+ TRPDI FAV+ + QF AP AH A
Sbjct: 1218 PMTPNIRLSKNDGLLLEDKEMYRRLVGKLMYLTITRPDITFAVNKLCQFSSAPRTAHLAA 1277
Query: 1266 VFRILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGGNLVTWRS 1325
V+++L+Y+KGT G+GL + + ++ YTDADW + RRST+G+ FVG +L++WRS
Sbjct: 1278 VYKVLQYIKGTVGQGLFYSAEDDLTLKGYTDADWGTCPDSRRSTTGFTMFVGSSLISWRS 1337
Query: 1326 KKQNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAISIAHNPV 1385
KKQ V+RSSAEAE+R++A CE+ W+ + L++ P+ +Y D+ AA+ IA NPV
Sbjct: 1338 KKQPTVSRSSAEAEYRALALASCEMAWLSTLLLALRVHSGVPI-LYSDSTAAVYIATNPV 1396
Query: 1386 LHDRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVADVLTKSLPKRQF 1433
H+RTKH+E+D H ++EK+D+G++ + ++ TK QVAD+LTK L QF
Sbjct: 1397 FHERTKHIEIDCHTVREKLDNGQLKLLHVKTKDQVADILTKPLFPYQF 1444
Score = 401 bits (1031), Expect = e-111
Identities = 262/863 (30%), Positives = 421/863 (48%), Gaps = 99/863 (11%)
Query: 1 ITTIRLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGADYDTWDAENSMVMTWLV 60
I + RL+ Y WS ++++ + + K+G++ G +P + ++ W NSMV +WL+
Sbjct: 78 IISHRLDETTYGDWSVAMRISLDAKNKLGFVDGSLPRPLESDPNFRLWSRCNSMVKSWLL 137
Query: 61 NSLTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGC 120
NS++ +I + L A D+W ++ ++ L N + Y LT ++ ++RQ S+++Y+
Sbjct: 138 NSVSPQIYRSILRLNDATDIWRDLFDRFN-LTNLPRTYNLTQEIQDLRQGTMSLSEYYTL 196
Query: 121 LKRIWQDLDLFNEYEWKSPEDC----KHYQKMVDVSRVFKFLAGLNVEFDEVRGRILGRN 176
LK +W LD + P C + YQK + +++ KFLAGLN + VR +I+ +
Sbjct: 197 LKTLWDQLDSTEALD--DPCTCGKAVRLYQK-AEKAKIMKFLAGLNESYAIVRRQIIAKK 253
Query: 177 PIPPIGEVFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVPQVNHKSFRNPR------G 230
+P + EV+ + ++ S++ G PP +A V +V+H +P G
Sbjct: 254 ALPSLAEVYHILDQDNSQK----GFFNVVAPP----AAFQVSEVSHSPITSPEIMYVQSG 305
Query: 231 GDKTHLVCDYCGRNRHTQETCFKLHGRPNN-SKAGKFGDRPMPTTSNAASSPFTKEQMDH 289
+K C +C R H E C+K HG P + GK D+P + AA + ++M
Sbjct: 306 PNKGRPTCSFCNRVGHIAERCYKKHGFPPGFTPKGKSSDKPPKPQAVAAQVTLSPDKMTG 365
Query: 290 LLKLLKFNSSPNTPIGTVA-------------QTGKDSWALSVQNH-------------- 322
L+ L N SP+ +A QT S
Sbjct: 366 QLETLAGNFSPDQIQNLIALFSSQLQPQIVSPQTASSQHEASSSQSVAPSGILFSPSTYC 425
Query: 323 -------------SNPWIIDSGASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKG 369
S+ W+IDSGA+ H+++ LF + S S V + G I+G G
Sbjct: 426 FIGILAVSHNSLSSDTWVIDSGATHHVSHDRKLFQTLDTSIVSF-VNLPTGPNVRISGVG 484
Query: 370 NIKISEQITLQSVLHVPKFACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTARE 429
+ I++ I LQ+VL +P+F NL+S+ L+ D V+F S C QD G T+G +
Sbjct: 485 TVLINKDIILQNVLFIPEFRLNLISISSLTTDLGTRVIFDPSCCQIQDLTKGLTLGEGKR 544
Query: 430 INGLYYFDETPLGNTAMFGSSRTSPPLVSNKIM---LWHKRLGHPSFSYLKHLFPEF--S 484
I LY D SP + N ++ +WHKRLGHPSFS L L +
Sbjct: 545 IGNLYVLDTQ-------------SPAISVNAVVDVSVWHKRLGHPSFSRLDSLSEVLGTT 591
Query: 485 KEISSSQFHCDACHLAKDHRVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSRKKWFVTFI 544
+ + +C CHLAK ++SF S + F L+H DVWGP ++T+ K+F+T +
Sbjct: 592 RHKNKKSAYCHVCHLAKQKKLSFPSANNICNSTFELLHIDVWGPFSVETVEGYKYFLTIV 651
Query: 545 DDHTRVCWVYLMEKKSEVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLVTN 604
DDH+R W+YL++ KS+V F F ++++NQ+ T + +RSDN E + F
Sbjct: 652 DDHSRATWIYLLKSKSDVLTVFPAFIDLVENQYDTRVKSVRSDNAKEL---AFTEFYKAK 708
Query: 605 GIIHQSTCRDTPQQNGIAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACHLINRMPSR 664
GI+ +C +TP+QN + ERK++H+L V RA+MF N+ WG+ +LTA LINR PS
Sbjct: 709 GIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSNMSLPYWGDCVLTAVFLINRTPSA 768
Query: 665 VLQYETPVQVLQNNFPTSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASN 724
+L +TP +VL P LK FGCLCY + R K P++ CVFLGY
Sbjct: 769 LLSNKTPFEVLTGKLPDYS-----QLKTFGCLCYSSTSSKQRHKFLPRSRACVFLGYPFG 823
Query: 725 KKGYKCFNPVTKKFFESMDVHFVEDQPFFRENSLQGESQSSNEEDNFWEILPVLDDIVTN 784
KGYK + ES VH + F E SQ S + ++ +D + +
