

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134967.10 + phase: 0 /pseudo
(624 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
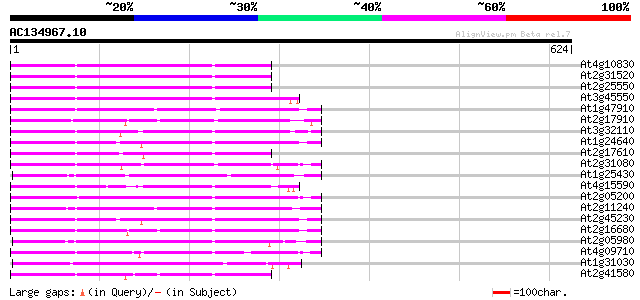
Score E
Sequences producing significant alignments: (bits) Value
At4g10830 putative protein 192 5e-49
At2g31520 putative non-LTR retroelement reverse transcriptase 188 9e-48
At2g25550 putative non-LTR retroelement reverse transcriptase 187 1e-47
At3g45550 putative protein 187 2e-47
At1g47910 reverse transcriptase, putative 187 2e-47
At2g17910 putative non-LTR retroelement reverse transcriptase 184 1e-46
At3g32110 non-LTR reverse transcriptase, putative 175 6e-44
At1g24640 hypothetical protein 175 6e-44
At2g17610 putative non-LTR retroelement reverse transcriptase 170 2e-42
At2g31080 putative non-LTR retroelement reverse transcriptase 169 3e-42
At1g25430 hypothetical protein 169 3e-42
At4g15590 reverse transcriptase like protein 164 1e-40
At2g05200 putative non-LTR retroelement reverse transcriptase 164 1e-40
At2g11240 pseudogene 163 2e-40
At2g45230 putative non-LTR retroelement reverse transcriptase 162 5e-40
At2g16680 putative non-LTR retroelement reverse transcriptase 161 9e-40
At2g05980 putative non-LTR retroelement reverse transcriptase 160 2e-39
At4g09710 RNA-directed DNA polymerase -like protein 158 8e-39
At1g31030 F17F8.5 158 1e-38
At2g41580 putative non-LTR retroelement reverse transcriptase 156 4e-38
>At4g10830 putative protein
Length = 1294
Score = 192 bits (488), Expect = 5e-49
Identities = 112/292 (38%), Positives = 168/292 (57%), Gaps = 5/292 (1%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W KWI V + SVLVNG P +RG+RQGDPLSP+LF+L A+ LN ++K+ V
Sbjct: 964 WIKWIMGAVKSVNYSVLVNGIPHGTIQPQRGIRQGDPLSPYLFILCADILNHLIKNRVAE 1023
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
G IG P V+HLQFADD+L + N +AL+ ++E SG K+N KSM
Sbjct: 1024 GDIRGIRIGNGVP-GVTHLQFADDSLFFCQSNVRNCQALKDVFDVYEYYSGQKINMSKSM 1082
Query: 121 LV-GVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWN 179
+ G + + ++L + YL +P ++ + I+ R+K R S W+
Sbjct: 1083 ITFGSRVHGTTQNRLKNILGIQSHGGGGKYLGLPEQFGRKKRDMFNYIIERVKKRTSSWS 1142
Query: 180 SRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKIS 239
+++LS G+ ++LKSV S+PVYA+S FK P I+S I++L +NF+ + R+I
Sbjct: 1143 AKYLSPAGKEIMLKSVAMSMPVYAMSCFKLPLNIVSEIEALLMNFW---WEKNAKKREIP 1199
Query: 240 WLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARY 291
W++W+ + K+ GGLG R L +FN ALL K WRM+ + L+ R++ ARY
Sbjct: 1200 WIAWKRLQYSKKEGGLGFRDLAKFNDALLAKQVWRMINNPNSLFARIMKARY 1251
>At2g31520 putative non-LTR retroelement reverse transcriptase
Length = 1524
Score = 188 bits (477), Expect = 9e-48
Identities = 111/292 (38%), Positives = 165/292 (56%), Gaps = 5/292 (1%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W WI V + SVL+NGSP RG+RQGDPLSP+LF+L + L+ ++ +
Sbjct: 758 WIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASS 817
