

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
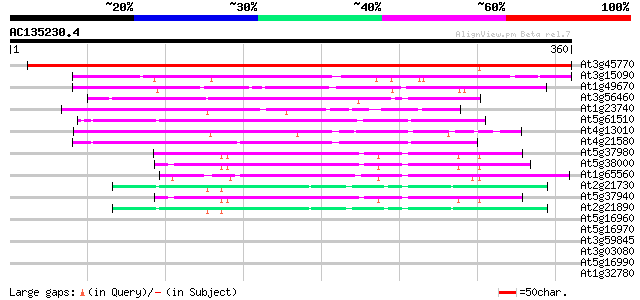
Query= AC135230.4 - phase: 0
(360 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g45770 nuclear receptor binding factor-like protein 513 e-146
At3g15090 zinc-binding dehydrogenase, putative 74 9e-14
At1g49670 ARP protein (At1g49670; F14J22.10) 74 9e-14
At3g56460 quinone reductase-like protein 73 2e-13
At1g23740 auxin-induced protein like 73 2e-13
At5g61510 quinone oxidoreductase - like protein 69 4e-12
At4g13010 unknown protein 67 2e-11
At4g21580 putative NADPH quinone oxidoreductase 64 1e-10
At5g37980 quinone oxidoreductase -like protein 51 1e-06
At5g38000 oxidoreductase-like protein 45 6e-05
At1g65560 putative oxido reductase (At1g65560) 45 6e-05
At2g21730 putative cinnamyl-alcohol dehydrogenase 44 1e-04
At5g37940 quinone oxidoreductase -like protein 42 5e-04
At2g21890 putative cinnamyl-alcohol dehydrogenase 42 5e-04
At5g16960 quinone oxidoreductase -like protein 41 0.001
At5g16970 quinone oxidoreductase -like protein 39 0.004
At3g59845 allyl alcohol dehydrogenase-like protein 38 0.007
At3g03080 putative NADP-dependent oxidoreductase 37 0.017
At5g16990 quinone oxidoreductase - like protein 37 0.022
At1g32780 alcohol dehydrogenase like protein 36 0.037
>At3g45770 nuclear receptor binding factor-like protein
Length = 375
Score = 513 bits (1321), Expect = e-146
Identities = 258/360 (71%), Positives = 297/360 (81%), Gaps = 11/360 (3%)
Query: 12 SSLFNFTSLRSRILNTHT-HAFSSAVSPPSKAIIYESHGQPDAVTKLVDIPATEVKENDL 70
SS NF S+R T +FS+ +SPPSKAI+YE HG PD+VT+LV++P EVKEND+
Sbjct: 16 SSTANFRSIRRGETPTLCIKSFSTIMSPPSKAIVYEEHGSPDSVTRLVNLPPVEVKENDV 75
Query: 71 CVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPP 130
CVKM+AAPINPSDINRI+GVYPVRP PAVGGYEGVGEV++VGS V FSPGDWVIPSPP
Sbjct: 76 CVKMIAAPINPSDINRIEGVYPVRPPVPAVGGYEGVGEVYAVGSNVNGFSPGDWVIPSPP 135
Query: 131 SFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGAT 190
S GTWQTY+VK+++VWHK++K PMEYAATITVNPLTAL MLED V LNSGD++VQNGAT
Sbjct: 136 SSGTWQTYVVKEESVWHKIDKECPMEYAATITVNPLTALRMLEDFVNLNSGDSVVQNGAT 195
Query: 191 SMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGG 250
S+VGQCVIQLA+ RGI IN+IRDR G E +E+LK LGADEVF+ES+L VKNVKSLLG
Sbjct: 196 SIVGQCVIQLARLRGISTINLIRDRAGSDEAREQLKALGADEVFSESQLNVKNVKSLLGN 255
Query: 251 IPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFK---------- 300
+PEPALGFNCVGGN+ASLVLK+LR GGTMVTYGGMSKKP+TVST+SFIFK
Sbjct: 256 LPEPALGFNCVGGNAASLVLKYLREGGTMVTYGGMSKKPITVSTTSFIFKDLALRGFWLQ 315
Query: 301 NWLSTDKAEEGRRMIDRLLGLVQDGKLKYKMELTPFNDFNTALDKALGKLGSQPKQVIKF 360
+WLS K +E R MID LLGL +DGKLKY+ EL PF +F ALDKALGKLG QPKQVI F
Sbjct: 316 SWLSMGKVKECREMIDYLLGLARDGKLKYETELVPFEEFPVALDKALGKLGRQPKQVITF 375
>At3g15090 zinc-binding dehydrogenase, putative
Length = 366
Score = 74.3 bits (181), Expect = 9e-14
Identities = 93/344 (27%), Positives = 143/344 (41%), Gaps = 34/344 (9%)
