

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135230.11 + phase: 0 /pseudo
(954 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
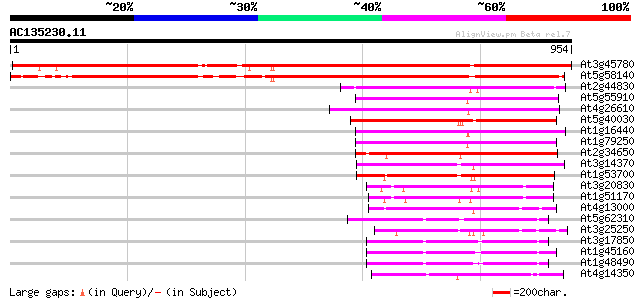
Score E
Sequences producing significant alignments: (bits) Value
At3g45780 nonphototropic hypocotyl 1 (NPH1) 1195 0.0
At5g58140 non phototropic hypocotyl 1-like (NPL1) 980 0.0
At2g44830 protein kinase like protein 349 5e-96
At5g55910 serine/threonine-specific protein kinase ATPK64 (pir||... 345 7e-95
At4g26610 putative protein kinase 345 7e-95
At5g40030 protein kinase -like protein 339 4e-93
At1g16440 putative protein kinase 337 1e-92
At1g79250 hypothetical protein 337 2e-92
At2g34650 protein kinase PINOID 298 1e-80
At3g14370 putative protein kinase 291 9e-79
At1g53700 auxin-induced protein kinase like protein 285 9e-77
At3g20830 unknown protein 229 4e-60
At1g51170 putative serine/threonine protein kinase 226 4e-59
At4g13000 unknown protein 220 3e-57
At5g62310 IRE (root hair elongation) 218 1e-56
At3g25250 protein kinase, putative 216 4e-56
At3g17850 protein kinase like protein 214 1e-55
At1g45160 similar to IRE homolog 1 dbj|BAA89784.1 213 4e-55
At1g48490 IRE - like protein kinase 203 3e-52
At4g14350 protein kinase 200 3e-51
>At3g45780 nonphototropic hypocotyl 1 (NPH1)
Length = 996
Score = 1195 bits (3092), Expect = 0.0
Identities = 644/1014 (63%), Positives = 750/1014 (73%), Gaps = 88/1014 (8%)
Query: 6 KSPSSSSQRPSFPRDPRGSLEVFNPTSNST---SPVRSPSHLKTWT-------------- 48
+ PS+ + PRD RGSLEVFNP++ T +PV P W
Sbjct: 5 EKPSTKPSSRTLPRDTRGSLEVFNPSTQLTRPDNPVFRPEP-PAWQNLSDPRGTSPQPRP 63
Query: 49 ETEEQHKDFISTDE--VTNTSWMAIKEGETG-----------------AAAQRAAEWGLV 89
+ E + + +D+ TSWMA+K+ AA QRAAEWGLV
Sbjct: 64 QQEPAPSNPVRSDQEIAVTTSWMALKDPSPETISKKTITAEKPQKSAVAAEQRAAEWGLV 123
Query: 90 LRTDAETGKPQGVGVRNSG--DDEQNGK-FSGKRNSNNSGRVSGDSSDGGDP---RGFPR 143
L+TD +TGKPQGVGVRNSG +++ NGK + +RNS NS R SG+ SDG P G PR
Sbjct: 124 LKTDTKTGKPQGVGVRNSGGTENDPNGKKTTSQRNSQNSCRSSGEMSDGDVPGGRSGIPR 183
Query: 144 VSEDLKDALSAFQQTFVVSDATKPDYPIMYASAGFFNMTGYTSKEVIGRNCRFLQGADTD 203
VSEDLKDALS FQQTFVVSDATKPDYPIMYASAGFFNMTGYTSKEV+GRNCRFLQG+ TD
Sbjct: 184 VSEDLKDALSTFQQTFVVSDATKPDYPIMYASAGFFNMTGYTSKEVVGRNCRFLQGSGTD 243
Query: 204 PQDVAKIREALEGGKSYCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLKLIGMLVEVNKH 263
++AKIRE L G +YCGR+LNYKKDGT FWNLLTI+PIKD+ G VLK IGM VEV+KH
Sbjct: 244 ADELAKIRETLAAGNNYCGRILNYKKDGTSFWNLLTIAPIKDESGKVLKFIGMQVEVSKH 303
Query: 264 TEGSKEKNLRPNGLPESLIRYDARQKEKASSSVSELLQAMKRPRALSESGQ-RPFIIKSG 322
TEG+KEK LRPNGLPESLIRYDARQK+ A++SV+EL++A+KRPRALSES PF+ KS
Sbjct: 304 TEGAKEKALRPNGLPESLIRYDARQKDMATNSVTELVEAVKRPRALSESTNLHPFMTKS- 362
Query: 323 GCSEEDQEIEKVEHKSRRKSDSVA-SFRPKSQRKSRSSMERISELPENANKNSHRHSFMG 381
E E+ K +RR S++V S R S R+SM+RI+E+PE ++ S SFMG
Sbjct: 363 ----ESDELPK--KPARRMSENVVPSGRRNSGGGRRNSMQRINEIPEKKSRKSSL-SFMG 415
Query: 382 LMKALIMKLL*I*ALRVRMMTGMIV-------LNLMTKKS*GKSEKVLILLLHLSV-LRK 433
+ K +L + G I ++ ++ +KV + + L
Sbjct: 416 IKKKSE-------SLDESIDDGFIEYGEEDDEISDRDERPESVDDKVRQKEMRKGIDLAT 468
Query: 434 TLSLLIQGFLT---------ILY----FLELTEYSREEILGRNCRFLQGPETDPATVRKI 480
TL + + F+ I++ FLELTEYSREEILGRNCRFLQGPETD TV+KI
Sbjct: 469 TLERIEKNFVITDPRLPDNPIIFASDSFLELTEYSREEILGRNCRFLQGPETDLTTVKKI 528
Query: 481 REAIDNQTEVTVQLINYTRTGKKFWNLFHLQPMRDHKGEVQYFIGVQLDGSQHVEPLHNC 540
R AIDNQTEVTVQLINYT++GKKFWN+FHLQPMRD KGEVQYFIGVQLDGS+HVEP+ N
Sbjct: 529 RNAIDNQTEVTVQLINYTKSGKKFWNIFHLQPMRDQKGEVQYFIGVQLDGSKHVEPVRNV 588
Query: 541 IKEDTAKEGEQLVKQTAENVGEAVRELPDANQKPDDLWLNHSKVVHPKPHRKDNDAWRAI 600
I+E KEGE LVK+TA N+ EAVRELPDAN P+DLW NHSKVVH KPHRKD+ W AI
Sbjct: 589 IEETAVKEGEDLVKKTAVNIDEAVRELPDANMTPEDLWANHSKVVHCKPHRKDSPPWIAI 648
Query: 601 QKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRA 660
QK++E+GE I LKHF+P+KPLGSGDTGSVHLVEL GT Q FAMKAMDK VMLNRNKVHRA
Sbjct: 649 QKVLESGEPIGLKHFKPVKPLGSGDTGSVHLVELVGTDQLFAMKAMDKAVMLNRNKVHRA 708
Query: 661 CTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFY 720
EREILD+LDHPFLPALYASFQTKTH+CLITDYYPGGELF+LLD+QP KVLKEDAVRFY
Sbjct: 709 RAEREILDLLDHPFLPALYASFQTKTHICLITDYYPGGELFMLLDRQPRKVLKEDAVRFY 768
Query: 721 AAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKK 780
AA+V++ALEYLHCQGIIYRDLKPENVLIQ NG +SL+DFDLSCLTSCKPQL+IP+ ++KK
Sbjct: 769 AAQVVVALEYLHCQGIIYRDLKPENVLIQGNGDISLSDFDLSCLTSCKPQLLIPSIDEKK 828
