

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135160.2 + phase: 0 /pseudo
(319 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
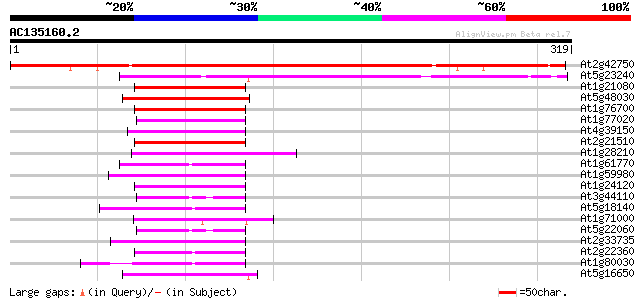
Score E
Sequences producing significant alignments: (bits) Value
At2g42750 unknown protein 373 e-104
At5g23240 unknown protein 121 6e-28
At1g21080 putative dnaJ protein 65 4e-11
At5g48030 DnaJ protein-like 65 6e-11
At1g76700 DnaJ like protein 64 8e-11
At1g77020 62 3e-10
At4g39150 dnaJ-like protein 58 6e-09
At2g21510 putative DnaJ protein 58 6e-09
At1g28210 AtJ1 (atj) 58 6e-09
At1g61770 unknown protein 54 9e-08
At1g59980 unknown protein 54 1e-07
At1g24120 DNAJ-like heatshock protein (At1g24120) 52 3e-07
At3g44110 dnaJ protein homolog atj3 51 7e-07
At5g18140 unknown protein 51 1e-06
At1g71000 51 1e-06
At5g22060 DNAJ PROTEIN HOMOLOG ATJ 50 1e-06
At2g33735 unknown protein 50 1e-06
At2g22360 DnaJ like protein 50 1e-06
At1g80030 unknown protein 50 1e-06
At5g16650 unknown protein 50 2e-06
>At2g42750 unknown protein
Length = 344
Score = 373 bits (957), Expect = e-104
Identities = 204/344 (59%), Positives = 245/344 (70%), Gaps = 32/344 (9%)
Query: 1 MAQLLTPLYPEALKIHNSSLCTRSSWQ-MLQQKT--AVSPWRFATPSCKRR--GFGRVRV 55
MAQ+L+P+ + LK NS+L +RS KT A S W + KRR GR+RV
Sbjct: 1 MAQILSPVCTDLLKFQNSALSSRSGASPRFSAKTTGASSSWYLPRYAGKRRTDSIGRLRV 60
Query: 56 ATEQESFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCT 115
ATE S ++ V A+DYYAVLGLLPDAT E+IKKAYYNCMK+CHPDLSGNDPETTNFC
Sbjct: 61 ATEDASSLSTGDV-ADDYYAVLGLLPDATQEEIKKAYYNCMKSCHPDLSGNDPETTNFCM 119
Query: 116 FINEVYEVLSDPVQRRVYDDIHGYSLTSINPFMDDSSPKDHVFVDEFSCIGCKNCANVAC 175
FIN++YE+LSDPVQR VYD+IHGY++T+INPF+DDS+P+DHVFVDEF+CIGCKNCANVA
Sbjct: 120 FINDIYEILSDPVQRMVYDEIHGYTVTAINPFLDDSTPRDHVFVDEFACIGCKNCANVAP 179
Query: 176 DVFGIEEDFGRARVYNQFGNPELIQTAIESCPVDCIHWTSAAQLSLLEDEMRRIERVNLY 235
D+F IEEDFGRAR NQ GNP+L+Q A+E+CPVDCIH TSAAQLSLLEDEMRR+ERVN+
Sbjct: 180 DIFQIEEDFGRARACNQRGNPDLVQQAVETCPVDCIHQTSAAQLSLLEDEMRRVERVNVA 239
Query: 236 F*KHKV*MGSALGDVFRM-----------TIITAGKYPMGEKTV------------KILD 272
MGS DVFRM + A M K K +
Sbjct: 240 LMLSG--MGSGAVDVFRMARSRWEKRQAKVLNQARSRMMKRKNTDETPSYWDNLWGKQNE 297
Query: 273 YENSEEEVKERAKRAAAAARKWREYSRKGGAVKPSTVKLPEATS 316
Y+ SEEEV+ERA+RAAAAAR+WREYSR+G +P T KLP++ S
Sbjct: 298 YQKSEEEVQERAQRAAAAARRWREYSRRGVDKRP-TFKLPDSAS 340
>At5g23240 unknown protein
Length = 465
Score = 121 bits (303), Expect = 6e-28
Identities = 86/265 (32%), Positives = 131/265 (48%), Gaps = 21/265 (7%)
Query: 63 STSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYE 122
S+S S+ D Y +LG+ + QIK AY K CHPD++G+ + +NE Y+
Sbjct: 41 SSSSSITDFDLYDLLGIDRSSDKSQIKSAYRALQKRCHPDIAGDPGH--DMAIILNEAYQ 98
Query: 123 VLSDPVQRRVYD-------DIHGYSLTSI-NPFMDDSSPKDHVFVDEFSCIGCKNCANVA 174
+LSDP+ R+ YD ++ GY+ I + + + + FVDE C+GC CA A
Sbjct: 99 LLSDPISRQAYDKEQAKLEELRGYTGKPIYSVWCGPETEQRAAFVDEVKCVGCLKCALCA 158