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVY---PVRPEP 97
+A+I G P+ ++P + N++ VK A +NP D RI+ Y +P
Sbjct: 33 RAVILPRFGGPEVFELRENVPVPNLNPNEVLVKAKAVSVNPLDC-RIRAGYGRSVFQPHL 91
Query: 98 PAVGGYEGVGEVHSVGSAVTCFSPGDWVIPS--PPSF-GTWQTYIVKDQNVWHKVNKGVP 154
P + G + GEV ++G++V G V + P + GT+ Y + ++ + +
Sbjct: 92 PIIVGRDVSGEVAAIGTSVKSLKVGQEVFGALHPTALRGTYTDYGILSEDELTEKPSSIS 151
Query: 155 MEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRD 214
A+ I LTA L+ + G ++ G VG IQLA + G H
Sbjct: 152 HVEASAIPFAALTAWRALKSNARITEGQRLLVFGGGGAVGFSAIQLAVASGCH-----VT 206
Query: 215 RPGVGEVKERLKDLGADEVF----TESELEVKN----VKSLLGGIPEPALGFNCV--GGN 264
VG+ K+R+ GA++ + EL VK V +GG +G N + GGN
Sbjct: 207 ASCVGQTKDRILAAGAEQAVDYTTEDIELAVKGKFDAVLDTIGGPETERIGINFLRKGGN 266
Query: 265 -------SASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKNWLSTDKAEEGRRMIDR 317
+ASL K+ G + + KK + S I W EG I R
Sbjct: 267 YMTLQGEAASLTDKYGFVVGLPLATSLLMKKKIQYQYSHGIDYWWTYMRADPEGLAEIQR 326
Query: 318 LLGLVQDGKLKYKMELT-PFNDFNTALDKALGKLGSQPKQVIKF 360
L+G GKLK +E T P D A +A K K V++F
Sbjct: 327 LVGA---GKLKIPVEKTFPITDV-VAAHEAKEKKQIPGKVVLEF 366
>At1g49670 ARP protein (At1g49670; F14J22.10)
Length = 629
Score = 74.3 bits (181), Expect = 9e-14
Identities = 81/323 (25%), Positives = 137/323 (42%), Gaps = 30/323 (9%)
Query: 41 KAIIYESHGQPDAVTKLVDIPAT-EVKENDLCVKMLAAPINPSDINRIQGVYPV--RPEP 97
K I++ + + T++V P + + + +K++ A +N SD+N G Y P+
Sbjct: 289 KMIVHTLSHKFRSATRIVRAPLQLPIGPHQVLLKIIYAGVNASDVNFSSGRYFTGGSPKL 348
Query: 98 PAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEY 157
P G+EGVG + +VG +V G + +FG + Y++ V + P E
Sbjct: 349 PFDAGFEGVGLIAAVGESVKNLEVG--TPAAVMTFGAYSEYMIVSSKHVLPVPRPDP-EV 405
Query: 158 AATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPG 217
A +T + LTAL+ LE + SG+ ++ A GQ +QLAK G N + G
Sbjct: 406 VAMLT-SGLTALIALEKAGQMKSGETVLVTAAAGGTGQFAVQLAKLSG----NKVIATCG 460
Query: 218 VGEVKERLKDLGADEVFTESELEVKNV--KSLLGGIPEPALGFNCVGGNSASLVLKFLRR 275
E + LK+LG D V +K V K G+ + + VGG + L L
Sbjct: 461 GSEKAKLLKELGVDRVIDYKSENIKTVLKKEFPKGV---NIIYESVGGQMFDMCLNALAV 517
Query: 276 GGTMVTYGGMSK-------KPV-------TVSTSSFIFKNWLSTDKAEEGRRMIDRLLGL 321
G ++ G +S+ +P + S + ++ ++ +D+L L
Sbjct: 518 YGRLIVIGMISQYQGEKGWEPAKYPGLCEKILAKSQTVAGFFLVQYSQLWKQNLDKLFNL 577
Query: 322 VQDGKLKYKMELTPFNDFNTALD 344
GKLK ++ F N D
Sbjct: 578 YALGKLKVGIDQKKFIGLNAVAD 600
>At3g56460 quinone reductase-like protein
Length = 348
Score = 73.2 bits (178), Expect = 2e-13
Identities = 69/256 (26%), Positives = 118/256 (45%), Gaps = 11/256 (4%)
Query: 51 PDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVH 110
P V+K IP+ + + V+++A +N ++ +I G Y +P P + G + G V
Sbjct: 23 PVEVSKTHPIPSLN-SDTSVRVRVIATSLNYANYLQILGKYQEKPPLPFIPGSDYSGIVD 81
Query: 111 SVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALL 170
++G AVT F GD V S G++ +IV DQ+ V + M AA + V T+ +