Query: 781 KRKKKKKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILL 840
K+K QQK+QQ P FMAEPMRASNSFVGTEEYIAPEII+G+GHTSAVDWWALGIL+
Sbjct: 829 KKK------QQKSQQTPIFMAEPMRASNSFVGTEEYIAPEIISGAGHTSAVDWWALGILM 882
Query: 841 YEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEG 900
YEMLYGYTPFRGKTRQKTF N+L KDLKFP S P S Q KQLI+ LL RDPK RLG EG
Sbjct: 883 YEMLYGYTPFRGKTRQKTFTNVLQKDLKFPASIPASLQVKQLIFRLLQRDPKKRLGCFEG 942
Query: 901 ANEIKSHPFFKNVNWALIRCMKPPELDAPILLENDEKKEAKDIDPGLDDLQKNI 954
ANE+K H FFK +NWALIRC PPEL+ PI E E K +DP L+DLQ N+
Sbjct: 943 ANEVKQHSFFKGINWALIRCTNPPELETPIFSGEAENGE-KVVDPELEDLQTNV 995
>At5g58140 non phototropic hypocotyl 1-like (NPL1)
Length = 915
Score = 980 bits (2533), Expect = 0.0
Identities = 531/960 (55%), Positives = 658/960 (68%), Gaps = 81/960 (8%)
Query: 1 MERLKKSPSSSSQRPSFPRDPRGSLEVFNPTSNSTSPVRSPSHLKTWTETEEQHKDFIST 60
MER + PS + S R SLE+FNP+S + + S K +
Sbjct: 1 MERPRAPPSPLNDAESLSE--RRSLEIFNPSSGKETHGSTSSSSKPPLD---------GN 49
Query: 61 DEVTNTSWMAIKEGETGAAAQRAAEWGLVLRTDAETGKPQGVGVRNSGDDEQNGKFSGK- 119
++ +++ WM ++ + +R AEWGL KP +SGDD + K S +
Sbjct: 50 NKGSSSKWMEFQD--SAKITERTAEWGL------SAVKP------DSGDDGISFKLSSEV 95
Query: 120 RNSNNSGRVSGDSSDGGDPRGFPRVSEDLKDALSAFQQTFVVSDATKPDYPIMYASAGFF 179
S N R S + S + FPRVS++LK ALS QQTFVVSDAT+P PI+YAS+GFF
Sbjct: 96 ERSKNMSRRSSEESTSSESGAFPRVSQELKTALSTLQQTFVVSDATQPHCPIVYASSGFF 155
Query: 180 NMTGYTSKEVIGRNCRFLQGADTDPQDVAKIREALEGGKSYCGRLLNYKKDGTPFWNLLT 239
MTGY+SKE++GRNCRFLQG DTD +VAKIR+ ++ GKSYCGRLLNYKKDGTPFWNLLT
Sbjct: 156 TMTGYSSKEIVGRNCRFLQGPDTDKNEVAKIRDCVKNGKSYCGRLLNYKKDGTPFWNLLT 215
Query: 240 ISPIKDDDGNVLKLIGMLVEVNKHTEGSKEKNLRPNGLPESLIRYDARQKEKASSSVSEL 299
++PIKDD GN +K IGM VEV+K+TEG +K LRPNGL +SLIRYDARQKEKA S++E+
Sbjct: 216 VTPIKDDQGNTIKFIGMQVEVSKYTEGVNDKALRPNGLSKSLIRYDARQKEKALDSITEV 275
Query: 300 LQAMK-RPRALSESGQRPFIIKSGGCSEEDQEIEKVEHKSRRKSDSVASFRPKSQRKSRS 358
+Q ++ R + ES ++K D + R+SD + KS
Sbjct: 276 VQTIRHRKSQVQESVSNDTMVKP------DSSTTPTPGRQTRQSDEAS--------KSFR 321
Query: 359 SMERISELPENANKNSH-RHSFMGLMKALIMKLL*I*ALRVRMMTGMIVLNLMTKKS*GK 417
+ R+S + K+S+ RH + M+ +M V+
Sbjct: 322 TPGRVSTPTGSKLKSSNNRHEDLLRMEP------------EELMLSTEVIGQRDSWDLSD 369
Query: 418 SEKVLILLLHLSVLRKTLSLLIQGFLT---------ILY----FLELTEYSREEILGRNC 464
E+ + + L+ TL + + F+ I++ FLELTEYSREEILGRNC
Sbjct: 370 RERDIRQGIDLAT---TLERIEKNFVISDPRLPDNPIIFASDSFLELTEYSREEILGRNC 426
Query: 465 RFLQGPETDPATVRKIREAIDNQTEVTVQLINYTRTGKKFWNLFHLQPMRDHKGEVQYFI 524
RFLQGPETD ATV+KIR+AI +Q E+TVQLINYT++GKKFWNLFHLQPMRD KGE+QYFI
Sbjct: 427 RFLQGPETDQATVQKIRDAIRDQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFI 486
Query: 525 GVQLDGSQHVEPLHNCIKEDTAKEGEQLVKQTAENVGEAVRELPDANQKPDDLWLNHSKV 584
GVQLDGS HVEPL N + E T + +LVK TA NV EAVRELPDAN +P+DLW HSK
Sbjct: 487 GVQLDGSDHVEPLQNRLSERTEMQSSKLVKATATNVDEAVRELPDANTRPEDLWAAHSKP 546
Query: 585 VHPKPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMK 644
V+P PH K++ +W+AI+KI +GE + L HF+PIKPLGSGDTGSVHLVEL+GTG+ +AMK
Sbjct: 547 VYPLPHNKESTSWKAIKKIQASGETVGLHHFKPIKPLGSGDTGSVHLVELKGTGELYAMK 606
Query: 645 AMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLL 704
AM+K +MLNRNK HRAC EREI+ +LDHPFLP LYASFQT THVCLITD+ PGGELF LL
Sbjct: 607 AMEKTMMLNRNKAHRACIEREIISLLDHPFLPTLYASFQTSTHVCLITDFCPGGELFALL 666
Query: 705 DQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCL 764
D+QP K+L ED+ RFYAAEV+I LEYLHC GI+YRDLKPEN+L++++GH+ L DFDLS +
Sbjct: 667 DRQPMKILTEDSARFYAAEVVIGLEYLHCLGIVYRDLKPENILLKKDGHIVLADFDLSFM 726
Query: 765 TSCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITG 824
T+C PQLIIPA K++R K+Q +PTF+AEP SNSFVGTEEYIAPEIITG
Sbjct: 727 TTCTPQLIIPAAPSKRRR--------SKSQPLPTFVAEPSTQSNSFVGTEEYIAPEIITG 778
Query: 825 SGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIY 884
+GHTSA+DWWALGILLYEMLYG TPFRGK RQKTFANILHKDL FP S PVS +QLI
Sbjct: 779 AGHTSAIDWWALGILLYEMLYGRTPFRGKNRQKTFANILHKDLTFPSSIPVSLVGRQLIN 838
Query: 885 WLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPELDAPI-LLENDEKKEAKDI 943
LL+RDP +RLGS GANEIK H FF+ +NW LIR M PP LDAP+ ++E D AKDI
Sbjct: 839 TLLNRDPSSRLGSKGGANEIKQHAFFRGINWPLIRGMSPPPLDAPLSIIEKD--PNAKDI 896
>At2g44830 protein kinase like protein
Length = 765
Score = 349 bits (895), Expect = 5e-96
Identities = 189/422 (44%), Positives = 247/422 (57%), Gaps = 42/422 (9%)
Query: 563 AVRELPDANQKPDDLWLNHSKVVHPKPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPLG 622