Sbjct: 824 FKGYKLLD------LESNVVHISRNVEFHEELFPLASSQQSATTAS--DVFTPMDPLSSG 875
Query: 785 DHLDAKITEPR-NPNLELLEHMV 806
+ + + + P+ +P+ ++ + +
Sbjct: 876 NSITSHLPSPQISPSTQISKRRI 898
>At4g17450 retrotransposon like protein
Length = 1433
Score = 504 bits (1299), Expect = e-142
Identities = 241/536 (44%), Positives = 357/536 (65%), Gaps = 6/536 (1%)
Query: 902 KRTRTCTKHPMSNYLSYDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEMNAL 961
KR + KH + ++ Y+ + A+++ I+N +P+ EA W A+KEE+ A+
Sbjct: 882 KRQKKPPKH-LQDFHCYNNTTEPFHAFINNITNAVIPQRYSEAKDFKAWCDAMKEEIGAM 940
Query: 962 NKNNTWCITDLPYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAP 1021
+ NTW + LP +KKA+GCKWVFT+K ADGS+ERYKARLVAKG+TQ G+DY+ETF+P
Sbjct: 941 VRTNTWSVVSLPPNKKAIGCKWVFTIKHNADGSIERYKARLVAKGYTQEEGLDYEETFSP 1000
Query: 1022 VAKINSIRILLSLAVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLGRG----KV 1077
VAK+ S+R++L LA W++HQ D+ NAFLNG+L EE+YM++PPG+ D +G +
Sbjct: 1001 VAKLTSVRMMLLLAAKMKWSVHQLDISNAFLNGDLDEEIYMKIPPGYADLVGEALPPHAI 1060
Query: 1078 CRLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDII 1137
CRL KS+YGLKQ+ R W+ + + +KG GF +S ADHT+F K++ G + ++VYVDDI+
Sbjct: 1061 CRLHKSIYGLKQASRQWYLKLSNTLKGMGFQKSNADHTLFIKYAN-GVLMGVLVYVDDIM 1119
Query: 1138 MTGDDIGEISDLKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGM 1197
+ + ++ L++ F ++DLG KYFLG+E ARS++GI + QRKYIL+LL+ TG
Sbjct: 1120 IVSNSDDAVAQFTAELKSYFKLRDLGAAKYFLGIEIARSEKGISICQRKYILELLSTTGF 1179
Query: 1198 TGCKAAETPMDPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHA 1257
G K + P+DP+VKL + D Y++L G+L+YL TRPDIA+AV+ + QF HA
Sbjct: 1180 LGSKPSSIPLDPSVKLNKEDGVPLTDSTSYRKLVGKLMYLQITRPDIAYAVNTLCQFSHA 1239
Query: 1258 PGPAHFEAVFRILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVG 1317
P H AV ++LRYLKGT G+GL + + YTD+D+ + RR + YC F+G
Sbjct: 1240 PTSVHLSAVHKVLRYLKGTVGQGLFYSADDKFDLRGYTDSDFGSCTDSRRCVAAYCMFIG 1299
Query: 1318 GNLVTWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAA 1377
LV+W+SKKQ+ V+ S+AEAEFR+++ G E++W+ + + K+ P +YCDN AA
Sbjct: 1300 DYLVSWKSKKQDTVSMSTAEAEFRAMSQGTKEMIWLSRLFDDFKVPFIPPAYLYCDNTAA 1359
Query: 1378 ISIAHNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVADVLTKSLPKRQF 1433
+ I +N V H+RTK VE+D + +E ++SG + ++ T QVAD LTK++ QF
Sbjct: 1360 LHIVNNSVFHERTKFVELDCYKTREAVESGFLKTMFVETGEQVADPLTKAIHPAQF 1415
Score = 323 bits (828), Expect = 4e-88
Identities = 245/854 (28%), Positives = 399/854 (46%), Gaps = 148/854 (17%)
Query: 6 LNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGADYDTWDAENSMVMTWLVNSLTE 65
L+G NY WS ++++ + + K+ ++ G +PD + W NSMV TWL+N +TE
Sbjct: 85 LDGTNYNNWSIAMRMSLDAKNKLSFVDGSLPRPDVSDRMFKIWSRCNSMVKTWLLNVVTE 144
Query: 66 EIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGCLKRIW 125
+W ++ + + N + Y+L + ++Q ++ Y+ K +W
Sbjct: 145 --------------MWNDLFSRFR-VSNLPRKYQLEQSIHTLKQGNLDLSTYYTKKKTLW 189
Query: 126 QDLDLFNEYEWKSPEDCKHYQKMVD---VSRVFKFLAGLNVEFDEVRGRILGRNPIPPIG 182
+ L + +C+H +++++ SR+ +FL GLN F +RG+IL P P +
Sbjct: 190 EQLANTRVLTVRKC-NCEHVKELLEEAETSRIIQFLMGLNDNFAHIRGQILNMKPRPGLT 248
Query: 183 EVFAEVRREESRRQV---MLGKKKAAVPPPVEGSALAVPQVN--HKSFRNPRGGDKTHLV 237
E++ + ++ES+R V L AA V+ S + QVN S++ P+
Sbjct: 249 EIYNMLDQDESQRLVGNPTLSNPTAAFQ--VQASPIIDSQVNMAQGSYKKPK-------- 298
Query: 238 CDYCGRNRHTQETCFKLHGRPNNSKAGK---FGD--------RPMPTTSNAASSPFTKEQ 286
C YC + H + C+K HG P SK K G +P+ T N + + +
Sbjct: 299 CSYCNKLGHLVDKCYKKHGYPPGSKWTKGQTIGSTNLASTQLQPVNETPNEKTDSYEEFS 358
Query: 287 MDHLLKLLKF---------------------NSSPNTPI-----GTVAQTGKDSW----- 315
D + ++ + ++SP+ P+ GT +++
Sbjct: 359 TDQIQTMISYLSTKLHIASASPMPTTSSASISASPSVPMISQISGTFLSLFSNAYYDMLI 418
Query: 316 -ALSVQNHSNP--WIIDSGASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKGNIK 372
++S + +P W+IDSGA+ H+T+ L+ + F S + V++ + IAG G I+
