

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
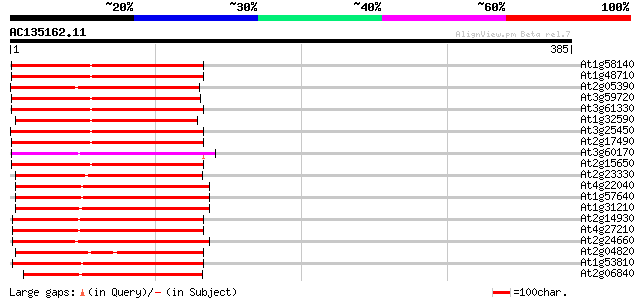
Query= AC135162.11 + phase: 0 /pseudo
(385 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At1g58140 hypothetical protein 126 2e-29
At1g48710 hypothetical protein 126 2e-29
At2g05390 putative retroelement pol polyprotein 124 9e-29
At3g59720 copia-type reverse transcriptase-like protein 124 1e-28
At3g61330 copia-type polyprotein 123 2e-28
At1g32590 hypothetical protein, 5' partial 123 2e-28
At3g25450 hypothetical protein 119 3e-27
At2g17490 putative retroelement pol polyprotein 115 4e-26
At3g60170 putative protein 114 7e-26
At2g15650 putative retroelement pol polyprotein 110 2e-24
At2g23330 putative retroelement pol polyprotein 106 2e-23
At4g22040 LTR retrotransposon like protein 100 2e-21
At1g57640 100 2e-21
At1g31210 putative reverse transcriptase 100 2e-21
At2g14930 pseudogene 100 2e-21
At4g27210 putative protein 99 4e-21
At2g24660 putative retroelement pol polyprotein 99 4e-21
At2g04820 putative retroelement pol polyprotein 99 4e-21
At1g53810 99 4e-21
At2g06840 putative retroelement pol polyprotein 98 9e-21
>At1g58140 hypothetical protein
Length = 1320
Score = 126 bits (317), Expect = 2e-29
Identities = 61/132 (46%), Positives = 88/132 (66%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + RL KALYGLKQ+ RAWN RI + +++F+KC E+ +Y+K K+ L+ CLYV
Sbjct: 942 EDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILI-ACLYV 1000
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI T +N + E FK M K+F MTD+G ++Y+LG+E + G+ Q+ YA ++LK
Sbjct: 1001 DDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLK 1060
Query: 122 RFNMLNCNPAST 133
+F M + NP T
Sbjct: 1061 KFKMDDSNPVCT 1072
>At1g48710 hypothetical protein
Length = 1352
Score = 126 bits (317), Expect = 2e-29
Identities = 61/132 (46%), Positives = 88/132 (66%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + RL KALYGLKQ+ RAWN RI + +++F+KC E+ +Y+K K+ L+ CLYV
Sbjct: 974 EDKVLRLKKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILI-ACLYV 1032
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI T +N + E FK M K+F MTD+G ++Y+LG+E + G+ Q+ YA ++LK
Sbjct: 1033 DDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLK 1092
Query: 122 RFNMLNCNPAST 133
+F M + NP T
Sbjct: 1093 KFKMDDSNPVCT 1104
>At2g05390 putative retroelement pol polyprotein
Length = 1307
Score = 124 bits (311), Expect = 9e-29
Identities = 60/130 (46%), Positives = 86/130 (66%), Gaps = 1/130 (0%)
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLY 60
NE +++LHKALYGLKQ+ RAWN ++ L + FVKC E +VY + ++ L+ + +Y
Sbjct: 916 NEGKVYKLHKALYGLKQAPRAWNTKLNKILQELNFVKCSKEPSVY-RRQEEKKLLIVAIY 974
Query: 61 VDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDIL 120
VDDL+VT S+L I FK M KF M+DLG+LTY+LG+E G++ Q++YA I+