A R + + W N + ++ KPH+ ++ W AI I + + HF+ +K LG
Sbjct: 312 ASRASDSSGLSEESSWSNFTGSLN-KPHKGNDPWWNAILAIRTRDGILGMSHFKLLKRLG 370
Query: 623 SGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYASF 682
GD GSV+L EL GT +FA+K MDK + +R K++RA TER+IL +LDHPFLP LY F
Sbjct: 371 CGDIGSVYLAELSGTRCHFAVKVMDKASLEDRKKLNRAQTERDILQLLDHPFLPTLYTHF 430
Query: 683 QTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLK 742
+T CL+ +Y PGG+L L +QP K E A RFYAAEVL+ALEYLH G++YRDLK
Sbjct: 431 ETDRFSCLVMEYCPGGDLHTLRQRQPGKHFSEYAARFYAAEVLLALEYLHMLGVVYRDLK 490
Query: 743 PENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKR-------------------- 782
PENVL++ +GH+ L+DFDLS + P LI + D +R
Sbjct: 491 PENVLVRDDGHIMLSDFDLSLRCAVSPTLIKTFDSDPSRRGAFCVQPACMEPTSACIIQP 550
Query: 783 ----------KKKKKKGQQKTQ---------QIPTFMAEPMRASNSFVGTEEYIAPEIIT 823
K KK +KTQ +P +AEP S SFVGT EY+APEII
Sbjct: 551 SCFLPRSIFPNKNKKNKSRKTQADFFKSHSGSLPELVAEPNTRSMSFVGTHEYLAPEIIK 610
Query: 824 GSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLI 883
G GH SAVDWW GI ++E+LYG TPF+G + T N++ + LKFP+S S + LI
Sbjct: 611 GEGHGSAVDWWTFGIFVHELLYGKTPFKGSGNRATLFNVVGEQLKFPESPATSYAGRDLI 670
Query: 884 YWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPELDAPILLENDEKKEAKDI 943
LL +DPKNRLG+ GA EIK HPFF+ VNWALIRC PPE+ P +E + + I
Sbjct: 671 QALLVKDPKNRLGTKRGATEIKQHPFFEGVNWALIRCSTPPEV--PRQMETEPPPKYGPI 728
Query: 944 DP 945
DP
Sbjct: 729 DP 730
>At5g55910 serine/threonine-specific protein kinase ATPK64
(pir||S20918)
Length = 498
Score = 345 bits (885), Expect = 7e-95
Identities = 181/388 (46%), Positives = 234/388 (59%), Gaps = 42/388 (10%)
Query: 588 KPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMD 647
KPH+ ++ W AIQ + + L HFR +K LG GD G+VHL EL GT YFAMK MD
Sbjct: 82 KPHKANDVRWEAIQAVRTKHGGLGLNHFRLLKRLGCGDIGTVHLAELNGTRCYFAMKVMD 141
Query: 648 KGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQ 707
K + +R K+ RA TEREIL LDHPFLP LY+ F+T+ CL+ ++ PGG+L L +Q
Sbjct: 142 KTALASRKKLLRAQTEREILQCLDHPFLPTLYSHFETEKFSCLVMEFCPGGDLHTLRQRQ 201
Query: 708 PTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSC 767
P K E A +FY AEVL+A+EYLH GIIYRDLKPENVL++ +GHV L+DFDLS +
Sbjct: 202 PGKRFTEQAAKFYVAEVLLAMEYLHMLGIIYRDLKPENVLVRDDGHVMLSDFDLSLRCTV 261
Query: 768 KPQLIIPAN-----------------------------------------EDKKKRKKKK 786
++ AN + KK +K K
Sbjct: 262 SLSIVRSANVGSEGLSKNSVSCSQQPACIQQPSCISMAPTSCFGPRFFSSKSKKDKKPKT 321
Query: 787 KKGQQKTQQIPTFMAEPMRA-SNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLY 845
+ G + +P +AEP A S SFVGT EY+APEII G GH SAVDWW GI LYE+L+
Sbjct: 322 ENGNHQVTPLPELVAEPTGARSMSFVGTHEYLAPEIIKGEGHGSAVDWWTFGIFLYELLF 381
Query: 846 GYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIK 905
G TPF+G + T N++ + L+FP+S VS A+ LI LL ++P++RL GA EIK
Sbjct: 382 GKTPFKGSGNRATLFNVVGQPLRFPESPVVSFAARDLIRSLLVKEPQHRLAYKRGATEIK 441
Query: 906 SHPFFKNVNWALIRCMKPPELDAPILLE 933
HPFF+ VNWAL+RC PPE+ P+ LE
Sbjct: 442 QHPFFEGVNWALVRCASPPEIPKPVDLE 469
>At4g26610 putative protein kinase
Length = 506
Score = 345 bits (885), Expect = 7e-95
Identities = 187/429 (43%), Positives = 252/429 (58%), Gaps = 38/429 (8%)
Query: 544 DTAKEGEQLVKQTAENVGEAVRELPDANQKPDDLWLNHSKVVHPKPHRKDNDAWRAIQKI 603
D+ GE + ++ G++ P + D S + KPH+ ++ W AIQ +
Sbjct: 52 DSKTGGEVKFNEKSDQSGKSNTCRPSTSSDISDESTCSSFSGNNKPHKANDVRWEAIQAV 111
Query: 604 IENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTE 663
+ L HFR +K LG GD G+VHL EL GT +FAMK MDKG + +R K+ RA TE
Sbjct: 112 RTKHGVLGLNHFRLLKRLGCGDIGTVHLAELHGTRCFFAMKVMDKGALASRKKLLRAQTE 171
Query: 664 REILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAE 723
REIL LDHPFLP LY+ F+T+ CL+ ++ PGG+L L +QP K E A +FY AE
Sbjct: 172 REILQCLDHPFLPTLYSHFETEKFSCLVMEFCPGGDLHTLRQRQPGKRFSEQAAKFYVAE 231
Query: 724 VLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLI---IPANEDK- 779
VL+A+EYLH GIIYRDLKPENVL++ +GHV L+DFDLS + P ++ + A+E +
Sbjct: 232 VLLAMEYLHMLGIIYRDLKPENVLVRDDGHVMLSDFDLSLRCTVSPTVVRSTVLASEGQK 291
Query: 780 ---------------------------------KKRKKKKKKGQQKTQQIPTFMAEPMRA 806
KK KK K + + +P +AEP A
Sbjct: 292 NSGYCAQPACIQQPSCISAPTTCFSPRYFSSKSKKDKKMKNETGNQVSPLPELVAEPTSA 351
Query: 807 -SNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHK 865
S SFVGT EY+APEII G GH SAVDWW GI LYE+L+G TPF+G + T N++ +
Sbjct: 352 RSMSFVGTHEYLAPEIIKGEGHGSAVDWWTFGIFLYELLFGKTPFKGSGNRATLFNVVGQ 411
Query: 866 DLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPE 925
