

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
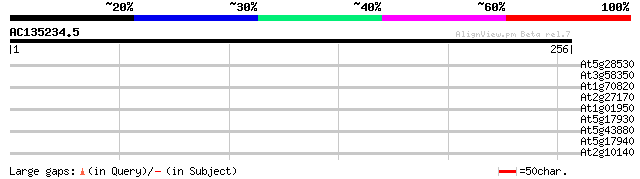
Query= AC135234.5 - phase: 0 /pseudo
(256 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At5g28530 far-red impaired response protein (FAR1) - like 35 0.039
At3g58350 putative protein 30 1.3
At1g70820 phosphoglucomutase precursor like protein 30 1.7
At2g27170 putative chromosome associated protein 29 2.8
At1g01950 unknown protein 28 3.7
At5g17930 unknown protein 28 4.8
At5g43880 unknown protein 28 6.3
At5g17940 putative protein 27 8.2
At2g10140 putative protein 27 8.2
>At5g28530 far-red impaired response protein (FAR1) - like
Length = 700
Score = 35.0 bits (79), Expect = 0.039
Identities = 34/142 (23%), Positives = 67/142 (46%), Gaps = 29/142 (20%)
Query: 92 KAGFGVCIDKSVLKRPYLTTQSERSGIYKPP---------KTRKKPNLEGIG---SMKCN 139
K+GF + R +T+S+ G+Y+ + RKK N+E S++C
Sbjct: 92 KSGFSI--------RKARSTESQNLGVYRRDFVCYRSGFNQPRKKANVEHPRERKSVRCG 143
Query: 140 CPFRLKGFFDKDTND----WWLAILCGMHNHDLEEKLQGHLIAG--RLSAEEKKKVIDMT 193
C +L + K+ D W+++ +HNH+L E Q L+ ++ ++++++ ++
Sbjct: 144 CDGKL--YLTKEVVDGVSHWYVSQFSNVHNHELLEDDQVRLLPAYRKIQQSDQERILLLS 201
Query: 194 KSLTVPRNILTNLKENNKESVT 215
K+ P N + L E K V+
Sbjct: 202 KA-GFPVNRIVKLLELEKGVVS 222
>At3g58350 putative protein
Length = 355
Score = 30.0 bits (66), Expect = 1.3
Identities = 17/61 (27%), Positives = 31/61 (49%), Gaps = 2/61 (3%)
Query: 144 LKGFFDKDTNDWWLAILCGMHNHDLEEKLQGHLIAGRLSAEEKKKVIDMTKSLTVPRNIL 203
L+ +FD+ T +W L+ +C + +++ K G L+ G L + KV++ L V
Sbjct: 147 LEQWFDEKTTNWGLSSMCPL--NEIHAKDSGFLLNGELKIVVEIKVLETIGKLDVTEETS 204
Query: 204 T 204
T
Sbjct: 205 T 205
>At1g70820 phosphoglucomutase precursor like protein
Length = 615
Score = 29.6 bits (65), Expect = 1.7
Identities = 32/126 (25%), Positives = 54/126 (42%), Gaps = 8/126 (6%)
Query: 86 VRRQGVKAGFGVCIDKSVLKRPYLTTQSERSGIYKPPKTRKKPNLEGIGSMKCNCPF-RL 144
+R G G G I+ L+ P L R I P+ K +E I + + +L
Sbjct: 453 MRLAGSNEGIGSLIED--LEEP-LEAVELRLNILSEPRDAKAKGIEAIETFRQYIEEGKL 509
Query: 145 KGFFDKDTNDWWLAILCGMHNHDLEEKLQGHLIAGRLSAEEKKKV---IDMTKSLTVPRN 201
KG+ D W+ C + ++D + H+ R+S EE + + M +S+ P N
Sbjct: 510 KGWELGTCGDCWVTEGCLVDSNDHPSAIDAHMYRARVSDEESGEEYGWVHMRQSIHNP-N 568
Query: 202 ILTNLK 207
I N++
Sbjct: 569 IALNMQ 574
>At2g27170 putative chromosome associated protein
Length = 1163
Score = 28.9 bits (63), Expect = 2.8
Identities = 20/65 (30%), Positives = 33/65 (50%), Gaps = 1/65 (1%)
Query: 166 HDLEEKLQGHLIAGRLSAEEKKKVID-MTKSLTVPRNILTNLKENNKESVTTIKQVYNVR 224
HD EKL+ +A ++EE K+ D + K+ +++ +LKE KE T K+ V