Sbjct: 419 SSVSQEPAVSPRGWVIDSGATHHVTHNRDLYLN-FRSLENTFVRLPNDCTVKIAGIGFIQ 477
Query: 373 ISEQITLQSVLHVPKFACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREING 432
+S+ I+L +VL++P+F NL+S + IG ++
Sbjct: 478 LSDAISLHNVLYIPEFKFNLIS----------------------ELTKELMIGRGSQVGN 515
Query: 433 LYYFDETPLGNT-AMFGSSRTSPPL-VSNKIML----WHKRLGHPSFSYLKHL------- 479
LY D +T ++ G++ P V + +++ WHKRLGHP++S + L
Sbjct: 516 LYVLDFNENNHTVSLKGTTSMCPEFSVCSSVVVDSVTWHKRLGHPAYSKIDLLSDVLNLK 575
Query: 480 FPEFSKEISSSQFHCDACHLAKDHRVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSRKKW 539
+ +KE S C CHL+K +SF SR S F L+H D WGP + T
Sbjct: 576 VKKINKEHSPVCHVCHVCHLSKQKHLSFQSRQNMCSAAFDLVHIDTWGPFSVPT------ 629
Query: 540 FVTFIDDHTRVCWVYLMEKKSEVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKYLST 599
+D T W+YL++ KS+V F F NM+ Q+ T + +RSDN E K+
Sbjct: 630 -----NDAT---WIYLLKNKSDVLHVFPAFINMVHTQYQTKLKSVRSDNAHEL--KFTDL 679
Query: 600 FLVTNGIIHQSTCRDTPQQNGIAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACHLIN 659
F +GI+ +C +TP+QN + ERK++H+L V RA++F N+P WG+ +LTA LIN
Sbjct: 680 F-AAHGIVAYHSCPETPEQNSVVERKHQHILNVARALLFQSNIPLEFWGDCVLTAVFLIN 738
Query: 660 RMPSRVLQYETPVQVLQNNFPTSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFL 719
R+P+ VL ++P + L+N P LK FGCLCY R K +P+A CVFL
Sbjct: 739 RLPTPVLNNKSPYEKLKNIPPAYE-----SLKTFGCLCYSSTSPKQRHKFEPRARACVFL 793
Query: 720 GYASNKKGYKCFNPVTKKFFESMDVHFVED-QPFFRENSLQGESQSSNEEDNFWEILPVL 778
GY KGYK + T S V F ED PF SS +D+ + P+L
Sbjct: 794 GYPLGYKGYKLLDIETHAVSISRHVIFHEDIFPFI----------SSTIKDDIKDFFPLL 843
Query: 779 DDIVTNDHLDAKIT 792
D L + T
Sbjct: 844 QFPARTDDLPLEQT 857
>At4g23160 putative protein
Length = 1240
Score = 485 bits (1248), Expect = e-137
Identities = 245/561 (43%), Positives = 372/561 (65%), Gaps = 26/561 (4%)
Query: 860 SSSSGNSSHSSNDLPDLSFPDINLPIAVRKNIPDLNIPIAERKRTR-------------T 906
S + ++S SS D+ P N ++ ++P+ ++ + R+ + +
Sbjct: 3 SDADASTSSSSIDI----MPSAN----IQNDVPEPSVHTSHRRTRKPAYLQDYYCHSVAS 54
Query: 907 CTKHPMSNYLSYDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNT 966
T H +S +LSY+K+S + +++ I+ P T EA W A+ +E+ A+ +T
Sbjct: 55 LTIHDISQFLSYEKVSPLYHSFLVCIAKAKEPSTYNEAKEFLVWCGAMDDEIGAMETTHT 114
Query: 967 WCITDLPYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKIN 1026
W I LP +KK +GCKWV+ +K +DG++ERYKARLVAKG+TQ GID+ ETF+PV K+
Sbjct: 115 WEICTLPPNKKPIGCKWVYKIKYNSDGTIERYKARLVAKGYTQQEGIDFIETFSPVCKLT 174
Query: 1027 SIRILLSLAVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFE----DKLGRGKVCRLKK 1082
S++++L+++ +N+TLHQ D+ NAFLNG+L EE+YM+LPPG+ D L VC LKK
Sbjct: 175 SVKLILAISAIYNFTLHQLDISNAFLNGDLDEEIYMKLPPGYAARQGDSLPPNAVCYLKK 234
Query: 1083 SLYGLKQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIMTGDD 1142
S+YGLKQ+ R WF +F + G GF QS +DHT F K + + +L VYVDDII+ ++
Sbjct: 235 SIYGLKQASRQWFLKFSVTLIGFGFVQSHSDHTYFLKITATLFLCVL-VYVDDIIICSNN 293
Query: 1143 IGEISDLKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMTGCKA 1202
+ +LK +L++ F ++DLG LKYFLG+E ARS GI + QRKY LDLL ETG+ GCK
Sbjct: 294 DAAVDELKSQLKSCFKLRDLGPLKYFLGLEIARSAAGINICQRKYALDLLDETGLLGCKP 353
Query: 1203 AETPMDPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHAPGPAH 1262
+ PMDP+V + + + +D + Y+RL GRL+YL TR DI+FAV+ +SQF AP AH
Sbjct: 354 SSVPMDPSVTFSAHSGGDFVDAKAYRRLIGRLMYLQITRLDISFAVNKLSQFSEAPRLAH 413
Query: 1263 FEAVFRILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGGNLVT 1322
+AV +IL Y+KGT G+GL + ++ +Q++ ++DA + + RRST+GYC F+G +L++
Sbjct: 414 QQAVMKILHYIKGTVGQGLFYSSQAEMQLQVFSDASFQSCKDTRRSTNGYCMFLGTSLIS 473
Query: 1323 WRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAISIAH 1382
W+SKKQ VV++SSAEAE+R+++ E++W+ +F +EL++ P ++CDN AAI IA
Sbjct: 474 WKSKKQQVVSKSSAEAEYRALSFATDEMMWLAQFFRELQLPLSKPTLLFCDNTAAIHIAT 533
Query: 1383 NPVLHDRTKHVEVDKHFIKEK 1403
N V H+RTKH+E D H ++E+