L+FP+S VS A+ LI LL ++P++RL GA E+K HPFF+ VNWAL+RC PPE
Sbjct: 412 PLRFPESPVVSFAARDLIRSLLVKEPQHRLAYKRGATEMKQHPFFEGVNWALVRCASPPE 471
Query: 926 LDAPILLEN 934
+ P+ E+
Sbjct: 472 IPKPVDYES 480
>At5g40030 protein kinase -like protein
Length = 499
Score = 339 bits (870), Expect = 4e-93
Identities = 187/390 (47%), Positives = 240/390 (60%), Gaps = 42/390 (10%)
Query: 580 NHSKVVHP-KPHRKDNDAWRAIQKI-IENGEQISLKHFRPIKPLGSGDTGSVHLVELEGT 637
N +V P KPH+ ++ W AIQ + E + L HFR +K LG GD GSV+L EL
Sbjct: 77 NFKRVFAPSKPHKGNDLRWDAIQNVKCSKNEDLGLGHFRLLKKLGCGDIGSVYLAELREM 136
Query: 638 GQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPG 697
G +FAMK MDKG+++ R K+ RA TEREIL +LDHPFLP LY+ F+T+ CL+ ++ G
Sbjct: 137 GCFFAMKVMDKGMLIGRKKLVRAQTEREILGLLDHPFLPTLYSHFETEKFSCLLMEFCSG 196
Query: 698 GELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLT 757
G+L +L +QP K E A RFYA+EVL+ALEYLH G++YRDLKPENV+++ +GH+ L+
Sbjct: 197 GDLHILRQKQPGKHFSELAARFYASEVLLALEYLHMMGVVYRDLKPENVMVREDGHIMLS 256
Query: 758 DFDLS-----------------------CL------TSCK-------PQLIIPANEDKKK 781
DFDLS C+ SCK P P + K
Sbjct: 257 DFDLSLQSFVSPTLIQSTSQPSCHIASYCIQPPCIDPSCKLPVACIQPSCFKPRFLNNKP 316
Query: 782 RKKKKKKGQQKTQQIPTFMAEPMRA-SNSFVGTEEYIAPEIITGSGHTSAVDWWALGILL 840
RK K +K + +P +AEP A S SFVGT EY+APEII G GH S+VDWW GI L
Sbjct: 317 RKAKTEKA--GSDSLPMLIAEPTAARSMSFVGTHEYLAPEIIRGDGHGSSVDWWTFGIFL 374
Query: 841 YEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEG 900
YE+L G TPF+G ++T N++ + LKFP+ +S AK LI LL +DPK RLG +G
Sbjct: 375 YELLTGKTPFKGNGNRETLFNVVGQPLKFPEGS-ISFAAKDLIRGLLTKDPKKRLGFKKG 433
Query: 901 ANEIKSHPFFKNVNWALIRCMKPPELDAPI 930
A EIK HPFF NVNWALIR PPE+ PI
Sbjct: 434 ATEIKQHPFFNNVNWALIRSTTPPEIPKPI 463
>At1g16440 putative protein kinase
Length = 431
Score = 337 bits (865), Expect = 1e-92
Identities = 179/386 (46%), Positives = 233/386 (59%), Gaps = 28/386 (7%)
Query: 588 KPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQ--YFAMKA 645
KPH + W A+ + G ++ + FR +K LG GD GSV+LVEL+G YFAMK
Sbjct: 18 KPHTGGDIRWDAVNSLKSRGIKLGISDFRVLKRLGYGDIGSVYLVELKGANPTTYFAMKV 77
Query: 646 MDKGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLD 705
MDK +++RNK+ RA TEREIL LDHPFLP LY+ F+T CL+ ++ GG L+ L
Sbjct: 78 MDKASLVSRNKLLRAQTEREILSQLDHPFLPTLYSHFETDKFYCLVMEFCSGGNLYSLRQ 137
Query: 706 QQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLT 765
+QP K EDA RF+A+EVL+ALEYLH GI+YRDLKPENVL++ +GH+ L+DFDLS
Sbjct: 138 KQPNKCFTEDAARFFASEVLLALEYLHMLGIVYRDLKPENVLVRDDGHIMLSDFDLSLRC 197
Query: 766 SCKPQLIIPAN--------EDK-----------------KKRKKKKKKGQQKTQQIPTFM 800
S P L+ N +D + KK +K +P M
Sbjct: 198 SVNPTLVKSFNGGGTTGIIDDNAAVQGCYQPSAFFPRMLQSSKKNRKSKSDFDGSLPELM 257
Query: 801 AEPMRA-SNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTF 859
AEP S SFVGT EY+APEII GH SAVDWW GI +YE+L+G TPF+G+ + T
Sbjct: 258 AEPTNVKSMSFVGTHEYLAPEIIKNEGHGSAVDWWTFGIFIYELLHGATPFKGQGNKATL 317
Query: 860 ANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIR 919
N++ + L+FP+ VS AK LI LL ++P+NR+ GA EIK HPFF+ VNWALIR
Sbjct: 318 YNVIGQPLRFPEYSQVSSTAKDLIKGLLVKEPQNRIAYKRGATEIKQHPFFEGVNWALIR 377
Query: 920 CMKPPELDAPILLENDEKKEAKDIDP 945
PP L P+ KKE + + P
Sbjct: 378 GETPPHLPEPVDFSCYVKKEKESLPP 403
>At1g79250 hypothetical protein
Length = 517
Score = 337 bits (863), Expect = 2e-92
Identities = 177/382 (46%), Positives = 226/382 (58%), Gaps = 39/382 (10%)
Query: 588 KPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMD 647
KPH + W A+ + G Q+ + FR +K LG GD GSV+LVEL GT YFAMK MD
Sbjct: 81 KPHTGGDIRWDAVNTLTSKGVQLGISDFRLLKRLGYGDIGSVYLVELRGTITYFAMKVMD 140
Query: 648 KGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQ 707
K + +RNK+ RA TEREIL LDHPFLP LY+ F+T CL+ ++ GG L+ L +Q
Sbjct: 141 KASLASRNKLLRAQTEREILSQLDHPFLPTLYSHFETDKFYCLVMEFCGGGNLYSLRQKQ 200
Query: 708 PTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSC 767
P K EDA RF+A+EVL+ALEYLH GI+YRDLKPENVL++ +GH+ L+DFDLS S
Sbjct: 201 PNKCFTEDAARFFASEVLLALEYLHMLGIVYRDLKPENVLVRDDGHIMLSDFDLSLRCSV 260
Query: 768 KPQLIIPAN--------------------------------------EDKKKRKKKKKKG 789
P L+ ++ KK RK K G
Sbjct: 261 SPTLVKSSSVHAAGGGSGSSRPVGLIDEDAAVQGCIQPSTFFPRILQSSKKNRKAKSDFG 320
Query: 790 QQKTQQIPTFMAEPMRA-SNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYT 848
+P MAEP S SFVGT EY+APEII G GH SAVDWW GI +YE+LYG T
Sbjct: 321 LFVNGSMPELMAEPTNVKSMSFVGTHEYLAPEIIRGEGHGSAVDWWTFGIFIYELLYGAT 380
