

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
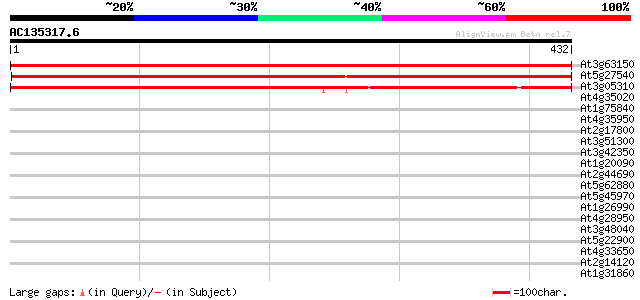
Query= AC135317.6 + phase: 0
(432 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g63150 rac-GTP binding protein like 457 e-129
At5g27540 unknown protein 441 e-124
At3g05310 unknown protein 363 e-100
At4g35020 Rho1Ps homolog/ Rac-like protein 40 0.002
At1g75840 rac-like GTP binding protein (ARAC5) 38 0.009
At4g35950 ras-related small GTP-binding protein 36 0.047
At2g17800 putative GTP-binding protein 36 0.047
At3g51300 rac-like GTP binding protein Arac11 35 0.062
At3g42350 putative protein 35 0.080
At1g20090 RAC-like GTP-binding protein ARAC4 35 0.080
At2g44690 rac-like protein ARAC9 (ARAC9) 35 0.10
At5g62880 Arac10 34 0.14
At5g45970 Rac-like gtp binding protein ARAC2 (sp|Q38903) 34 0.18
At1g26990 polyprotein, putative 32 0.52
At4g28950 rac GTP binding protein Arac7 32 0.68
At3g48040 rac GTP binding protein Arac8 (Arac8) 32 0.89
At5g22900 Na+/H+ antiporter-like protein 30 2.6
At4g33650 dynamin like protein 2a (ADL2a) 30 3.4
At2g14120 dynamin like protein 2b (ADL2b) 29 4.4
At1g31860 unknown protein 29 5.8
>At3g63150 rac-GTP binding protein like
Length = 643
Score = 457 bits (1176), Expect = e-129
Identities = 224/433 (51%), Positives = 311/433 (71%), Gaps = 1/433 (0%)
Query: 1 MDGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFL 60
+DG L D E+N+FQ+ FGA L ++ +K +V P+GV LGLT PGF+ + ++F+
Sbjct: 211 LDGALNDAELNDFQVNCFGAPLDPVELMGVKKVVQERQPDGVTDLGLTLPGFLFLFSLFI 270
Query: 61 KKGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTD 120
++GR ET WA+LRK GY + L+L + LPVP+K++ DQS+EL+ A++FL G+F+L D D
Sbjct: 271 ERGRPETAWAILRKCGYNDSLELHAELLPVPAKQSPDQSIELTNEAMDFLSGIFQLYDLD 330
Query: 121 KDQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSL 180
D L+PAE+D LF AP+SPW + PY +AAE T G +++ GFLS WALMTLLDPR SL
Sbjct: 331 NDGALQPAELDDLFQTAPDSPWLEDPYKEAAEKTPGGSLTINGFLSEWALMTLLDPRKSL 390
Query: 181 ANLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPF 240
ANL IGY +P++ +T +RS DRK Q+TERNVFQC+VFG K +GKSALL + LGR F
Sbjct: 391 ANLTYIGYGHDPASTFSVTRKRSVDRKKQRTERNVFQCFVFGPKKSGKSALLDSFLGRKF 450
Query: 241 SNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSS 300
SN Y T ERYAAN+I+ G+KKTLILREIPED V FL+NK+ LAACDVA V+DSS
Sbjct: 451 SNSYKATMGERYAANVIDQPGGSKKTLILREIPEDRVKKFLTNKESLAACDVAVVVYDSS 510
Query: 301 DGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQELKIDAPI 360
D YSW+++ ++L +V +GE G+ P LL+AAKDDL P+P +V +S +V EL ID P+
Sbjct: 511 DVYSWRKAREILMEVARRGEERGYGTPCLLVAAKDDLDPYPMSVQESDRVCMELGIDIPV 570
Query: 361 RLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLH-TFIFALAGAAMALAG 419
LS+K + N+++S+I++ AE+PH+SIPETE R+ + +QL++ + +F G A+ AG
Sbjct: 571 SLSMKLGEPNSLFSRIVSTAENPHMSIPETESGRRSRNIRQLVNSSLLFVSVGTAVGFAG 630
Query: 420 FTARRARANRNSS 432
A RA + R ++
Sbjct: 631 LAAYRAYSARKNA 643
>At5g27540 unknown protein
Length = 648
