

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135230.3 + phase: 0 /pseudo
(605 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
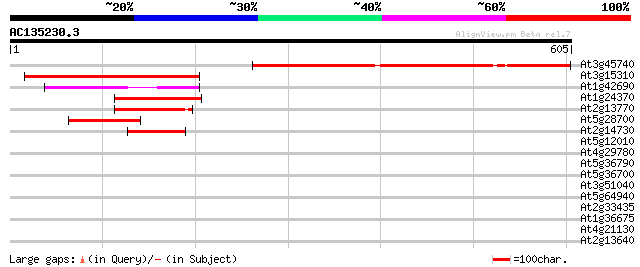
Score E
Sequences producing significant alignments: (bits) Value
At3g45740 unknown protein 483 e-136
At3g15310 unknown protein 180 2e-45
At1g42690 unknown protein 144 1e-34
At1g24370 hypothetical protein 124 1e-28
At2g13770 hypothetical protein 90 3e-18
At5g28700 putative protein 83 5e-16
At2g14730 hypothetical protein 63 5e-10
At5g12010 putative protein 42 0.001
At4g29780 unknown protein 35 0.12
At5g36790 p-nitrophenylphosphatase-like protein 33 0.35
At5g36700 N-glyceraldehyde-2-phosphotransferase-like 33 0.35
At3g51040 unknown protein 30 3.9
At5g64940 ABC transporter-like 30 5.1
At2g33435 RRM-containing protein 30 5.1
At1g36675 putative protein 30 5.1
At4g21130 hypothetical protein 29 6.6
At2g13640 hypothetical protein 29 6.6
>At3g45740 unknown protein
Length = 376
Score = 483 bits (1243), Expect = e-136
Identities = 242/343 (70%), Positives = 282/343 (81%), Gaps = 10/343 (2%)
Query: 262 RPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASEL 321
R SFGIAFDIDGVILLG++PVGGSP+ALR+LY+ G LK P++FLTNGGG+PE+KRASE+
Sbjct: 36 RSSFGIAFDIDGVILLGSSPVGGSPSALRRLYDDSGALKIPFLFLTNGGGLPESKRASEM 95
Query: 322 SELLGLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYGFKNVISIDAYAS 381
S LLG+ VS QV+Q HSPFR+LVNRFE++L+VAAGKGEPA VMS YGFKNVIS+D YAS
Sbjct: 96 SHLLGVQVSPLQVIQAHSPFRKLVNRFENELVVAAGKGEPAAVMSNYGFKNVISMDEYAS 155
Query: 382 RFENIDPLAPYKKWTTKLATTQNPKFDESGPQIDVFSERVQAAFIVSDPVDWSRDVQVLC 441
F+NIDPLAPYKK + E + DV S+RVQAAFIVSDPVDWSRD+QVLC
Sbjct: 156 YFDNIDPLAPYKK-----LMFRQDGHKELRSREDVLSQRVQAAFIVSDPVDWSRDIQVLC 210
Query: 442 DILKTGGLPGRNVGTQPHLYFANDDLEYQTKFPSERLGMGAFRIALESIFNRTHHHSLEY 501
DIL+TGGLPG+ +G QPHLY ANDDL+YQT+FP+ERLGMGAFRIALESIFNR H LEY
Sbjct: 211 DILRTGGLPGKEIGPQPHLYIANDDLDYQTEFPTERLGMGAFRIALESIFNRIHEKPLEY 270
Query: 502 TCFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNAQPFQTLYMIGDNPAVDIRGARQT 561
T FGKP+P VFKNAE VL++ + Y N + + F+TLYMIGDNP +DIRGARQ
Sbjct: 271 TSFGKPNPFVFKNAEDVLKE----IATSPYSSN-QGSHHFKTLYMIGDNPKIDIRGARQA 325
Query: 562 GHPWFSILTRTGVFKGKENHDKFPADLVVDTVEEAVDYILAKE 604
G PWFSILTRTGVFKGK+NH +FPADLVVDTVEEAVD+IL +E
Sbjct: 326 GTPWFSILTRTGVFKGKDNHSEFPADLVVDTVEEAVDFILTRE 368
>At3g15310 unknown protein
Length = 415
Score = 180 bits (457), Expect = 2e-45
Identities = 87/188 (46%), Positives = 120/188 (63%)
Query: 17 DTYIVNRFIQRRKKLEEGSASRTRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRM 76
D N F++ E +R R YF+R + L +DYF++ + FRRR+RM
Sbjct: 22 DQQFENVFVELSNSEEVAKKNRKRAYFDRKREEGHVLLWNDYFSDNAIFPLQTFRRRFRM 81
Query: 77 QKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKIG 136