Query: 849 PFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHP 908
PF+G+ + T N++ + L+FP+ VS A+ LI LL ++P+ R+ GA EIK HP
Sbjct: 381 PFKGQGNRATLHNVIGQALRFPEVPHVSSAARDLIKGLLVKEPQKRIAYKRGATEIKQHP 440
Query: 909 FFKNVNWALIRCMKPPELDAPI 930
FF+ VNWALIR PP + P+
Sbjct: 441 FFEGVNWALIRSATPPHVPEPV 462
>At2g34650 protein kinase PINOID
Length = 438
Score = 298 bits (762), Expect = 1e-80
Identities = 164/368 (44%), Positives = 227/368 (61%), Gaps = 26/368 (7%)
Query: 588 KPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQ-----YFA 642
KPHR + A+ I++ + G ++ + FR ++ +G+GD G+V+L L G + YFA
Sbjct: 50 KPHRSSDFAYAEIRRRKKQG--LTFRDFRLMRRIGAGDIGTVYLCRLAGDEEESRSSYFA 107
Query: 643 MKAMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFL 702
MK +DK + + K+HRA E+ IL MLDHPFLP LYA F+ C++ +Y GG+L
Sbjct: 108 MKVVDKEALALKKKMHRAEMEKTILKMLDHPFLPTLYAEFEASHFSCIVMEYCSGGDLHS 167
Query: 703 LLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLS 762
L +QP + + RFYAAEVL+ALEYLH GIIYRDLKPEN+L++ +GH+ L+DFDLS
Sbjct: 168 LRHRQPHRRFSLSSARFYAAEVLVALEYLHMLGIIYRDLKPENILVRSDGHIMLSDFDLS 227
Query: 763 CL---------TSCKP---QLIIPANEDKKKRKKKKKKGQQKTQQI-PT--FMAEPMRA- 806
+S P QL P + R ++ +K Q + PT F+AEP+ A
Sbjct: 228 LCSDSIAAVESSSSSPENQQLRSPRRFTRLARLFQRVLRSKKVQTLEPTRLFVAEPVTAR 287
Query: 807 SNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKD 866
S SFVGT EY+APE+ +G H +AVDWWA G+ LYEM+YG TPF T NI+ +
Sbjct: 288 SGSFVGTHEYVAPEVASGGSHGNAVDWWAFGVFLYEMIYGKTPFVAPTNDVILRNIVKRQ 347
Query: 867 LKFPKSKPVSP---QAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKP 923
L FP P + A+ LI LL++DP RLGS GA E+K HPFFK +N+ALIR + P
Sbjct: 348 LSFPTDSPATMFELHARNLISGLLNKDPTKRLGSRRGAAEVKVHPFFKGLNFALIRTLTP 407
Query: 924 PELDAPIL 931
PE+ + ++
Sbjct: 408 PEIPSSVV 415
>At3g14370 putative protein kinase
Length = 480
Score = 291 bits (746), Expect = 9e-79
Identities = 155/375 (41%), Positives = 223/375 (59%), Gaps = 27/375 (7%)
Query: 590 HRKDNDAWRAIQ--KIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMD 647
HR+ + W AI+ K++ + I L+H + I+ LG+G+ G V L L + FA+K +D
Sbjct: 61 HRRHDPHWSAIKSAKLLSSDGNIHLRHLKLIRHLGTGNLGRVFLCNLRDSSARFALKVID 120
Query: 648 KGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQ 707
+ + K+ + TE EIL +LDHPFLP LYA + CL+ DY P G+L LL +Q
Sbjct: 121 RNCLTTEKKLSQVETEAEILSLLDHPFLPTLYARIDESHYTCLLIDYAPNGDLHSLLRKQ 180
Query: 708 PTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSC 767
P L VRF+AAEVL+ALEYLH GI+YRDLKPENVL++ +GHV L+DFDL C
Sbjct: 181 PGNRLPIQPVRFFAAEVLVALEYLHAMGIVYRDLKPENVLLREDGHVMLSDFDL-----C 235
Query: 768 KPQLIIPANEDKKKRKKKK----------------KKGQQKTQQIPTFMAEPMRA-SNSF 810
++P + ++ R+ +K ++ + + F AEP+ A S S
Sbjct: 236 FKSDVVPTFKSRRYRRSSSSPSLRRRRSGCFSVAAEKKYEREEIVSEFAAEPVTAFSRSC 295
Query: 811 VGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILH--KDLK 868
VGT EY+APE+++G+GH S VDWWA GI LYE+LYG TPF+G+++++T NI+ K
Sbjct: 296 VGTHEYLAPELVSGNGHGSGVDWWAFGIFLYELLYGTTPFKGESKEQTLRNIVSTTKTAS 355
Query: 869 FPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPELDA 928
F + +A+ LI LL +DP+ RLG GA +IK HPFF + W LIR KPPE
Sbjct: 356 FHMDGDLD-EARDLIEKLLVKDPRKRLGCARGAQDIKRHPFFDGIKWPLIRHYKPPEEVR 414
Query: 929 PILLENDEKKEAKDI 943
++++ + A +
Sbjct: 415 GLVIKKSTRPHASHV 429
>At1g53700 auxin-induced protein kinase like protein
Length = 476
Score = 285 bits (729), Expect = 9e-77
Identities = 157/359 (43%), Positives = 220/359 (60%), Gaps = 30/359 (8%)
Query: 590 HRKDNDAWRAIQKI--IENGEQISLKHFRPIKPLGSGDTGSVHLVELEG----TGQYFAM 643
HR+ + W +I+ + + ++ L+HF+ ++ LG+G+ G V L L TG FA+
Sbjct: 66 HRRYDPHWTSIRAATTLSSDGRLHLRHFKLVRHLGTGNLGRVFLCHLRDCPNPTG--FAL 123
Query: 644 KAMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLL 703
K +D+ V L K+ TE EIL +LDHPFLP LYA + CL+ DY P G+L L
Sbjct: 124 KVIDRDV-LTAKKISHVETEAEILSLLDHPFLPTLYARIDASHYTCLLIDYCPNGDLHSL 182
Query: 704 LDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSC 763
L +QP L VRF+AAEVL+ALEYLH GI+YRDLKPEN+LI+ +GH+ L+DFDL
Sbjct: 183 LRKQPNNRLPISPVRFFAAEVLVALEYLHALGIVYRDLKPENILIREDGHIMLSDFDL-- 240
Query: 764 LTSCKPQLIIPANEDKKKRK-----KKKKKG---------QQKTQQIPTFMAEPMRA-SN 808
C ++P ++ R+ +K ++G ++ + + F AEP+ A S
Sbjct: 241 ---CFKADVVPTFRSRRFRRTSSSPRKTRRGGGCFSTEVEYEREEIVAEFAAEPVTAFSK 297
Query: 809 SFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANIL-HKDL 867
S VGT EY+APE++ G+GH S VDWWA GI LYEMLYG TPF+G T+++T NI+ + D+
