

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
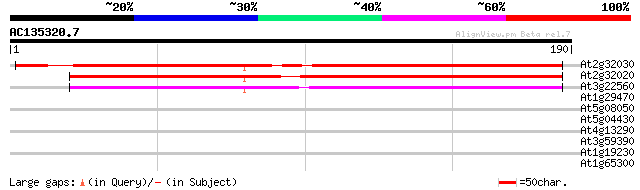
Query= AC135320.7 + phase: 0
(190 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g32030 putative alanine acetyl transferase 171 2e-43
At2g32020 putative alanine acetyl transferase 164 3e-41
At3g22560 hypothetical protein 120 6e-28
At1g29470 unknown protein 28 2.2
At5g08050 putative protein 27 5.0
At5g04430 putative RNA-binding protein 27 6.5
At4g13290 cytochrome p450 - like protein 27 6.5
At3g59390 putative protein 27 6.5
At1g19230 hypothetical protein 27 6.5
At1g65300 27 8.5
>At2g32030 putative alanine acetyl transferase
Length = 188
Score = 171 bits (434), Expect = 2e-43
Identities = 90/186 (48%), Positives = 124/186 (66%), Gaps = 15/186 (8%)
Query: 3 ITSISSKPGAKEERIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINF 62
++++SS P K I LRP+ LSD+DD M+W TD V +FC+WE YTS++ I +
Sbjct: 14 LSTVSSSPPEK--------IHLRPMTLSDVDDFMVWATDSNVTRFCTWEPYTSREAAIAY 65
Query: 63 IENIATKFLWCKAICI-NDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCV 121
+ + W +AIC+ NDR IG +S++ P D+ R E+GYVLGSKYWGKG+AT
Sbjct: 66 LNDALLPHPWLRAICLDNDRPIGSISVT---PVDEIRG---EIGYVLGSKYWGKGIATEA 119
Query: 122 VKQVVKVAFCELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIIS 181
V+ V F E ++RLEALVDV+N GSQ+VLEK GF KEGV+RK++ +KG RDM++
Sbjct: 120 VRLVAGEIFKEKPEMQRLEALVDVDNVGSQKVLEKVGFVKEGVMRKFMYLKGNVRDMVMF 179
Query: 182 SVLFTD 187
S L +D
Sbjct: 180 SFLPSD 185
>At2g32020 putative alanine acetyl transferase
Length = 183
Score = 164 bits (414), Expect = 3e-41
Identities = 84/168 (50%), Positives = 112/168 (66%), Gaps = 7/168 (4%)
Query: 21 QITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCKAICI-N 79
+I+LRP+ LSD+DD M+W TD KVA+FC+WE TS+D+ I +I + W +AIC+ +
Sbjct: 19 RISLRPMTLSDVDDYMVWATDPKVARFCTWEPCTSRDEAIKYITDRVLTHPWLRAICLED 78
Query: 80 DRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLERL 139
DR IG + + + N E+GYVL KYWGKG AT V+ V F E +ERL
Sbjct: 79 DRPIGYILIMAVD------NIRKEIGYVLARKYWGKGFATEAVRLVTAEVFEEFPEIERL 132
Query: 140 EALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTD 187
EALVDV+N GSQRVLEK GF +EGV+RK++ +KG RD ++ S L TD
Sbjct: 133 EALVDVDNVGSQRVLEKVGFTREGVMRKFICIKGSVRDTVMFSFLSTD 180
>At3g22560 hypothetical protein
Length = 175
Score = 120 bits (300), Expect = 6e-28
Identities = 66/169 (39%), Positives = 99/169 (58%), Gaps = 5/169 (2%)
Query: 21 QITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLWCKAICI-- 78
+I LRP NLSD +D+ W D+ V ++ W+ S ++ I N A W ++I +
Sbjct: 7 RIFLRPFNLSDAEDVFKWAGDDDVTRYLRWDSVNSLEEAKQHILNKAIPHPWRRSISLLQ 66
Query: 79 NDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLER 138
+ +IG VS+ S + R A+L Y + ++WG+G+AT V+ V+ A + + R
Sbjct: 67 DGHSIGYVSVKPDSGDGRCR---ADLAYAVAKEFWGRGIATAAVRMAVEQALEDFPEVVR 123
Query: 139 LEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTD 187