Sbjct: 534 NAVFHERTKHIESDCHSVRER 554
>At4g22040 LTR retrotransposon like protein
Length = 1109
Score = 474 bits (1220), Expect = e-133
Identities = 228/525 (43%), Positives = 362/525 (68%), Gaps = 3/525 (0%)
Query: 908 TKHPMSNYLSYDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTW 967
T+ P S +S ++ + SHKA+++ ++ P T EA+ D W+ A+ E+ +L N T+
Sbjct: 571 TEFPYSK-MSCNRFTSSHKAFLAAVTAGMEPTTYNEAMVDKAWREAMSAEIESLRVNQTF 629
Query: 968 CITDLPYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINS 1027
I +LP K+A+G KWV+ +K ++DG++ERYKARLV G Q G+DY ETFAPVAK+++
Sbjct: 630 SIVNLPPGKRALGNKWVYKIKYRSDGAIERYKARLVVLGNCQKEGVDYDETFAPVAKMST 689
Query: 1028 IRILLSLAVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLGRGKVCRLKKSLYGL 1087
+R+ L +A +W +HQ DV NAFL+G+L EEVYM+LP GF+ KVCRL KSLYGL
Sbjct: 690 VRLFLGVAAARDWHVHQMDVHNAFLHGDLKEEVYMKLPQGFQCD-DPSKVCRLHKSLYGL 748
Query: 1088 KQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIMTGDDIGEIS 1147
KQ+PR WF + S +K +GFTQS +D+++F ++ +G ++VYVDD+I++G ++
Sbjct: 749 KQAPRCWFSKLSSALKQYGFTQSLSDYSLF-SYNNDGVFVHVLVYVDDLIISGSCPDAVA 807
Query: 1148 DLKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMTGCKAAETPM 1207
K LE+ F +KDLG LKYFLG+E +R+ +G +L+QRKY+LD+++E G+ G + + P+
Sbjct: 808 QFKSYLESCFHMKDLGLLKYFLGIEVSRNAQGFYLSQRKYVLDIISEMGLLGARPSAFPL 867
Query: 1208 DPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHAPGPAHFEAVF 1267
+ N KL + D RY+RL GRLIYL+ TRP+++++V ++QFM P H+ A
Sbjct: 868 EQNHKLSLSTSPLLSDSSRYRRLVGRLIYLAVTRPELSYSVHTLAQFMQNPRQDHWNAAI 927
Query: 1268 RILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGGNLVTWRSKK 1327
R++RYLK PG+G++ + +Q+ + D+D+A RRS +GY +G ++W++KK
Sbjct: 928 RVVRYLKSNPGQGILLSSTSTLQINGWCDSDYAACPLTRRSLTGYFVQLGDTPISWKTKK 987
Query: 1328 QNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAISIAHNPVLH 1387
Q ++RSSAEAE+R++A E++W+K+ + +L ++ M+++ D+K+AI+++ NPV H
Sbjct: 988 QPTISRSSAEAEYRAMAFLTQELMWLKRVLYDLGVSHVQAMRIFSDSKSAIALSVNPVQH 1047
Query: 1388 DRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVADVLTKSLPKRQ 1432
+RTKHVEVD HFI++ I G I S++P+ Q+AD+LTK+L +++
Sbjct: 1048 ERTKHVEVDCHFIRDAILDGIIATSFVPSHKQLADILTKALGEKE 1092
Score = 254 bits (648), Expect = 3e-67
Identities = 148/422 (35%), Positives = 226/422 (53%), Gaps = 21/422 (4%)
Query: 332 ASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKGNIKISEQITLQSVLHVPKFACN 391
AS HMT L S S + +ADG+ +G +++ + L+SV +V + +
Sbjct: 169 ASHHMTGNLELLSG-MRSMSPVLIILADGNKRVAVSEGTVRLGSHLILKSVFYVKELESD 227
Query: 392 LLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYYFDETPLGNTAMFGSSR 451
L+SV ++ ++++C V V QD+ + G + NG + F + N A +S
Sbjct: 228 LISVGQMMDENHCVVQLADHFLVIQDRTTRMVTGIGKRENGSFCF--RGMENAAAVHTSV 285
Query: 452 TSPPLVSNKIMLWHKRLGHPSFSYLKHLFPEF---SKEISSSQFHCDACHLAKDHRVSFN 508
+P LWH+RLGH S + L E KEI + CD C AK R +F
Sbjct: 286 KAP------FDLWHRRLGHASDKIVNLLPRELLSSGKEILENV--CDTCMRAKQTRDTFP 337
Query: 509 SRGYSASKPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDHTRVCWVYLMEKKSEVEQRFQD 568
R + F LIH DVWGP + + S ++F+T +DD++R WVYLM KSE ++ +D
Sbjct: 338 LRDNRSMDSFQLIHCDVWGPYRTPSYSGARYFLTIVDDYSRGVWVYLMTDKSETQKHLKD 397
Query: 569 FFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLVTNGIIHQSTCRDTPQQNGIAERKNRH 628
F +++ QF T I +RSDNGTE+ + + + GI H+++C TP QNG ERK+RH
Sbjct: 398 FMALVERQFDTEIKTVRSDNGTEFL--CMREYFLHKGIAHETSCVGTPHQNGRVERKHRH 455
Query: 629 LLEVTRAIMFSMNVPKYLWGNALLTACHLINRMPSRVLQYETPVQVLQNNFPTSRIITNI 688
+L + RA+ F +P WG +L+A +LINR PS +LQ ++P ++L P
Sbjct: 456 ILNIARALRFQSYLPIQFWGECILSAAYLINRTPSMLLQGKSPYEMLYKTAPKYS----- 510
Query: 689 PLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYKCFNPVTKKFFESMDVHFVE 748
L+VFG LCY + N K ++ +CVF+GY +KG++ F+ +KFF S DV F E
Sbjct: 511 HLRVFGSLCYAHNQNHKGDKFAARSRRCVFVGYPHGQKGWRLFDLEEQKFFVSRDVIFQE 570