Sbjct: 82 AIGPAVTKFRVGDRVC-SFADLGSFAQFIVADQSRLFLVPERCDMVAAAALPVAFGTSHV 140
Query: 171 MLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVK----ERLK 226
L L SG ++ GA VG +Q+ K G I + R + +K + +
Sbjct: 141 ALVHRARLTSGQVLLVLGAAGGVGLAAVQIGKVCGAIVIAVARGTEKIQLLKSMGVDHVV 200
Query: 227 DLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMS 286
DLG + V + + +K K L G+ + ++ VGG +K L+ G ++ G S
Sbjct: 201 DLGTENVISSVKEFIKTRK--LKGVD---VLYDPVGGKLTKESMKVLKWGAQILVIGFAS 255
Query: 287 KKPVTVSTSSFIFKNW 302
+ + + + KNW
Sbjct: 256 GEIPVIPANIALVKNW 271
>At1g23740 auxin-induced protein like
Length = 386
Score = 73.2 bits (178), Expect = 2e-13
Identities = 71/268 (26%), Positives = 115/268 (42%), Gaps = 27/268 (10%)
Query: 34 SAVSPPSKAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPV 93
+++ KA +Y +G D + +I E+KE+ + +K++AA +NP D R QG +
Sbjct: 72 ASIPKEMKAWVYSDYGGVDVLKLESNIVVPEIKEDQVLIKVVAAALNPVDAKRRQGKFKA 131
Query: 94 RPEP-PAVGGYEGVGEVHSVGSAVTCFSPGDWV--------IPSPPSFGTWQTYIVKDQN 144
P P V GY+ G V VGSAV GD V + P FG+ Y ++
Sbjct: 132 TDSPLPTVPGYDVAGVVVKVGSAVKDLKEGDEVYANVSEKALEGPKQFGSLAEYTAVEEK 191
Query: 145 VWHKVNKGVPMEYAATITVNPLTALLMLEDCV--TLNSGDAIVQNGATSMVGQCVIQLAK 202
+ K + AA + PL E V ++G +I+ VG VIQLAK
Sbjct: 192 LLALKPKNIDFAQAAGL---PLAIETADEGLVRTEFSAGKSILVLNGAGGVGSLVIQLAK 248
Query: 203 SRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEP-ALGFNCV 261
++ + + E E ++ LGAD L + K + +P+ + F+ +
Sbjct: 249 H--VYGASKVAATAST-EKLELVRSLGAD-------LAIDYTKENIEDLPDKYDVVFDAI 298
Query: 262 GGNSASLVLKFLRRGGTMVTYGGMSKKP 289
G +K ++ GG +V G P
Sbjct: 299 G--MCDKAVKVIKEGGKVVALTGAVTPP 324
>At5g61510 quinone oxidoreductase - like protein
Length = 406
Score = 68.9 bits (167), Expect = 4e-12
Identities = 71/262 (27%), Positives = 111/262 (42%), Gaps = 7/262 (2%)
Query: 44 IYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGY 103
+YE HG P+ V K D+ E KE ++ VK A +N D+ +GVY P G
Sbjct: 89 VYE-HGGPE-VLKWEDVEVGEPKEGEIRVKNKAIGLNFIDVYFRKGVYKPA-SMPFTPGM 145
Query: 104 EGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATITV 163
E VGEV +VGS +T GD V + G + + + V + AA+I +
Sbjct: 146 EAVGEVVAVGSGLTGRMIGDLVAYAGNPMGAYAEEQILPADKVVPVPSSIDPIVAASIML 205
Query: 164 NPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKE 223
+TA +L C + G I+ + A VG + Q A + G I + + KE
Sbjct: 206 KGMTAQFLLRRCFKVEPGHTILVHAAAGGVGSLLCQWANALGATVIGTVSTNEKAAQAKE 265
Query: 224 RLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYG 283
D + ++E V V + G + ++ VG ++ L L+ G MV++G
Sbjct: 266 ---DGCHHVIMYKNEDFVSRVNDITSGKGVNVV-YDSVGKDTFKGSLACLKSRGYMVSFG 321
Query: 284 GMSKKPVTVSTSSFIFKNWLST 305
S P + S K+ T
Sbjct: 322 QSSGSPDPIPLSDLAPKSLFLT 343
>At4g13010 unknown protein
Length = 329
Score = 66.6 bits (161), Expect = 2e-11
Identities = 85/306 (27%), Positives = 129/306 (41%), Gaps = 35/306 (11%)
Query: 42 AIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQG-VYPVRPEP-PA 99
A+ Y S+G A + V +P K N++C+K+ A +NP D +G + P P P