L+A+V+VEN SQRVLEKAGF+KEG+L KY KG RDM + S + D
Sbjct: 124 LQAVVEVENKASQRVLEKAGFRKEGLLEKYGFSKGVIRDMFLYSYVKDD 172
>At1g29470 unknown protein
Length = 768
Score = 28.5 bits (62), Expect = 2.2
Identities = 12/29 (41%), Positives = 16/29 (54%)
Query: 33 DDLMIWTTDEKVAKFCSWELYTSKDDGIN 61
+D+ IW K+ K WEL T K D +N
Sbjct: 473 EDVGIWKAMSKLTKAMCWELMTIKKDELN 501
>At5g08050 putative protein
Length = 158
Score = 27.3 bits (59), Expect = 5.0
Identities = 25/90 (27%), Positives = 38/90 (41%), Gaps = 12/90 (13%)
Query: 85 CVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCELSYLERLEALVD 144
C SL+ S +K + A + K AT VK + +V E+L+ L
Sbjct: 7 CSSLTVHSMANKKPSPSAATRTITSKK----STATPQVKLLTRV--------EQLKLLTK 54
Query: 145 VENAGSQRVLEKAGFQKEGVLRKYLVMKGK 174
E AG + EK+GF + R L+ K +
Sbjct: 55 AEKAGLLSLAEKSGFSLSTIERLGLLTKAE 84
>At5g04430 putative RNA-binding protein
Length = 313
Score = 26.9 bits (58), Expect = 6.5
Identities = 12/29 (41%), Positives = 19/29 (65%)
Query: 17 IDLTQITLRPLNLSDLDDLMIWTTDEKVA 45
+++TQ+T + +SD D M TTD KV+
Sbjct: 257 MEITQMTGARIKISDRGDFMSGTTDRKVS 285
>At4g13290 cytochrome p450 - like protein
Length = 490
Score = 26.9 bits (58), Expect = 6.5
Identities = 16/45 (35%), Positives = 26/45 (57%), Gaps = 4/45 (8%)
Query: 26 PLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKF 70
P+NLS L + T++ + + Y+SK+DGI+ +ENI F
Sbjct: 171 PVNLSQL---FMTLTNDIICRAALGRKYSSKEDGID-VENIVRAF 211
>At3g59390 putative protein
Length = 273
Score = 26.9 bits (58), Expect = 6.5
Identities = 17/79 (21%), Positives = 32/79 (39%)
Query: 1 MEITSISSKPGAKEERIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGI 60
+ +++I G EER T + R L D ++ + E + C W+ G
Sbjct: 23 LRVSAIRKDIGFLEERSCRTTVQGRYLISDDEGNVCDALSLESRTRCCPWKGERFSCHGC 82
Query: 61 NFIENIATKFLWCKAICIN 79
N + + +C + C+N
Sbjct: 83 NILSQCCNSYEFCVSCCLN 101
>At1g19230 hypothetical protein
Length = 926
Score = 26.9 bits (58), Expect = 6.5
Identities = 11/31 (35%), Positives = 19/31 (60%)
Query: 114 GKGVATCVVKQVVKVAFCELSYLERLEALVD 144
GK ++TCV+ + A+C + + R + LVD
Sbjct: 683 GKDLSTCVIGRSKFSAYCNIDMINRPKLLVD 713
>At1g65300
Length = 274
Score = 26.6 bits (57), Expect = 8.5
Identities = 16/33 (48%), Positives = 20/33 (60%), Gaps = 1/33 (3%)
Query: 130 FCELSYLERLEALVDVENAGSQRVL-EKAGFQK 161
F ELS L+R + +VD E SQR+ EK QK
Sbjct: 66 FMELSVLDRTKKMVDQETFISQRIAKEKEQLQK 98
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,198,965
Number of Sequences: 26719
Number of extensions: 166228
Number of successful extensions: 397
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 387
Number of HSP's gapped (non-prelim): 10
length of query: 190
length of database: 11,318,596
effective HSP length: 94
effective length of query: 96
effective length of database: 8,807,010
effective search space: 845472960
effective search space used: 845472960
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC135320.7