Query: 749 DQ 750
+
Sbjct: 571 TE 572
Score = 63.2 bits (152), Expect = 1e-09
Identities = 31/122 (25%), Positives = 62/122 (50%), Gaps = 1/122 (0%)
Query: 10 NYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGADYDTWDAENSMVMTWLVNSLTEEIGA 69
NY W+ + +R R K G++ G QP D + W N+++++W+ ++ E+
Sbjct: 42 NYEEWACGFKTALRSRKKFGFLDGTIPQPLDGSPDLEDWLTINALLVSWMKMTIDSELLT 101
Query: 70 NYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGCLKRIWQDLD 129
N A+DLWE + + +S + N + ++ L +Q +V Y+G L +IW +++
Sbjct: 102 NISHRDVARDLWEQIRKRFS-VSNGPKNQKMKADLATCKQEGMTVEGYYGKLNKIWDNIN 160
Query: 130 LF 131
+
Sbjct: 161 SY 162
>At1g47650 hypothetical protein
Length = 1409
Score = 469 bits (1207), Expect = e-132
Identities = 234/509 (45%), Positives = 341/509 (66%), Gaps = 17/509 (3%)
Query: 924 SHKAYVSRISNLFVPRTIQEALGDPNWKLAVK---------EEMNALNKNNTWCITDLPY 974
SH ++SR N+ + DP + ++K E+ A+ +TW IT LP
Sbjct: 710 SHTVFISR--NVVFHEEVFPLAVDPKSESSLKLFTPMVPVSSEIGAMETTHTWEITTLPA 767
Query: 975 DKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILLSL 1034
KKAVGCKWVF++K ADGS+ERYKARLVAKG+TQ G+DY +TF+PVAK+ +I++LL +
Sbjct: 768 GKKAVGCKWVFSLKFLADGSLERYKARLVAKGYTQKEGLDYTDTFSPVAKMTTIKLLLKI 827
Query: 1035 AVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLG----RGKVCRLKKSLYGLKQS 1090
+ + W L Q DV NAFLNGEL EE+YM+LP G+ + G VCRLK+S+YGLKQ+
Sbjct: 828 SASKKWFLKQLDVSNAFLNGELEEEIYMKLPEGYAARKGITLPPNSVCRLKRSIYGLKQA 887
Query: 1091 PRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIMTGDDIGEISDLK 1150
R WF++F + + GF ++ DHT+F K +G+ I++VYVDDI + G L
Sbjct: 888 SRQWFKKFSASLLDLGFKKTHGDHTLFIKEY-DGEFVIVLVYVDDIAIASTSEGAAIQLT 946
Query: 1151 RRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMTGCKAAETPMDPN 1210
+ L+ F ++DLG LKYFLG+E AR++ GI + QRKY L+LL TGM CK PM PN
Sbjct: 947 QDLQQRFKLRDLGDLKYFLGLEIARTEAGISICQRKYALELLASTGMVNCKPVSVPMIPN 1006
Query: 1211 VKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMHAPGPAHFEAVFRIL 1270
VK+ D + DRE+Y+R+ G+L+YL+ TR DI FAV+ + QF AP +H +A +R+L
Sbjct: 1007 VKMMKTDGDLLDDREQYRRIVGKLMYLTITRLDITFAVNKLCQFSSAPRTSHLQAAYRVL 1066
Query: 1271 RYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNV 1330
+Y+KG+ G+GL + + ++ + D+DWA + RRST+G+ FVG +L++WRSKKQ+V
Sbjct: 1067 QYIKGSVGQGLFYSASADLTLKGFADSDWASCPDSRRSTTGFSMFVGDSLISWRSKKQHV 1126
Query: 1331 VARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAISIAHNPVLHDRT 1390
V+RSSAEAE+R++A CE++W+ + LK + P+ +Y D+ AI IA NPV H+RT
Sbjct: 1127 VSRSSAEAEYRALALVTCELVWLHTLLMSLKASYLVPV-LYSDSTTAIYIATNPVFHERT 1185
Query: 1391 KHVEVDKHFIKEKIDSGEICMSYIPTKSQ 1419
KH+E+D H ++EK+D+GE+ + ++ T+ Q
Sbjct: 1186 KHIELDCHTVREKLDNGELKLLHVRTEDQ 1214
Score = 252 bits (643), Expect = 1e-66
Identities = 143/429 (33%), Positives = 228/429 (52%), Gaps = 43/429 (10%)
Query: 327 IIDSGASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKGNIKISEQITLQSVLHVP 386
+IDSGA H+++ LF + +S V + G I+G G +++++ I L+++L +P
Sbjct: 333 VIDSGAIHHVSHDKSLFLN-LDTSVVSAVNLPAGPTVRISGVGTLRLNDDILLKNILFIP 391
Query: 387 KFACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYYFDETPLGNTAM 446
+F NL+S+ L++D V+F +++C QD + +G R I LY D
Sbjct: 392 EFRLNLISISSLTDDIGSRVIFDKTSCEIQDPIKARMLGQGRRIANLYVLD--------- 442
Query: 447 FGSSRTSPPLVSNK----IMLWHKRLGHPSFSYLKHLFPEF--SKEISSSQFHCDACHLA 500
P+VS I +WH+RLGH S L + +K + +C CHLA
Sbjct: 443 -----IEDPIVSVNAVVDISMWHRRLGHASLQRLDVISDSLGTTKPKNKGSDYCHVCHLA 497
Query: 501 KDHRVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDHTRVCWVYLMEKKS 560
K ++ F S+ ++ F L+H D+WGP ++T+ K+F+T +DDH+R W+YL+ KS
Sbjct: 498 KQRKLPFPSQNKVCNEIFDLLHIDIWGPFSVETVDGYKYFLTIVDDHSRATWIYLLRNKS 557
Query: 561 EVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLVTNGIIHQSTCRDTPQQNG 620
EV F F ++NQ+ + +RSDN E K+ S + GI+ +C +TP+QN