Sbjct: 8 ALQYNSYGGGAAGLEHVQVPVPTPKSNEVCLKLEATSLNPVDWKIQKGMIRPFLPRKFPC 67
Query: 100 VGGYEGVGEVHSVGSAVTCFSPGDWVIP--SPPSFGTWQTYIVKDQNVWHKVNKGVPMEY 157
+ + GEV VGS V F GD V+ S G + V + + K + V
Sbjct: 68 IPATDVAGEVVEVGSGVKNFKAGDKVVAVLSHLGGGGLAEFAVATEKLTVKRPQEVGAAE 127
Query: 158 AATITVNPLTALLMLEDCVTLNSGDA-----IVQNGATSMVGQCVIQLAKSRGIHNINII 212
AA + V LTAL L + L I+ A+ VG +QLAK H +
Sbjct: 128 AAALPVAGLTALQALTNPAGLKLDGTGKKANILVTAASGGVGHYAVQLAKLANAH----V 183
Query: 213 RDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKF 272
G + E +K LGADEV E +KS G + + +C G S+
Sbjct: 184 TATCGARNI-EFVKSLGADEVLDYKTPEGAALKSPSGKKYDAVV--HCANGIPFSVFEPN 240
Query: 273 LRRGGTMV----------TYGGMSKKPVTVSTSSFIFKNWLSTDKAEEGRRMIDRLLGLV 322
L G ++ TY + K +T+S + L KAE ++ ++ LV
Sbjct: 241 LSENGKVIDITPGPNAMWTY---AVKKITMSKKQLV--PLLLIPKAEN----LEFMVNLV 291
Query: 323 QDGKLK 328
++GK+K
Sbjct: 292 KEGKVK 297
>At4g21580 putative NADPH quinone oxidoreductase
Length = 325
Score = 64.3 bits (155), Expect = 1e-10
Identities = 66/261 (25%), Positives = 113/261 (43%), Gaps = 8/261 (3%)
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAV 100
KAI+ G+P+ V +L D+ EVK++++ +++LA +N +D + G+Y P
Sbjct: 2 KAIVISEPGKPE-VLQLRDVADPEVKDDEVLIRVLATALNRADTLQRLGLYNPPPGSSPY 60
Query: 101 GGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAAT 160
G E G + SVG V+ + GD V G + V ++ + G+ ++ AA
Sbjct: 61 LGLECSGTIESVGKGVSRWKVGDQVCALLSGGGYAEKVSVPAGQIF-PIPAGISLKDAAA 119
Query: 161 ITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGE 220
T + L+ G++ + +G +S +G IQ+AK G+ + G E
Sbjct: 120 FPEVACTVWSTVFMMGRLSVGESFLIHGGSSGIGTFAIQIAKHLGVR----VFVTAGSDE 175
Query: 221 VKERLKDLGADEVFT-ESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTM 279
K+LGAD ++E V VK+ G + +C+G L L G +
Sbjct: 176 KLAACKELGADVCINYKTEDFVAKVKAETDGKGVDVI-LDCIGAPYLQKNLDSLNFDGRL 234
Query: 280 VTYGGMSKKPVTVSTSSFIFK 300
G M + SS + K
Sbjct: 235 CIIGLMGGANAEIKLSSLLPK 255
>At5g37980 quinone oxidoreductase -like protein
Length = 353
Score = 50.8 bits (120), Expect = 1e-06
Identities = 60/263 (22%), Positives = 110/263 (41%), Gaps = 32/263 (12%)
Query: 93 VRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGT---------WQTY--IVK 141
+R E P V +S+G ++ F + P++ T W+ Y I
Sbjct: 63 IRMEKPDPSSPASVARAYSIGKPISGFGVAKAIDSCHPNYKTGDLLWGRVGWEEYSVITP 122
Query: 142 DQNVWHKVNK-GVPME-YAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQ 199
+ K++ VP+ Y + + LTA + + + G+ + + A+ VGQ V Q
Sbjct: 123 TPSSHFKIHHTDVPLSFYTGLLGIPGLTAYVGFYEICSPKKGETVFVSAASGAVGQLVGQ 182
Query: 200 LAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFT---ESELEVKNVKSLLGGIPEPAL 256
AK G + + + V +K + G DE F E +L + GI +
Sbjct: 183 FAKMAGCYVVGSASSKEKVDLLKTK---FGYDEAFNYKEEHDLSAALKRCFPEGID---I 236
Query: 257 GFNCVGGNSASLVLKFLRRGGTMVTYGGMS----KKPVTV-STSSFIFK-----NWLSTD 306
F VGG VL+ +R G + G +S K+P V + +S ++K + + +
