

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
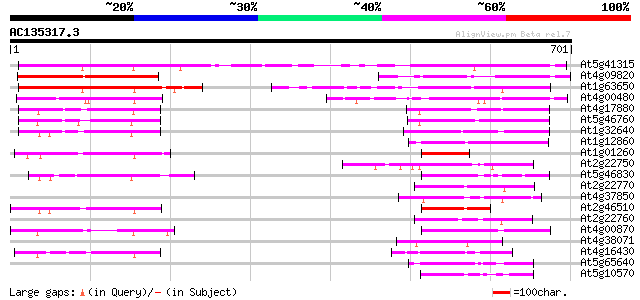
Query= AC135317.3 + phase: 0
(701 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At5g41315 bHLH transcription factor-like protein 288 9e-78
At4g09820 putative protein 261 9e-70
At1g63650 putative transcription factor (BHLH2) 199 4e-51
At4g00480 putative transcription factor BHLH12 137 2e-32
At4g17880 putative transcription factor BHLH4 99 6e-21
At5g46760 putative transcription factor BHLH5 99 1e-20
At1g32640 transcription factor like protein, BHLH6 98 1e-20
At1g12860 bHLH transcription factor (bHLH033) 92 1e-18
At1g01260 transcription factor MYC7E, putative bHLH transcriptio... 88 1e-17
At2g22750 putative bHLH transcription factor (bHLH018) 84 2e-16
At5g46830 putative transcription factor bHLH28 82 8e-16
At2g22770 putative bHLH transcription factor (bHLH020) 82 1e-15
At4g37850 putative bHLH transcription factor (bHLH025) 81 2e-15
At2g46510 putative bHLH transcription factor 79 9e-15
At2g22760 putative bHLH transcription factor (bHLH019) 78 1e-14
At4g00870 putative bHLH transcription factor (bHLH014) 76 7e-14
At4g38071 putative bHLH transcription factor (bHLH131) 73 5e-13
At4g16430 putative transcription factor BHLH3 73 6e-13
At5g65640 putative bHLH transcription factor (bHLH093) 72 8e-13
At5g10570 putative bHLH transcription factor (bHLH061) 71 2e-12
>At5g41315 bHLH transcription factor-like protein
Length = 637
Score = 288 bits (736), Expect = 9e-78
Identities = 221/724 (30%), Positives = 338/724 (46%), Gaps = 142/724 (19%)
Query: 11 LQNMLQAAVQSVQWTYSLFWQLCPQQL-ILVWGDGYYNGSIKTRKTVQPMEVSAEEASLQ 69
L+ L +V+++QW+Y +FW + Q +L WGDGYYNG IKTRKT+Q E+ A++ L+
Sbjct: 14 LKKHLAVSVRNIQWSYGIFWSVSASQSGVLEWGDGYYNGDIKTRKTIQASEIKADQLGLR 73
Query: 70 RSQQLRELYESLSAGETNPP--------TRRP-CASLSPEDLTESEWFYLMCVSFSFPPG 120
RS+QL ELYESLS E++ TRR A+LSPEDL ++EW+YL+C+SF F G
Sbjct: 74 RSEQLSELYESLSVAESSSSGVAAGSQVTRRASAAALSPEDLADTEWYYLVCMSFVFNIG 133
Query: 121 VGLPGRAYTKRQHIWLTGANEVDSKIFSRAILA-----KTVVCIPVLDGVVEFGTTDKVQ 175
G+PGR + + IWL A+ DSK+FSR++LA KTVVC P L GVVE GTT+ +
Sbjct: 134 EGMPGRTFANGEPIWLCNAHTADSKVFSRSLLAKSAAVKTVVCFPFLGGVVEIGTTEHIT 193
Query: 176 EDLNFIKHVKSFFLDGHS--LPPKPALSEHSTSNPTSST----DHIPTIMYTMADPPSTN 229
ED+N I+ VK+ FL+ PA S++ N D I M++ P+ +
Sbjct: 194 EDMNVIQCVKTSFLEAPDPYATILPARSDYHIDNVLDPQQILGDEIYAPMFSTEPFPTAS 253
Query: 230 PNQDDMDEDEEEEEEEDEEDDEVESESEDETGGRIRRATSMTAIVEPSELMQIEMPDDIR 289
P++ D+E E+ D+ D + +E TGG S++ ++ DD
Sbjct: 254 PSRTTNGFDQEHEQVADDHDSFM---TERITGG-------------ASQVQSWQLMDDEL 297
Query: 290 IGSPNDGSNHLDSDFHLLVVSNQGNPLGHVDSYKTDLTQRWGPIEEPVGNLLQVQL-PSS 348
+ N D V G + QR G I+E N+ + P +
Sbjct: 298 SNCVHQSLNSSDCVSQTFVEGAAGRV---AYGARKSRVQRLGQIQEQQRNVKTLSFDPRN 354
Query: 349 DPYIYMWSILINSIVYPIFAGSTSMQNRTQFNAQPPKSKNIHTPHVLHHQLEDLTQEDTH 408
D Y S++ IF + + QF
Sbjct: 355 DDVHY------QSVISTIFKTNHQLILGPQFR---------------------------- 380
Query: 409 YSQTVSTILQNQSTQWTIDSSPSINYITSSNQSSFTNWTNHHNFHPLPPPETTTTTSQCL 468
+ Q+ T+W SS+ SS T T T SQ +
Sbjct: 381 -----NCDKQSSFTRWK----------KSSSSSSGT--------------ATVTAPSQGM 411
Query: 469 LKYILFTVPYLHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLN 528
LK I+F VP +H K KL + + NH + E++RREKLN
Sbjct: 412 LKKIIFDVPRVHQK-----------------EKLMLDSPEARDETGNHAVLEKKRREKLN 454
Query: 529 ERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQIETEQQ--------- 579
ERF+ LR ++P + K+DK SIL DTIEYL++L R++Q+LE+ +TE +
Sbjct: 455 ERFMTLRKIIPSINKIDKVSILDDTIEYLQELERRVQELESCRESTDTETRGTMTMKRKK 514
Query: 580 -----SRSGVTVLVGPT-DKKKVRIVEECGATRAKAVETEVVSSVQVSIIESDALLEIEC 633
R+ T + KKV + A A T + ++++ ++ ++E+ C