Sbjct: 558 EVLTVFPAFIEQVENQYKVRVKAVRSDNAPEL--KFTSLY-QQKGIVSFHSCPETPEQNL 614
Query: 621 IAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACHLINRMPSRVLQYETPVQVLQNNFP 680
+ ERK++H+L V+RA+MF VP LWG+ +LTA LINR PS++L +TP ++L P
Sbjct: 615 VVERKHQHILNVSRALMFQSQVPLSLWGDCVLTAVFLINRTPSQLLSNKTPYEILTGTVP 674
Query: 681 TSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYKCFNPVTKKFFE 740
+ + R K P+++ CVFLGY + KGYK + + F
Sbjct: 675 -----------------FYFAKK--RHKFQPRSKACVFLGYPAGYKGYKLMDLESHTVFI 715
Query: 741 SMDVHFVED 749
S +V F E+
Sbjct: 716 SRNVVFHEE 724
Score = 104 bits (259), Expect = 4e-22
Identities = 73/281 (25%), Positives = 135/281 (47%), Gaps = 23/281 (8%)
Query: 1 ITTIRLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGADYDTWDAENSMVMTWLV 60
I + RL+ NY W+ ++ + + + K G+I G +P + ++ W SMV +WL+
Sbjct: 77 IISHRLDETNYGDWNVAMLISLDAKNKSGFIDGTLPRPLESDKNFRLWSRCKSMVKSWLL 136
Query: 61 NSLTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGC 120
NS++ +I + L A D+W ++ + ++ N + Y LT ++ + RQ S+++Y+
Sbjct: 137 NSVSPQIYRSILRMNDASDIWRDLHSRF-NVTNLPRTYNLTQEIQDFRQGTLSLSEYYTR 195
Query: 121 LKRIWQDLDLFNEYEWKSPEDC---KHYQKMVDVSRVFKFLAGLNVEFDEVRGRILGRNP 177
LK +W L+ E + P C Q+ + +++ K LAGLN + +R +I+ +
Sbjct: 196 LKTLWDQLESTEELD--DPCICGKAMRLQQKAEQAKIVKLLAGLNDSYAIIRRQIIAKKA 253
Query: 178 IPPIGEVFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVPQVNHKSFRNPRGGDKTHLV 237
+P + EV+ + ++ S++ G PP + A+ N+ D L
Sbjct: 254 LPSLAEVYHILDQDNSQQ----GFSNVVAPPAAFQVSEAMMATNN---------DVNMLC 300
Query: 238 CDYCGRNRHTQETCFKLHGRP-NNSKAGKFGDRPMPTTSNA 277
++ G H E C+K HG P + GK GD+ S A
Sbjct: 301 SEWVG---HIAERCYKKHGFPLGFTPKGKGGDKSQVIDSGA 338
>At2g15920 putative retroelement pol polyprotein
Length = 1100
Score = 446 bits (1146), Expect = e-125
Identities = 231/540 (42%), Positives = 337/540 (61%), Gaps = 26/540 (4%)
Query: 902 KRTRTCTKHPMSNYLSYDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEMNAL 961
K T++ K+P+S+ +SY ++S SH Y+ I+ + + EA G W AV E+ A+
Sbjct: 564 KNTQSDQKYPISSTISYSQISPSHMCYIDNITKILISTNFPEAQGTKEWCEAVDVEIGAM 623
Query: 962 NKNNTWCITDLPYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAP 1021
N W IT LP DKKAVGCKWVFT+K ADGS+ERYKARLVAKG+TQ G+DY TF+P
Sbjct: 624 ESTNKWEITTLPKDKKAVGCKWVFTLKILADGSLERYKARLVAKGYTQKEGLDYTNTFSP 683
Query: 1022 VAKINSIRILLSLAVNFNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLG----RGKV 1077
VAK+ +I++LL + + W L Q DV N+FLNGEL EE+YM+LP G ++ G V
Sbjct: 684 VAKMTTIKLLLKIYASKKWFLKQLDVSNSFLNGELEEEIYMKLPEGSAERKGIILPSNAV 743
Query: 1078 CRLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADHTMF-FKHSREGKIAILIVYVDDI 1136
CRLK+S+YGLKQ+ R WF++F + + GF + DHT+F E + +++VYVDDI
Sbjct: 744 CRLKRSIYGLKQASRQWFKKFSASLLSLGFFKIHGDHTLFLIDCGGEYVVVLVLVYVDDI 803
Query: 1137 IMTGDDIGEISDLKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETG 1196
+ I+ L F ++DLG LKYFL +E +R++ GI L RK +LL G
Sbjct: 804 V--------IASTNEDLHNLFKLRDLGDLKYFLCLEISRTEVGISLCHRKNAWELLASIG 855
Query: 1197 MTGCKAAETPMDPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFMH 1256
M CK PM NVKL + + DRE+Y+R+ G+L+ L+ TRPDI +
Sbjct: 856 MINCKPVSVPMITNVKLMKADGELLEDREQYRRIVGKLMDLTITRPDITLRTT------- 908
Query: 1257 APGPAHFEAVFRILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFV 1316
H +A R+L+Y+KG G+GL + + ++ + D+DW + RRST+G+ FV
Sbjct: 909 -----HLQAAHRVLQYIKGIVGQGLFYSASSDLTLKGFADSDWVSCADSRRSTTGFTMFV 963
Query: 1317 GGNLVTWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKA 1376
G +L++ RSKKQ VV+RSSAEAE+R++A CE++ + + L A P+ ++ ++
Sbjct: 964 GDSLISLRSKKQRVVSRSSAEAEYRALALATCELVSLHTLLVSLTAATSVPI-LFSNSTT 1022
Query: 1377 AISIAHNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVADVLTKSLPKRQFDDM 1436
AI IA NPV H+RTKH E+D + ++EK+D GE+ + ++ T+ QVAD++TK L QF+ +