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
G IG P ++HLQFADD+L + N +AL+ ++E SG K+N KSM
Sbjct: 818 GDLRGVRIGNGAP-AITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSM 876
Query: 121 LV-GVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWN 179
+ G + S + +L YL +P ++ +E I++R+K R S W+
Sbjct: 877 ITFGSRVYGSTQSKLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWS 936
Query: 180 SRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKIS 239
+RFLS G+ ++LKSV ++PVYA+S FK P GI+S I+SL +NF+ + + R I
Sbjct: 937 ARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFW---WEKASNQRGIP 993
Query: 240 WLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARY 291
W++W+ + K+ GGLG R L +FN ALL K WR++ L+ RV+ ARY
Sbjct: 994 WVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARY 1045
>At2g25550 putative non-LTR retroelement reverse transcriptase
Length = 1750
Score = 187 bits (476), Expect = 1e-47
Identities = 111/292 (38%), Positives = 164/292 (56%), Gaps = 5/292 (1%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W WI V + SVL+NGSP RG+RQGDPLSP+LF+L + L+ ++ +
Sbjct: 984 WIGWIMAAVKSVHYSVLINGSPHGYITPTRGIRQGDPLSPYLFILCGDILSHLINGRASS 1043
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
G IG P ++HLQFADD+L + N +AL+ ++E SG K+N KSM
Sbjct: 1044 GDLRGVRIGNGAP-AITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSM 1102
Query: 121 LV-GVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWN 179
+ G + S +L YL +P ++ +E I++R+K R S W+
Sbjct: 1103 ITFGSRVYGSTQSRLKQILEIPNQGGGGKYLGLPEQFGRKKKEMFEYIIDRVKKRTSTWS 1162
Query: 180 SRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKIS 239
+RFLS G+ ++LKSV ++PVYA+S FK P GI+S I+SL +NF+ + + R I
Sbjct: 1163 ARFLSPAGKEIMLKSVALAMPVYAMSCFKLPKGIVSEIESLLMNFW---WEKASNQRGIP 1219
Query: 240 WLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARY 291
W++W+ + K+ GGLG R L +FN ALL K WR++ L+ RV+ ARY
Sbjct: 1220 WVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRLIQYPNSLFARVMKARY 1271
>At3g45550 putative protein
Length = 851
Score = 187 bits (475), Expect = 2e-47
Identities = 119/332 (35%), Positives = 182/332 (53%), Gaps = 14/332 (4%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W WI V + SVL+NGSP RG+RQGDPLSP+LF+L + L+ ++K +
Sbjct: 223 WIGWIMAAVKSVHYSVLINGSPHGYISPTRGIRQGDPLSPYLFILCGDILSHLIKVKASS 282
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
G IG P ++HLQFADD+L + N +AL+ ++E SG K+N KS+
Sbjct: 283 GDIRGVRIGNGAP-AITHLQFADDSLFFCQANVRNCQALKDVFDVYEYYSGQKINVQKSL 341
Query: 121 LV-GVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWN 179
+ G + S ++L+ YL +P ++ + I++R+K R + W+
Sbjct: 342 ITFGSRVYGSTQTRLKTLLNIPNQGGGGKYLGLPEQFGRKKKEMFNYIIDRVKERTASWS 401
Query: 180 SRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKIS 239
++FLS G+ +LLKSV ++PVYA+S FK P GI+S I+SL +NF+ + + R I
Sbjct: 402 AKFLSPAGKEILLKSVALAMPVYAMSCFKLPQGIVSEIESLLMNFW---WEKASNKRGIP 458
