

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135160.3 - phase: 0
(239 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
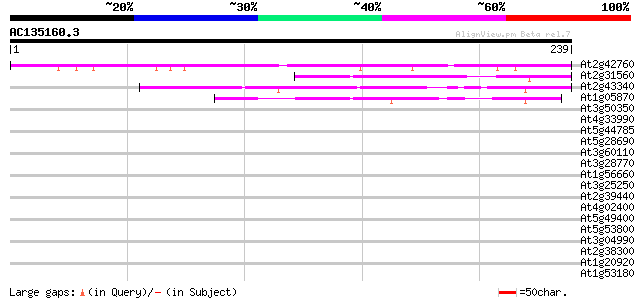
Score E
Sequences producing significant alignments: (bits) Value
At2g42760 unknown protein 150 4e-37
At2g31560 unknown protein 52 4e-07
At2g43340 unknown protein 48 5e-06
At1g05870 unknown protein 47 7e-06
At3g50350 hypothetical protein 37 0.007
At4g33990 putative protein 36 0.016
At5g44785 unknown protein 35 0.027
At5g28690 unknown protein 35 0.027
At3g60110 unknown protein 35 0.046
At3g28770 hypothetical protein 34 0.079
At1g56660 hypothetical protein 34 0.079
At3g25250 protein kinase, putative 33 0.10
At2g39440 hypothetical protein 33 0.13
At4g02400 33 0.18
At5g49400 putative protein 32 0.23
At5g53800 unknown protein 32 0.30
At3g04990 hypothetical protein 32 0.30
At2g38300 unknown protein 32 0.30
At1g20920 putative RNA helicase 32 0.30
At1g53180 putative protein 32 0.39
>At2g42760 unknown protein
Length = 267
Score = 150 bits (380), Expect = 4e-37
Identities = 114/271 (42%), Positives = 151/271 (55%), Gaps = 37/271 (13%)
Query: 1 MSAEQILTLLDSFWFETTI-------LTNKNPSK--VEEILPQ---DTNLLLVPTPNLDV 48
M+ E++L L + W E I L K+ K +EIL + + L P L
Sbjct: 1 MAGEELLKLFEQNWSERPIFKKDKENLNGKSREKRGEKEILEERREEEALKNFPVSFLVE 60
Query: 49 RFYSEQNLSIIGS---VFSDSP-----SPNSVL----TSSKLRTIPSEREFGEFSTGTSS 96
R S++ + S +FS S SP SVL T KL+TI S +E F+
Sbjct: 61 RAMSDETMMTTSSKTSLFSSSSDDLFLSPRSVLPVKPTPMKLQTILSGKEVNAFTIAERE 120
Query: 97 ENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEED-KDSKLVS 155
+KE+ KKK+S R RK +S+S+LE++ELKGFMDLGFVFSE+D KDS LVS
Sbjct: 121 RLLSEKEEQRKKKKKKSNV---RTRKGKSMSDLEYEELKGFMDLGFVFSEDDHKDSDLVS 177
Query: 156 LIPGLQRLGRENDDA--EEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERK---NPLV 210
++PGLQRL +++D EE E+EE KI N + +PYLSEAWD R+ K P +
Sbjct: 178 ILPGLQRLVKKDDGVTKEEEEEEEEDKIGGNRAA--RPYLSEAWDHCGGRKGKKQITPEI 235
Query: 211 NWRV--PDKGSEIDMKDNLKFWAHAVASIAR 239
WRV P SE+D+KDNL+ WAHAVAS R
Sbjct: 236 KWRVPAPAAASEVDLKDNLRLWAHAVASTIR 266
>At2g31560 unknown protein
Length = 202
Score = 51.6 bits (122), Expect = 4e-07
Identities = 34/119 (28%), Positives = 58/119 (48%), Gaps = 14/119 (11%)
Query: 122 KSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEHKKI 181
+++SL++ + +ELKG +DLGF FS D+ +L + +P L+ + + + + H K
Sbjct: 94 RAKSLTDDDLEELKGCLDLGFGFS-YDEIPELCNTLPALELCYSMSQKFLDDKQQNHHKS 152
Query: 182 DENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPDKGSE-IDMKDNLKFWAHAVASIAR 239
E S P + P+ NW++ G + D+K LK+WA VA R
Sbjct: 153 QEEDDSSPPPTTTA------------PIANWKISSPGDDPDDVKARLKYWAQTVACTVR 199