Sbjct: 1023 AIYIAINPVFHERTKHTEIDCNTVREKVDDGELKLLHVRTEDQVADIMTKPLSPHQFEHL 1082
Score = 156 bits (394), Expect = 9e-38
Identities = 118/476 (24%), Positives = 215/476 (44%), Gaps = 68/476 (14%)
Query: 1 ITTIRLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGADYDTWDAENSMVMTWLV 60
I + RL+ NY W+ ++ + + + K G+I G +P + ++ W NSM+
Sbjct: 30 IISHRLDETNYGDWNVAMLISLDVKNKSGFIDGTLPRPLETDKNFCLWSRCNSMICR--- 86
Query: 61 NSLTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGC 120
+ L A D+W +++ ++ + N + Y LT ++ ++RQ S+++Y+
Sbjct: 87 ---------SILRMNDASDIWRDLNSRFN-MTNLPRTYNLTQEIQDLRQGTLSLSEYYTR 136
Query: 121 LKRIWQDLDLFNEYEWKSPEDCKHYQKMVDVSRVFKFLAGLNVEFDEVRGRILGRNPIPP 180
LK +W LD E + + C ++ +++ KFLAGLN + +R +I+ + +P
Sbjct: 137 LKTLWDQLDSTEELD----DPCTRQKR----AKIVKFLAGLNESYAIIRRQIIAKKILPS 188
Query: 181 IGEVFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVPQ-VNHKSFRNPRGGDKTHLVCD 239
+ EV+ V ++ S++ A P + S + + ++ + G +K +C
Sbjct: 189 LAEVYHIVDQDNSQQGF---SNVVARPAAFQVSEVTITNSIDPTIYYVQNGPNKGRSMCS 245
Query: 240 YCGRNRHTQETCFKLHGRPNN-SKAGKFGDR---PMPTTSNAASSP-------------- 281
+C R H E C+K HG P + GK GD+ P P SN A +
Sbjct: 246 FCNRVGHIAERCYKKHGFPPRFTPKGKVGDKTQKPKPVASNVALATTESNDTHSGLKSLV 305
Query: 282 --FTKEQMDHLLKLLKFNSSP----NTPIGTVAQTGKDSWA-----------LSVQNH-- 322
+KEQ+ + + P N+ + + +Q + L+V H
Sbjct: 306 GNLSKEQLQQFIAMFSSQLQPQPHSNSAVASSSQADNIGISFSPSTYSFIGILTVAQHTL 365
Query: 323 -SNPWIIDSGASEHMTNCSHLFSSYFPSSGSEKVKIADGSYSSIAGKGNIKISEQITLQS 381
S W+IDSGA H+ L +S S V + G I+G G +++++ L++
Sbjct: 366 SSKTWVIDSGAIHHVNLLLTLNTSVLSS-----VNLPAGPTVKISGVGTLRLNDDNLLKN 420
Query: 382 VLHVPKFACNLLSVHKLSEDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYYFD 437
VL +P+F NL+S+ L++D V+F + C QD G+ +G R + LY D
Sbjct: 421 VLFIPEFCLNLMSISSLTDDIGSRVIFNQHACEIQDLIKGRMLGHGRRVANLYVMD 476
>At1g58140 hypothetical protein
Length = 1320
Score = 440 bits (1131), Expect = e-123
Identities = 212/507 (41%), Positives = 314/507 (61%), Gaps = 1/507 (0%)
Query: 930 SRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTWCITDLPYDKKAVGCKWVFTVKC 989
S+I P QEA+ W+ A+ EE+ ++ KN+TW +T LP KA+G KWV+ K
Sbjct: 798 SQIEEKCEPMDFQEAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHKAIGVKWVYKAKK 857
Query: 990 KADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILLSLAVNFNWTLHQYDVKN 1049
+ G VERYKARLVAKG++Q GIDY E FAPVA++ ++R+++SLA W +HQ DVK+
Sbjct: 858 NSKGEVERYKARLVAKGYSQRAGIDYDEVFAPVARLETVRLIISLAAQNKWKIHQMDVKS 917
Query: 1050 AFLNGELHEEVYMRLPPGFEDKLGRGKVCRLKKSLYGLKQSPRAWFERFGSVVKGHGFTQ 1109
AFLNG+L EEVY+ P G+ K KV RLKK+LYGLKQ+PRAW R K F +
Sbjct: 918 AFLNGDLEEEVYIEQPQGYIVKGEEDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIK 977
Query: 1110 SQADHTMFFKHSREGKIAILIVYVDDIIMTGDDIGEISDLKRRLEAEFDIKDLGKLKYFL 1169
+H ++ K +E I I +YVDD+I TG++ + K+ + EF++ D+G + Y+L
Sbjct: 978 CPYEHALYIKIQKED-ILIACLYVDDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYL 1036
Query: 1170 GMEFARSKEGIFLNQRKYILDLLTETGMTGCKAAETPMDPNVKLKSVAEDEIIDRERYQR 1229
G+E + GIF+ Q Y ++L + M TPM+ +KL E E +D ++
Sbjct: 1037 GIEVKQEDNGIFITQEGYAKEVLKKFKMDDSNPVCTPMECGIKLSKKEEGEGVDPTTFKS 1096
Query: 1230 LAGRLIYLSHTRPDIAFAVSVISQFMHAPGPAHFEAVFRILRYLKGTPGKGLMFRNRGHI 1289
L G L YL+ TRPDI +AV V+S++M P HF+A RILRY+KGT GL +
Sbjct: 1097 LVGSLRYLTCTRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTSDY 1156
Query: 1290 QVEAYTDADWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNVVARSSAEAEFRSVAHGFCE 1349
++ Y+D+DW G+++DR+STSG+ ++G TW SKKQ +V S+ EAE+ + C
Sbjct: 1157 KLVGYSDSDWGGDVDDRKSTSGFVFYIGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCH 1216
Query: 1350 VLWIKKFMQELKIAGPTPMKVYCDNKAAISIAHNPVLHDRTKHVEVDKHFIKEKIDSGEI 1409
+W++ ++EL + P K++ DNK+AI++A NPV HDR+KH++ H+I+E + ++
Sbjct: 1217 AIWLRNLLKELSLPQEEPTKIFVDNKSAIALAKNPVFHDRSKHIDTRYHYIRECVSKKDV 1276