Sbjct: 237 YFENVGGKMLDAVLENMRTHGRIAACGMISQYNLKEPEGVHNLASIVYKRIRVQGFAAVE 296
Query: 307 KAEEGRRMIDRLLGLVQDGKLKY 329
++ + +D +L V++GK+ Y
Sbjct: 297 FFDKYSKFLDFILPYVREGKITY 319
>At5g38000 oxidoreductase-like protein
Length = 353
Score = 45.1 bits (105), Expect = 6e-05
Identities = 58/267 (21%), Positives = 111/267 (40%), Gaps = 35/267 (13%)
Query: 94 RPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGT---------WQTY--IVKD 142
+P+P + + ++G ++ F + P++ T W+ Y I
Sbjct: 67 KPDPSSPAS---MAHAFTIGKPISGFGVAKAIDSGHPNYKTGDLLWGRVGWEEYSVITPT 123
Query: 143 QNVWHKVNK-GVPME-YAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQL 200
+ K++ VP+ Y + + LTA + + + G+ + + A+ VGQ V Q
Sbjct: 124 PSSHFKIHHTDVPLSFYTGLLGIPGLTAYIGFYEICSPKKGETVFVSAASGAVGQLVGQF 183
Query: 201 AKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFT---ESELEVKNVKSLLGGIPEPALG 257
AK G + + V +K + G D+ F E +L + GI +
Sbjct: 184 AKMAGCYVVGSASSEEKVDLLKTK---FGYDDAFNYKEEKDLSAALKRCFPEGID---IY 237
Query: 258 FNCVGGNSASLVLKFLRRGGTMVTYGGMS----KKP-VTVSTSSFIFK-----NWLSTDK 307
F VGG VL+ +R G + G +S KKP V +T++ + K + + +
Sbjct: 238 FENVGGKMLEAVLENMRTHGRIAACGMISQYNLKKPEVLHNTATIVHKRIRVQGFAAVEF 297
Query: 308 AEEGRRMIDRLLGLVQDGKLKYKMELT 334
+ + +D +L V++GKL Y +++
Sbjct: 298 FDRYSKFLDFILPHVREGKLTYVEDIS 324
>At1g65560 putative oxido reductase (At1g65560)
Length = 350
Score = 45.1 bits (105), Expect = 6e-05
Identities = 66/287 (22%), Positives = 120/287 (40%), Gaps = 36/287 (12%)
Query: 97 PPAVGGY--EGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIV-------KDQNVWH 147
PP V G EG G + S T + PGD V W+ Y + + +N+
Sbjct: 72 PPFVPGQRIEGFGLARVIDSDDTNYKPGDIV----SGIIGWEEYSLLRSSDNLQLRNI-- 125
Query: 148 KVNKGVPMEY-AATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGI 206
+++ +P+ Y + + TA + GD++ + A VGQ V QLAK G
Sbjct: 126 QLDDDIPLSYHLGLLGMAGFTAYAGFNEICCPKKGDSVFVSAACGAVGQLVGQLAKLHGC 185
Query: 207 HNINIIRDRPGVGEVKERLKDLGADEVFT---ESELEVKNVKSLLGGIPEPALGFNCVGG 263
+ + + V +K +LG DE F E++L+ + GI + F+ VGG
Sbjct: 186 YVVGSAGSKQKVEILK---NELGYDEAFNYKEEADLDTALKRYFPEGID---IYFDNVGG 239
Query: 264 NSASLVLKFLRRGGTMVTYGGMSKKPVTVSTS------SFIFK-----NWLSTDKAEEGR 312
+ L ++ G + G +S + ++ S+ S I+K +L +D
Sbjct: 240 SMLDAALLNMKVRGRIALCGMVSLQSLSTSSQGIKNLYSAIYKRLRLEGFLQSDYLHIFP 299
Query: 313 RMIDRLLGLVQDGKLKYKMELTPFNDFNTALDKALGKLGSQPKQVIK 359
+ ++ + ++GK+ Y +++ D A L + KQV++
Sbjct: 300 QFLENVKRYYKEGKIVYVEDISEGLDLAPAALVGLFSGKNIGKQVVR 346
>At2g21730 putative cinnamyl-alcohol dehydrogenase
Length = 376
Score = 43.9 bits (102), Expect = 1e-04
Identities = 70/316 (22%), Positives = 121/316 (38%), Gaps = 48/316 (15%)
Query: 67 ENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWV- 125
END+ VK+L + SD++ I+ + P + G+E VG VG VT F GD V
Sbjct: 31 ENDVTVKILFCGVCHSDLHTIKNHWGFS-RYPIIPGHEIVGIATKVGKNVTKFKEGDRVG 89
Query: 126 ----------------------------IPSPPSFGT------WQTYIVKDQNVWHKVNK 151
S S GT + IV D +
Sbjct: 90 VGVIIGSCQSCESCNQDLENYCPKVVFTYNSRSSDGTSRNQGGYSDVIVVDHRFVLSIPD 149
