

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135311.2 + phase: 0
(164 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
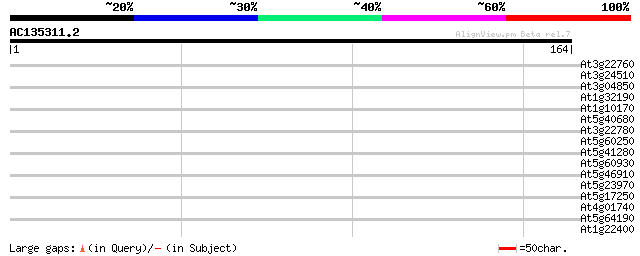
Sequences producing significant alignments: (bits) Value
At3g22760 unknown protein 29 0.99
At3g24510 unknown protein 29 1.3
At3g04850 unknown protein 29 1.3
At1g32190 unknown protein 29 1.3
At1g10170 hypothetical protein 29 1.3
At5g40680 putative protein 28 2.2
At3g22780 DNA binding protein like (tso1) 28 2.9
At5g60250 unknown protein 27 3.8
At5g41280 predicted GPI-anchored protein 27 3.8
At5g60930 microtubule-associated motor - like 27 4.9
At5g46910 putative protein 27 4.9
At5g23970 acetyl-CoA:benzylalcohol acetyltranferase-like protein 27 6.4
At5g17250 unknown protein 27 6.4
At4g01740 putative CHP-rich zinc finger protein 27 6.4
At5g64190 putative protein 26 8.4
At1g22400 Putative UDP-glucose glucosyltransferase 26 8.4
>At3g22760 unknown protein
Length = 609
Score = 29.3 bits (64), Expect = 0.99
Identities = 12/32 (37%), Positives = 17/32 (52%), Gaps = 3/32 (9%)
Query: 88 SACKNCECGWSSPL---CTCYDITDFCYKPCN 116
S+CK C C S L C C+ +C +PC+
Sbjct: 325 SSCKRCNCKKSKCLKLYCECFAAGFYCIEPCS 356
>At3g24510 unknown protein
Length = 86
Score = 28.9 bits (63), Expect = 1.3
Identities = 24/89 (26%), Positives = 39/89 (42%), Gaps = 15/89 (16%)
Query: 13 MKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNY------DAKSTATACCNTC 66
+KL LL+F+L TS + A + + VI G++ + Y D + + +C
Sbjct: 6 LKLPLLIFILVITSN-LGAEARKLTGVADVIVEGEAASPQYGIIDENDTLNKSPSCKRDI 64
Query: 67 LCSVTPELTQCRCADVGKTCHSACKNCEC 95
C+ QCR G C+ + CEC
Sbjct: 65 DCTF-----QCR---RGGFCNLILQKCEC 85
>At3g04850 unknown protein
Length = 639
Score = 28.9 bits (63), Expect = 1.3
Identities = 12/31 (38%), Positives = 17/31 (54%), Gaps = 3/31 (9%)
Query: 89 ACKNCECGWSSPL---CTCYDITDFCYKPCN 116
+CK C+C S L C C+ FC +PC+
Sbjct: 451 SCKRCKCRKSQCLKLYCECFSAGLFCGEPCS 481
>At1g32190 unknown protein
Length = 422
Score = 28.9 bits (63), Expect = 1.3
Identities = 15/60 (25%), Positives = 25/60 (41%), Gaps = 5/60 (8%)
Query: 56 KSTATACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWSSPLCTCYDITDFCYKPC 115
+ +T CC + LC + + RC +C C +C C C+C + C+ C
Sbjct: 289 RDESTGCCCSGLCRPSCSCPKPRCPKPSCSCGCGCGDCGCF----KCSCPTLKG-CFSCC 343
>At1g10170 hypothetical protein
Length = 1188
Score = 28.9 bits (63), Expect = 1.3
Identities = 20/70 (28%), Positives = 26/70 (36%), Gaps = 20/70 (28%)
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSA----C-----KNCECGWSSPLCTCYDITD-- 109
+C N C +L CR + CH+ C + C CG +S CY T
Sbjct: 502 SCSNVCR-----KLLPCRLHTCNEMCHAGDCPPCLVQVNQKCRCGSTSRAVECYITTSSE 556
Query: 110 ----FCYKPC 115
C KPC
Sbjct: 557 AEKFVCAKPC 566
>At5g40680 putative protein
Length = 415
Score = 28.1 bits (61), Expect = 2.2
Identities = 14/51 (27%), Positives = 24/51 (46%), Gaps = 1/51 (1%)
Query: 41 QVISNGDSTTNNYDAKSTATACCN-TCLCSVTPELTQCRCADVGKTCHSAC 90
+++ G S T ++D K + CC + +T E T C C +G + C
Sbjct: 361 RILVIGASVTKSWDNKMSVYTCCPFPKVEKITWEETSCDCVQLGHFIRNCC 411
>At3g22780 DNA binding protein like (tso1)
Length = 695
Score = 27.7 bits (60), Expect = 2.9
Identities = 11/31 (35%), Positives = 16/31 (51%), Gaps = 3/31 (9%)
Query: 89 ACKNCECGWSSPL---CTCYDITDFCYKPCN 116
+CK C C S L C C+ +C +PC+
Sbjct: 399 SCKRCNCKKSKCLKLYCECFAAGVYCIEPCS 429
>At5g60250 unknown protein
Length = 655
Score = 27.3 bits (59), Expect = 3.8
Identities = 13/45 (28%), Positives = 18/45 (39%), Gaps = 13/45 (28%)
Query: 77 CRCADVGKTCHSACKNCECGWSSPLCTCYDITDFCYKPCNVWQQQ 121
CRC H C NC GW+ + TC + C W ++
Sbjct: 488 CRCG------HEFCYNCGGGWNKIMGTCLN-------RCPTWNEE 519