Sbjct: 515 PCDAGERTSANCANNETGNGKKVSVNNVGEAEPADTGFTGLTDNLRIGSFGNEVVIELRC 574
Query: 634 LHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKVKENGGNGKKVSIV-EVKRA 692
REG+LL++M ++ +L ++ VQSS +G+ + K K G K++ +K A
Sbjct: 575 AWREGVLLEIMDVISDLHLDSHSVQSSTGDGLLCLTVNCKHK-----GSKIATPGMIKEA 629
Query: 693 LNQI 696
L ++
Sbjct: 630 LQRV 633
>At4g09820 putative protein
Length = 379
Score = 261 bits (667), Expect = 9e-70
Identities = 124/177 (70%), Positives = 148/177 (83%), Gaps = 1/177 (0%)
Query: 10 KLQNMLQAAVQSVQWTYSLFWQLCPQQLILVWGDGYYNGSIKTRKTVQPMEVSAEEASLQ 69
+LQ +L+ AVQSV WTYS+FWQ CPQQ +LVWG+GYYNG+IKTRKT QP EV+AEEA+L+
Sbjct: 19 ELQGLLKTAVQSVDWTYSVFWQFCPQQRVLVWGNGYYNGAIKTRKTTQPAEVTAEEAALE 78
Query: 70 RSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFPPGVGLPGRAYT 129
RSQQLRELYE+L AGE+ R C +LSPEDLTE+EWFYLMCVSFSFPP G+PG+AY
Sbjct: 79 RSQQLRELYETLLAGESTSEARA-CTALSPEDLTETEWFYLMCVSFSFPPPSGMPGKAYA 137
Query: 130 KRQHIWLTGANEVDSKIFSRAILAKTVVCIPVLDGVVEFGTTDKVQEDLNFIKHVKS 186
+R+H+WL+GANEVDSK FSRAILAKTVVCIP+LDGVVE GTT K ++ +K S
Sbjct: 138 RRKHVWLSGANEVDSKTFSRAILAKTVVCIPMLDGVVELGTTKKNGKEHQQVKTAPS 194
Score = 168 bits (425), Expect = 1e-41
Identities = 103/240 (42%), Positives = 141/240 (57%), Gaps = 54/240 (22%)
Query: 461 TTTTSQCLLKYILFTVPYLHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAE 520
T +SQ +LK ++F VP+LH D K P+++LS HV+AE
Sbjct: 191 TAPSSQWVLKQMIFRVPFLHDNTKD-------------------KRLPREDLS--HVVAE 229
Query: 521 RRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQIETEQQS 580
RRRREKLNE+FI LRS+VPFVTKMDK SILGDTI Y+ LR+++ +LE + EQQ
Sbjct: 230 RRRREKLNEKFITLRSMVPFVTKMDKVSILGDTIAYVNHLRKRVHELENTHH----EQQH 285
Query: 581 RSGVTVLVGPTDKKKVRIVEECGATRAKAVETEVVSSVQVSIIESDALLEIECLHREGLL 640
+ R + + + V+VSIIE+D LLE+ C +R+GLL
Sbjct: 286 K------------------------RTRTCKRKTSEEVEVSIIENDVLLEMRCEYRDGLL 321
Query: 641 LDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKVKENGGNGKKVSIVEVKRALNQIIPHN 700
LD++ +L EL IE V +S+N+ F AE+RAKV+ GKK SI EVKRA++Q+I H+
Sbjct: 322 LDILQVLHELGIETTAVHTSVNDHDFEAEIRAKVR-----GKKASIAEVKRAIHQVIIHD 376
>At1g63650 putative transcription factor (BHLH2)
Length = 596
Score = 199 bits (506), Expect = 4e-51
Identities = 109/246 (44%), Positives = 154/246 (62%), Gaps = 20/246 (8%)
Query: 11 LQNMLQAAVQSVQWTYSLFWQLCPQQL-ILVWGDGYYNGSIKTRKTVQPMEVSAEEASLQ 69
L+ L +V+++QW+Y +FW + Q +L WGDGYYNG IKTRKT+Q EV ++ L+
Sbjct: 13 LKKQLAVSVRNIQWSYGIFWSVSASQPGVLEWGDGYYNGDIKTRKTIQAAEVKIDQLGLE 72
Query: 70 RSQQLRELYESLSAGETNPP-----TRRP-CASLSPEDLTESEWFYLMCVSFSFPPGVGL 123
RS+QLRELYESLS E++ TRR A+LSPEDLT++EW+YL+C+SF F G G+
Sbjct: 73 RSEQLRELYESLSLAESSASGSSQVTRRASAAALSPEDLTDTEWYYLVCMSFVFNIGEGI 132
Query: 124 PGRAYTKRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPVLDGVVEFGTTDKVQEDL 178
PG A + + IWL A DSK+F+R++LAK TVVC P L GV+E GTT+ ++ED+
Sbjct: 133 PGGALSNGEPIWLCNAETADSKVFTRSLLAKSASLQTVVCFPFLGGVLEIGTTEHIKEDM 192
Query: 179 NFIKHVKSFFLDGHSLPPKPALSEHS----TSNPTSSTDHIPTIMYTMADPPSTNPNQDD 234
N I+ VK+ FL+ PP +S S +P S + P + T A P ++ +
Sbjct: 193 NVIQSVKTLFLEA---PPYTTISTRSDYQEIFDPLSDDKYTP-VFITEAFPTTSTSGFEQ 248
Query: 235 MDEDEE 240
ED +
Sbjct: 249 EPEDHD 254
Score = 131 bits (330), Expect = 1e-30
Identities = 102/361 (28%), Positives = 169/361 (46%), Gaps = 64/361 (17%)
Query: 328 QRWGPIEEPVGNLLQVQLPSSDPYIYMWSILINSIVYPIFAGSTSMQNRTQFNAQPPKSK 387
Q W + E + N + L SSD V F G+T + P KS+
Sbjct: 266 QSWQFVGEEISNCIHQSLNSSD------------CVSQTFVGTTG-----RLACDPRKSR 308
Query: 388 NIHTPHVLHHQLEDLTQEDTHYSQTVSTILQNQSTQWTIDSSPSINYITSSNQSSFTNWT 447
+ +D HY +STI + +T I N+ +SSFT W
Sbjct: 309 IQRLGQIQEQSNHVNMDDDVHYQGVISTIFK--TTHQLILGPQFQNF---DKRSSFTRWK 363
Query: 448 NHHNFHPLPPPETTTTTSQCLLKYILFTVPYLHTKNHDETSPQTHDAGVDPSSKLRGKGT 507