Sbjct: 975 VDDLLVTGSSLDLILCFKKDMAGKFEMSDLGQLTYYLGIEVLHRKNGIILRQERYAMKII 1034
Query: 121 KRFNMLNCNP 130
+ M NCNP
Sbjct: 1035 EEAGMSNCNP 1044
>At3g59720 copia-type reverse transcriptase-like protein
Length = 1272
Score = 124 bits (310), Expect = 1e-28
Identities = 59/130 (45%), Positives = 87/130 (66%), Gaps = 1/130 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + RL K LYGLKQ+ RAWN RI + +++F+KC E+ +Y+K K+ L+ CLYV
Sbjct: 974 EDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILI-ACLYV 1032
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI T +N + E FK M K+F MTD+G ++Y+LG+E + G+ Q+ YA ++LK
Sbjct: 1033 DDLIFTGNNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLK 1092
Query: 122 RFNMLNCNPA 131
+F M + NP+
Sbjct: 1093 KFKMDDSNPS 1102
>At3g61330 copia-type polyprotein
Length = 1352
Score = 123 bits (308), Expect = 2e-28
Identities = 59/132 (44%), Positives = 87/132 (65%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + RL K LYGLKQ+ RAWN RI + +++F+KC E+ +Y+K K+ L+ CLYV
Sbjct: 974 EDKVLRLKKVLYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALYIKIQKEDILI-ACLYV 1032
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI T +N + E FK M K+F MTD+G ++Y+LG+E + G+ Q+ YA ++LK
Sbjct: 1033 DDLIFTGNNPSIFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLK 1092
Query: 122 RFNMLNCNPAST 133
+F + + NP T
Sbjct: 1093 KFKIDDSNPVCT 1104
>At1g32590 hypothetical protein, 5' partial
Length = 1263
Score = 123 bits (308), Expect = 2e-28
Identities = 59/125 (47%), Positives = 86/125 (68%), Gaps = 1/125 (0%)
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+++L KALYGLKQ+ RAW RI F ++ F KC E+ ++VK + LV + +YVDDL
Sbjct: 926 VYKLKKALYGLKQAPRAWYSRIEEFFGKEGFEKCYCEHTLFVKKERSDFLV-VSVYVDDL 984
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFN 124
I T S++ IE FK MM++F MTDLGK+ YFLG+E + G+ +Q+KYA++I+K++
Sbjct: 985 IYTGSSMEMIEGFKNSMMEEFAMTDLGKMKYFLGVEVIQDERGIFINQRKYAAEIIKKYG 1044
Query: 125 MLNCN 129
M CN
Sbjct: 1045 MEGCN 1049
>At3g25450 hypothetical protein
Length = 1343
Score = 119 bits (298), Expect = 3e-27
Identities = 59/133 (44%), Positives = 90/133 (67%), Gaps = 1/133 (0%)
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLY 60
+++ +++LHKALYGL+Q+ RAWN ++ L + +F KC E ++Y K ++ LV + +Y
Sbjct: 955 SKEKVYKLHKALYGLRQAPRAWNTKLNEILKELKFEKCHKEPSLYRKQEGENILV-VAVY 1013
Query: 61 VDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDIL 120
VDDL+VT SNL I FK M+ KF M+DLGKLTY+LG+E ++ +G+ Q++YA IL
Sbjct: 1014 VDDLLVTGSNLDIILNFKKGMVGKFEMSDLGKLTYYLGIEVLQSKDGITLKQERYAKKIL 1073
Query: 121 KRFNMLNCNPAST 133
+ M CN +T
Sbjct: 1074 EEAGMSKCNTVNT 1086
>At2g17490 putative retroelement pol polyprotein
Length = 822
Score = 115 bits (288), Expect = 4e-26
Identities = 61/132 (46%), Positives = 82/132 (61%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + RLHKALYGLKQ+ RAW RI F ++ F + ++ +Y K LV +C+YV