>At2g43340 unknown protein
Length = 241
Score = 47.8 bits (112), Expect = 5e-06
Identities = 51/194 (26%), Positives = 86/194 (44%), Gaps = 24/194 (12%)
Query: 56 LSIIGSVFSDSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQS-- 113
LS S +D S + + TSS SE E E + G +K+ KKK +
Sbjct: 7 LSRCSSYEADVDSDSEISTSSFSSCSYSESEEEEINNGFGVVQS-KKQTKKLEKKKSNVL 65
Query: 114 -------QSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRE 166
+ ++++SL++ + +ELKG +DLGF F+ E+ +L + +P L+
Sbjct: 66 LEGYVVDSAVNDDLKRTKSLTDDDLEELKGCVDLGFGFNYEE-IPELCNTLPALELCYSM 124
Query: 167 NDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPDKG-SEIDMKD 225
+ + + H S + P + V D +P+ +W++ G + D+K
Sbjct: 125 SQKFIDQDHHHH--------SSSSP--EKKSSVLD--SPVSPIASWKISSPGDNPDDVKA 172
Query: 226 NLKFWAHAVASIAR 239
LKFWA AVA R
Sbjct: 173 RLKFWAQAVACTMR 186
>At1g05870 unknown protein
Length = 189
Score = 47.4 bits (111), Expect = 7e-06
Identities = 38/152 (25%), Positives = 73/152 (48%), Gaps = 34/152 (22%)
Query: 88 GEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEE 147
G T +SS QK+D+ +S+SL++ + ++L+G +DLGF FS
Sbjct: 61 GYVETASSSSVDDQKDDLT---------------RSKSLTDDDLEDLRGCLDLGFGFS-Y 104
Query: 148 DKDSKLVSLIPGLQ---RLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRE 204
D+ +L + +P L+ + ++ D ++ + E +++ + P ++
Sbjct: 105 DEIPELCNTLPALELCYSMSQKFLDDKQNKSPETSSVED---CPSPPLVT---------- 151
Query: 205 RKNPLVNWRVPDKG-SEIDMKDNLKFWAHAVA 235
P+ NW++ G + D+K LK+WA AVA
Sbjct: 152 -ATPIANWKISSPGDNPDDVKARLKYWAQAVA 182
>At3g50350 hypothetical protein
Length = 181
Score = 37.4 bits (85), Expect = 0.007
Identities = 20/53 (37%), Positives = 32/53 (59%), Gaps = 1/53 (1%)
Query: 110 KKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVF-SEEDKDSKLVSLIPGLQ 161
K+Q S R R+ +SL++ + ELK +LGF F S E+ D +L + +P L+
Sbjct: 57 KRQDISRHRHLRRGKSLTDEDLDELKASFELGFGFGSPENADPRLSNTLPALE 109
>At4g33990 putative protein
Length = 844
Score = 36.2 bits (82), Expect = 0.016
Identities = 22/57 (38%), Positives = 35/57 (60%), Gaps = 3/57 (5%)
Query: 106 AYTKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEE--DKDSKLVSLIPGL 160
A+ +KK QS R R S+S+++ + +ELKG ++LGF F + D D +L +P L
Sbjct: 748 AWLRKKGKQSLGRLGR-SKSVTDEDLEELKGCIELGFGFEPDSPDLDPRLSETLPAL 803
>At5g44785 unknown protein
Length = 440
Score = 35.4 bits (80), Expect = 0.027
Identities = 17/63 (26%), Positives = 32/63 (49%)
Query: 53 EQNLSIIGSVFSDSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQ 112
E + + G +F DSP PN + S ++ + F + +T T+ +++V KKK
Sbjct: 142 EDRIHVSGKLFIDSPPPNVTYSQSNVQVMVQNLNFVQAATSTTKTISPPEKEVTSIKKKP 201
Query: 113 SQS 115
++S
Sbjct: 202 ARS 204
>At5g28690 unknown protein
Length = 192