+ +T SQ ++K ILF VP ++ K +E P T
Sbjct: 364 RSSSV------KTLGEKSQKMIKKILFEVPLMNKK--EELLPDT---------------- 399
Query: 508 PQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDL 567
E + NH L+E++RREKLNERF+ LRS++P ++K+DK SIL DTIEYL+ L++++Q+L
Sbjct: 400 --PEETGNHALSEKKRREKLNERFMTLRSIIPSISKIDKVSILDDTIEYLQDLQKRVQEL 457
Query: 568 ETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEV------------- 614
E+ +TE +R + P D+++ R C ++ K + V
Sbjct: 458 ESCRESADTE--TRITMMKRKKPDDEEE-RASANCMNSKRKGSDVNVGEDEPADIGYAGL 514
Query: 615 VSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKV 674
++++S + ++ ++E+ C REG+LL++M ++ +L ++ VQSS +G+ + K
Sbjct: 515 TDNLRISSLGNEVVIELRCAWREGILLEIMDVISDLNLDSHSVQSSTGDGLLCLTVNCKH 574
Query: 675 K 675
K
Sbjct: 575 K 575
>At4g00480 putative transcription factor BHLH12
Length = 526
Score = 137 bits (346), Expect = 2e-32
Identities = 78/202 (38%), Positives = 117/202 (57%), Gaps = 23/202 (11%)
Query: 9 SKLQNMLQAAVQSVQWTYSLFWQLC-PQQLILVWGDGYYNGSIKTRKTVQPMEVSAEEAS 67
S L+ L AV+SVQW+Y++FW Q +L WG+G YNG +K RK S +
Sbjct: 21 SLLRKQLALAVRSVQWSYAIFWSSSLTQPGVLEWGEGCYNGDMKKRKKSYE---SHYKYG 77
Query: 68 LQRSQQLRELYESLSAGETNPPTRRP----------CAS----LSPEDLTESEWFYLMCV 113
LQ+S++LR+LY S+ G++ C S LSP+DL++ EW+YL+ +
Sbjct: 78 LQKSKELRKLYLSMLEGDSGTTVSTTHDNLNDDDDNCHSTSMMLSPDDLSDEEWYYLVSM 137
Query: 114 SFSFPPGVGLPGRAYTKRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPVLDGVVEF 168
S+ F P LPGRA + IWL A ++K+FSR++LA+ TVVC P L GV+E
Sbjct: 138 SYVFSPSQCLPGRASATGETIWLCNAQYAENKLFSRSLLARSASIQTVVCFPYLGGVIEL 197
Query: 169 GTTDKVQEDLNFIKHVKSFFLD 190
G T+ + ED N ++++KS ++
Sbjct: 198 GVTELISEDHNLLRNIKSCLME 219
Score = 99.8 bits (247), Expect = 5e-21
Identities = 87/315 (27%), Positives = 148/315 (46%), Gaps = 47/315 (14%)
Query: 397 HQLE-DLTQEDTHYSQTVSTILQNQSTQWTIDSSPSI-----NYITSSNQSSFTNWTNHH 450
HQL ++ ED HY +T+ST+L N S + + +I N +TS SSF W
Sbjct: 241 HQLPLGISDEDLHYKRTISTVL-NYSADRSGKNDKNIRHRQPNIVTSEPGSSFLRWKQCE 299
Query: 451 NFHPLPPPETTTTTSQCLLKYILFTVPYLHTKNHDETSPQTHDAGVDPSSKLRGKGTPQD 510
SQ +L+ IL VP +HTK + PS + G QD
Sbjct: 300 Q---QVSGFVQKKKSQNVLRKILHDVPLMHTKR------------MFPS---QNSGLNQD 341
Query: 511 ELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETR 570
+ S R K NE+F +LR++VP V ++DK SIL +TI+YL++L ++++LE+
Sbjct: 342 DPSD---------RRKENEKFSVLRTMVPTVNEVDKESILNNTIKYLQELEARVEELESC 392
Query: 571 NRQIETEQQSRSGV-----TVLVGPTD---KKKVRIVEECGATRAKAVETEVVSSVQVSI 622
+ ++ R +VL+ T +I + G T V + + ++V +
Sbjct: 393 MGSVNFVERQRKTTENLNDSVLIEETSGNYDDSTKIDDNSGETEQVTVFRD-KTHLRVKL 451
Query: 623 IESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKVKENGGNGK 682
E++ ++E+ C +R+ ++ D+M L L ++ V+S N L+AK +
Sbjct: 452 KETEVVIEVRCSYRDYIVADIMETLSNLHMDAFSVRSHTLNKFLTLNLKAKFR----GAA 507
Query: 683 KVSIVEVKRALNQII 697
S+ +KR L ++I
Sbjct: 508 VASVGMIKRELRRVI 522
>At4g17880 putative transcription factor BHLH4
Length = 589
Score = 99.4 bits (246), Expect = 6e-21
Identities = 60/195 (30%), Positives = 98/195 (49%), Gaps = 24/195 (12%)
Query: 11 LQNMLQAAVQSVQ--WTYSLFWQLCP----------QQLILVWGDGYYNGSIKTRKTVQP 58
LQ LQA ++ WTY++FWQ ++L WGDGYY G + + +
Sbjct: 63 LQQRLQALIEGANENWTYAVFWQSSHGFAGEDNNNNNTVLLGWGDGYYKGEEEKSRKKKS 122
Query: 59 MEVSAEEASLQRSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFP 118
SA E R + +REL +S G E++T++EWF+L+ ++ SF
Sbjct: 123 NPASAAEQE-HRKRVIRELNSLISGGVGGGD------EAGDEEVTDTEWFFLVSMTQSFV 175
Query: 119 PGVGLPGRAYTKRQHIWLTGANEVDSKIFSRAILA-----KTVVCIPVLDGVVEFGTTDK 173
G GLPG+A++ IWL+G+N + RA +T+VC+ +GVVE G+++
Sbjct: 176 KGTGLPGQAFSNSDTIWLSGSNALAGSSCERARQGQIYGLQTMVCVATENGVVELGSSEI 235
Query: 174 VQEDLNFIKHVKSFF 188
+ + + + V +FF
Sbjct: 236 IHQSSDLVDKVDTFF 250