+K +FLRIV LSS +F + DA + G SP+ KCT A+R+LAYG A+DAVDEY+++G
Sbjct: 82 KKPLFLRIVDRLSSELMFFQHRRDATGRFGHSPIQKCTAAIRLLAYGYASDAVDEYLRMG 141
Query: 137 SSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWKN 196
+T + CL F KGII + YLRAPT + R+ ++ + RGFP MIGS+DCMHWEWKN
Sbjct: 142 ETTAMSCLENFTKGIISFFGDEYLRAPTATNLRRLLNIGKIRGFPGMIGSLDCMHWEWKN 201
Query: 197 YPKAWEGK 204
P AW+G+
Sbjct: 202 CPTAWKGQ 209
>At1g42690 unknown protein
Length = 333
Score = 144 bits (363), Expect = 1e-34
Identities = 74/167 (44%), Positives = 95/167 (56%), Gaps = 30/167 (17%)
Query: 38 RTRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQ 97
+ + Y R+ + +L++DYF PTY +FRRR+RM K +F+RIV S+ YF Q
Sbjct: 39 KKKLYIERNREEGHIQLVNDYFTENPTYPPHIFRRRFRMNKSLFMRIVERFSNEVPYFKQ 98
Query: 98 QVDAANKKGISPLAKCTTAMRMLAYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQ 157
+ DA + G S L K T A+RMLAYG+AADA
Sbjct: 99 RRDATRRLGFSALQKSTAAIRMLAYGIAADA----------------------------- 129
Query: 158 VYLRAPTQDDPHRIQHVSETRGFPWMIGSIDCMHWEWKNYPKAWEGK 204
YLR PT+DD R+ H+ E RGFP MIGSIDCMHWEWKN P AW+G+
Sbjct: 130 -YLRRPTRDDLIRLLHIGEQRGFPGMIGSIDCMHWEWKNCPTAWKGQ 175
>At1g24370 hypothetical protein
Length = 413
Score = 124 bits (312), Expect = 1e-28
Identities = 55/93 (59%), Positives = 69/93 (74%)
Query: 114 TTAMRMLAYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQH 173
T A RMLAYGV AD+ DEYIKIG ST LE L RFC+ I+ ++ YLR+P +D R+ H
Sbjct: 2 TAAFRMLAYGVPADSTDEYIKIGESTALESLKRFCRAIVEVFACRYLRSPDANDVARLLH 61
Query: 174 VSETRGFPWMIGSIDCMHWEWKNYPKAWEGKLA 206
+ E+RGFP M+GS+DCMHW+WKN P AW G+ A
Sbjct: 62 IGESRGFPRMLGSLDCMHWKWKNCPTAWGGQYA 94
>At2g13770 hypothetical protein
Length = 244
Score = 90.1 bits (222), Expect = 3e-18
Identities = 43/84 (51%), Positives = 54/84 (64%), Gaps = 2/84 (2%)
Query: 114 TTAMRMLAYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQH 173
T A RMLAYGV AD+ DEYIKIG ST LE L RFC+ I+ ++ YLR+P +D R+ H
Sbjct: 2 TAAFRMLAYGVPADSTDEYIKIGESTALESLKRFCRAIVEVFACCYLRSPDANDVARLLH 61
Query: 174 VSETRGFPWMIGSIDCMHWEWKNY 197
+ E+RGFP + D W W Y
Sbjct: 62 IGESRGFPEAVADYDL--WIWHAY 83
>At5g28700 putative protein
Length = 292
Score = 82.8 bits (203), Expect = 5e-16
Identities = 42/78 (53%), Positives = 53/78 (67%)
Query: 64 TYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRMLAYG 123
TY +FR R+RM K +F+RIV S+ YF Q+ DA + S L K T A+RMLAYG
Sbjct: 33 TYPPHIFRHRFRMNKPLFMRIVERFSNEVPYFKQRRDATGRLDFSALQKSTAAIRMLAYG 92
Query: 124 VAADAVDEYIKIGSSTTL 141
+AADAVDEY++IG ST L
Sbjct: 93 IAADAVDEYLRIGESTLL 110
>At2g14730 hypothetical protein
Length = 117
Score = 62.8 bits (151), Expect = 5e-10
Identities = 28/62 (45%), Positives = 41/62 (65%)
Query: 128 AVDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVSETRGFPWMIGSI 187
AVD+Y++IG +T + C++ + II L+ + YLR PT+ D R+ + E RGF M GSI
Sbjct: 19 AVDKYLRIGENTLMSCMIHSVEAIIYLFGKEYLRRPTRQDLKRLLRIGELRGFLGMTGSI 78
Query: 188 DC 189
DC
Sbjct: 79 DC 80
>At5g12010 putative protein
Length = 502
Score = 41.6 bits (96), Expect = 0.001
Identities = 33/128 (25%), Positives = 56/128 (42%), Gaps = 5/128 (3%)
Query: 65 YDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRMLAYGV 124
Y + F++ +RM K F I +L+S+ + D A + I + + LA G
Sbjct: 170 YPEEDFKKAFRMSKSTFELICDELNSA----VAKEDTALRNAIPVRQRVAVCIWRLATGE 225
Query: 125 AADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVSET-RGFPWM 183
V + +G ST + +L CK I + YL+ P + I+ E+ G P +
Sbjct: 226 PLRLVSKKFGLGISTCHKLVLEVCKAIKDVLMPKYLQWPDDESLRNIRERFESVSGIPNV 285
Query: 184 IGSIDCMH 191
+GS+ H
Sbjct: 286 VGSMYTTH 293
>At4g29780 unknown protein
Length = 540
Score = 35.0 bits (79), Expect = 0.12
Identities = 30/132 (22%), Positives = 55/132 (40%), Gaps = 5/132 (3%)
Query: 61 NEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRML 120
+ P + + FRR +RM K F I +L ++ + + + I + + L
Sbjct: 204 SRPDFPEDEFRREFRMSKSTFNLICEELDTT----VTKKNTMLRDAIPAPKRVGVCVWRL 259
Query: 121 AYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQDDPHRIQHVSET-RG 179
A G V E +G ST + ++ C+ I + YL P+ + + + E+
Sbjct: 260 ATGAPLRHVSERFGLGISTCHKLVIEVCRAIYDVLMPKYLLWPSDSEINSTKAKFESVHK 319
Query: 180 FPWMIGSIDCMH 191
P ++GSI H
Sbjct: 320 IPNVVGSIYTTH 331
>At5g36790 p-nitrophenylphosphatase-like protein
Length = 362
Score = 33.5 bits (75), Expect = 0.35
Identities = 31/104 (29%), Positives = 45/104 (42%), Gaps = 8/104 (7%)
Query: 258 DLLIRPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKR 317
D LI FD DGVI G+ + G P L L L VF+TN K+
Sbjct: 73 DQLIDSVETFIFDCDGVIWKGDKLIEGVPETLDMLRAKGKRL----VFVTN-NSTKSRKQ 127
Query: 318 ASELSELLGLNVSASQVLQGH---SPFRQLVNRFEDKLIVAAGK 358
+ E LGLNV+ ++ + + Q +N +DK + G+
Sbjct: 128 YGKKFETLGLNVNEEEIFASSFAAAAYLQSINFPKDKKVYVIGE 171
>At5g36700 N-glyceraldehyde-2-phosphotransferase-like
Length = 289
Score = 33.5 bits (75), Expect = 0.35
Identities = 31/104 (29%), Positives = 45/104 (42%), Gaps = 8/104 (7%)
Query: 258 DLLIRPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKR 317
D LI FD DGVI G+ + G P L L L VF+TN K+
Sbjct: 20 DQLIDSVETFIFDCDGVIWKGDKLIEGVPETLDMLRAKGKRL----VFVTN-NSTKSRKQ 74
Query: 318 ASELSELLGLNVSASQVLQGH---SPFRQLVNRFEDKLIVAAGK 358
+ E LGLNV+ ++ + + Q +N +DK + G+
Sbjct: 75 YGKKFETLGLNVNEEEIFASSFAAAAYLQSINFPKDKKVYVIGE 118
>At3g51040 unknown protein
Length = 231
Score = 30.0 bits (66), Expect = 3.9
Identities = 19/61 (31%), Positives = 31/61 (50%), Gaps = 4/61 (6%)
Query: 21 VNRFIQRRKKLEEGSASRTRKYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRMQKHV 80
V+R+IQ K++E S S + FN + + +D EPT+DDA+ + Q H