>At5g41280 predicted GPI-anchored protein
Length = 286
Score = 27.3 bits (59), Expect = 3.8
Identities = 18/54 (33%), Positives = 23/54 (42%), Gaps = 8/54 (14%)
Query: 35 STSFITQVISNGDSTTNNYDAKSTATAC-------CNTCLCSVTPELTQCRCAD 81
S+SF + + + N YD +S C TCL ELTQC C D
Sbjct: 175 SSSFTPYYLMDTTTFDNLYDLESVVQCSPHLDPKNCTTCLKLALQELTQC-CGD 227
>At5g60930 microtubule-associated motor - like
Length = 647
Score = 26.9 bits (58), Expect = 4.9
Identities = 17/57 (29%), Positives = 27/57 (46%), Gaps = 6/57 (10%)
Query: 47 DSTTNNYDAKSTATACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWSSPLCT 103
+S T + DA + CC TC S + + +C+C +C + CG SS C+
Sbjct: 391 NSETPSDDAVKSDVCCC-TCSKSSSCKTMKCQCRATKGSCGPS-----CGCSSVKCS 441
>At5g46910 putative protein
Length = 707
Score = 26.9 bits (58), Expect = 4.9
Identities = 11/29 (37%), Positives = 18/29 (61%), Gaps = 1/29 (3%)
Query: 79 CADVGKTCHSACKNCECGWSSPLCTCYDI 107
C+ + C+ A NCEC +S P+C +D+
Sbjct: 419 CSLCKRDCYLAFINCEC-YSHPVCLRHDV 446
>At5g23970 acetyl-CoA:benzylalcohol acetyltranferase-like protein
Length = 428
Score = 26.6 bits (57), Expect = 6.4
Identities = 13/66 (19%), Positives = 26/66 (38%), Gaps = 3/66 (4%)
Query: 44 SNGDSTTNNYDAKSTATACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWSSPLCT 103
++ + ++ +AK + + LC ++PE+ V C + GW P+
Sbjct: 319 NDSEDAKSSVEAKERIASAMLSSLCEISPEM---ETYAVSSWCRMSFYEANFGWGKPVWV 375
Query: 104 CYDITD 109
D D
Sbjct: 376 APDSVD 381
>At5g17250 unknown protein
Length = 884
Score = 26.6 bits (57), Expect = 6.4
Identities = 19/63 (30%), Positives = 29/63 (45%), Gaps = 6/63 (9%)
Query: 1 MELMNKNKKA------MVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYD 54
ME KNKK ++ +A+L+F GF T + F ST + S +N+D
Sbjct: 1 METYLKNKKLTALGFLLIHAIAILIFTRGFLLTRTELPFHSTCSDVSLSPCLASPRSNHD 60
Query: 55 AKS 57
+ S
Sbjct: 61 SSS 63
>At4g01740 putative CHP-rich zinc finger protein
Length = 652
Score = 26.6 bits (57), Expect = 6.4
Identities = 10/38 (26%), Positives = 17/38 (44%)
Query: 53 YDAKSTATACCNTCLCSVTPELTQCRCADVGKTCHSAC 90
Y K++ C+ C PE+ C D + H++C
Sbjct: 521 YGEKASGICWCDICEAKADPEIWFYTCKDQRASLHTSC 558
>At5g64190 putative protein
Length = 502
Score = 26.2 bits (56), Expect = 8.4
Identities = 23/90 (25%), Positives = 35/90 (38%), Gaps = 24/90 (26%)
Query: 98 SSPLCTCYDITDFCYKPCNVWQQQQKVQVVVT---IVLAQDHSHL--------------- 139
S P C C +DF N+ Q+ VVT IV + +HS L
Sbjct: 25 SLPFCICPSTSDFPNSTLNLTAQKSPSPKVVTFSIIVQSNNHSPLYLWTTKQELSINPNS 84
Query: 140 ------IVLVMMLNHFVTQIASLAGASSHY 163
+ ++ +L +FV I + SS+Y
Sbjct: 85 PNPFDELTIISLLFNFVETILTYTSNSSNY 114
>At1g22400 Putative UDP-glucose glucosyltransferase
Length = 489
Score = 26.2 bits (56), Expect = 8.4
Identities = 20/82 (24%), Positives = 32/82 (38%), Gaps = 9/82 (10%)
Query: 12 VMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTCLCSVT 71
+M++A L+ GF T V+ ++ F+ SN ++ +S A
Sbjct: 28 MMRVAKLLHARGFYVTFVNTVYNHNRFLRSRGSNALDGLPSFRFESIADGL--------- 78
Query: 72 PELTQCRCADVGKTCHSACKNC 93
PE D+ C S KNC
Sbjct: 79 PETDMDATQDITALCESTMKNC 100
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.131 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,436,942
Number of Sequences: 26719
Number of extensions: 128622
Number of successful extensions: 503
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 500
Number of HSP's gapped (non-prelim): 17
length of query: 164
length of database: 11,318,596
effective HSP length: 92
effective length of query: 72
effective length of database: 8,860,448
effective search space: 637952256
effective search space used: 637952256
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC135311.2