Score = 92.8 bits (229), Expect = 6e-19
Identities = 62/182 (34%), Positives = 96/182 (52%), Gaps = 11/182 (6%)
Query: 496 VDPSSKLRGKGTPQD---ELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGD 552
V+P K R +G E NHV AER+RREKLN+RF LR++VP V+KMDKAS+LGD
Sbjct: 394 VEPEKKPRKRGRKPANGREEPLNHVEAERQRREKLNQRFYSLRAVVPNVSKMDKASLLGD 453
Query: 553 TIEYLKQLRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVET 612
I Y+ +L+ K+Q E+ +++ + + V+ K + + + +V
Sbjct: 454 AISYISELKSKLQKAESDKEELQKQ------IDVMNKEAGNAKSSVKDRKCLNQESSVLI 507
Query: 613 EVVSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRA 672
E+ V V II DA++ I+C R M L+EL +EV S+ N + + +
Sbjct: 508 EM--EVDVKIIGWDAMIRIQCSKRNHPGAKFMEALKELDLEVNHASLSVVNDLMIQQATV 565
Query: 673 KV 674
K+
Sbjct: 566 KM 567
>At5g46760 putative transcription factor BHLH5
Length = 592
Score = 98.6 bits (244), Expect = 1e-20
Identities = 63/198 (31%), Positives = 99/198 (49%), Gaps = 34/198 (17%)
Query: 11 LQNMLQAAVQSV--QWTYSLFWQLC--------PQQLILVWGDGYYNGSIKTRKTVQPME 60
LQ LQA ++S WTY++FWQ+ +IL WGDGYY G K
Sbjct: 52 LQQRLQALIESAGENWTYAIFWQISHDFDSSTGDNTVILGWGDGYYKGEEDKEKKKNNTN 111
Query: 61 VSAEEASLQRSQQLRELYESLSAG-----ETNPPTRRPCASLSPEDLTESEWFYLMCVSF 115
+ +E R + +REL +S G E+N E++T++EWF+L+ ++
Sbjct: 112 TAEQE---HRKRVIRELNSLISGGIGVSDESND-----------EEVTDTEWFFLVSMTQ 157
Query: 116 SFPPGVGLPGRAYTKRQHIWLTGANEVDSKIFSRA-----ILAKTVVCIPVLDGVVEFGT 170
SF GVGLPG ++ + IWL+G+ + RA KT+VCI +GVVE G+
Sbjct: 158 SFVNGVGLPGESFLNSRVIWLSGSGALTGSGCERAGQGQIYGLKTMVCIATQNGVVELGS 217
Query: 171 TDKVQEDLNFIKHVKSFF 188
++ + + + + V + F
Sbjct: 218 SEVISQSSDLMHKVNNLF 235
Score = 89.4 bits (220), Expect = 6e-18
Identities = 58/180 (32%), Positives = 93/180 (51%), Gaps = 7/180 (3%)
Query: 498 PSSKLRGKGTPQD---ELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTI 554
P K R +G E NHV AER+RREKLN+RF LR++VP V+KMDKAS+LGD I
Sbjct: 395 PEKKPRKRGRKPANGREEPLNHVEAERQRREKLNQRFYSLRAVVPNVSKMDKASLLGDAI 454
Query: 555 EYLKQLRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEV 614
Y+ +L+ K+Q E+ +I+ + S G K +E ++ + + +
Sbjct: 455 SYINELKSKLQQAESDKEEIQKKLDGMS----KEGNNGKGCGSRAKERKSSNQDSTASSI 510
Query: 615 VSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKV 674
+ V II D ++ ++C ++ M L+EL +EV S+ N + + + K+
Sbjct: 511 EMEIDVKIIGWDVMIRVQCGKKDHPGARFMEALKELDLEVNHASLSVVNDLMIQQATVKM 570
>At1g32640 transcription factor like protein, BHLH6
Length = 623
Score = 98.2 bits (243), Expect = 1e-20
Identities = 60/193 (31%), Positives = 98/193 (50%), Gaps = 23/193 (11%)
Query: 11 LQNMLQAAVQSVQ--WTYSLFWQLCPQ---QLILVWGDGYYNG-----SIKTRKTVQPME 60
LQ LQA ++ WTY++FWQ +L WGDGYY G + + R + P
Sbjct: 68 LQQRLQALIEGTHEGWTYAIFWQPSYDFSGASVLGWGDGYYKGEEDKANPRRRSSSPPFS 127
Query: 61 VSAEEASLQRSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFPPG 120
A++ R + LREL +S G P E++T++EWF+L+ ++ SF G
Sbjct: 128 TPADQE--YRKKVLRELNSLISGGVA------PSDDAVDEEVTDTEWFFLVSMTQSFACG 179
Query: 121 VGLPGRAYTKRQHIWLTGANEVDSKIFSRA-----ILAKTVVCIPVLDGVVEFGTTDKVQ 175
GL G+A+ +W++G++++ RA T+ CIP +GVVE G+T+ ++
Sbjct: 180 AGLAGKAFATGNAVWVSGSDQLSGSGCERAKQGGVFGMHTIACIPSANGVVEVGSTEPIR 239
Query: 176 EDLNFIKHVKSFF 188
+ + I V+ F
Sbjct: 240 QSSDLINKVRILF 252
Score = 86.3 bits (212), Expect = 5e-17
Identities = 58/184 (31%), Positives = 95/184 (51%), Gaps = 10/184 (5%)
Query: 493 DAGVDPSSKLRGKGTPQD-ELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILG 551
+ V+ K RG+ E NHV AER+RREKLN+RF LR++VP V+KMDKAS+LG
Sbjct: 429 EVAVEKRPKKRGRKPANGREEPLNHVEAERQRREKLNQRFYALRAVVPNVSKMDKASLLG 488
Query: 552 DTIEYLKQLRRKIQDLETRNRQIETEQQSRSGVTVLVG-PTDKKKVRIVEECGATRAKAV 610
D I Y+ +L+ K+ ++T + +++ + Q L G + C + + +
Sbjct: 489 DAIAYINELKSKV--VKTESEKLQIKNQLEEVKLELAGRKASASGGDMSSSCSSIKPVGM 546