Query: 240 WLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGGRLE 299
W++W+ + K+ GGLG R L +FN ALL K WR++ L+ RV+ ARY + ++
Sbjct: 459 WVAWKRLQYSKKEGGLGFRDLAKFNDALLAKQAWRIIQYPNSLFARVMKARYFKDNSIID 518
Query: 300 VGGRSVSCW-WSE----VRRLRDG----VGDG 322
RS + WS + LR G +GDG
Sbjct: 519 AKTRSQQSYGWSSLLSGIALLRKGTRYVIGDG 550
>At1g47910 reverse transcriptase, putative
Length = 1142
Score = 187 bits (474), Expect = 2e-47
Identities = 116/350 (33%), Positives = 181/350 (51%), Gaps = 18/350 (5%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W WI C++T VL+NG P +RGLRQGDPLSP+LF+L E L ++
Sbjct: 386 WISWIMWCITTVQYKVLINGQPKGLIIPERGLRQGDPLSPYLFILCTEVLIANIRKAERQ 445
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
L +G + +P VSHL FADD+L + + L +E++SG ++NF KS
Sbjct: 446 NLITGIKVATPSP-AVSHLLFADDSLFFCKANKEQCGIILEILKQYESVSGQQINFSKSS 504
Query: 121 L-VGVNIGSSWLYEAASVLSCK--VGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSG 177
+ G + S + +L G+ +L L +GG+ ++ + + +R++SR++G
Sbjct: 505 IQFGHKVEDSIKADIKLILGIHNLGGMGSYLGLPESLGGSKTKV--FSFVRDRLQSRING 562
Query: 178 WNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRK 237
W+++FLS GG+ V++KSV +LP Y +S F+ P I S + S F+ G D+R
Sbjct: 563 WSAKFLSKGGKEVMIKSVAATLPRYVMSCFRLPKAITSKLTSAVAKFWWSSNG---DSRG 619
Query: 238 ISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGGR 297
+ W++W +C K GGLG R + +FN ALL K WR++ L+ +V RY
Sbjct: 620 MHWMAWDKLCSSKSDGGLGFRNVDDFNSALLAKQLWRLITAPDSLFAKVFKGRYFRKSNP 679
Query: 298 LE-VGGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
L+ + S S W + R V G +++RVG G S W+D
Sbjct: 680 LDSIKSYSPSYGWRSMISARSLVYKG--------LIKRVGSGASISVWND 721
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 184 bits (468), Expect = 1e-46
Identities = 122/358 (34%), Positives = 186/358 (51%), Gaps = 32/358 (8%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W WI CV++ + SVL+NG P RG+RQGDPLSP LF+L E L ++ +A
Sbjct: 580 WISWIMNCVTSVSYSVLINGQPYGHIIPTRGIRQGDPLSPALFVLCTEALIHILNKAEQA 639
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
+G + V+HL FADDTLL+ + L L + +SG +N +KS
Sbjct: 640 GKITGIQF-QDKKVSVNHLLFADDTLLMCKATKQECEELMQCLSQYGQLSGQMINLNKSA 698
Query: 121 LV-GVNIG---SSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLS 176
+ G N+ W+ ++ S +S + G +L L + G+ R L + I +++SRL+
Sbjct: 699 ITFGKNVDIQIKDWI-KSRSGISLEGGTGKYLGLPECLSGSKRDLFGF--IKEKLQSRLT 755
Query: 177 GWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNR 236
GW ++ LS GG+ VLLKS+ +LPVY +S FK P + + ++ ++F+ + R
Sbjct: 756 GWYAKTLSQGGKEVLLKSIALALPVYVMSCFKLPKNLCQKLTTVMMDFWWNS---MQQKR 812
Query: 237 KISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGG 296
KI WLSWQ + L K+ GG G + L+ FN ALL K WR+L ++G L+ RV +RY
Sbjct: 813 KIHWLSWQRLTLPKDQGGFGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNSD 872
Query: 297 RLEV--GGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRR-----VGDGVVTSFWSDR 347