Sbjct: 298 SCVGTHEYLAPELVAGNGHGSGVDWWAFGIFLYEMLYGTTPFKGGTKEQTLRNIVSNDDV 357
Query: 868 KFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPEL 926
F + +AK LI LL +DP+ RLG GA +IK H FF+ + W LIR KPPE+
Sbjct: 358 AFTLEEEGMVEAKDLIEKLLVKDPRKRLGCARGAQDIKRHEFFEGIKWPLIRNYKPPEI 416
>At3g20830 unknown protein
Length = 408
Score = 229 bits (585), Expect = 4e-60
Identities = 146/350 (41%), Positives = 197/350 (55%), Gaps = 40/350 (11%)
Query: 608 EQISLKHFRPIKPLGSGDTGSVHL----VELEGTGQYFAMKAMDKGVMLNRNKVHRACTE 663
E + L + +K LG G TG+V L V + FA+K + K + + + RA E
Sbjct: 14 EILDLDSIKALKILGKGATGTVFLAHDVVSTSSSSSPFAVKLVPKS---SASSLRRARWE 70
Query: 664 REIL-----DMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVR 718
E+L D +PFLP L ASF++ + Y GG+L +LL +Q V +R
Sbjct: 71 IEVLRRLSVDSNQNPFLPRLLASFESPEYFAWAVPYCSGGDLNVLLHRQNDGVFSSSVIR 130
Query: 719 FYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDL--SCLTSCKPQLIIPAN 776
FY AE++ ALE+LH GI YRDLKPEN+LIQ++GHV+LTDFDL S +P P
Sbjct: 131 FYVAEIVCALEHLHTMGIAYRDLKPENILIQQSGHVTLTDFDLSRSLKKPLRPHFYQPDP 190
Query: 777 E---DKKKRK----------KKKKKGQQKTQQI--------PTFMAEPMRASNSFVGTEE 815
E D+KK + +K K G +KT+ T + R SNSFVGT+E
Sbjct: 191 ELIIDRKKSRSFSRLISPTAEKNKTGLKKTRSARVNPINRRKTSFSSGER-SNSFVGTDE 249
Query: 816 YIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPV 875
Y++PE+I G GH AVDWWALG+L YEM+YG TPF+GK++++TF N+L K+ +F KP
Sbjct: 250 YVSPEVIRGDGHDFAVDWWALGVLTYEMMYGETPFKGKSKKETFRNVLMKEPEF-AGKP- 307
Query: 876 SPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALI-RCMKPP 924
LI LL +DP RLG GA EIK FF V W L+ ++PP
Sbjct: 308 -NDLTDLIRRLLVKDPNRRLGCHRGAAEIKELAFFAGVRWDLLTEVLRPP 356
>At1g51170 putative serine/threonine protein kinase
Length = 404
Score = 226 bits (577), Expect = 4e-59
Identities = 140/345 (40%), Positives = 190/345 (54%), Gaps = 36/345 (10%)
Query: 610 ISLKHFRPIKPLGSGDTGSVHLVELE----GTGQYFAMKAMDKGVMLNRNKVHRACTERE 665
++L + +K LG G TG+V LV FA+K +DK + + + RA E +
Sbjct: 17 LNLDRLKVLKLLGKGATGTVFLVHDSVSDSSVSSPFALKLVDKS---SASSLRRARWEIQ 73
Query: 666 ILDMLD-----HPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFY 720
IL L +PFLP L AS ++ + Y GG+L +L +Q V ++FY
Sbjct: 74 ILRRLSDDTNPNPFLPKLLASSESSEFIAWALPYCSGGDLNVLRQRQNDGVFSSSVIKFY 133
Query: 721 AAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSC------------LTSCK 768
AE++ AL++LH GI YRDLKPEN+L+Q +GHV+LTDFDLSC L+ +
Sbjct: 134 LAEIVCALDHLHTMGIAYRDLKPENILLQESGHVTLTDFDLSCSLNKPTRPEFYHLSDPE 193
Query: 769 PQLIIPANEDKKK------RKKKKKKGQQKTQQIP--TFMAEPMRASNSFVGTEEYIAPE 820
P +N K R+KKKK + I SNSFVGT+EYI+PE
Sbjct: 194 PDPNPESNLSHNKKSLRIFRQKKKKTKSARVNPITRRRLSFSGGERSNSFVGTDEYISPE 253
Query: 821 IITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAK 880
+I G GH AVDWWALG+L YEM+YG TPF+G+ +++TF N+L K+ +F KP
Sbjct: 254 VIRGDGHDFAVDWWALGVLTYEMMYGETPFKGRNKKETFRNVLVKEPEF-AGKP--SDLT 310
Query: 881 QLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALI-RCMKPP 924
LI LL +DP R G GA EIK H FFK V W L+ ++PP
Sbjct: 311 DLIRRLLVKDPTKRFGFWRGAAEIKEHAFFKGVRWELLTEVLRPP 355
>At4g13000 unknown protein
Length = 372
Score = 220 bits (560), Expect = 3e-57
Identities = 131/337 (38%), Positives = 188/337 (54%), Gaps = 26/337 (7%)
Query: 610 ISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNR---NKVHRACTEREI 666
++ H LG G G V LV+ + ++ A+K + + + ++ ++ R E+ +
Sbjct: 15 LNFDHLEIFSALGRGSKGVVFLVKADN--KWLALKVILRESIESKKAKDEYKRISFEQGV 72
Query: 667 LDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLI 726
L DHP P L+ T + DY PG +L L +Q ++ ++ +RFYAAE++I
Sbjct: 73 LSRFDHPLFPRLHGVISTDKVIGYAIDYCPGRDLNSLRKKQSEEMFSDEIIRFYAAELVI 132
Query: 727 ALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSC-LTSCKPQLIIPANEDKKKRKKK 785
ALEYLH QGI+YRDLKP+NV+IQ NGH+ L DFDLS L PQ ++ KK
Sbjct: 133 ALEYLHNQGIVYRDLKPDNVMIQENGHLMLVDFDLSTNLPPRTPQSSFSSSPRLSTATKK 192
Query: 786 KK----------KGQQKTQQIPTFMAEPM--RASNSFVGTEEYIAPEIITGSGHTSAVDW 833
++ G + SNSFVGTEEY+APE+ITGSGH AVDW
Sbjct: 193 ERSIFAFSGLCNSGISPDDSVSRSSESEFSGEKSNSFVGTEEYVAPEVITGSGHDFAVDW 252
Query: 834 WALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKN 893
W+LG++LYEMLYG TPFRG R++TF IL + P + + L+ LL +DP
Sbjct: 253 WSLGVVLYEMLYGATPFRGSNRKETFLKILTEP---PSLVGETTSLRDLVRKLLEKDPSR 309
Query: 894 RLGSLEGANEIKSHPFFKNVNWALI-RCMKPPELDAP 929
R+ ++EG IK H FFK ++W L+ + +PP + AP
Sbjct: 310 RI-NVEG---IKGHDFFKGLDWDLVLKVSRPPYIPAP 342