Query: 611 ETEVVSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAEL 670
E ++V II DA++ +E R +M L +L +EV S+ N + + +
Sbjct: 547 E------IEVKIIGWDAMIRVESSKRNHPAARLMSALMDLELEVNHASMSVVNDLMIQQA 600
Query: 671 RAKV 674
K+
Sbjct: 601 TVKM 604
>At1g12860 bHLH transcription factor (bHLH033)
Length = 450
Score = 91.7 bits (226), Expect = 1e-18
Identities = 64/176 (36%), Positives = 97/176 (54%), Gaps = 9/176 (5%)
Query: 499 SSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLK 558
++K + KG P A +++AERRRR+KLN+R +LRS+VP ++KMD+ASILGD I+YLK
Sbjct: 256 NNKGKKKGMP-----AKNLMAERRRRKKLNDRLYMLRSVVPKISKMDRASILGDAIDYLK 310
Query: 559 QLRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEVVSSV 618
+L ++I DL T ++E+ S S + L R+ EE + + V
Sbjct: 311 ELLQRINDLHT---ELESTPPSSSSLHPLTPTPQTLSYRVKEELCPSSSLPSPKGQQPRV 367
Query: 619 QVSIIESDAL-LEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAK 673
+V + E A+ + + C R GLLL M L L ++V S NG + RA+
Sbjct: 368 EVRLREGKAVNIHMFCGRRPGLLLSTMRALDNLGLDVQQAVISCFNGFALDVFRAE 423
>At1g01260 transcription factor MYC7E, putative bHLH transcription
factor (AtbHLH013)
Length = 590
Score = 88.2 bits (217), Expect = 1e-17
Identities = 70/214 (32%), Positives = 107/214 (49%), Gaps = 27/214 (12%)
Query: 7 LGS--KLQNMLQAAVQ-----SVQWTYSLFWQLCPQQ---LILVWGDGYYNGSIKTRKT- 55
LGS LQN L V+ + W Y++FWQ+ + L+L WGDGY + K+
Sbjct: 42 LGSDENLQNKLSDLVERPNASNFSWNYAIFWQISRSKAGDLVLCWGDGYCREPKEGEKSE 101
Query: 56 -VQPMEVSAEEASLQ--RSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMC 112
V+ + + EE + Q R + L++L++ E CA L + +T++E F L
Sbjct: 102 IVRILSMGREEETHQTMRKRVLQKLHDLFGGSE-----EENCA-LGLDRVTDTEMFLLSS 155
Query: 113 VSFSFPPGVGLPGRAYTKRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPVLDGVVE 167
+ FSFP G G PG+ + + +WL+ S R+ LAK TVV +P GVVE
Sbjct: 156 MYFSFPRGEGGPGKCFASAKPVWLSDVVNSGSDYCVRSFLAKSAGIQTVVLVPTDLGVVE 215
Query: 168 FGTTDKVQEDLNFIKHVKSFFLDGHSLPPKPALS 201
G+T + E + I ++S F SLPP A++
Sbjct: 216 LGSTSCLPESEDSILSIRSLFTS--SLPPVRAVA 247
Score = 72.4 bits (176), Expect = 8e-13
Identities = 32/60 (53%), Positives = 46/60 (76%)
Query: 515 NHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQI 574
NHV AER+RREKLN+RF LRS+VP ++KMDKAS+LGD + Y+ +L K++ +E ++
Sbjct: 433 NHVEAERQRREKLNQRFYALRSVVPNISKMDKASLLGDAVSYINELHAKLKVMEAERERL 492
>At2g22750 putative bHLH transcription factor (bHLH018)
Length = 304
Score = 84.3 bits (207), Expect = 2e-16
Identities = 67/262 (25%), Positives = 123/262 (46%), Gaps = 38/262 (14%)
Query: 417 LQNQSTQWTIDSSPSINYITSSNQSSFTNWTNHHNFHPL----PPPETTTTTSQCLLKYI 472
L + +T + +D+S + + + N HP PPP +S+ L
Sbjct: 15 LDDTTTCYNLDASCNKSLVEERPSKILKTTHISPNLHPFSSSNPPPPKHQPSSRILS--- 71
Query: 473 LFTVPYLHTKNHDET----SPQTHDAGVDPSSK----LRGKGTPQD-----ELSANHVLA 519
F LH NH+ SP+ + G+ K +RG Q + +H+LA
Sbjct: 72 -FEKTGLHVMNHNSPNLIFSPKDEEIGLPEHKKAELIIRGTKRAQSLTRSQSNAQDHILA 130
Query: 520 ERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQIETEQQ 579
ER+RREKL +RF+ L +L+P + KMDKAS+LGD I+++K L+ +++ E + ++ E
Sbjct: 131 ERKRREKLTQRFVALSALIPGLKKMDKASVLGDAIKHIKYLQESVKEYEEQKKEKTMES- 189
Query: 580 SRSGVTVLVGPTDKKKVRIVEE-------CGATRAKAVETEVVSSVQVSIIESDALLEIE 632
VLV KK +++E + + + + ++V + D L++I
Sbjct: 190 -----VVLV----KKSSLVLDENHQPSSSSSSDGNRNSSSSNLPEIEVRVSGKDVLIKIL 240
Query: 633 CLHREGLLLDVMVMLRELRIEV 654
C ++G ++ +M + +L + +
Sbjct: 241 CEKQKGNVIKIMGEIEKLGLSI 262
>At5g46830 putative transcription factor bHLH28
Length = 511
Score = 82.4 bits (202), Expect = 8e-16
Identities = 57/226 (25%), Positives = 111/226 (48%), Gaps = 33/226 (14%)
Query: 24 WTYSLFWQLCPQ----QLILVWGDGYYNGS--------IKTRKTVQPMEVSAEEASLQRS 71
W+Y++FW+ + +L WGDG Y G ++ +KT+ +S+ E +RS