Score = 35.4 bits (80), Expect = 0.027
Identities = 24/67 (35%), Positives = 39/67 (57%), Gaps = 12/67 (17%)
Query: 104 DVAYTKKKQSQSYRRRRRKSRSLSELE----------FKELKGFMDLGFVFSEEDKDSKL 153
DVA+ K+++ Q + + +K +S+SE + +ELKG ++LGF FSEE KL
Sbjct: 50 DVAWEKRRR-QMLKIQEKKQKSVSENDNDSPDLTDEDLRELKGSIELGFGFSEE-AGQKL 107
Query: 154 VSLIPGL 160
+ +P L
Sbjct: 108 CNTLPAL 114
>At3g60110 unknown protein
Length = 644
Score = 34.7 bits (78), Expect = 0.046
Identities = 35/165 (21%), Positives = 68/165 (41%), Gaps = 17/165 (10%)
Query: 64 SDSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKS 123
S++ S S SS L +P +++ + S + QK++ + + + S
Sbjct: 413 SEAESSVSRQKSSVLSLVPCKKKSSALKKTSPSSSSRQKDEKKSQEVSEEKIVTTTATTS 472
Query: 124 RSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEHKKIDE 183
S KE+ V +++ K + + I E+ D ++ E EE+ K ++
Sbjct: 473 ARSSRRTSKEIA-------VVAKDTKTGRAKNNIKKQTDTKTESSDDDDDEKEENSKTEK 525
Query: 184 NVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPDKGSEIDMKDNLK 228
++D K +++ F +R +KN P KG E K+ K
Sbjct: 526 KTVADKKKSVAD----FLKRIKKNS------PQKGKETTSKNQKK 560
>At3g28770 hypothetical protein
Length = 2081
Score = 33.9 bits (76), Expect = 0.079
Identities = 37/145 (25%), Positives = 62/145 (42%), Gaps = 20/145 (13%)
Query: 65 DSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSR 124
D+ + +SKL+ + + + S ++S+N +KE Y +KK ++ K +
Sbjct: 976 DNKKETTKSENSKLKEENKDNKEKKESEDSASKNREKKE---YEEKKSKTKEEAKKEKKK 1032
Query: 125 SLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEH---KKI 181
S + +E K SEE K K L+ +E + E+ E E H KK
Sbjct: 1033 SQDKK--REEKD--------SEERKSKKEKEESRDLKAKKKEEETKEKKESENHKSKKKE 1082
Query: 182 DENVLSDNKPYLSEAWDVFDQRERK 206
D+ DNK E D++E+K
Sbjct: 1083 DKKEHEDNKSMKKEE----DKKEKK 1103
>At1g56660 hypothetical protein
Length = 522
Score = 33.9 bits (76), Expect = 0.079
Identities = 36/145 (24%), Positives = 70/145 (47%), Gaps = 8/145 (5%)
Query: 89 EFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEED 148
E S +N +++D + ++K+ + + ++ K S +E E K+LKG G +ED
Sbjct: 241 EMKEKDSKKNKKKEKDESCAEEKKKKPDKEKKEKDES-TEKEDKKLKGKKGKGEKPEKED 299
Query: 149 KDSKLVSLIPGLQRLGRENDDAEEGEDEEHK---KIDENVLSD--NKPYLSEAWDVFDQR 203
+ K Q + E D +EG+ +++K K E V+ + K + D + +
Sbjct: 300 EGKKTKEHDATEQEMDDEAADHKEGKKKKNKDKAKKKETVIDEVCEKETKDKDDDEGETK 359
Query: 204 ERKNPLVNWRVPDKGSEIDMKDNLK 228
++KN + +KG E D+K++ K
Sbjct: 360 QKKNKKKE-KKSEKG-EKDVKEDKK 382
>At3g25250 protein kinase, putative
Length = 421
Score = 33.5 bits (75), Expect = 0.10
Identities = 24/71 (33%), Positives = 39/71 (54%), Gaps = 4/71 (5%)
Query: 121 RKSRSL--SELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEH 178