Sbjct: 600 EDHVLRLHKALYGLKQAPRAWYSRIDEFFQRENFTRSDNDHALYTKEVLGKLLV-VCIYV 658
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLIVT + +E FK M +F M+DLG L YFLGME ++ EG+ Q+ YA +LK
Sbjct: 659 DDLIVTGDDEEMVEEFKTAMKNEFEMSDLGLLNYFLGMEIVQSLEGIFLSQECYARKLLK 718
Query: 122 RFNMLNCNPAST 133
+FNM + ST
Sbjct: 719 KFNMEDSKIMST 730
>At3g60170 putative protein
Length = 1339
Score = 114 bits (286), Expect = 7e-26
Identities = 61/160 (38%), Positives = 96/160 (59%), Gaps = 21/160 (13%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
E+ +++L KALYGLKQ+ RAW RI + +++EF +C +E+ ++ K ++ N++ + LYV
Sbjct: 963 EEKVYKLRKALYGLKQAPRAWYSRIEAYFLKEEFERCPSEHTLFTK-TRVGNILIVSLYV 1021
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI T S+ A + FK MM +F M+DLGK+ +FLG+E ++ G+ Q++YA ++L
Sbjct: 1022 DDLIFTGSDKAMCDEFKKSMMLEFEMSDLGKMKHFLGIEVKQSDGGIFICQRRYAREVLA 1081
Query: 122 RFNMLNCNPA--------------------STLFKQIVGS 141
RF M N T+FKQ+VGS
Sbjct: 1082 RFGMDESNAVKNPIVPGTKLTKDENGEKVDETMFKQLVGS 1121
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 110 bits (274), Expect = 2e-24
Identities = 56/132 (42%), Positives = 81/132 (60%), Gaps = 1/132 (0%)
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
E+ + RL+KALYGLKQ+ RAW +RI ++ +Q F + + +Y K + L+ + LYV
Sbjct: 973 EEKVLRLYKALYGLKQAPRAWYERIDSYFIQNGFARSMNDAALYSKKKGEDVLI-VSLYV 1031
Query: 62 DDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
DDLI+T +N I FK +M +F MTDLG L YFLGME + G+ Q+KYA+ ++
Sbjct: 1032 DDLIITGNNTHLINTFKKNMKDEFEMTDLGLLNYFLGMEVNQDDSGIFLSQEKYANKLID 1091
Query: 122 RFNMLNCNPAST 133
+F M ST
Sbjct: 1092 KFGMKESKSVST 1103
>At2g23330 putative retroelement pol polyprotein
Length = 1496
Score = 106 bits (264), Expect = 2e-23
Identities = 52/128 (40%), Positives = 81/128 (62%), Gaps = 1/128 (0%)
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+ RL K+LYGLKQ+ R W ++ L F + +Y+++ + D+ ++H+ +YVDDL
Sbjct: 1125 VCRLRKSLYGLKQAPRCWFSKLSTALRNIGFTQSYEDYSLFSLKNGDT-IIHVLVYVDDL 1183
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFN 124
IV +NL I+ FK + K F+M DLGKL YFLG+E ++ +G Q+KYA DI+K
Sbjct: 1184 IVAGNNLDAIDRFKSQLHKCFHMKDLGKLKYFLGLEVSRGPDGFCLSQRKYALDIVKETG 1243
Query: 125 MLNCNPAS 132
+L C P++
Sbjct: 1244 LLGCKPSA 1251
>At4g22040 LTR retrotransposon like protein
Length = 1109
Score = 100 bits (248), Expect = 2e-21
Identities = 49/133 (36%), Positives = 81/133 (60%), Gaps = 1/133 (0%)
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+ RLHK+LYGLKQ+ R W ++ + L Q F + ++Y+++ N+ D VH+ +YVDDL
Sbjct: 738 VCRLHKSLYGLKQAPRCWFSKLSSALKQYGFTQSLSDYSLFSYNN-DGVFVHVLVYVDDL 796
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFN 124
I++ S + FK ++ F+M DLG L YFLG+E ++ ++G Q+KY DI+
Sbjct: 797 IISGSCPDAVAQFKSYLESCFHMKDLGLLKYFLGIEVSRNAQGFYLSQRKYVLDIISEMG 856
Query: 125 MLNCNPASTLFKQ 137
+L P++ +Q