L G R W S + FG ++L + +G+G T W+D+
Sbjct: 873 FLSATRGSRPSYAWRSIL--------------FGRELLMQGLRTVIGNGQKTFVWTDK 916
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 175 bits (444), Expect = 6e-44
Identities = 123/355 (34%), Positives = 178/355 (49%), Gaps = 23/355 (6%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W +WI +CV + +L NG TD F RGLRQGDPLSP+LF+L E L +++S + A
Sbjct: 1141 WVQWIMKCVEGPSMRLLWNGEKTDAFKPLRGLRQGDPLSPYLFVLCIERLCHLIESSIAA 1200
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
+ + I + P +SH+ FADD +L S +R LR L F SG KV+ KS
Sbjct: 1201 KKWKPIKISQSGP-RLSHICFADDLILFAEASIDQIRVLRGVLEKFCGASGQKVSLEKSK 1259
Query: 121 L-----VGVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRL 175
+ V ++G E+ + +G YL +PI + ++ R SRL
Sbjct: 1260 IYFSKNVLRDLGKRISEESGMKATKDLG----KYLGVPILQKRINKETFGEVIKRFSSRL 1315
Query: 176 SGWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDN 235
+GW R LSF GRL L K+VLTS+ V+ +S K P + +D + F G + +
Sbjct: 1316 AGWKGRMLSFAGRLTLTKAVLTSILVHTMSTIKLPQSTLDGLDKVSRAFL---WGSTLEK 1372
Query: 236 RKISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARY--GE 293
+K ++W VCL + GGLG+R N AL+ K WR+L D LW +V+ ++Y G+
Sbjct: 1373 KKQHLVAWTRVCLPRREGGLGIRSATAMNKALIAKVGWRVLNDGSSLWAQVVRSKYKVGD 1432
Query: 294 LGGR-LEVGGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSDR 347
+ R V + S W V GVG W + +GDG FW+DR
Sbjct: 1433 VHDRNWTVAKSNWSSTWRSV-----GVGLRDVIWREQHWV--IGDGRQIRFWTDR 1480
>At1g24640 hypothetical protein
Length = 1270
Score = 175 bits (444), Expect = 6e-44
Identities = 119/351 (33%), Positives = 174/351 (48%), Gaps = 20/351 (5%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W WI CVS+ T SVL+N P L+RGLRQGDPLSPFLF+L EGL ++
Sbjct: 565 WVNWIMVCVSSVTYSVLINDCPFGLIILQRGLRQGDPLSPFLFVLCTEGLTHLLNKAQWE 624
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
G P +V HL FADD+L L S L+ L ++ +G +N +KS
Sbjct: 625 GALEGIQFSENGP-MVHHLLFADDSLFLCKASREQSLVLQKILKVYGNATGQTINLNKS- 682
Query: 121 LVGVNIGSSWLYEAASVLSCKVGIV----PFLYLDMPIGGNPRRLCFWEPIVNRIKSRLS 176
+ G + + +GI YL +P + ++ + +R+K +L
Sbjct: 683 --SITFGEKVDEQLKGTIRTCLGIFTEGGAGTYLGLPECFSGSKVDMLHYLKDRLKEKLD 740
Query: 177 GWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNR 236
W +R LS GG+ VLLKSV ++PV+A+S FK P +++S +F+ + +R
Sbjct: 741 VWFTRCLSQGGKEVLLKSVALAMPVFAMSCFKLPITTCENLESAMASFW---WDSCDHSR 797
Query: 237 KISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGG 296
KI W SW+ +CL K+ GGLG R ++ FN ALL K WR+L L R+L +RY +
Sbjct: 798 KIHWQSWERLCLPKDSGGLGFRDIQSFNQALLAKQAWRLLHFPDCLLSRLLKSRYFDATD 857
Query: 297 RLEVG-GRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
L+ + S W + R+ + G + +RVGDG W D
Sbjct: 858 FLDAALSQRPSFGWRSILFGRELLSKG--------LQKRVGDGASLFVWID 900
>At2g17610 putative non-LTR retroelement reverse transcriptase
Length = 773
Score = 170 bits (430), Expect = 2e-42
