

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135311.8 + phase: 0
(189 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
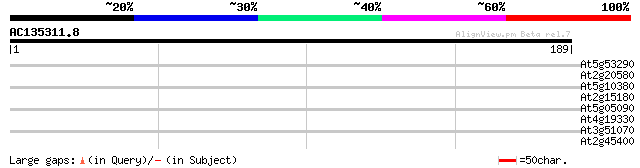
Score E
Sequences producing significant alignments: (bits) Value
At5g53290 unknown protein 29 1.7
At2g20580 putative 26S proteasome regulatory subunit S2 29 1.7
At5g10380 putative protein 28 2.9
At2g15180 hypothetical protein 27 4.9
At5g05090 unknown protein 27 6.5
At4g19330 puatative protein 27 6.5
At3g51070 putative protein 27 8.4
At2g45400 putative flavonol reductase 27 8.4
>At5g53290 unknown protein
Length = 354
Score = 28.9 bits (63), Expect = 1.7
Identities = 21/67 (31%), Positives = 30/67 (44%), Gaps = 6/67 (8%)
Query: 119 RCLNNITISANGRNVVAMVVDECDSRKGCDEQHDYQPPC-----PNNIVDASKAVWKALG 173
R +N IT+ + NVV V + D ++ + Q P PNN V S + K G
Sbjct: 69 RFVNEITVEPSCNNVVTGVSMK-DRKRLSSSSDETQSPASSRQRPNNKVSVSGQIKKFRG 127
Query: 174 VPKEQWG 180
V + WG
Sbjct: 128 VRQRPWG 134
>At2g20580 putative 26S proteasome regulatory subunit S2
Length = 891
Score = 28.9 bits (63), Expect = 1.7
Identities = 13/35 (37%), Positives = 19/35 (54%)
Query: 74 TKAYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVA 108
T A +L Q G S+ DK +HS+D P++A
Sbjct: 413 TSAAASLGMIQLWDVDSGLSQLDKYFHSNDNPIIA 447
>At5g10380 putative protein
Length = 301
Score = 28.1 bits (61), Expect = 2.9
Identities = 27/104 (25%), Positives = 43/104 (40%), Gaps = 12/104 (11%)
Query: 8 VSFIFLAITLIFTSFVYSEAQNCRPSGRIRGKKAPPGKCNQENDSDCCVQGKMYTTYVC- 66
V I L + L+F +++ Q RI PG NQ+ D D + +
Sbjct: 49 VGSIILFLFLVFFLYLHITQQR-----RISAASVTPGDTNQQEDEDETEERDFSDFHHVW 103
Query: 67 -SPSVSTHTKAY--LTLNSFQKGG---DGGGPSECDKQYHSDDT 104
P+V H A +T+ F+KG DG S C ++ D++
Sbjct: 104 QIPTVGLHRSAINSITVVGFKKGEGIIDGTECSVCLNEFEEDES 147
>At2g15180 hypothetical protein
Length = 474
Score = 27.3 bits (59), Expect = 4.9
Identities = 21/62 (33%), Positives = 28/62 (44%), Gaps = 7/62 (11%)
Query: 47 NQENDSDCCVQGKMY--TTYVCSPSVSTHTKAYLTLNSFQKGGDGGGPS-ECDKQYHSDD 103
+ E DS C Q Y T +V S H Y + + F++ DGG P E +Y S
Sbjct: 9 SDEEDSYGCSQDSYYNETRFV----ESWHDDGYSSSSDFEEEPDGGAPEPEPPDRYSSHA 64
Query: 104 TP 105
TP
Sbjct: 65 TP 66
>At5g05090 unknown protein
Length = 266
Score = 26.9 bits (58), Expect = 6.5
Identities = 13/38 (34%), Positives = 20/38 (52%), Gaps = 1/38 (2%)
Query: 69 SVSTHTKAY-LTLNSFQKGGDGGGPSECDKQYHSDDTP 105
+V++H + Y L L + GG GGG + D + S P
Sbjct: 124 NVASHLQKYRLYLKRMKSGGGGGGSGDSDHLFASSPVP 161
>At4g19330 puatative protein
Length = 537
Score = 26.9 bits (58), Expect = 6.5
Identities = 12/35 (34%), Positives = 17/35 (48%)
Query: 39 KKAPPGKCNQENDSDCCVQGKMYTTYVCSPSVSTH 73
+KAP ++N C + GK+Y C STH
Sbjct: 318 RKAPSMMVARKNAFTCVLDGKLYVMGGCEADESTH 352
>At3g51070 putative protein
Length = 895
Score = 26.6 bits (57), Expect = 8.4
Identities = 16/55 (29%), Positives = 26/55 (47%), Gaps = 4/55 (7%)
Query: 117 KSRCLNNITISANGRNVVAMVVDECDSRKGCDE--QHDYQPPCPNNIVDASKAVW 169
KS C +TI+ + N + + + + C E +H+ P C NN D + A W
Sbjct: 619 KSLCWELVTINKDKLNGIGAAIYQKPATNECYEKRKHNKPPLCKNN--DDANAAW 671
>At2g45400 putative flavonol reductase
Length = 364
Score = 26.6 bits (57), Expect = 8.4
Identities = 19/77 (24%), Positives = 30/77 (38%), Gaps = 2/77 (2%)
Query: 28 QNCRPSGRIRGKKAPPGKCNQENDSDCCVQGKMYTTYVCSPSVSTHTKAYLT--LNSFQK 85
+ C+ + P +E + VQG M C + + Y + + F
Sbjct: 110 EGCKAVFHVAHPMDPNSNETEETVTKRTVQGLMGILKSCLDAKTVKRFFYTSSAVTVFYS 169
Query: 86 GGDGGGPSECDKQYHSD 102
GG+GGG E D+ SD
Sbjct: 170 GGNGGGGGEVDESVWSD 186
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.134 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,976,872
Number of Sequences: 26719
Number of extensions: 218438
Number of successful extensions: 401
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 398
Number of HSP's gapped (non-prelim): 8
length of query: 189
length of database: 11,318,596
effective HSP length: 94
effective length of query: 95
effective length of database: 8,807,010
effective search space: 836665950
effective search space used: 836665950
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC135311.8