Score = 441 bits (1134), Expect = e-124
Identities = 222/433 (51%), Positives = 303/433 (69%), Gaps = 3/433 (0%)
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DG L++ E+N+FQ++ F A LQ +I +K +V +PEGVN GLT GF+ +H +F++
Sbjct: 215 DGALSEAELNDFQVKCFHAPLQPSEIEGVKRVVQEKLPEGVNERGLTVTGFLFLHALFIE 274
Query: 62 KGRTETFWAVLRKFGYGNDLKLRDDFLPVPS-KKASDQSVELSGAAIEFLKGVFRLVDTD 120
KGR ET W VLRKFGY ND++L ++ LP K+A DQS EL+ AAI+FLKG++ L D D
Sbjct: 275 KGRLETTWTVLRKFGYNNDIRLAEELLPSAIFKRAPDQSFELTNAAIDFLKGMYMLFDDD 334
Query: 121 KDQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSL 180
+D LRP E++ LF APESPWK+APY DAAE T +G +S FLS W+LMTLL+P S+
Sbjct: 335 QDNNLRPQEIEDLFSTAPESPWKEAPYEDAAEKTALGGLSFDAFLSMWSLMTLLEPARSV 394
Query: 181 ANLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPF 240
NLI IG+ +PS A+R+T RR DRK Q+ ER VFQC+VFG AGKSALL LGR +
Sbjct: 395 ENLIYIGFPGDPSTAIRVTRRRRLDRKKQQCERKVFQCFVFGPNNAGKSALLNCFLGRSY 454
Query: 241 SNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSS 300
+++ TT ERYA N+++ +G KKTLI+REIPED V S+K+ LAACD+A FV+DSS
Sbjct: 455 TDNQESTTDERYAVNMVD-ESGAKKTLIMREIPEDGVQGLFSSKESLAACDIAVFVYDSS 513
Query: 301 DGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQELKIDAPI 360
D SWKR+ LL +V N GE TG+ P L+++AKDDL P ++ +S ++ Q++ I+ P+
Sbjct: 514 DESSWKRATQLLVEVANYGEATGYEVPCLMVSAKDDLDSSPISIQESTRMTQDMGIEPPV 573
Query: 361 RLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALA-GAAMALAG 419
+S K D NN++ KI+ AA+HPHLSIPETE + RK + +L++ + A++ GAA + G
Sbjct: 574 SISSKLGDFNNLFRKILTAAQHPHLSIPETEAGKSRKHYNRLINRSLMAVSIGAAAVVVG 633
Query: 420 FTARRARANRNSS 432
A R A R SS
Sbjct: 634 LAAYRVYATRKSS 646
>At3g05310 unknown protein
Length = 648
Score = 363 bits (931), Expect = e-100
Identities = 198/441 (44%), Positives = 288/441 (64%), Gaps = 12/441 (2%)
Query: 1 MDGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFL 60
MDG L+D E+NE Q + F L +I Q+K ++ P+GVN GLT GF+ ++ +
Sbjct: 211 MDGILSDEELNELQKKCFDTPLVPCEIKQMKNVMQVTFPQGVNERGLTLDGFLFLNTRLI 270
Query: 61 KKGRTETFWAVLRKFGYGNDLKLRDDFLPVPS-KKASDQSVELSGAAIEFLKGVFRLVDT 119
++ R +T W +LRKFGY NDL+L DD +P S K+ +DQSVEL+ AIEFL+ V+ D+
Sbjct: 271 EEARIQTLWTMLRKFGYSNDLRLGDDLVPYSSFKRQADQSVELTNVAIEFLREVYEFFDS 330
Query: 120 DKDQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYS 179
+ D L P E+ LF+ APESPW Y D E G +SL+ FLS W+LMTL+DP S
Sbjct: 331 NGDNNLEPHEMGYLFETAPESPWTKPLYKDVTEENMDGGLSLEAFLSLWSLMTLIDPPRS 390
Query: 180 LANLICIGY-RDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGR 238
L L+ I + D+PS+A+R+T +R DRK +K+ER V QC+VFG K AGKSALL +GR
Sbjct: 391 LEYLMYIRFPSDDPSSAVRVTRKRVLDRKEKKSERKVVQCFVFGPKNAGKSALLNQFIGR 450
Query: 239 PF---SNDYTPTTVERYAANIIE---LIAGTKKTLILREIPEDEVSNFLSNKDCLAACDV 292
+ SN+ +T E YA N+++ +I+ T KTL+L+E+ + F+ +K+ LAACDV
Sbjct: 451 SYDDDSNNNNGSTDEHYAVNMVKEPGVISDTDKTLVLKEVRIKD-DGFMLSKEALAACDV 509
Query: 293 AAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQ 352