Sbjct: 49 WSYAIFWKPSYDDFSGEAVLKWGDGVYTGGNEEKTRGRLRRKKTI----LSSPEEKERRS 104
Query: 72 QQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFPPGVGLPGRAYTKR 131
+REL +S GE P + ++T+ EWF+L+ +++SF G GL G+A+
Sbjct: 105 NVIRELNLMIS-GEAFPVVEDDVSDDDDVEVTDMEWFFLVSMTWSFGNGSGLAGKAFASY 163
Query: 132 QHIWLTGANEVDSKIFSRA-----ILAKTVVCIPVLDGVVEFGTTDKVQEDLNFIKHVKS 186
+ +TG++ + RA + +T++CIP +GV+E +T++++ + + ++
Sbjct: 164 NPVLVTGSDLIYGSGCDRAKQGGDVGLQTILCIPSHNGVLELASTEEIRPNSDLFNRIRF 223
Query: 187 FFLDGHSLPPKPALSEHSTSNPTSSTDHIPTIMYTMADPPST-NPN 231
F S++ + P S+++ P + + T NPN
Sbjct: 224 LF----------GGSKYFSGAPNSNSELFPFQLESSCSSTVTGNPN 259
Score = 72.8 bits (177), Expect = 6e-13
Identities = 55/163 (33%), Positives = 84/163 (50%), Gaps = 16/163 (9%)
Query: 515 NHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQI 574
NHV AER RREKLN RF LR++VP V+KMDK S+L D + Y+ +L+ K +++E I
Sbjct: 343 NHVEAERMRREKLNHRFYALRAVVPNVSKMDKTSLLEDAVCYINELKSKAENVELEKHAI 402
Query: 575 ETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEVVSSVQVSIIES-DALLEIEC 633
E + + ++ I C KA E + ++V I+ES DA++ +E
Sbjct: 403 EIQFNELKEIA-------GQRNAIPSVC-KYEEKASE---MMKIEVKIMESDDAMVRVES 451
Query: 634 L--HREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKV 674
H G L M L +L +EV S+ N + + + K+
Sbjct: 452 RKDHHPGARL--MNALMDLELEVNHASISVMNDLMIQQANVKM 492
>At2g22770 putative bHLH transcription factor (bHLH020)
Length = 320
Score = 81.6 bits (200), Expect = 1e-15
Identities = 48/159 (30%), Positives = 85/159 (53%), Gaps = 11/159 (6%)
Query: 506 GTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQ 565
G + L HVLAER+RR+KLNER I L +L+P + K DKA++L D I++LKQL+ +++
Sbjct: 123 GRREPHLLKEHVLAERKRRQKLNERLIALSALLPGLKKTDKATVLEDAIKHLKQLQERVK 182
Query: 566 DLETRNRQIETEQQSRSGVTVLVGPT--DKKKVRIVEECGATRAKAVETEVVS------- 616
LE ++ T++ +S + V D C A + ++ VS
Sbjct: 183 KLE--EERVVTKKMDQSIILVKRSQVYLDDDSSSYSSTCSAASPLSSSSDEVSIFKQTMP 240
Query: 617 SVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVI 655
++ + + D L+ + C +G ++ ++ L + R+EV+
Sbjct: 241 MIEARVSDRDLLIRVHCEKNKGCMIKILSSLEKFRLEVV 279
>At4g37850 putative bHLH transcription factor (bHLH025)
Length = 304
Score = 81.3 bits (199), Expect = 2e-15
Identities = 58/186 (31%), Positives = 99/186 (53%), Gaps = 18/186 (9%)
Query: 486 ETSPQTHDAGVDPSSKLRGKGTPQDELSAN---HVLAERRRREKLNERFIILRSLVPFVT 542
E Q H + + K + P +N H++AER+RREKL +RF+ L +LVP +
Sbjct: 96 EAQVQPHQKSDEFNRKGTKRAQPFSRNQSNAQDHIIAERKRREKLTQRFVALSALVPGLK 155
Query: 543 KMDKASILGDTIEYLKQLRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEEC 602
KMDKAS+LGD ++++K L+ ++ +LE E +++ R VLV KK I+++
Sbjct: 156 KMDKASVLGDALKHIKYLQERVGELE------EQKKERRLESMVLV----KKSKLILDDN 205
Query: 603 GATRAKAVETEV----VSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQ 658
+ + + E + ++V + D L++I C ++G L +M + +L I +I
Sbjct: 206 NQSFSSSCEDGFSDLDLPEIEVRFSDEDVLIKILCEKQKGHLAKIMAEIEKLHI-LITNS 264
Query: 659 SSLNNG 664
S LN G
Sbjct: 265 SVLNFG 270
>At2g46510 putative bHLH transcription factor
Length = 662
Score = 79.0 bits (193), Expect = 9e-15
Identities = 57/205 (27%), Positives = 98/205 (47%), Gaps = 22/205 (10%)
Query: 1 MAAPSPLGSKLQNMLQ-AAVQSVQWTYSLFWQLCPQ---QLILVWGDGYYNG-----SIK 51
M L KL +++ ++ W Y++FWQ Q +L WGDG K
Sbjct: 42 MGTDDTLNKKLSSLVDWPNSENFSWNYAIFWQQTMSRSGQQVLGWGDGCCREPNEEEESK 101
Query: 52 TRKTVQPMEVSAEEASLQ--RSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFY 109
++ + AEE + Q R + L++L+ + + +LS E +T +E F+
Sbjct: 102 VVRSYNFNNMGAEEETWQDMRKRVLQKLHRLFGGSDEDN------YALSLEKVTATEIFF 155
Query: 110 LMCVSFSFPPGVGLPGRAYTKRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPVLDG 164
L + F F G G PGR Y+ +H+WL+ A +S R+ +AK T+V +P G