++SR+L + LE L G G VF D D+KL++L L+ + E ED+E+
Sbjct: 7 KQSRALDFNRLEVLSLLGRGAKGVVFLVRDDDAKLLALKVILKEAIEKKKKGRESEDDEY 66
Query: 179 KKI--DENVLS 187
K++ ++ VLS
Sbjct: 67 KRVSFEQGVLS 77
>At2g39440 hypothetical protein
Length = 773
Score = 33.1 bits (74), Expect = 0.13
Identities = 41/165 (24%), Positives = 65/165 (38%), Gaps = 29/165 (17%)
Query: 42 PTPNLDVRFYSEQNLSIIGSVFSDS---PSPNSVLTSSKLRTIPSEREFGEFSTGTSSEN 98
P L+ FY E NL + DS P PN + ++L T+ SE E +S G+ E
Sbjct: 534 PVSVLEPMFY-EDNLDDSEDILDDSEDLPYPNFLSLENQLETLKSESE--SYSDGSGMEV 590
Query: 99 HHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIP 158
+E S + KE K +GF+ ++E +DS + I
Sbjct: 591 SSDEE---------------------SALDSAIKESKESEPIGFLDTQESRDSSYIDDIL 629
Query: 159 GLQRLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQR 203
LG +N + + KI E + + K Y +W D++
Sbjct: 630 AEVLLGDKNCVPGKRDLVITPKIFEKL--EKKYYTETSWKRSDRK 672
>At4g02400
Length = 870
Score = 32.7 bits (73), Expect = 0.18
Identities = 26/118 (22%), Positives = 52/118 (44%), Gaps = 9/118 (7%)
Query: 112 QSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGRENDDAE 171
+ Q + R + SE E +++ G + S + ++ S +QR +D E
Sbjct: 625 KGQDKKESRDEESEDSESEAEQMVD----GILTSASKETYEIPSQAELIQRAFAGDDVVE 680
Query: 172 EGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERKNPLVNWRV-----PDKGSEIDMK 224
E E ++ + +++ V KP L W + ++K L +W V +K ++D+K
Sbjct: 681 EFEKDKQEVLNQEVPEPEKPVLVPGWGQWTNVQKKRGLPSWMVREHEDANKKRKLDLK 738
>At5g49400 putative protein
Length = 280
Score = 32.3 bits (72), Expect = 0.23
Identities = 28/90 (31%), Positives = 39/90 (43%), Gaps = 6/90 (6%)
Query: 64 SDSPSPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAY-TKKKQSQSYRRRRRK 122
SDS S + S E+S+ + SE+ ++ A +KKKQ Q RRRR
Sbjct: 184 SDSESDSEASVFETDSDGSSGESSSEYSSSSDSEDERRRRRKAKKSKKKQKQRKERRRRY 243
Query: 123 SRSLSELEFKELKGFMDLGFVFSEEDKDSK 152
S S SE E D S+ED+ +
Sbjct: 244 SSSSSESSESESASDSD-----SDEDRSRR 268
>At5g53800 unknown protein
Length = 339
Score = 32.0 bits (71), Expect = 0.30
Identities = 25/113 (22%), Positives = 48/113 (42%), Gaps = 23/113 (20%)
Query: 85 REFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVF 144
R++ S+ + SE+ + D ++ + + R+R+RK R E E K +
Sbjct: 101 RDYSSSSSDSESESESEYSDSEESESEDERRRRKRKRKEREEEEKERKRRR--------- 151
Query: 145 SEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSEAW 197
+KD K + + + D ++ E+ KK E V K ++E+W
Sbjct: 152 --REKDKK---------KRNKSDKDGDKKRKEKKKKKSEKV---KKGAVTESW 190
>At3g04990 hypothetical protein
Length = 227
Score = 32.0 bits (71), Expect = 0.30
Identities = 26/98 (26%), Positives = 48/98 (48%), Gaps = 7/98 (7%)
Query: 147 EDKDSKLVSLIPGLQRLGRENDDAE----EGEDEEHKKIDENVLSDNKPYLSEAWDVFDQ 202
