

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135232.7 + phase: 0
(221 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
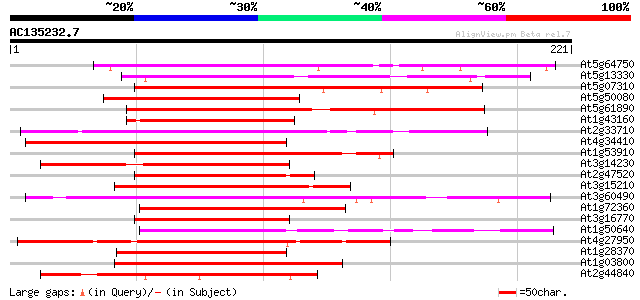
Score E
Sequences producing significant alignments: (bits) Value
At5g64750 unknown protein 149 1e-36
At5g13330 putative protein 129 9e-31
At5g07310 putative transcription factor 128 2e-30
At5g50080 putative protein 123 7e-29
At5g61890 putative protein 120 6e-28
At1g43160 RAP2.6 (At1g43160) 120 6e-28
At2g33710 putative AP2 domain transcription factor 117 4e-27
At4g34410 unknown protein 111 3e-25
At1g53910 unknown protein 105 1e-23
At3g14230 transcription factor EREBP like protein 104 3e-23
At2g47520 putative AP2 domain transcription factor 104 4e-23
At3g15210 ethylene responsive element binding factor 4 102 2e-22
At3g60490 transcription factor - like protein 100 8e-22
At1g72360 putative AP2 domain transcription factor 100 1e-21
At3g16770 AP2 domain containing protein RAP2.3 99 1e-21
At1g50640 ethylene responsive element binding factor 3 99 2e-21
At4g27950 unknown protein 98 4e-21
At1g28370 putative ethylene responsive element binding factor 4 ... 98 4e-21
At1g03800 unknown protein 98 4e-21
At2g44840 putative ethylene response element binding protein (ER... 97 5e-21
>At5g64750 unknown protein
Length = 391
Score = 149 bits (375), Expect = 1e-36
Identities = 96/222 (43%), Positives = 123/222 (55%), Gaps = 44/222 (19%)
Query: 34 YEYRT-----DNNNVKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDA 88
YEY T + + DQP+R+YRGVRQRPWGKWAAEIRDPFKA RVWLGTF+ AE A
Sbjct: 161 YEYTTTATASSETSSFSGDQPRRRYRGVRQRPWGKWAAEIRDPFKAARVWLGTFDNAESA 220
Query: 89 AKAYDQASLRFRGNKAKLNFPENVKLKQQQST------------PTHLNISHSNSALLSH 136
A+AYD+A+LRFRGNKAKLNFPENVKL + ST PT S S + LL
Sbjct: 221 ARAYDEAALRFRGNKAKLNFPENVKLVRPASTEAQPVHQTAAQRPTQSRNSGSTTTLLPI 280
Query: 137 QPRTDPDPIVHNETLHTLQSSNKYY------------------DYFNGQNFPMASLQT-- 176
+P ++ VH++ L +QS N Y D ++ +FP+ T
Sbjct: 281 RPASNQS--VHSQPL--MQSYNLSYSEMARQQQQFQQHHQQSLDLYDQMSFPLRFGHTGG 336
Query: 177 SVSPSASYSSSYTTTTFASSFSSPQTTSSIPSGL---TAWPS 215
S+ S S SSS++ F+ + P S+ +G WPS
Sbjct: 337 SMMQSTSSSSSHSRPLFSPAAVQPPPESASETGYLQDIQWPS 378
>At5g13330 putative protein
Length = 212
Score = 129 bits (325), Expect = 9e-31
Identities = 78/163 (47%), Positives = 99/163 (59%), Gaps = 16/163 (9%)
Query: 45 NEDQPKRK-YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNK 103
++DQP+R+ YRGVRQRPWGKWAAEIRDP KA RVWLGTFETAE+AA AYD+A+L+F+G K
Sbjct: 30 DQDQPRRRHYRGVRQRPWGKWAAEIRDPKKAARVWLGTFETAEEAALAYDRAALKFKGTK 89
Query: 104 AKLNFPENVKLKQQQSTPTHLNISHSNSALLSHQPRTDPDPIVHNETLHTLQSSNKYYDY 163
AKLNFPE V Q T ISH+ + P P + T + +
Sbjct: 90 AKLNFPERV-----QGPTTTTTISHAPRGVSESMNSPPPRPGPPSTT------TTSWPMT 138
Query: 164 FNGQNFPMASLQTSVSP-SASYSSSYTTTTFASSFSSPQTTSS 205