A F++DSSD YSW R++D+L +V + +G+ FP L++AAK DL PFP A+ +S +V Q
Sbjct: 510 AIFIYDSSDEYSWNRAVDMLAEVATIAKDSGYVFPCLMVAAKTDLDPFPVAIQESTRVTQ 569
Query: 353 ELKIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALA- 411
++ IDAPI +S K D +N++ KI+ AAE+PHL+IPE E K+K+ +L + + A++
Sbjct: 570 DIGIDAPIPISSKLGDVSNLFRKILTAAENPHLNIPEIE--SKKKRSCKLNNRSLMAVSI 627
Query: 412 GAAMALAGFTARRARANRNSS 432
G A+ +AG + R R S
Sbjct: 628 GTAVLIAGLASFRLYTARKQS 648
>At4g35020 Rho1Ps homolog/ Rac-like protein
Length = 198
Score = 40.0 bits (92), Expect = 0.002
Identities = 51/194 (26%), Positives = 76/194 (38%), Gaps = 18/194 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN+I + G L L + E
Sbjct: 8 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVI--VDGNTINLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K V + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFLLAFSLVSKASYE-----NVSKKWVPELRHYAPGVPIILVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV S +ELK I AP + + NV + +AA L
Sbjct: 121 DLRDDKQFFAEHPGAVPISTAQGEELKKLIGAPAYIECSAKTQQNV-KAVFDAAIKVVLQ 179
Query: 387 IPETEFVRKRKQHQ 400
P+ + +KRK +
Sbjct: 180 PPKNKKKKKRKSQK 193
>At1g75840 rac-like GTP binding protein (ARAC5)
Length = 196
Score = 38.1 bits (87), Expect = 0.009
Identities = 47/191 (24%), Positives = 75/191 (38%), Gaps = 18/191 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ +L + F DY PT + ++AN++ + G L L + E
Sbjct: 8 KCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV--VDGNTVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVAKKWIPELRHYAPGVPIILVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +ELK I +PI + S NV + +AA L
Sbjct: 121 DLRDDKQFFIDHPGAVPITTNQGEELKKLIGSPIYIECSSKTQQNV-KAVFDAAIKVVLQ 179
Query: 387 IPETEFVRKRK 397
P+ + +K K
Sbjct: 180 PPKQKKKKKNK 190
>At4g35950 ras-related small GTP-binding protein
Length = 197
Score = 35.8 bits (81), Expect = 0.047
Identities = 47/192 (24%), Positives = 75/192 (38%), Gaps = 18/192 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G L L + E
Sbjct: 8 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGATVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHYAPGVPIVLVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +ELK I AP + S NV + +AA L
Sbjct: 121 DLRDDKQFFIDHPGAVPITTVQGEELKKLIGAPAYIECSSKSQENV-KGVFDAAIRVVLQ 179
Query: 387 IPETEFVRKRKQ 398
P+ + + + Q
Sbjct: 180 PPKQKKKKNKAQ 191
>At2g17800 putative GTP-binding protein
Length = 197
Score = 35.8 bits (81), Expect = 0.047
Identities = 47/192 (24%), Positives = 75/192 (38%), Gaps = 18/192 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G L L + E
Sbjct: 8 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGATVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHYAPGVPIVLVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +ELK I AP + S NV + +AA L
Sbjct: 121 DLRDDKQFFIDHPGAVPITTAQGEELKKLIGAPAYIECSSKTQENV-KGVFDAAIRVVLQ 179
Query: 387 IPETEFVRKRKQ 398