Sbjct: 79 VSRYIQINKEMES-SRSSSSGMFNGERRYEQE---EDSHEKEPTWDDALRKSTQEYQHHS 134
Query: 81 F 81
+
Sbjct: 135 Y 135
>At5g64940 ABC transporter-like
Length = 761
Score = 29.6 bits (65), Expect = 5.1
Identities = 16/52 (30%), Positives = 29/52 (55%), Gaps = 1/52 (1%)
Query: 486 ALESIFNRTHHHSLEYTCFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKN 537
++E IF+R + + G+ H + K E+VL+ P + +D +DI+ KN
Sbjct: 281 SVEDIFDRFDYEPIAAASLGQVHRARLKGQEVVLKVQRPGL-KDLFDIDLKN 331
>At2g33435 RRM-containing protein
Length = 979
Score = 29.6 bits (65), Expect = 5.1
Identities = 31/100 (31%), Positives = 44/100 (44%), Gaps = 12/100 (12%)
Query: 28 RKKLEEGSASRTRKYFNRDHVAANQ-------RLIDDYFANEPTYDDAMFRRRYRMQKHV 80
RKK E S+SR K DHV A Q ++ DY E YD + R K
Sbjct: 462 RKKEEAISSSRGEKPIKEDHVGAAQLLGNDLVEMVSDYHETEKGYDRSKKLSREERVKDS 521
Query: 81 FLRIVGDLSSS--DNYFTQQVD--AANKKGISPLAKCTTA 116
+ +SSS +N Q+ D +N+K + +C+TA
Sbjct: 522 SRKKEEAISSSREENLDKQKKDESTSNRKRKAE-GECSTA 560
>At1g36675 putative protein
Length = 268
Score = 29.6 bits (65), Expect = 5.1
Identities = 12/19 (63%), Positives = 16/19 (84%)
Query: 87 DLSSSDNYFTQQVDAANKK 105
DLSSSDNYFT++ +A K+
Sbjct: 175 DLSSSDNYFTKRFEATKKE 193
>At4g21130 hypothetical protein
Length = 537
Score = 29.3 bits (64), Expect = 6.6
Identities = 24/84 (28%), Positives = 40/84 (47%), Gaps = 6/84 (7%)
Query: 14 DVEDTYIVNRFIQRRKKLEEGSASRTRKYFNRDHVAANQRLIDDYFANE---PTYDDAMF 70
+VED + ++RK+L E + +R + R+H N+ DD F + T
Sbjct: 58 EVEDEFAHETVGEKRKRLAEDTLNRIEEAKQREHEEDNEE--DDDFRDSLVAKTLMQEQL 115
Query: 71 RRRYRMQKHVFLRIVGDLSSSDNY 94
+ R+++ LR V DL SSD +
Sbjct: 116 EKSGRVRRANALR-VQDLQSSDKF 138
>At2g13640 hypothetical protein
Length = 384
Score = 29.3 bits (64), Expect = 6.6
Identities = 22/120 (18%), Positives = 53/120 (43%), Gaps = 13/120 (10%)
Query: 13 RDVEDTYIVNRFIQRRKKLEEGSASRTRKYFNRD--------HVAANQRLIDDYFANEPT 64
R+ E N F+ ++K + S + KYF ++ ++ + ++D +P
Sbjct: 182 RETEKNKARNSFLALKQKEDHRSETCVEKYFKKETKTNLKQLNMKTRRVSLEDVTTKKP- 240
Query: 65 YDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQVDAANKKGI----SPLAKCTTAMRML 120
++D ++ + +K + + V ++ + D AN KG+ +++C A+ +L
Sbjct: 241 HEDGSLIKKTKTEKELKKKKVDEMVKLFEAAKKAADVANAKGVLSGKPEVSRCIDALSLL 300
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,655,655
Number of Sequences: 26719
Number of extensions: 591924
Number of successful extensions: 1246
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 1231
Number of HSP's gapped (non-prelim): 22
length of query: 605
length of database: 11,318,596
effective HSP length: 105
effective length of query: 500
effective length of database: 8,513,101
effective search space: 4256550500
effective search space used: 4256550500
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC135230.3