Query: 175 CDVFGIEEDFGRARVYNQFGNPE-LIQTAIESCPVDCIHWTSAAQLSLLEDEMRRIERVN 233
F IE +GRARV Q+ +PE I+ AIE+CPVDCI + L+ LE M + R N
Sbjct: 159 EKTFAIETAYGRARVVAQWADPESKIKEAIEACPVDCISMVERSDLAPLEFLMSKQPRGN 218
Query: 234 LYF*KHKV*MGSALGDVFRMTIITAGKY-PMGEKTVKILDYENSEEEVKERAKRAAAAAR 292
+ ++ +G+ +G+ + K+ K + E S+ EV+ A A +
Sbjct: 219 V-----RIGVGNTVGERVSNVFVDVKKFQERYAKAMSRTTKETSQREVQISAVEAIRSIS 273
Query: 293 KWREYSRKGGAVKPSTVKLPEATSS 317
W Y R KP + PE+ S
Sbjct: 274 NWL-YWRSSPYTKPLS---PESNMS 294
>At1g21080 putative dnaJ protein
Length = 391
Score = 65.5 bits (158), Expect = 4e-11
Identities = 30/63 (47%), Positives = 41/63 (64%)
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRR 131
++Y VLG+ P AT +IKKAYY + HPD + NDP+ + + E Y+VLSDP QR+
Sbjct: 6 EFYDVLGVSPTATEAEIKKAYYIKARQVHPDKNPNDPQAAHNFQVLGEAYQVLSDPGQRQ 65
Query: 132 VYD 134
YD
Sbjct: 66 AYD 68
>At5g48030 DnaJ protein-like
Length = 456
Score = 64.7 bits (156), Expect = 6e-11
Identities = 29/72 (40%), Positives = 45/72 (62%)
Query: 65 SDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVL 124
S + A+DYY+VLG+ +A +IKKAYY K HPD++ +DPE +++ YE+L
Sbjct: 87 SSFMSAKDYYSVLGVSKNAQEGEIKKAYYGLAKKLHPDMNKDDPEAETKFQEVSKAYEIL 146
Query: 125 SDPVQRRVYDDI 136
D +R +YD +
Sbjct: 147 KDKEKRDLYDQV 158
>At1g76700 DnaJ like protein
Length = 398
Score = 64.3 bits (155), Expect = 8e-11
Identities = 30/63 (47%), Positives = 40/63 (62%)
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRR 131
+YY VLG+ P AT +IKKAYY + HPD + NDP+ + + E Y+VLSD QR+
Sbjct: 6 EYYDVLGVSPTATESEIKKAYYIKARQVHPDKNPNDPQAAHNFQVLGEAYQVLSDSGQRQ 65
Query: 132 VYD 134
YD
Sbjct: 66 AYD 68
>At1g77020
Length = 358
Score = 62.4 bits (150), Expect = 3e-10
Identities = 29/62 (46%), Positives = 37/62 (58%)
Query: 73 YYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRRV 132
YY VLG+ P A+ E+I+KAYY + HPD + DP + E Y+VLSDPV R
Sbjct: 7 YYDVLGVTPSASEEEIRKAYYIKARQVHPDKNQGDPLAAEKFQVLGEAYQVLSDPVHREA 66
Query: 133 YD 134
YD
Sbjct: 67 YD 68
>At4g39150 dnaJ-like protein
Length = 345
Score = 58.2 bits (139), Expect = 6e-09
Identities = 27/67 (40%), Positives = 38/67 (56%)
Query: 68 VGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDP 127
V +YY +LG+ DA+ +IKKAYY + HPD + DP+ + E Y+VL DP
Sbjct: 2 VKESEYYDILGVKIDASGAEIKKAYYVQARQVHPDKNPGDPQAAKNFQILGEAYQVLGDP 61
Query: 128 VQRRVYD 134
+R YD
Sbjct: 62 EKRTAYD 68
>At2g21510 putative DnaJ protein
Length = 346
Score = 58.2 bits (139), Expect = 6e-09
Identities = 26/63 (41%), Positives = 38/63 (60%)
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRR 131
+YY +LG+ DA+ +IKKAYY + HPD + DP+ + E Y+VLS+P +R
Sbjct: 6 EYYEILGVKTDASDAEIKKAYYLKARKVHPDKNPGDPQAAKNFQVLGEAYQVLSNPDKRA 65
Query: 132 VYD 134
YD
Sbjct: 66 AYD 68
>At1g28210 AtJ1 (atj)
Length = 427
Score = 58.2 bits (139), Expect = 6e-09