Sbjct: 156 LASMYFFFNHGEGGPGRCYSSGKHVWLSDAVNSESDYCFRSFMAKSAGIRTIVMVPTDAG 215
Query: 165 VVEFGTTDKVQEDLNFIKHVKSFFL 189
V+E G+ + E++ +K V++ F+
Sbjct: 216 VLELGSVWSLPENIGLVKSVQALFM 240
Score = 77.4 bits (189), Expect = 3e-14
Identities = 38/87 (43%), Positives = 61/87 (69%), Gaps = 2/87 (2%)
Query: 515 NHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQI 574
NHV AER+RREKLN+RF LRS+VP ++KMDKAS+LGD I Y+K+L+ K++ +E + ++
Sbjct: 395 NHVEAERQRREKLNQRFYALRSVVPNISKMDKASLLGDAISYIKELQEKVKIME--DERV 452
Query: 575 ETEQQSRSGVTVLVGPTDKKKVRIVEE 601
T++ T+ V + + ++ + E
Sbjct: 453 GTDKSLSESNTITVEESPEVDIQAMNE 479
>At2g22760 putative bHLH transcription factor (bHLH019)
Length = 295
Score = 78.2 bits (191), Expect = 1e-14
Identities = 53/153 (34%), Positives = 84/153 (54%), Gaps = 13/153 (8%)
Query: 506 GTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQ 565
GT L+ HVLAER+RREKL+E+FI L +L+P + K DK +IL D I +KQL+ +
Sbjct: 110 GTRSPVLAKEHVLAERKRREKLSEKFIALSALLPGLKKADKVTILDDAISRMKQLQ---E 166
Query: 566 DLETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETE---VVSSVQVSI 622
L T + E +Q S + V K KV EE + + +V E + ++ I
Sbjct: 167 QLRTLKEEKEATRQMESMILV-----KKSKVFFDEEPNLSCSPSVHIEFDQALPEIEAKI 221
Query: 623 IESDALLEIECLHREGLLLDVMVMLR--ELRIE 653
++D L+ I C +G +++++ + +LRIE
Sbjct: 222 SQNDILIRILCEKSKGCMINILNTIENFQLRIE 254
>At4g00870 putative bHLH transcription factor (bHLH014)
Length = 423
Score = 75.9 bits (185), Expect = 7e-14
Identities = 51/163 (31%), Positives = 86/163 (52%), Gaps = 7/163 (4%)
Query: 515 NHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQI 574
+HV AE++RREKLN RF LR++VP V++MDKAS+L D + Y++ L+ KI DLET +++
Sbjct: 249 SHVEAEKQRREKLNHRFYALRAIVPKVSRMDKASLLSDAVSYIESLKSKIDDLETEIKKM 308
Query: 575 E-TEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEVVSSVQVSIIESDALLEIEC 633
+ TE + P+ + + + R +E VQV I+ +A++ ++
Sbjct: 309 KMTETDKLDNSSSNTSPSSVEYQVNQKPSKSNRGSDLE------VQVKIVGEEAIIRVQT 362
Query: 634 LHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKVKE 676
+ +M L E+ V +S + V V ++ V E
Sbjct: 363 ENVNHPTSALMSALMEMDCRVQHANASRLSQVMVQDVVVLVPE 405
Score = 64.3 bits (155), Expect = 2e-10
Identities = 59/226 (26%), Positives = 92/226 (40%), Gaps = 55/226 (24%)
Query: 1 MAAPSPLGSKLQNMLQAAVQSV--QWTYSLFWQLC----PQQLILVWGDGYYNGSIKTRK 54
+ + SP LQ L+ V++ +W Y +FWQ + LVW DG++ G+
Sbjct: 25 IVSSSPPDLVLQQKLRFVVETSPDRWAYVIFWQKMFDDQSDRSYLVWVDGHFCGNKNNNS 84
Query: 55 TVQPMEVSAE-EASLQRSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCV 113
S E E + L Y + GE P + +S E L
Sbjct: 85 QENYTTNSIECELMMDGGDDLELFYAASFYGEDRSPRKE----VSDESL----------- 129
Query: 114 SFSFPPGVGLPGRAYTKRQHIWLTGANEVDSKIFSRAILA-----KTVVCIPVLDGVVEF 168
+WLTG +E+ + RA A T+V IP+ +G++E
Sbjct: 130 --------------------VWLTGPDELRFSNYERAKEAGFHGVHTLVSIPINNGIIEL 169
Query: 169 GTTDKVQEDLNFIKHVKSFFLDGHSLP--------PKPALSEHSTS 206
G+++ + ++ NFI VKS F G + PKPA+S+HS S
Sbjct: 170 GSSESIIQNRNFINRVKSIFGSGKTTKHTNQTGSYPKPAVSDHSKS 215
>At4g38071 putative bHLH transcription factor (bHLH131)
Length = 256
Score = 73.2 bits (178), Expect = 5e-13
Identities = 47/150 (31%), Positives = 82/150 (54%), Gaps = 17/150 (11%)
Query: 484 HDETSPQTHDAGVDPSSKLRGKG---------TPQDELSAN-HVLAERRRREKLNERFII 533
+ T ++ AG SSKL +G T E++A H AERRRR ++N +F
Sbjct: 54 YSTTMGRSFFAGAATSSKLFSRGFSVTKPKSKTESKEVAAKKHSDAERRRRLRINSQFAT 113
Query: 534 LRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETR-----NRQIETEQQSRSGVTVLV 588
LR+++P + K DKAS+LG+T+ Y +L++ +QD+ T N +++ +R V+
Sbjct: 114 LRTILPNLVKQDKASVLGETVRYFNELKKMVQDIPTTPSLEDNLRLDHCNNNRDLARVVF 173
Query: 589 GPTDKKKV--RIVEECGATRAKAVETEVVS 616
+D++ + + E A +AKAV E+++
Sbjct: 174 SCSDREGLMSEVAESMKAVKAKAVRAEIMT 203