Sbjct: 233 HDAREKLEQVEVARTKASEESTKMYDRVEKAQDDSKSLDESLKELTKELQTLYKEKETVE 292
Query: 225 TRWCK 229
+ K
Sbjct: 293 AQQTK 297
>At1g01950 unknown protein
Length = 893
Score = 28.5 bits (62), Expect = 3.7
Identities = 23/90 (25%), Positives = 45/90 (49%), Gaps = 8/90 (8%)
Query: 161 CGMHNHDLEEKLQGHLIAGRLSAEEKKKVIDMTKSLTVP-----RNILTN---LKENNKE 212
C M + +KL+ LI+ + + E K+ ++ +T + L N L+++ +E
Sbjct: 468 CQMEYMESVKKLEEKLISNQRNHENGKRNGEVNGVVTASEFTRLKESLENEMKLRKSAEE 527
Query: 213 SVTTIKQVYNVRTRWCKGERGDMTELQFLI 242
V+ +K ++TR +GE +T LQ L+
Sbjct: 528 EVSKVKSQSTLKTRSGEGEDAGITRLQKLL 557
>At5g17930 unknown protein
Length = 707
Score = 28.1 bits (61), Expect = 4.8
Identities = 18/81 (22%), Positives = 40/81 (49%), Gaps = 3/81 (3%)
Query: 166 HDLEEKLQGHLIAGRLSAEEKKKVIDMTKSLTVPRNILTNLKENNKESVTT-IKQVYNVR 224
H+ + + +A L ++ K + ++TK T + +L + E+N E++T + +Y R
Sbjct: 252 HERKPESSSKYVAPHLRSQAKSESEELTKMRTRIKGLLNKMAESNVETITAELATIY--R 309
Query: 225 TRWCKGERGDMTELQFLISKL 245
+ K + + + L+S L
Sbjct: 310 DEYQKEDSLSLNGISLLLSYL 330
>At5g43880 unknown protein
Length = 836
Score = 27.7 bits (60), Expect = 6.3
Identities = 19/67 (28%), Positives = 30/67 (44%), Gaps = 9/67 (13%)
Query: 88 RQGVKAGFGVCIDKSVLKRPYLTTQSERSGIYKPPKTRKKPNLEGIGSMKCNCPFRLKGF 147
R G K+G G+ K ++ Y T QS R + KP G + +CP +GF
Sbjct: 246 RDGSKSGKGLDFFKWPVEEEYPTKQSTRIVVLKP---------NGQVTKASSCPTSPRGF 296
Query: 148 FDKDTND 154
+++ D
Sbjct: 297 EGRESRD 303
>At5g17940 putative protein
Length = 427
Score = 27.3 bits (59), Expect = 8.2
Identities = 12/50 (24%), Positives = 27/50 (54%)
Query: 166 HDLEEKLQGHLIAGRLSAEEKKKVIDMTKSLTVPRNILTNLKENNKESVT 215
H+ + + +A L ++ K + ++TK T + +L + E+N E++T
Sbjct: 272 HERKPESSSKYVAPHLRSQAKSESEELTKMRTRIKGLLNKMAESNVETIT 321
>At2g10140 putative protein
Length = 663
Score = 27.3 bits (59), Expect = 8.2
Identities = 11/27 (40%), Positives = 15/27 (54%)
Query: 74 ARWREREELLTWVRRQGVKAGFGVCID 100
A WR +EL TW ++ G+CID
Sbjct: 269 AEWRYFQELHTWFAKEPRNVYLGLCID 295
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,093,226
Number of Sequences: 26719
Number of extensions: 262467
Number of successful extensions: 683
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 680
Number of HSP's gapped (non-prelim): 9
length of query: 256
length of database: 11,318,596
effective HSP length: 97
effective length of query: 159
effective length of database: 8,726,853
effective search space: 1387569627
effective search space used: 1387569627
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC135234.5