E KD++LV ++ L+R E + E EDE K E LS + E+ ++
Sbjct: 103 ELKDNQLVQVMAELKRRYSEARHVQKRKREMEDETATKKKE--LSMTVDQIQESGKQLEK 160
Query: 203 RERKNPLVNWRVPDKGSEIDM-KDNLKFWAHAVASIAR 239
+ R+ L + + +KG E+D+ K +K W + +++
Sbjct: 161 KSREVELKDKEIEEKGKELDLVKSQVKAWERKLIQLSK 198
>At2g38300 unknown protein
Length = 340
Score = 32.0 bits (71), Expect = 0.30
Identities = 17/60 (28%), Positives = 33/60 (54%), Gaps = 5/60 (8%)
Query: 156 LIPGLQRLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVP 215
+IP ++ G+ N + EE E+EE ++ +E+ +S N + D++ + P V +VP
Sbjct: 1 MIPSMEGGGKTNREEEEEEEEEEEEGEESKVSSNSTV-----EESDKKTKVRPYVRSKVP 55
>At1g20920 putative RNA helicase
Length = 1166
Score = 32.0 bits (71), Expect = 0.30
Identities = 32/122 (26%), Positives = 54/122 (44%), Gaps = 11/122 (9%)
Query: 71 SVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELE 130
SV SS+ P + + G E ++E++ +KK + +RRR+ + EL+
Sbjct: 236 SVGRSSRHEDSPKRKSVED--NGEKKEKKTREEELEDEQKKLDEEVEKRRRRVQEWQELK 293
Query: 131 FKELKGFMD-LGFVFSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEHKKIDENVLSDN 189
K+ + + G E K K + L E+DD EEG EE + + +V +
Sbjct: 294 RKKEEAESESKGDADGNEPKAGKAWT-------LEGESDD-EEGHPEEKSETEMDVDEET 345
Query: 190 KP 191
KP
Sbjct: 346 KP 347
Score = 27.7 bits (60), Expect = 5.7
Identities = 20/97 (20%), Positives = 43/97 (43%), Gaps = 6/97 (6%)
Query: 84 EREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFV 143
+R + S +++ + +D K+K+ + RRRR K R E ++ D V
Sbjct: 48 DRRKKRVKSSDSEDDYDRDDDEEREKRKEKERERRRRDKDRVKRRSERRKSSDSED--DV 105
Query: 144 FSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEHKK 180
E+++D + V+ + G + + G+D + +
Sbjct: 106 EEEDERDKRRVN----EKERGHREHERDRGKDRKRDR 138
>At1g53180 putative protein
Length = 358
Score = 31.6 bits (70), Expect = 0.39
Identities = 16/27 (59%), Positives = 19/27 (70%), Gaps = 1/27 (3%)
Query: 213 RVPDKGSEIDMKDNLKFWAHAVASIAR 239
RVP K S +MKD +KFWA AVA+ R
Sbjct: 330 RVP-KDSSKEMKDQIKFWARAVATNVR 355
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.130 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,778,398
Number of Sequences: 26719
Number of extensions: 277460
Number of successful extensions: 1359
Number of sequences better than 10.0: 94
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 59
Number of HSP's that attempted gapping in prelim test: 1276
Number of HSP's gapped (non-prelim): 124
length of query: 239
length of database: 11,318,596
effective HSP length: 96
effective length of query: 143
effective length of database: 8,753,572
effective search space: 1251760796
effective search space used: 1251760796
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135160.3