Sbjct: 857 LLGARPSAFPLEQ 869
>At1g57640
Length = 1444
Score = 100 bits (248), Expect = 2e-21
Identities = 49/133 (36%), Positives = 81/133 (60%), Gaps = 1/133 (0%)
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+ RLHK+LYGLKQ+ R W ++ + L Q F + ++Y+++ N+ D VH+ +YVDDL
Sbjct: 1073 VCRLHKSLYGLKQAPRCWFSKLSSALKQYGFTQSLSDYSLFSYNN-DGIFVHVLVYVDDL 1131
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFN 124
I++ S + FK ++ F+M DLG L YFLG+E ++ ++G Q+KY DI+
Sbjct: 1132 IISGSCPDAVAQFKSYLESCFHMKDLGLLKYFLGIEVSRNAQGFYLSQRKYVLDIISEMG 1191
Query: 125 MLNCNPASTLFKQ 137
+L P++ +Q
Sbjct: 1192 LLGARPSAFPLEQ 1204
>At1g31210 putative reverse transcriptase
Length = 1415
Score = 100 bits (248), Expect = 2e-21
Identities = 52/133 (39%), Positives = 83/133 (62%), Gaps = 1/133 (0%)
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+ RL KA+YGLKQ+ RAW NFL+ FV K++ +++V + +D ++++ LYVDD+
Sbjct: 999 VCRLTKAIYGLKQAPRAWFDTFSNFLLDYGFVCSKSDPSLFVCH-QDGKILYLLLYVDDI 1057
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFN 124
++T S+ + +E + +F+M DLG YFLG++ + GL HQ YA+DIL++
Sbjct: 1058 LLTGSDQSLLEDLLQALKNRFSMKDLGPPRYFLGIQIEDYANGLFLHQTAYATDILQQAG 1117
Query: 125 MLNCNPASTLFKQ 137
M +CNP T Q
Sbjct: 1118 MSDCNPMPTPLPQ 1130
>At2g14930 pseudogene
Length = 1149
Score = 99.8 bits (247), Expect = 2e-21
Identities = 51/131 (38%), Positives = 83/131 (62%), Gaps = 1/131 (0%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D + LHKALYGLKQ+ RAW + FL+ FV ++ +++V K+ +++ + LYVD
Sbjct: 991 DHVCLLHKALYGLKQAPRAWFDKFSKFLLSFGFVCSMSDPSLFVC-VKNKDVIMLLLYVD 1049
Query: 63 DLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKR 122
D+++T ++ + + + K+F M DLG+L+YFLG++ S+GL Q+KYA D+L
Sbjct: 1050 DMVITGNSSKLLSSLLSELNKQFKMKDLGRLSYFLGIQAQFHSQGLFLSQQKYAEDLLAT 1109
Query: 123 FNMLNCNPAST 133
M NC+P +T
Sbjct: 1110 AAMSNCSPVAT 1120
>At4g27210 putative protein
Length = 1318
Score = 99.0 bits (245), Expect = 4e-21
Identities = 48/131 (36%), Positives = 83/131 (62%), Gaps = 1/131 (0%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D + LHK++YGLKQS RAW + FL++ F K++ ++++ + ++NL+ + LYVD
Sbjct: 851 DHVCLLHKSIYGLKQSPRAWFDKFSTFLLEFGFFCSKSDPSLFIY-AHNNNLILLLLYVD 909
Query: 63 DLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKR 122
D+++T ++ + + + K+F MTD+G+L YFLG++ + GL Q+KYA D+L
Sbjct: 910 DMVITGNSSQTLTSLLAALNKEFRMTDMGQLHYFLGIQVQRQQNGLFMSQQKYAEDLLIA 969
Query: 123 FNMLNCNPAST 133
+M +C P T
Sbjct: 970 ASMEHCTPLPT 980
>At2g24660 putative retroelement pol polyprotein
Length = 1156
Score = 99.0 bits (245), Expect = 4e-21
Identities = 51/135 (37%), Positives = 86/135 (62%), Gaps = 1/135 (0%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D + RL K+LYGLKQ+ R W K++ + L++ FV+ +Y+ + +++ + + +YVD
Sbjct: 775 DKVCRLRKSLYGLKQAPRCWFKKLSDALLRFGFVQGHEDYSFF-SYTRNGIELRVLVYVD 833