Identities = 100/295 (33%), Positives = 156/295 (51%), Gaps = 11/295 (3%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W WI C++T T S+L+NG P KRG+RQGDP+SP+L+LL EGL+ ++++ ++A
Sbjct: 57 WCNWIMTCITTTTYSILINGQPVRRIIPKRGIRQGDPISPYLYLLCTEGLSALIQASIKA 116
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
+ G+ P +SHL FA D+L+ + L L L+E SG VNF KS
Sbjct: 117 KQLHGFKASRNGP-AISHLLFAHDSLVFCKATLEECMTLVNVLKLYEKASGQAVNFQKSA 175
Query: 121 LVGVNIGSSWLYEAASVLSCKVGIVPF----LYLDMPIGGNPRRLCFWEPIVNRIKSRLS 176
++ G + + LS +GI YL +P + + I + ++
Sbjct: 176 IL---FGKGLDFRTSEQLSQLLGIYKTEGFGRYLGLPEFVGRNKTNAFSFIAQTMDQKMD 232
Query: 177 GWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNR 236
W ++ LS G+ VL+KS++T++P Y++S F P +I I S F+ ++
Sbjct: 233 NWYNKLLSPAGKEVLIKSIVTAIPTYSMSCFLLPMRLIHQITSAMRWFW---WSNTKVKH 289
Query: 237 KISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARY 291
KI W++W + K+ GGL +R LK+FN+ALL K WR+L L RV A+Y
Sbjct: 290 KIPWVAWSKLNDPKKMGGLAIRDLKDFNIALLAKQSWRILQQPFSLMARVFKAKY 344
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 169 bits (429), Expect = 3e-42
Identities = 121/355 (34%), Positives = 176/355 (49%), Gaps = 23/355 (6%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W I V+ + SVL NG TD F RGLRQGDPLSP+LF+L E L ++++ V
Sbjct: 465 WTSRIMAGVTDPSMSVLWNGERTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIEASVGK 524
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
+ + ++ +SH+ FADD +L S A +R +R L F SG KV+ KS
Sbjct: 525 REWKPIAVSCGGS-KLSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQKVSLEKSK 583
Query: 121 LV---GVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSG 177
+ V+ L S + C + YL MPI + ++ R+ +RL+G
Sbjct: 584 IFFSHNVSREMEQLISEESGIGCTKELGK--YLGMPILQKRMNKETFGEVLERVSARLAG 641
Query: 178 WNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRK 237
W R LS GR+ L K+VL+S+PV+ +S P + ++D F G + + +K
Sbjct: 642 WKGRSLSLAGRITLTKAVLSSIPVHVMSAILLPVSTLDTLDRYSRTFLWGS---TMEKKK 698
Query: 238 ISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGG- 296
LSW+ +C K GG+G+R ++ N AL+ K WR+L D+ LW RV+ +Y ++GG
Sbjct: 699 QHLLSWRKICKPKAEGGIGLRSARDMNKALVAKVGWRLLQDKESLWARVVRKKY-KVGGV 757
Query: 297 ----RLEVGGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSDR 347
L+ R S W S LR+ V G GW GDG FW DR
Sbjct: 758 QDTSWLKPQPRWSSTWRSVAVGLREVVVK-GVGWV-------PGDGCTIRFWLDR 804
>At1g25430 hypothetical protein
Length = 1213
Score = 169 bits (429), Expect = 3e-42
Identities = 118/347 (34%), Positives = 174/347 (50%), Gaps = 21/347 (6%)
Query: 4 WIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLF 63
WI +C+ST T +V +NG F +GLRQGDPLSP+LF+LA E + ++ S E+ L
Sbjct: 622 WISQCISTPTFTVSINGGNGGFFKSTKGLRQGDPLSPYLFVLAMEAFSNLLHSRYESGLI 681
Query: 64 SGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKS--ML 121
Y A+N + +SHL FADD ++ ++ + L F + SGLKVN KS L
Sbjct: 682 H-YHPKASN-LSISHLMFADDVMIFFDGGSFSLHGICETLDDFASWSGLKVNKDKSHLYL 739