Query: 152 GVPMEYAATITVNPLTALLMLEDC-VTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNIN 210
G+P + A + +T ++ +T SG + NG + G +++ K+ G+
Sbjct: 150 GLPSDSGAPLLCAGITVYSPMKYYGMTKESGKRLGVNGLGGL-GHIAVKIGKAFGLRVTV 208
Query: 211 IIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVL 270
I R + +E + LGAD ++ + +K +G + + V A L L
Sbjct: 209 ISRSSE---KEREAIDRLGADSFLVTTDSQ--KMKEAVGTMD---FIIDTVSAEHALLPL 260
Query: 271 -KFLRRGGTMVTYGGMSKKPVTVSTSSFIFKNWLSTDKAEEGRRMIDRLLGLVQDGKLKY 329
L+ G +V G + +KP+ + S + + G + +L K+
Sbjct: 261 FSLLKVNGKLVALG-LPEKPLDLPIFSLVLGRKMVGGSQIGGMKETQEMLEFCAKHKIVS 319
Query: 330 KMELTPFNDFNTALDK 345
+EL +D N+A+D+
Sbjct: 320 DIELIKMSDINSAMDR 335
>At5g37940 quinone oxidoreductase -like protein
Length = 353
Score = 42.0 bits (97), Expect = 5e-04
Identities = 55/262 (20%), Positives = 110/262 (40%), Gaps = 35/262 (13%)
Query: 94 RPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGT---------WQTY--IVKD 142
+P+P + + ++G ++ F + P++ T W+ Y I
Sbjct: 67 KPDPSSQAS---MAHAFTIGKPISGFGVAKAIDSGHPNYKTGDLLWGRVGWEEYSVITPT 123
Query: 143 QNVWHKVNK-GVPME-YAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQL 200
+ K++ VP+ Y + + LTA + + + G+ + + A+ VGQ V Q
Sbjct: 124 PSSHFKIHHTDVPLSFYTGLLGIPGLTAYVGFYEICSPKKGETVFVSAASGAVGQLVGQF 183
Query: 201 AKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFT---ESELEVKNVKSLLGGIPEPALG 257
AK G + + + V +K + G D+ F E +L + GI +
Sbjct: 184 AKMAGCYVVGSASSKEKVDLLKTK---FGYDDAFNYKEEKDLSAALKRCFPEGID---IY 237
Query: 258 FNCVGGNSASLVLKFLRRGGTMVTYGGMS----KKPVTV-STSSFIFK-----NWLSTDK 307
F VGG VL+ +R G + G +S K+P + +T++ + K ++ + +
Sbjct: 238 FENVGGKMLDAVLQNMRTHGRIAACGMISQYNLKEPEGLHNTATIVHKRIRVQDFAAVEF 297
Query: 308 AEEGRRMIDRLLGLVQDGKLKY 329
+ + +D +L V++GK+ Y
Sbjct: 298 FDRYSKFLDFILPHVREGKITY 319
>At2g21890 putative cinnamyl-alcohol dehydrogenase
Length = 375
Score = 42.0 bits (97), Expect = 5e-04
Identities = 69/315 (21%), Positives = 120/315 (37%), Gaps = 47/315 (14%)
Query: 67 ENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWV- 125
END+ VK+L + SD++ I+ + P + G+E VG VG VT F GD V
Sbjct: 31 ENDVTVKILFCGVCHSDLHTIKNHWGFS-RYPIIPGHEIVGIATKVGKNVTKFKEGDRVG 89
Query: 126 ----------------------------IPSPPSFGT-----WQTYIVKDQNVWHKVNKG 152
S S GT + IV D + G
Sbjct: 90 VGVIIGSCQSCESCNQDLENYCPKVVFTYNSRSSDGTRNQGGYSDVIVVDHRFVLSIPDG 149
Query: 153 VPMEYAATITVNPLTALLMLEDC-VTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINI 211
+P + A + +T ++ +T SG + NG + G +++ K+ G+ I
Sbjct: 150 LPSDSGAPLLCAGITVYSPMKYYGMTKESGKRLGVNGLGGL-GHIAVKIGKAFGLRVTVI 208
Query: 212 IRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVL- 270
R + +E + LGAD ++ + +K +G + + V A L L
Sbjct: 209 SRSSE---KEREAIDRLGADSFLVTTDSQ--KMKEAVGTMD---FIIDTVSAEHALLPLF 260
Query: 271 KFLRRGGTMVTYGGMSKKPVTVSTSSFIFKNWLSTDKAEEGRRMIDRLLGLVQDGKLKYK 330
L+ G +V G + +KP+ + + + G + +L K+