+N A L TS + SY YT+T F+ FS+P ++SS
Sbjct: 139 YNQDILQYAQLLTSNNEVDLSY---YTSTLFSQPFSTPSSSSS 178
>At5g07310 putative transcription factor
Length = 263
Score = 128 bits (322), Expect = 2e-30
Identities = 75/150 (50%), Positives = 91/150 (60%), Gaps = 13/150 (8%)
Query: 50 KRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
KR YRGVRQRPWGKWAAEIRDP KA RVWLGTFETAE AA AYD A+L+F+G+KAKLNFP
Sbjct: 89 KRHYRGVRQRPWGKWAAEIRDPQKAARVWLGTFETAEAAALAYDNAALKFKGSKAKLNFP 148
Query: 110 ENVKLKQQQSTPT-HLNISHSNSALLSHQPRTDPDPI------VHNETLHTLQSSNKYYD 162
E +L ST T N SN+ + P+T+P I +N+ LH +SN
Sbjct: 149 ERAQLASNTSTTTGPPNYYSSNNQIYYSNPQTNPQTIPYFNQYYYNQYLHQGGNSNDALS 208
Query: 163 Y------FNGQNFPMASLQTSVSPSASYSS 186
Y G + +L T+ S S+ SS
Sbjct: 209 YSLAGGETGGSMYNHQTLSTTNSSSSGGSS 238
>At5g50080 putative protein
Length = 220
Score = 123 bits (309), Expect = 7e-29
Identities = 54/77 (70%), Positives = 68/77 (88%)
Query: 38 TDNNNVKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASL 97
+ +N ++ + KR+YRGVRQRPWGKWAAEIRDP +A RVWLGTF+TAE AA+AYD+A+L
Sbjct: 62 SSHNPIEEAREKKRRYRGVRQRPWGKWAAEIRDPHRAARVWLGTFDTAEAAARAYDEAAL 121
Query: 98 RFRGNKAKLNFPENVKL 114
RFRGNKAKLNFPE+V++
Sbjct: 122 RFRGNKAKLNFPEDVRI 138
>At5g61890 putative protein
Length = 248
Score = 120 bits (301), Expect = 6e-28
Identities = 65/143 (45%), Positives = 92/143 (63%), Gaps = 9/143 (6%)
Query: 47 DQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKL 106
D +R YRGVRQRPWGKWAAEIRDP KA RVWLGTFETAE AA AYD+A+L+F+G+KAKL
Sbjct: 84 DLRRRHYRGVRQRPWGKWAAEIRDPKKAARVWLGTFETAESAALAYDEAALKFKGSKAKL 143
Query: 107 NFPENVKLKQQQSTPTHLNISHSNSALLSHQPRTDP--DPIVHNETLHTLQSSNKYYDYF 164
NFPE V+L + +S++ + +P++ P + H+ + + S N Y
Sbjct: 144 NFPERVQLGSNST-------YYSSNQIPQMEPQSIPNYNQYYHDASSGDMLSFNLGGGYG 196
Query: 165 NGQNFPMASLQTSVSPSASYSSS 187
+G + M+ ++ + + + SSS
Sbjct: 197 SGTGYSMSHDNSTTTAATTSSSS 219
>At1g43160 RAP2.6 (At1g43160)
Length = 192
Score = 120 bits (301), Expect = 6e-28
Identities = 57/66 (86%), Positives = 62/66 (93%), Gaps = 1/66 (1%)
Query: 47 DQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKL 106
++PK KYRGVRQRPWGKWAAEIRDP KATRVWLGTFETAE AA+AYD A+LRFRG+KAKL
Sbjct: 56 ERPK-KYRGVRQRPWGKWAAEIRDPHKATRVWLGTFETAEAAARAYDAAALRFRGSKAKL 114
Query: 107 NFPENV 112
NFPENV
Sbjct: 115 NFPENV 120
>At2g33710 putative AP2 domain transcription factor
Length = 218
Score = 117 bits (294), Expect = 4e-27
Identities = 71/184 (38%), Positives = 106/184 (57%), Gaps = 11/184 (5%)
Query: 5 LCNVGVECSSNWTNTVTTTTGRSQVEEQIYEYRTDNNNVKNEDQPKRKYRGVRQRPWGKW 64
LC S+ ++ V+ TG+ + I + + + + +R YRGVRQRPWGKW
Sbjct: 23 LCCCSTLSESDVSDFVSELTGQP-IPSSIDDQSSSLTLQEKSNSRQRNYRGVRQRPWGKW 81
Query: 65 AAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPENVKLKQQQSTPTHL 124
AAEIRDP KA RVWLGTF+TAE+AA AYD+A+ FRG+KAKLNFPE++++ Q P+
Sbjct: 82 AAEIRDPNKAARVWLGTFDTAEEAALAYDKAAFEFRGHKAKLNFPEHIRVNPTQLYPSPA 141
Query: 125 NISHSNSALLSHQPRTDPDPIVHNETLHTLQSSNKYYDYFNGQNFPMASLQTSVSPSASY 184
SH + P + P PI + L ++Y + + + A+L ++ S+S
Sbjct: 142 T-SHDRIIV---TPPSPPPPIAPDILL------DQYGHFQSRSSDSSANLSMNMLSSSSS 191
Query: 185 SSSY 188
S ++
Sbjct: 192 SLNH 195
>At4g34410 unknown protein
Length = 268
Score = 111 bits (277), Expect = 3e-25
Identities = 52/103 (50%), Positives = 70/103 (67%)
Query: 7 NVGVECSSNWTNTVTTTTGRSQVEEQIYEYRTDNNNVKNEDQPKRKYRGVRQRPWGKWAA 66
N +E + +T+++ R + + + ++ K YRGVRQRPWGK+AA
Sbjct: 90 NQRIEKNHQQEEEITSSSNRRRESSPVAKKAEGGGKIRKRKNKKNGYRGVRQRPWGKFAA 149
Query: 67 EIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
EIRDP +ATRVWLGTFETAEDAA+AYD+A++ FRG +AKLNFP
Sbjct: 150 EIRDPKRATRVWLGTFETAEDAARAYDRAAIGFRGPRAKLNFP 192
>At1g53910 unknown protein
Length = 358
Score = 105 bits (263), Expect = 1e-23
Identities = 51/103 (49%), Positives = 67/103 (64%), Gaps = 6/103 (5%)
Query: 50 KRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
K +YRG+RQRPWGKWAAEIRDP + R+WLGTF+TAE+AA+AYD A+ R RG+KAK+NFP
Sbjct: 122 KNQYRGIRQRPWGKWAAEIRDPREGARIWLGTFKTAEEAARAYDAAARRIRGSKAKVNFP 181
Query: 110 ENVKLKQQQSTPTHLNISHSNSALLSHQPRTDPDP-IVHNETL 151
E Q N+ + +P +P P +V N +
Sbjct: 182 EENMKANSQKRSVKANLQKPVA-----KPNPNPSPALVQNSNI 219
>At3g14230 transcription factor EREBP like protein
Length = 375
Score = 104 bits (260), Expect = 3e-23
Identities = 51/98 (52%), Positives = 66/98 (67%), Gaps = 6/98 (6%)
Query: 13 SSNWTNTVTTTTGRSQVEEQIYEYRTDNNNVKNEDQPKRKYRGVRQRPWGKWAAEIRDPF 72
+S + +TV + + VE ++ KN+ YRG+RQRPWGKWAAEIRDP
Sbjct: 90 ASAFVSTVGSAYAKKTVESAEQAEKSSKRKRKNQ------YRGIRQRPWGKWAAEIRDPR 143
Query: 73 KATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPE 110
K +R WLGTF+TAE+AA+AYD A+ R RG KAK+NFPE
Sbjct: 144 KGSREWLGTFDTAEEAARAYDAAARRIRGTKAKVNFPE 181
>At2g47520 putative AP2 domain transcription factor
Length = 171
Score = 104 bits (259), Expect = 4e-23
Identities = 46/71 (64%), Positives = 59/71 (82%), Gaps = 1/71 (1%)
Query: 50 KRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
K YRG+RQRPWGKWAAEIRDP K RVWLGTF+TA++AA+AYD A+++ RG KAKLNFP
Sbjct: 47 KNLYRGIRQRPWGKWAAEIRDPSKGVRVWLGTFKTADEAARAYDVAAIKIRGRKAKLNFP 106
Query: 110 ENVKLKQQQST 120
N +++++ T
Sbjct: 107 -NTQVEEEADT 116
>At3g15210 ethylene responsive element binding factor 4
Length = 222
Score = 102 bits (253), Expect = 2e-22
Identities = 52/93 (55%), Positives = 67/93 (71%), Gaps = 1/93 (1%)
Query: 42 NVKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRG 101
N + + + +YRGVR+RPWG++AAEIRDP K TRVWLGTF+TAE+AA+AYD A+ FRG
Sbjct: 14 NQTHNNAKEIRYRGVRKRPWGRYAAEIRDPGKKTRVWLGTFDTAEEAARAYDTAARDFRG 73
Query: 102 NKAKLNFPENVKLKQQQSTPTHLNISHSNSALL 134
KAK NFP ++L Q+ PT S S S+ L