>At5g62310 IRE (root hair elongation)
Length = 1168
Score = 218 bits (555), Expect = 1e-56
Identities = 125/347 (36%), Positives = 194/347 (55%), Gaps = 15/347 (4%)
Query: 575 DDLWLNHSKVVHPKPHRKDNDAWRAIQKIIENG---EQISLKHFRPIKPLGSGDTGSVHL 631
DD ++ S + + D D R+++ N ++ S++ F IKP+ G G V L
Sbjct: 711 DDEKVDSSNAMPDEESSADEDTVRSLRASPLNPRAKDRTSIEDFEIIKPISRGAFGRVFL 770
Query: 632 VELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLI 691
+ TG FA+K + K M+ +N V ER IL + +PF+ + SF + ++ L+
Sbjct: 771 AKKRATGDLFAIKVLKKADMIRKNAVESILAERNILISVRNPFVVRFFYSFTCRENLYLV 830
Query: 692 TDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRN 751
+Y GG+LF LL + L ED R Y AEV++ALEYLH II+RDLKP+N+LI ++
Sbjct: 831 MEYLNGGDLFSLL--RNLGCLDEDMARIYIAEVVLALEYLHSVNIIHRDLKPDNLLINQD 888
Query: 752 GHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKKG--QQKTQQIPTFMAEPMRASNS 809
GH+ LTDF LS + +I + +D G + + + R ++
Sbjct: 889 GHIKLTDFGLSKVG------LINSTDDLSGESSLGNSGFFAEDGSKAQHSQGKDSRKKHA 942
Query: 810 FVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKF 869
VGT +Y+APEI+ G GH DWW++G++L+E+L G PF +T Q+ F NI+++D+ +
Sbjct: 943 VVGTPDYLAPEILLGMGHGKTADWWSVGVILFEVLVGIPPFNAETPQQIFENIINRDIPW 1002
Query: 870 PK-SKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNW 915
P + +S +A LI LL +P RLG+ GA E+K H FFK++NW
Sbjct: 1003 PNVPEEISYEAHDLINKLLTENPVQRLGA-TGAGEVKQHHFFKDINW 1048
>At3g25250 protein kinase, putative
Length = 421
Score = 216 bits (551), Expect = 4e-56
Identities = 136/356 (38%), Positives = 188/356 (52%), Gaps = 42/356 (11%)
Query: 621 LGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNK--------VHRACTEREILDMLDH 672
LG G G V LV + + A+K + K + + K R E+ +L DH
Sbjct: 23 LGRGAKGVVFLVR-DDDAKLLALKVILKEAIEKKKKGRESEDDEYKRVSFEQGVLSRFDH 81
Query: 673 PFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLH 732
P P+L+ T + DY PG L L Q + ++ +RFYAAE+++AL+YLH
Sbjct: 82 PLFPSLHGVLATDKVIGYAIDYCPGQNLNSLRKMQSESMFSDEIIRFYAAELVLALDYLH 141
Query: 733 CQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKK-----KRKKKK- 786
QGI+YRDLKP+NV+IQ NGH+ L DFDLS T+ P+ P+ K KRKK+
Sbjct: 142 NQGIVYRDLKPDNVMIQENGHLMLIDFDLS--TNLAPRTPQPSPSLSKPSPTMKRKKRLF 199
Query: 787 ------KKGQQKTQQIPTFMAEPM-------RASNSFVGTEEYIAPEIITGSGHTSAVDW 833
G + I + + SNSFVGTEEY+APE+I+G GH AVDW
Sbjct: 200 RFTSFCNSGISPQESISVHSSSTLAVSDSSGEKSNSFVGTEEYVAPEVISGDGHDFAVDW 259
Query: 834 WALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKN 893
W+LG++LYEMLYG TPFRG R++TF IL K P + + LI LL +DP
Sbjct: 260 WSLGVVLYEMLYGATPFRGSNRKETFYRILSKP---PNLTGETTSLRDLIRRLLEKDPSR 316
Query: 894 RLGSLEGANEIKSHPFFKNVNW-ALIRCMKPPELDAPILLENDEKKEAKDIDPGLD 948
R+ EIK H FF+ V+W +I +PP + AP +D + D++ +D
Sbjct: 317 RI----NVEEIKGHDFFRGVDWEKVILVSRPPYIPAP----DDGGDKGTDVNTKMD 364
>At3g17850 protein kinase like protein
Length = 1296
Score = 214 bits (546), Expect = 1e-55
Identities = 118/311 (37%), Positives = 183/311 (57%), Gaps = 12/311 (3%)
Query: 608 EQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREIL 667
++ S+ F IKP+ G G V L + TG FA+K + K M+ +N V ER+IL
Sbjct: 875 DRTSIDDFEIIKPISRGAFGRVFLAKKRTTGDLFAIKVLKKADMIRKNAVESILAERDIL 934
Query: 668 DMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIA 727
+ +PF+ + SF + ++ L+ +Y GG+L+ LL + L+ED VR Y AEV++A
Sbjct: 935 INVRNPFVVRFFYSFTCRDNLYLVMEYLNGGDLYSLL--RNLGCLEEDIVRVYIAEVVLA 992
Query: 728 LEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLS--CLTSCKPQLIIPANEDKKKRKKK 785
LEYLH +G+++RDLKP+N+LI +GH+ LTDF LS L + L PA ++
Sbjct: 993 LEYLHSEGVVHRDLKPDNLLIAHDGHIKLTDFGLSKVGLINSTDDLAGPAVSGTSLLDEE 1052
Query: 786 KKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLY 845
+ + +Q+ R S VGT +Y+APEI+ G+GH + DWW++GI+L+E++
Sbjct: 1053 ESRLAASEEQL------ERRKKRSAVGTPDYLAPEILLGTGHGATADWWSVGIILFELIV 1106
Query: 846 GYTPFRGKTRQKTFANILHKDLKFPK-SKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEI 904
G PF + Q+ F NIL++ + +P + +S +A +I L DP RLG+ GA E+
Sbjct: 1107 GIPPFNAEHPQQIFDNILNRKIPWPHVPEEMSAEAHDIIDRFLTEDPHQRLGA-RGAAEV 1165
Query: 905 KSHPFFKNVNW 915
K H FFK++NW
Sbjct: 1166 KQHIFFKDINW 1176
>At1g45160 similar to IRE homolog 1 dbj|BAA89784.1
Length = 1067
Score = 213 bits (542), Expect = 4e-55
Identities = 122/323 (37%), Positives = 181/323 (55%), Gaps = 12/323 (3%)
Query: 608 EQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREIL 667
++IS+ F IKP+ G G V L TG +FA+K + K M+ +N + R ER IL