Sbjct: 261 SLLKVSGKLVALG-LLEKPLDLPIFPLVLGRKMVGGSQIGGMKETQEMLEFCAKHKIVSD 319
Query: 331 MELTPFNDFNTALDK 345
+EL +D N+A+D+
Sbjct: 320 IELIKMSDINSAMDR 334
>At5g16960 quinone oxidoreductase -like protein
Length = 346
Score = 40.8 bits (94), Expect = 0.001
Identities = 50/210 (23%), Positives = 87/210 (40%), Gaps = 19/210 (9%)
Query: 135 WQTY--IVKDQNVWHKVNK-GVPMEYAATITVNP-LTALLMLEDCVTLNSGDAIVQNGAT 190
W+ Y I N+ K++ P+ Y + P +TA + + T GD + + A+
Sbjct: 107 WEEYSVITPIPNLHFKIHHTNFPLSYYTGLLGMPGMTAYVGFYEICTPKKGDTVFVSAAS 166
Query: 191 SMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGG 250
VGQ V Q AK G + + + V +K + G D+ F E E + +L
Sbjct: 167 GAVGQLVGQFAKLMGCYVVGSAGSKEKVDLLKNK---FGFDDAFNYKE-EHNLIGALKRC 222
Query: 251 IPEPA-LGFNCVGGNSASLVLKFLRRGGTMVTYGGMS----KKPVTVSTSSFI------F 299
PE + F VGG V+ +R G + G +S K P + S I
Sbjct: 223 FPEGIDIYFENVGGKMLDAVILNMRPHGRIAACGMISQYNLKNPEGIYGLSLITYKRIRI 282
Query: 300 KNWLSTDKAEEGRRMIDRLLGLVQDGKLKY 329
+ + D + ++ ++ +++GK+KY
Sbjct: 283 EGFNCFDYFHKYSEFLEFVVPYIKEGKIKY 312
>At5g16970 quinone oxidoreductase -like protein
Length = 345
Score = 38.9 bits (89), Expect = 0.004
Identities = 44/191 (23%), Positives = 83/191 (43%), Gaps = 20/191 (10%)
Query: 153 VPMEYAATITVNP-LTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINI 211
VP+ Y + P +TA + + G+ + + A+ VGQ V QLAK G + +
Sbjct: 127 VPLSYYTGLLGMPGMTAYAGFYEVCSPKEGETVYVSAASGAVGQLVGQLAKMMGCYVVGS 186
Query: 212 IRDRPGVGEVKERLKDLGADEVFT---ESELEVKNVKSLLGGIPEPALGFNCVGGNSASL 268
+ V +K + G D+ F ES+L + GI + F VGG
Sbjct: 187 AGSKEKVDLLKTK---FGFDDAFNYKEESDLTAALKRCFPNGID---IYFENVGGKMLDA 240
Query: 269 VLKFLRRGGTMVTYGGMSK-----KPVTVSTSSFIFK-----NWLSTDKAEEGRRMIDRL 318
VL + G + G +S+ + + S+ I+K ++ +D ++ + ++ +
Sbjct: 241 VLVNMNMHGRIAVCGMISQYNLENQEGVHNLSNIIYKRIRIQGFVVSDFYDKYSKFLEFV 300
Query: 319 LGLVQDGKLKY 329
L +++GK+ Y
Sbjct: 301 LPHIREGKITY 311
>At3g59845 allyl alcohol dehydrogenase-like protein
Length = 346
Score = 38.1 bits (87), Expect = 0.007
Identities = 61/265 (23%), Positives = 103/265 (38%), Gaps = 33/265 (12%)
Query: 81 PSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGD--WVIPSPPSFGTWQTY 138
PS + + P +P G+G + S F+ GD W I + F T
Sbjct: 67 PSTLALLDAFIPGKP-------IAGIGVSQVIDSDDPSFTKGDMIWGIVNWEEFSTINPA 119
Query: 139 IVKDQNVWHKVNKGVPMEYAATIT-VNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCV 197
+ +V N VP+ Y I + LTA + + GD + + A+ VGQ V
Sbjct: 120 AIFKIDV----NINVPLSYYTGILGMIGLTAYAGFFEICSPKKGDTVFVSAASGAVGQLV 175
Query: 198 IQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVFT---ESELEVKNVKSLLGGIPEP 254
Q AK G + ++ +V L G D+ F E +L+ + + GI
Sbjct: 176 GQFAKLMGCY---VVGSAGSKQKVDLLLNKFGYDDAFNYKEEPDLDSALKRCVPKGID-- 230
Query: 255 ALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVTV-----STSSFIFK-----NWLS 304
+ F VGG VL ++ G + G +S+ + + IFK + S
Sbjct: 231 -IYFENVGGKMLDAVLLNMKTYGRIAVCGMISQYHLETRDRLQNLPDIIFKKIRMQGFAS 289
Query: 305 TDKAEEGRRMIDRLLGLVQDGKLKY 329