Identities = 32/94 (34%), Positives = 46/94 (48%)
Query: 70 AEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQ 129
A +YY VLG+ P AT E+IKK+++ K HPD + N+P I E YE L + +
Sbjct: 46 ARNYYDVLGVSPKATREEIKKSFHELAKKFHPDTNRNNPSAKRKFQEIREAYETLGNSER 105
Query: 130 RRVYDDIHGYSLTSINPFMDDSSPKDHVFVDEFS 163
R YD + + +N DS + FS
Sbjct: 106 REEYDKLQYRNSDYVNNDGGDSERFRRAYQSNFS 139
>At1g61770 unknown protein
Length = 300
Score = 54.3 bits (129), Expect = 9e-08
Identities = 30/72 (41%), Positives = 37/72 (50%), Gaps = 1/72 (1%)
Query: 63 STSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYE 122
ST+ GAED YA+LG+ DA IK++YY HPD DPE+ I YE
Sbjct: 25 STAIYCGAEDCYALLGVAQDANASDIKRSYYKLSLQHHPD-KNPDPESRKLFVKIATAYE 83
Query: 123 VLSDPVQRRVYD 134
+L D R YD
Sbjct: 84 ILKDNTTRAQYD 95
>At1g59980 unknown protein
Length = 414
Score = 53.9 bits (128), Expect = 1e-07
Identities = 27/78 (34%), Positives = 40/78 (50%)
Query: 57 TEQESFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTF 116
+E + D + + Y VLG+ ++T ++IK AY HPD + +DP
Sbjct: 8 SENKDAGEEDELRRRNPYEVLGIPSNSTDQEIKSAYRRMALRYHPDKNPDDPVAAEMFKE 67
Query: 117 INEVYEVLSDPVQRRVYD 134
+ YEVLSDP RR+YD
Sbjct: 68 VTFAYEVLSDPENRRLYD 85
>At1g24120 DNAJ-like heatshock protein (At1g24120)
Length = 436
Score = 52.4 bits (124), Expect = 3e-07
Identities = 25/63 (39%), Positives = 36/63 (56%)
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRR 131
D Y VLG+L ++T ++IK AY HPD + NDP + + Y +LSDP +RR
Sbjct: 20 DPYEVLGVLRNSTDQEIKSAYRKLALKYHPDKTANDPVAADMFKEVTFSYNILSDPEKRR 79
Query: 132 VYD 134
+D
Sbjct: 80 QFD 82
>At3g44110 dnaJ protein homolog atj3
Length = 420
Score = 51.2 bits (121), Expect = 7e-07
Identities = 27/62 (43%), Positives = 36/62 (57%), Gaps = 4/62 (6%)
Query: 73 YYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRRV 132
+Y +LG+ A+PE +KKAY HPD G DPE + + YEVLSDP +R +
Sbjct: 15 FYEILGVPKSASPEDLKKAYKKAAIKNHPD-KGGDPEKFK---ELAQAYEVLSDPEKREI 70
Query: 133 YD 134
YD
Sbjct: 71 YD 72
>At5g18140 unknown protein
Length = 333
Score = 50.8 bits (120), Expect = 1e-06
Identities = 28/83 (33%), Positives = 44/83 (52%), Gaps = 1/83 (1%)
Query: 52 RVRVATEQESFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETT 111
R + +FS+S + G +++YAVLG+ +AT IK+AY + HPD++ D +
Sbjct: 57 RSKTTITSAAFSSSSNTGGQNHYAVLGIARNATQGDIKRAYRLLARKFHPDVN-KDSKAG 115
Query: 112 NFCTFINEVYEVLSDPVQRRVYD 134
+ YEVLS+ R YD
Sbjct: 116 ELFKSVRCSYEVLSNEATRTQYD 138
>At1g71000
Length = 146
Score = 50.8 bits (120), Expect = 1e-06
Identities = 32/93 (34%), Positives = 44/93 (46%), Gaps = 13/93 (13%)
Query: 71 EDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDP----ETTNFCTFINEVYEVLSD 126
+ YY +LG+ D++ EQI++AY+ K HPD DP E I E Y VLSD
Sbjct: 7 QTYYEILGVAVDSSAEQIRRAYHKLAKIWHPDRWTKDPFRSGEAKRRFQQIQEAYSVLSD 66
Query: 127 PVQRRVY---------DDIHGYSLTSINPFMDD 150
+R Y D+ YSL + +DD
Sbjct: 67 ERKRSSYDVGLYDSGEDEEKQYSLEELQTMVDD 99