P+ + + + Q
Sbjct: 180 PPKQKKKKSKAQ 191
>At3g51300 rac-like GTP binding protein Arac11
Length = 197
Score = 35.4 bits (80), Expect = 0.062
Identities = 46/192 (23%), Positives = 76/192 (38%), Gaps = 18/192 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G+ L L + E
Sbjct: 8 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGSTVNLGLWDTAGQE 65
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + + P +L+ K
Sbjct: 66 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHYAPGVPIVLVGTKL 120
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +EL+ I AP + S NV + +AA L
Sbjct: 121 DLRDDKQFFIDHPGAVPITTAQGEELRKQIGAPTYIECSSKTQENV-KAVFDAAIRVVLQ 179
Query: 387 IPETEFVRKRKQ 398
P+ + + + Q
Sbjct: 180 PPKQKKKKSKAQ 191
>At3g42350 putative protein
Length = 139
Score = 35.0 bits (79), Expect = 0.080
Identities = 16/55 (29%), Positives = 29/55 (52%)
Query: 270 REIPEDEVSNFLSNKDCLAACDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGH 324
+++ +D S F ++ + V S D +SWK++ +L +V N GE TG+
Sbjct: 14 KKVKDDFGSRFSESESTPLVHTIEFIVFSSYDEFSWKKATQMLVEVANYGEATGY 68
>At1g20090 RAC-like GTP-binding protein ARAC4
Length = 195
Score = 35.0 bits (79), Expect = 0.080
Identities = 45/191 (23%), Positives = 74/191 (38%), Gaps = 18/191 (9%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ +L + F DY PT + ++AN++ + G L L + E
Sbjct: 7 KCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV--VDGNTVNLGLWDTAGQE 64
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + P +L+ K
Sbjct: 65 DYNRLRPLSYRGADVFILAFSLISKASYE-----NIAKKWIPELRHYAPGVPIILVGTKL 119
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +ELK I + + + S NV + +AA L
Sbjct: 120 DLRDDKQFFIDHPGAVPITTNQGEELKKLIGSAVYIECSSKTQQNV-KAVFDAAIKVVLQ 178
Query: 387 IPETEFVRKRK 397
P+ + +K K
Sbjct: 179 PPKQKKKKKNK 189
>At2g44690 rac-like protein ARAC9 (ARAC9)
Length = 209
Score = 34.7 bits (78), Expect = 0.10
Identities = 19/65 (29%), Positives = 28/65 (42%), Gaps = 6/65 (9%)
Query: 193 SAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERY 252
SA++ TS S T +C G GK+ LL + F DY PT + +
Sbjct: 2 SASMAATSTSSA------TATTFIKCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNF 55
Query: 253 AANII 257
AN++
Sbjct: 56 NANVL 60
>At5g62880 Arac10
Length = 215
Score = 34.3 bits (77), Expect = 0.14
Identities = 19/65 (29%), Positives = 29/65 (44%), Gaps = 2/65 (3%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ +L F DY PT + ++AN++ + GT L L + E
Sbjct: 10 KCVTVGDGAVGKTCMLICYTSNKFPTDYIPTVFDNFSANVV--VEGTTVNLGLWDTAGQE 67
Query: 277 VSNFL 281
N L
Sbjct: 68 DYNRL 72
>At5g45970 Rac-like gtp binding protein ARAC2 (sp|Q38903)
Length = 201
Score = 33.9 bits (76), Expect = 0.18
Identities = 12/41 (29%), Positives = 21/41 (50%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
+C G GK+ +L + F DY PT + ++AN++
Sbjct: 8 KCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV 48
>At1g26990 polyprotein, putative