Sbjct: 74 AKAKTNFPTFLELSDQK-VPTGFARSPSQSSTL 105
>At3g60490 transcription factor - like protein
Length = 256
Score = 100 bits (248), Expect = 8e-22
Identities = 70/219 (31%), Positives = 106/219 (47%), Gaps = 25/219 (11%)
Query: 7 NVGVECSSNWTNTVTTTTGRSQVEEQIYEYRTDNNNVKNEDQPKRKYRGVRQRPWGKWAA 66
+V S +W+ T+ RS V++ + +NV ++++ YRGVR R WGKW +
Sbjct: 30 SVVTSSSDSWS-----TSKRSLVQDNDSGGKRRKSNVSDDNKNPTSYRGVRMRSWGKWVS 84
Query: 67 EIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPENVKL--KQQQSTPTHL 124
EIR+P K +R+WLGT+ TAE AA+A+D A+L +GN LNFPE L + +P +
Sbjct: 85 EIREPRKKSRIWLGTYPTAEMAARAHDVAALAIKGNSGFLNFPELSGLLPRPVSCSPKDI 144
Query: 125 NISHSNSALLS--HQPRTD---PDPIVHNETLHTLQSSNKYYDYFNGQNFPMASLQTSVS 179
+ + +A + H+P D D + H+E L T QSS F S TS +
Sbjct: 145 QAAATKAAEATTWHKPVIDKKLADELSHSELLSTAQSSTSSSFVF--------SSDTSET 196
Query: 180 PSASYSSSYTTT-----TFASSFSSPQTTSSIPSGLTAW 213
S S+ T F +P + +G W
Sbjct: 197 SSTDKESNEETVFDLPDLFTDGLMNPNDAFCLCNGTFTW 235
>At1g72360 putative AP2 domain transcription factor
Length = 262
Score = 99.8 bits (247), Expect = 1e-21
Identities = 44/81 (54%), Positives = 57/81 (70%)
Query: 52 KYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPEN 111
+Y+G+R+RPWG+WAAEIRDP K RVWLGTF TAE+AA+AYD + R RG KAKLNFP
Sbjct: 75 RYKGIRRRPWGRWAAEIRDPIKGVRVWLGTFNTAEEAARAYDLEAKRIRGAKAKLNFPNE 134
Query: 112 VKLKQQQSTPTHLNISHSNSA 132
K++ T + ++ A
Sbjct: 135 SSGKRKAKAKTVQQVEENHEA 155
>At3g16770 AP2 domain containing protein RAP2.3
Length = 248
Score = 99.4 bits (246), Expect = 1e-21
Identities = 44/61 (72%), Positives = 51/61 (83%)
Query: 50 KRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
K YRG+R+RPWGKWAAEIRDP K RVWLGTF TAE+AA AYD A+ + RG+KAKLNFP
Sbjct: 76 KNVYRGIRKRPWGKWAAEIRDPRKGVRVWLGTFNTAEEAAMAYDVAAKQIRGDKAKLNFP 135
Query: 110 E 110
+
Sbjct: 136 D 136
>At1g50640 ethylene responsive element binding factor 3
Length = 225
Score = 99.0 bits (245), Expect = 2e-21
Identities = 60/163 (36%), Positives = 89/163 (53%), Gaps = 30/163 (18%)
Query: 52 KYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPEN 111
++RGVR+RPWG++AAEIRDP+K RVWLGTF++AE+AA+AYD A+ RG KAK NFP
Sbjct: 27 RFRGVRKRPWGRFAAEIRDPWKKARVWLGTFDSAEEAARAYDSAARNLRGPKAKTNFP-- 84
Query: 112 VKLKQQQSTPTHLNISHSNSALLSHQPRTDPDPIVHNETLHTLQSSNKYYDYFNGQNFPM 171
+ P +L + + +Q + DP + H L + ++ Q FP+
Sbjct: 85 --IDSSSPPPPNLRFNQ-----IRNQNQNQVDPFMD----HRLFTDHQ-------QQFPI 126
Query: 172 ASLQTSVSPSASYSSSYTTTTFASSFSSPQTTSSIPSGLTAWP 214
+ TS S S++ SFS P+ T+ P+ +P
Sbjct: 127 VNRPTSSSMSST----------VESFSGPRPTTMKPATTKRYP 159
>At4g27950 unknown protein
Length = 335
Score = 97.8 bits (242), Expect = 4e-21
Identities = 58/152 (38%), Positives = 92/152 (60%), Gaps = 10/152 (6%)
Query: 4 ILCNVGVECSSNWTNTVTTTTGRSQVEEQIYEYRTDNNNVKNEDQPKRKYRGVRQRPWGK 63