>At4g16430 putative transcription factor BHLH3
Length = 467
Score = 72.8 bits (177), Expect = 6e-13
Identities = 50/151 (33%), Positives = 84/151 (55%), Gaps = 9/151 (5%)
Query: 478 YLHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSL 537
Y + + DET T + P + R ++E + NHV AER+RREKLN+RF LR++
Sbjct: 286 YGYEQGKDETLYLTDEQ--KPRKRGRKPANGREE-ALNHVEAERQRREKLNQRFYALRAV 342
Query: 538 VPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKKKVR 597
VP ++KMDKAS+L D I Y+ +++KI+ ET +QI ++S + P + +
Sbjct: 343 VPNISKMDKASLLADAITYITDMQKKIRVYET-EKQIMKRRESNQ-----ITPAEVDYQQ 396
Query: 598 IVEECGATRAKAVETEVVSSVQVSIIESDAL 628
++ + +ET VS V ++ E++ +
Sbjct: 397 RHDDAVVRLSCPLETHPVSKVIQTLRENEVM 427
Score = 63.9 bits (154), Expect = 3e-10
Identities = 56/194 (28%), Positives = 86/194 (43%), Gaps = 23/194 (11%)
Query: 6 PLGSKLQNMLQAAVQSVQWTYSLFWQLC----PQQLILVWGDGYYNGSIKTRKTVQPMEV 61
P S LQ L+ V+ W Y+LFW +L+WGDG+ + +K +
Sbjct: 45 PSDSNLQQGLRHVVEGSDWDYALFWLASNVNSSDGCVLIWGDGH----CRVKKGASGEDY 100
Query: 62 SAEEASLQRSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFPPGV 121
S ++ +R LR+L+ S + + + A LT+ + FYL + FSF
Sbjct: 101 SQQDEIKRRV--LRKLHLSFVGSDEDHRLVKSGA------LTDLDMFYLASLYFSFRCDT 152
Query: 122 GLPGRA--YTKRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPVLDGVVEFGTTDKV 174
G A Y + +W S R+ LA+ TV+ +PV GVVE G+ +
Sbjct: 153 NKYGPAGTYVSGKPLWAADLPSCLSYYRVRSFLARSAGFQTVLSVPVNSGVVELGSLRHI 212
Query: 175 QEDLNFIKHVKSFF 188
ED + I+ VKS F
Sbjct: 213 PEDKSVIEMVKSVF 226
>At5g65640 putative bHLH transcription factor (bHLH093)
Length = 351
Score = 72.4 bits (176), Expect = 8e-13
Identities = 50/156 (32%), Positives = 81/156 (51%), Gaps = 20/156 (12%)
Query: 499 SSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLK 558
S KL G+ + +++AERRRR++LN+R +LRS+VP ++KMD+ SILGD I+Y+K
Sbjct: 169 SKKLEGQ-------PSKNLMAERRRRKRLNDRLSMLRSIVPKISKMDRTSILGDAIDYMK 221
Query: 559 QLRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEVVSSV 618
+L KI L+ +++ S + L G D K + E K
Sbjct: 222 ELLDKINKLQDEEQELGNSNNSHH--SKLFG--DLKDLNANEPLVRNSPK---------F 268
Query: 619 QVSIIESDALLEIECLHREGLLLDVMVMLRELRIEV 654
++ + D ++I C + GLLL + L L +E+
Sbjct: 269 EIDRRDEDTRVDICCSPKPGLLLSTVNTLETLGLEI 304
>At5g10570 putative bHLH transcription factor (bHLH061)
Length = 315
Score = 70.9 bits (172), Expect = 2e-12
Identities = 48/141 (34%), Positives = 81/141 (57%), Gaps = 22/141 (15%)
Query: 514 ANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQ 573
+ +++AERRRR++LN+R +LRS+VP +TKMD+ SILGD I+Y+K+L KI L+ +
Sbjct: 150 SKNLMAERRRRKRLNDRLSLLRSIVPKITKMDRTSILGDAIDYMKELLDKINKLQ----E 205
Query: 574 IETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEVVSSVQVSIIESDALLEIEC 633
E E S S ++ L+ T++ VR +++ EV E + ++I C
Sbjct: 206 DEQELGSNSHLSTLI--TNESMVR----------NSLKFEVDQR------EVNTHIDICC 247
Query: 634 LHREGLLLDVMVMLRELRIEV 654
+ GL++ + L L +E+
Sbjct: 248 PTKPGLVVSTVSTLETLGLEI 268
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.313 0.131 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,712,854
Number of Sequences: 26719
Number of extensions: 788146
Number of successful extensions: 14415
Number of sequences better than 10.0: 568
Number of HSP's better than 10.0 without gapping: 408
Number of HSP's successfully gapped in prelim test: 166
Number of HSP's that attempted gapping in prelim test: 7607
Number of HSP's gapped (non-prelim): 2929
length of query: 701
length of database: 11,318,596
effective HSP length: 106
effective length of query: 595
effective length of database: 8,486,382
effective search space: 5049397290
effective search space used: 5049397290
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC135317.3