D + + ++ +L +++ KL Y
Sbjct: 290 YDFIDRFPKFLEFVLPYIKEEKLAY 314
>At3g03080 putative NADP-dependent oxidoreductase
Length = 350
Score = 37.0 bits (84), Expect = 0.017
Identities = 46/211 (21%), Positives = 83/211 (38%), Gaps = 22/211 (10%)
Query: 135 WQTY--IVKDQNVWHKVNKGVPMEYAATITVNP-LTALLMLEDCVTLNSGDAIVQNGATS 191
W Y I D + + + VP+ Y + P +TA + + G+ + + A+
Sbjct: 112 WGEYSLITPDFSHYKIQHTDVPLSYYTGLLGMPGMTAYAGFYEICSPKKGETVFVSAASG 171
Query: 192 MVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGADEVF---TESELEVKNVKSLL 248
VGQ V Q AK G + + V +K + G D+ F E +L +
Sbjct: 172 AVGQLVGQFAKIMGCYVVGSAGSNEKVDLLKNK---FGFDDAFNYKAEPDLNAALKRCFP 228
Query: 249 GGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSK----------KPVTVSTSSFI 298
GI + F VGG VL ++ G + G +S+ V
Sbjct: 229 EGID---IYFENVGGKMLDAVLLNMKLHGRIAVCGMISQYNLEDQEGVHNLANVIYKRIR 285
Query: 299 FKNWLSTDKAEEGRRMIDRLLGLVQDGKLKY 329
K ++ +D ++ + +D +L +++GK+ Y
Sbjct: 286 IKGFVVSDYFDKHLKFLDFVLPYIREGKITY 316
>At5g16990 quinone oxidoreductase - like protein
Length = 343
Score = 36.6 bits (83), Expect = 0.022
Identities = 43/191 (22%), Positives = 81/191 (41%), Gaps = 20/191 (10%)
Query: 153 VPMEYAATITVNP-LTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINI 211
VP+ Y + P +TA + + G+ + + A+ VGQ V Q AK G + +
Sbjct: 125 VPLSYYTGLLGMPGMTAYAGFYEVCSPKKGETVYVSAASGAVGQLVGQFAKMMGCYVVGS 184
Query: 212 IRDRPGVGEVKERLKDLGADEVFT---ESELEVKNVKSLLGGIPEPALGFNCVGGNSASL 268
+ V +K + G D+ F ES+L + GI + F VGG
Sbjct: 185 AGSKEKVDLLKTK---FGFDDAFNYKEESDLSAALKRCFPKGID---MYFENVGGKMLDA 238
Query: 269 VLKFLRRGGTMVTYGGMSK-----KPVTVSTSSFIFK-----NWLSTDKAEEGRRMIDRL 318
VL + G + G +S+ + + S+ I+K ++ D ++ + ++ +
Sbjct: 239 VLLNMNPHGRIAVCGMISQYNLENQEGVHNLSNIIYKRIRIQGFVVADFYDKYPKFLELV 298
Query: 319 LGLVQDGKLKY 329
L +++GK+ Y
Sbjct: 299 LPRIKEGKITY 309
>At1g32780 alcohol dehydrogenase like protein
Length = 394
Score = 35.8 bits (81), Expect = 0.037
Identities = 21/57 (36%), Positives = 29/57 (50%)
Query: 72 VKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVIPS 128
VK+L + I +D+ G P + G+E VG V SVG V GD+VIP+
Sbjct: 40 VKILYSSICHTDLGCWNGTNEAERAFPRILGHEAVGIVESVGEGVKDVKEGDYVIPT 96
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.135 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,358,279
Number of Sequences: 26719
Number of extensions: 370194
Number of successful extensions: 944
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 20
Number of HSP's successfully gapped in prelim test: 29
Number of HSP's that attempted gapping in prelim test: 908
Number of HSP's gapped (non-prelim): 51
length of query: 360
length of database: 11,318,596
effective HSP length: 101
effective length of query: 259
effective length of database: 8,619,977
effective search space: 2232574043
effective search space used: 2232574043
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC135230.4