Sbjct: 663 DRISIDDFEIIKPISRGAFGKVFLARKRTTGDFFAIKVLKKLDMIRKNDIERILQERNIL 722
Query: 668 DMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIA 727
+ +PFL + SF + ++ L+ +Y GG+L+ LL Q L E+ R Y AE+++A
Sbjct: 723 ITVRYPFLVRFFYSFTCRDNLYLVMEYLNGGDLYSLL--QKVGCLDEEIARIYIAELVLA 780
Query: 728 LEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKK 787
LEYLH I++RDLKP+N+LI NGH+ LTDF LS + + + +E +
Sbjct: 781 LEYLHSLKIVHRDLKPDNLLIAYNGHIKLTDFGLSKIGLINNTIDLSGHESDVSPRTNSH 840
Query: 788 KGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGY 847
Q+ E R +S VGT +Y+APEI+ G+ H A DWW+ GI+L+E+L G
Sbjct: 841 HFQKN--------QEEERIRHSAVGTPDYLAPEILLGTEHGYAADWWSAGIVLFELLTGI 892
Query: 848 TPFRGKTRQKTFANILHKDLKFPK-SKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKS 906
PF +K F NIL+ + +P +S +A+ LI LL +P+ RLG+ GA E+KS
Sbjct: 893 PPFTASRPEKIFDNILNGKMPWPDVPGEMSYEAQDLINRLLVHEPEKRLGA-NGAAEVKS 951
Query: 907 HPFFKNVNWALIRCMKPPELDAP 929
HPFF+ V+W + K + P
Sbjct: 952 HPFFQGVDWENLALQKAAFVPQP 974
>At1g48490 IRE - like protein kinase
Length = 1235
Score = 203 bits (517), Expect = 3e-52
Identities = 118/311 (37%), Positives = 178/311 (56%), Gaps = 18/311 (5%)
Query: 608 EQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREIL 667
++IS+ F +K + G G V L TG FA+K + K M+ +N V ER+IL
Sbjct: 821 DRISIDDFEVMKSISRGAFGHVILARKNTTGDLFAIKVLRKADMIRKNAVESILAERDIL 880
Query: 668 DMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIA 727
+PF+ + SF ++ L+ +Y GG+ + +L + L E R Y AEV++A
Sbjct: 881 INARNPFVVRFFYSFTCSENLYLVMEYLNGGDFYSML--RKIGCLDEANARVYIAEVVLA 938
Query: 728 LEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSC--LTSCKPQLIIPANEDKKKRKKK 785
LEYLH +G+++RDLKP+N+LI +GHV LTDF LS L + L P + ++
Sbjct: 939 LEYLHSEGVVHRDLKPDNLLIAHDGHVKLTDFGLSKVGLINNTDDLSGPVSSATSLLVEE 998
Query: 786 KKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLY 845
K K +PT + S VGT +Y+APEI+ G+GH + DWW++GI+LYE L
Sbjct: 999 KPK-------LPT-----LDHKRSAVGTPDYLAPEILLGTGHGATADWWSVGIILYEFLV 1046
Query: 846 GYTPFRGKTRQKTFANILHKDLKFPK-SKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEI 904
G PF Q+ F NIL++++++P + +S +A+ LI LL DP RLG+ GA E+
Sbjct: 1047 GIPPFNADHPQQIFDNILNRNIQWPPVPEDMSHEARDLIDRLLTEDPHQRLGA-RGAAEV 1105
Query: 905 KSHPFFKNVNW 915
K H FFK+++W
Sbjct: 1106 KQHSFFKDIDW 1116
>At4g14350 protein kinase
Length = 551
Score = 200 bits (509), Expect = 3e-51
Identities = 119/339 (35%), Positives = 184/339 (54%), Gaps = 15/339 (4%)
Query: 615 FRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPF 674
F P+ +G G G V + +GTG +AMK + K ML R +V ER +L +D
Sbjct: 119 FEPLTMIGKGAFGEVRICREKGTGNVYAMKKLKKSEMLRRGQVEHVKAERNLLAEVDSNC 178
Query: 675 LPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQ 734
+ LY SFQ + ++ LI +Y PGG++ LL ++ T L ED RFY E ++A+E +H
Sbjct: 179 IVKLYCSFQDEEYLYLIMEYLPGGDMMTLLMRKDT--LTEDEARFYIGETVLAIESIHKH 236
Query: 735 GIIYRDLKPENVLIQRNGHVSLTDF----DLSCLTSCKPQLIIPANEDKKKRKKKKKKGQ 790
I+RD+KP+N+L+ ++GH+ L+DF L C + + N + +
Sbjct: 237 NYIHRDIKPDNLLLDKDGHMKLSDFGLCKPLDCSNLQEKDFTVARNVSGALQSDGRPVAT 296
Query: 791 QKTQ--QIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYT 848
++TQ Q+ + + S VGT +YIAPE++ G+ DWW+LG ++YEML G+
Sbjct: 297 RRTQQEQLLNWQRNRRMLAYSTVGTPDYIAPEVLLKKGYGMECDWWSLGAIMYEMLVGFP 356
Query: 849 PFRGKTRQKTFANILH--KDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKS 906
PF T I++ LKFP +SP+AK LI LL + + RLG+ +GA+EIK
Sbjct: 357 PFYSDDPMTTCRKIVNWRNYLKFPDEVRLSPEAKDLICRLL-CNVEQRLGT-KGADEIKG 414
Query: 907 HPFFKNVNWALIRCMKP---PELDAPILLENDEKKEAKD 942
HP+F+ W + MK P+++ + +N EK E D
Sbjct: 415 HPWFRGTEWGKLYQMKAAFIPQVNDELDTQNFEKFEETD 453
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.135 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,763,959
Number of Sequences: 26719
Number of extensions: 1060464
Number of successful extensions: 6465
Number of sequences better than 10.0: 919
Number of HSP's better than 10.0 without gapping: 525
Number of HSP's successfully gapped in prelim test: 395
Number of HSP's that attempted gapping in prelim test: 4520
Number of HSP's gapped (non-prelim): 1824
length of query: 954
length of database: 11,318,596
effective HSP length: 109
effective length of query: 845
effective length of database: 8,406,225
effective search space: 7103260125
effective search space used: 7103260125
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC135230.11