Query: 63 DLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKR 122
DL++ ++ ++ FK ++ + F+M DLGKL YFLG+E ++ SEG+ Q+KYA DI+
Sbjct: 834 DLLICGNDGYMLQKFKEYLGRCFSMKDLGKLKYFLGIEVSRGSEGIFLSQRKYALDIITD 893
Query: 123 FNMLNCNPASTLFKQ 137
L C PA T +Q
Sbjct: 894 SGNLGCRPALTPLEQ 908
>At2g04820 putative retroelement pol polyprotein
Length = 387
Score = 99.0 bits (245), Expect = 4e-21
Identities = 52/130 (40%), Positives = 88/130 (67%), Gaps = 4/130 (3%)
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+ RL+KALYGLKQ+ RAW + + N+L+ F A+ ++++ +S L ++ +YVDD+
Sbjct: 18 VCRLNKALYGLKQAPRAWYQELRNYLLAAGFTNSLADTSLFIYKRANSYL-YVLVYVDDI 76
Query: 65 IVT-DSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRF 123
I+T D++L ++ F + +F++ DLG L+YFLG+E T+T +GL Q+KY +D+L +
Sbjct: 77 IITGDASL--VKNFNVSLADRFSLKDLGALSYFLGIEATRTKKGLHLMQRKYITDLLAKT 134
Query: 124 NMLNCNPAST 133
+ML+ P ST
Sbjct: 135 HMLDAKPIST 144
>At1g53810
Length = 1522
Score = 99.0 bits (245), Expect = 4e-21
Identities = 50/131 (38%), Positives = 82/131 (62%), Gaps = 1/131 (0%)
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D + LHK+LYGLKQS RAW R NFL++ F+ + +++V +S +++++ + LYVD
Sbjct: 1053 DHVCLLHKSLYGLKQSPRAWFDRFSNFLLEFGFICSLFDPSLFVYSS-NNDVILLLLYVD 1111
Query: 63 DLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKR 122
D+++T +N + + K+F M D+G++ YFLG++ GL Q+KYA D+L
Sbjct: 1112 DMVITGNNSQSLTHLLAALNKEFRMKDMGQVHYFLGIQIQTYDGGLFMSQQKYAEDLLIT 1171
Query: 123 FNMLNCNPAST 133
+M NC+P T
Sbjct: 1172 ASMANCSPMPT 1182
>At2g06840 putative retroelement pol polyprotein
Length = 1102
Score = 97.8 bits (242), Expect = 9e-21
Identities = 47/123 (38%), Positives = 77/123 (62%), Gaps = 1/123 (0%)
Query: 10 KALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDLIVTDS 69
++LYGLKQ+ R W ++ L + F + +Y+++ N +D ++H +YVDD I+ +
Sbjct: 740 ESLYGLKQAPRCWFAKLSTALRKLGFTQSYEDYSLFSLN-RDGTVIHFLVYVDDFIIVGN 798
Query: 70 NLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFNMLNCN 129
NL I+ FK H+ K F+M DLGKL YFLG+E ++ ++G Q+KYA DI+ +L
Sbjct: 799 NLKAIDHFKEHLHKCFHMKDLGKLKYFLGLEVSRGADGFCLSQQKYALDIINEAGLLGYK 858
Query: 130 PAS 132
P++
Sbjct: 859 PSA 861
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.366 0.163 0.562
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,128,217
Number of Sequences: 26719
Number of extensions: 209186
Number of successful extensions: 1596
Number of sequences better than 10.0: 89
Number of HSP's better than 10.0 without gapping: 85
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 1435
Number of HSP's gapped (non-prelim): 97
length of query: 385
length of database: 11,318,596
effective HSP length: 101
effective length of query: 284
effective length of database: 8,619,977
effective search space: 2448073468
effective search space used: 2448073468
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC135162.11