Query: 1410 CMSYIPTKSQVADVLTKSLPKRQFDDM 1436
+ Y+ T QVAD+ TK L + F M
Sbjct: 1277 QLEYVKTHDQVADIFTKPLKREDFIKM 1303
Score = 234 bits (596), Expect = 4e-61
Identities = 207/805 (25%), Positives = 346/805 (42%), Gaps = 87/805 (10%)
Query: 6 LNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGADYDTWD-------AENSMVMTW 58
L +NY WS ++ + + +P+ G+ T + +
Sbjct: 13 LTKSNYDNWSLRMKAILGAHDVWEIVEKGFIEPENEGSLSQTQKDGLRDSRKRDKKALCL 72
Query: 59 LVNSLTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAE----DSV 114
+ L E+ + +AK+ WE + Y ++ TL+ GE + + V
Sbjct: 73 IYQGLDEDTFEKVVEATSAKEAWEKLRTSYKGADQVKKVRLQTLR-GEFEALQMKEGELV 131
Query: 115 TKYFGCLKRIWQDLDLFNEYEWKSPEDCKHYQKMVDVSRVFKFLAGLNVEFDEVRGRILG 174
+ YF + + +L ++ +K+ DV + K L L+++F+ + I
Sbjct: 132 SDYFSRVLTVTNNLK-------------RNGEKLDDVRIMEKVLRSLDLKFEHIVTVIEE 178
Query: 175 RNPIPP--IGEVFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVPQVNHKSFRNPRGGD 232
+ I ++ ++ E ++ KKK + V + + N +S++ GG
Sbjct: 179 TKDLEAMTIEQLLGSLQAYEEKK-----KKKEDIVEQVLNMQITKEE-NGQSYQRRGGGQ 232
Query: 233 KTHLVCDYCGRNRHTQETCFKLHGRPNNSKAGKFGDRPMPTTSNAASSPFTKEQMDHLLK 292
G R + + R NS G+ P ++ + + H
Sbjct: 233 VRGRGRGGYGNGRGWRPHEDNTNQRGENSSRGRGKGHPKSRYDKSSVKCYNCGKFGHYAS 292
Query: 293 LLK------FNSSPNTPIGTVAQTGK---DSWALSVQNHSNPWIIDSGASEHMTNCSHLF 343
K F N + + S+ Q ++ W +DSGAS HM +F
Sbjct: 293 ECKAPSNKKFEEKANYVEEKIQEEDMLLMASYKKDEQEENHKWYLDSGASNHMCGRKSMF 352
Query: 344 SSYFPSSGSEKVKIADGSYSSIAGKGNIKI----SEQITLQSVLHVPKFACNLLSVHKLS 399
+ S V + D S + GKGNI I + + +V ++P N+LS+ +L
Sbjct: 353 AE-LDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQFISNVYYIPSMKTNILSLGQLL 411
Query: 400 EDSNCSVLFCRSTCVFQDQNSGKTIGTAREINGLYYFDETPLGNTAMF------GSSRTS 453
E + + +DQ S + P+ MF ++
Sbjct: 412 E-KGYDIRLKDNNLSIRDQESN-------------LITKVPMSKNRMFVLNIRNDIAQCL 457
Query: 454 PPLVSNKIMLWHKRLGHPSFSYLKHLFPEFSKE----ISSSQFHCDACHLAKDHRVSFNS 509
+ LWH R GH +F L+ L + I+ C+ C L K ++SF
Sbjct: 458 KMCYKEESWLWHLRFGHLNFGGLELLSRKEMVRGLPCINHPNQVCEGCLLGKQFKMSFPK 517
Query: 510 RGYS-ASKPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDHTRVCWVYLMEKKSEVEQRFQD 568
S A KP LIH+DV GP K K+L + +F+ FIDD +R WVY +++KSEV + F+
Sbjct: 518 ESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRKTWVYFLKEKSEVFEIFKK 577
Query: 569 FFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLVTNGIIHQSTCRDTPQQNGIAERKNRH 628
F ++ + I +RSD G E+ +K + NGI Q T +PQQNG+AERKNR
Sbjct: 578 FKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQLTVPRSPQQNGVAERKNRT 637
Query: 629 LLEVTRAIMFSMNVPKYLWGNALLTACHLINRMPSRVLQYETPVQVLQNNFPTSRIITNI 688
+LE+ R+++ S +PK LW A+ A +L+NR P++ + +TP + P
Sbjct: 638 ILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKTPQEAWSGRKPGVS----- 692
Query: 689 PLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYKCFNPVTKKFFESMDVHFVE 748
L+VFG + + ++P+ RSKLD K+EK +F+GY +N KGYK +NP TKK S ++ F E
Sbjct: 693 HLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDTKKTIISRNIVFDE 752
Query: 749 DQPFFRENSLQGESQSSNEED-NFW 772
+ + +SNEED NF+
Sbjct: 753 EGEW---------DWNSNEEDYNFF 768
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.136 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,503,012
Number of Sequences: 26719
Number of extensions: 1583065
Number of successful extensions: 5156
Number of sequences better than 10.0: 154
Number of HSP's better than 10.0 without gapping: 130
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 4230
Number of HSP's gapped (non-prelim): 386
length of query: 1436
length of database: 11,318,596
effective HSP length: 112
effective length of query: 1324
effective length of database: 8,326,068
effective search space: 11023714032
effective search space used: 11023714032
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC135396.1