>At5g22060 DNAJ PROTEIN HOMOLOG ATJ
Length = 419
Score = 50.4 bits (119), Expect = 1e-06
Identities = 27/62 (43%), Positives = 35/62 (55%), Gaps = 4/62 (6%)
Query: 73 YYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRRV 132
+Y +LG+ A PE +KKAY HPD G DPE + + YEVLSDP +R +
Sbjct: 15 FYEILGVPKTAAPEDLKKAYKKAAIKNHPD-KGGDPEKFK---ELAQAYEVLSDPEKREI 70
Query: 133 YD 134
YD
Sbjct: 71 YD 72
>At2g33735 unknown protein
Length = 119
Score = 50.4 bits (119), Expect = 1e-06
Identities = 26/77 (33%), Positives = 40/77 (51%)
Query: 58 EQESFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFI 117
+ F D +D+Y VL L DA+ ++I+ ++ HPD + T+ I
Sbjct: 8 DSHGFDFFDFEDYKDHYKVLELNCDASDDEIRSSFIRLALKWHPDKFKEEDSATSRFQEI 67
Query: 118 NEVYEVLSDPVQRRVYD 134
NE Y+VLSDP+ R+ YD
Sbjct: 68 NEAYQVLSDPIARQEYD 84
>At2g22360 DnaJ like protein
Length = 442
Score = 50.4 bits (119), Expect = 1e-06
Identities = 26/63 (41%), Positives = 37/63 (58%), Gaps = 1/63 (1%)
Query: 72 DYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVLSDPVQRR 131
DYY+VLG+ +AT +IK AY + HPD++ DP I+ YEVLSD ++
Sbjct: 86 DYYSVLGVSKNATKAEIKSAYRKLARNYHPDVN-KDPGAEEKFKEISNAYEVLSDDEKKS 144
Query: 132 VYD 134
+YD
Sbjct: 145 LYD 147
>At1g80030 unknown protein
Length = 500
Score = 50.4 bits (119), Expect = 1e-06
Identities = 34/94 (36%), Positives = 46/94 (48%), Gaps = 13/94 (13%)
Query: 41 ATPSCKRRGFGRVRVATEQESFSTSDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCH 100
+TP RRG V AT + DYYA LG+ A ++IK AY + H
Sbjct: 56 STPYRNRRGRSLVVFAT------------SGDYYATLGVSKSANNKEIKAAYRRLARQYH 103
Query: 101 PDLSGNDPETTNFCTFINEVYEVLSDPVQRRVYD 134
PD++ +P T I+ YEVLSD +R +YD
Sbjct: 104 PDVN-KEPGATEKFKEISAAYEVLSDEQKRALYD 136
>At5g16650 unknown protein
Length = 128
Score = 50.1 bits (118), Expect = 2e-06
Identities = 31/82 (37%), Positives = 39/82 (46%), Gaps = 5/82 (6%)
Query: 65 SDSVGAEDYYAVLGLLPDATPEQIKKAYYNCMKTCHPDLSGNDPETTNFCTFINEVYEVL 124
+D +DYY +L + DAT E I+ Y HPD D T INE Y VL
Sbjct: 4 NDKSPPKDYYKILEVDYDATEELIRLNYRKLALKWHPDKHKGDSAATEKFQEINEAYNVL 63
Query: 125 SDPVQRRVYD-----DIHGYSL 141
DP +R YD +IH Y+L
Sbjct: 64 MDPAKRFEYDFTGIYEIHKYTL 85
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.135 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,326,326
Number of Sequences: 26719
Number of extensions: 309892
Number of successful extensions: 955
Number of sequences better than 10.0: 89
Number of HSP's better than 10.0 without gapping: 70
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 836
Number of HSP's gapped (non-prelim): 95
length of query: 319
length of database: 11,318,596
effective HSP length: 99
effective length of query: 220
effective length of database: 8,673,415
effective search space: 1908151300
effective search space used: 1908151300
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135160.2