++ + VE SS+ T V+ + + + + + +V ++Q +KYRGVRQRPWGK
Sbjct: 72 LINEIRVEPSSSSTGDVSASPTKDRKRINV-DSTVQKPSVSGQNQ--KKYRGVRQRPWGK 128
Query: 64 WAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNF-----PENVKLKQQQ 118
WAAEIRDP + R+WLGTF TAE+AA YD A+++ RG A NF PE V+ +Q+Q
Sbjct: 129 WAAEIRDPEQRRRIWLGTFATAEEAAIVYDNAAIKLRGPDALTNFTVQPEPEPVQ-EQEQ 187
Query: 119 STPTHLNISHSNSALLSHQPRTDPDPIVHNET 150
+++++S S S + Q + P +++ +T
Sbjct: 188 EPESNMSVSISES-MDDSQHLSSPTSVLNYQT 218
>At1g28370 putative ethylene responsive element binding factor 4
protein
Length = 166
Score = 97.8 bits (242), Expect = 4e-21
Identities = 43/67 (64%), Positives = 55/67 (81%)
Query: 43 VKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGN 102
VK + +YRGVR+RPWG++AAEIRDPFK +RVWLGTF+T E+AA+AYD+ ++ FRG
Sbjct: 10 VKTNEGNGVRYRGVRKRPWGRYAAEIRDPFKKSRVWLGTFDTPEEAARAYDKRAIEFRGA 69
Query: 103 KAKLNFP 109
KAK NFP
Sbjct: 70 KAKTNFP 76
>At1g03800 unknown protein
Length = 245
Score = 97.8 bits (242), Expect = 4e-21
Identities = 47/90 (52%), Positives = 62/90 (68%)
Query: 42 NVKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRG 101
N E + +YRGVR+RPWG++AAEIRDP K RVWLG+F T E+AA+AYD A++RFRG
Sbjct: 42 NDGGEKSKEVRYRGVRRRPWGRYAAEIRDPVKKKRVWLGSFNTGEEAARAYDSAAIRFRG 101
Query: 102 NKAKLNFPENVKLKQQQSTPTHLNISHSNS 131
+KA NFP +TP + N+S + S
Sbjct: 102 SKATTNFPLIGYYGISSATPVNNNLSETVS 131
>At2g44840 putative ethylene response element binding protein
(EREBP)
Length = 226
Score = 97.4 bits (241), Expect = 5e-21
Identities = 55/121 (45%), Positives = 74/121 (60%), Gaps = 18/121 (14%)
Query: 13 SSNWTNTVTTTTGRSQVEEQIYEYRTDNNNVKNEDQPKRK---YRGVRQRPWGKWAAEIR 69
SS WT +V T ++ E + P++K YRGVR+RPWGK+AAEIR
Sbjct: 55 SSGWTPSVPPVTSPAE------ENKPPATKASGSHAPRQKGMQYRGVRRRPWGKFAAEIR 108
Query: 70 DPFK-ATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP--------ENVKLKQQQST 120
DP K RVWLGT+ET EDAA AYD+A+ + RG+KAKLNFP E V+++ ++ +
Sbjct: 109 DPKKNGARVWLGTYETPEDAAVAYDRAAFQLRGSKAKLNFPHLIGSCKYEPVRIRPRRRS 168
Query: 121 P 121
P
Sbjct: 169 P 169
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.310 0.124 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,342,071
Number of Sequences: 26719
Number of extensions: 227124
Number of successful extensions: 1053
Number of sequences better than 10.0: 198
Number of HSP's better than 10.0 without gapping: 164
Number of HSP's successfully gapped in prelim test: 34
Number of HSP's that attempted gapping in prelim test: 797
Number of HSP's gapped (non-prelim): 252
length of query: 221
length of database: 11,318,596
effective HSP length: 95
effective length of query: 126
effective length of database: 8,780,291
effective search space: 1106316666
effective search space used: 1106316666
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135232.7