Length = 1436
Score = 32.3 bits (72), Expect = 0.52
Identities = 28/81 (34%), Positives = 41/81 (50%), Gaps = 12/81 (14%)
Query: 329 LLIAAKDDLTPFPRAVLDSVKVAQELKIDAPIRLSIKSDDS-----NNVYSKIINAAEHP 383
LL D LT FP + L V+ K+ S++SD++ N +++K A+HP
Sbjct: 652 LLKQKSDVLTVFP-SFLKMVETQYHTKV-----CSVRSDNAHELKFNELFAKEGIKADHP 705
Query: 384 HLSIPETEFVRKRKQHQQLLH 404
PE FV +RK HQ LL+
Sbjct: 706 CPETPEQNFVVERK-HQHLLN 725
>At4g28950 rac GTP binding protein Arac7
Length = 209
Score = 32.0 bits (71), Expect = 0.68
Identities = 12/40 (30%), Positives = 19/40 (47%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANI 256
+C G GK+ +L F DY PT + ++AN+
Sbjct: 8 KCVTVGDGAVGKTCMLICYTSNKFPTDYIPTVFDNFSANV 47
>At3g48040 rac GTP binding protein Arac8 (Arac8)
Length = 208
Score = 31.6 bits (70), Expect = 0.89
Identities = 11/41 (26%), Positives = 19/41 (45%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
+C G GK+ +L F DY PT + ++ N++
Sbjct: 10 KCVTVGDGAVGKTCMLICYTSNKFPTDYIPTVFDNFSVNVV 50
>At5g22900 Na+/H+ antiporter-like protein
Length = 822
Score = 30.0 bits (66), Expect = 2.6
Identities = 25/97 (25%), Positives = 48/97 (48%), Gaps = 12/97 (12%)
Query: 287 LAACDVAAFVHDSSDGYSWKRSIDLLEKVVN--QGELTGHRFPSLLIAAKDDLTPFPRAV 344
+A C V FV+ SS G +++I K +N L+ + + + KDD AV
Sbjct: 638 VAPCSVGVFVYRSSSG---RKNISSGRKTINGTVPNLSSYNICMIFLGGKDD----REAV 690
Query: 345 LDSVKVAQELKIDAPIRLSIKSDD---SNNVYSKIIN 378
+ ++A++ +I+ I I +D+ N V+ K+++
Sbjct: 691 TLATRMARDPRINITIVRLITTDEKARENTVWDKMLD 727
>At4g33650 dynamin like protein 2a (ADL2a)
Length = 808
Score = 29.6 bits (65), Expect = 3.4
Identities = 13/24 (54%), Positives = 19/24 (79%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPF 240
Q V GS+++GKS++L AL+GR F
Sbjct: 61 QVVVVGSQSSGKSSVLEALVGRDF 84
>At2g14120 dynamin like protein 2b (ADL2b)
Length = 780
Score = 29.3 bits (64), Expect = 4.4
Identities = 13/24 (54%), Positives = 19/24 (79%)
Query: 217 QCYVFGSKTAGKSALLRALLGRPF 240
Q V GS+++GKS++L AL+GR F
Sbjct: 45 QVAVVGSQSSGKSSVLEALVGRDF 68
>At1g31860 unknown protein
Length = 281
Score = 28.9 bits (63), Expect = 5.8
Identities = 19/59 (32%), Positives = 28/59 (47%), Gaps = 4/59 (6%)
Query: 120 DKDQLLRPAEVDKLFDAAPESPWKDAPYMDA-AETTDMGYISLKGFLSRWALMTLLDPR 177
D + A+VD L D W D A A+ D G + ++GF++R AL T + R
Sbjct: 42 DNKNIALQAKVDNLLDRIK---WDDKGLAVAIAQNVDTGAVLMQGFVNREALSTTISSR 97
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,355,072
Number of Sequences: 26719
Number of extensions: 393650
Number of successful extensions: 1042
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1019
Number of HSP's gapped (non-prelim): 24
length of query: 432
length of database: 11,318,596
effective HSP length: 102
effective length of query: 330
effective length of database: 8,593,258
effective search space: 2835775140
effective search space used: 2835775140
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC135317.6