Query: 122 VGVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWNSR 181
G+N S A + +G +P YL +P+ R+ +EP++ +I +R W ++
Sbjct: 740 AGLNQLES---NANAAYGFPIGTLPIRYLGLPLMNRKLRIAEYEPLLEKITARFRSWVNK 796
Query: 182 FLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISWL 241
LSF GR+ L+ SV+ + +S F P G I I+SL F G K+SW
Sbjct: 797 CLSFAGRIQLISSVIFGSINFWMSTFLLPKGCIKRIESLCSRFLWSGNIEQAKGIKVSWA 856
Query: 242 SWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGGRLEV- 300
+ +CL K GGLG+R+L E+N L + WR+ V + LW + G V
Sbjct: 857 A---LCLPKSEGGLGLRRLLEWNKTLSMRLIWRLFVAKDSLWADWQHLHHLSRGSFWAVE 913
Query: 301 GGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVL-RRVGDGVVTSFWSD 346
GG+S S W + LR Q L +VG+G+ +W D
Sbjct: 914 GGQSDSWTWKRLLSLRP---------LAHQFLVCKVGNGLKADYWYD 951
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 164 bits (415), Expect = 1e-40
Identities = 117/333 (35%), Positives = 164/333 (49%), Gaps = 38/333 (11%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W K I ECV+ S+L NG TD F +RGLRQGDP+SP+LF+L E L +++ V
Sbjct: 417 WIKRIMECVAGPEMSLLWNGEKTDSFTPERGLRQGDPISPYLFVLCIERLCHQIETAVGR 476
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
+ SI P VSH+ FADD +L S A +L + + F + + + +
Sbjct: 477 GDWKSISISQGGP-KVSHVCFADDLILFAEASVAQKVSLEKSKIFFS--NNVSRDLEGLI 533
Query: 121 LVGVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWNS 180
IGS+ ++G YL MP+ + ++ R+ SRLSGW S
Sbjct: 534 TAETGIGSTR----------ELG----KYLGMPVLQKRINKDTFGEVLERVSSRLSGWKS 579
Query: 181 RFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISW 240
R LS GR+ L K+VL S+P++ +S P ++ +D + NF G + + RK
Sbjct: 580 RSLSLAGRITLTKAVLMSIPIHTMSSILLPASLLEQLDKVSRNFL---WGSTVEKRKQHL 636
Query: 241 LSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGGRLEV 300
LSW+ VC K GGLG+R K+ N ALL K WR+L D+ LW RVL +Y +V
Sbjct: 637 LSWKKVCRPKAAGGLGLRASKDMNRALLAKVGWRLLNDKVSLWARVLRRKY-------KV 689
Query: 301 GGRSVSCW------WSEVRR-----LRDGVGDG 322
S W WS R LR+GV G
Sbjct: 690 TDVHDSSWLVPKATWSSTWRSIGVGLREGVAKG 722
>At2g05200 putative non-LTR retroelement reverse transcriptase
Length = 1229
Score = 164 bits (415), Expect = 1e-40
Identities = 111/348 (31%), Positives = 168/348 (47%), Gaps = 14/348 (4%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W W+ ECV++ + S L+NG+P + RGLRQGDPLSP LF+L E L+ +
Sbjct: 497 WIDWVLECVTSVSYSFLINGTPQGKVVPTRGLRQGDPLSPCLFILCTEVLSGLCTRAQRL 556
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
+ G + P V +HL FADDT+ + L L + SG +NFHKS
Sbjct: 557 RQLPGVRVSINGPRV-NHLLFADDTMFFSKSDPESCNKLSEILSRYGKASGQSINFHKSS 615
Query: 121 LV-GVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWN 179
+ S + +L + YL +P R+ + I+++I+ + W
Sbjct: 616 VTFSSKTPRSVKGQVKRILKIRKEGGTGKYLGLPEHFGRRKRDIFGAIIDKIRQKSHSWA 675
Query: 180 SRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKIS 239
SRFLS G+ V+LK+VL S+P+Y++S FK P+ + I SL F+ D RK S
Sbjct: 676 SRFLSQAGKQVMLKAVLASMPLYSMSCFKLPSALCRKIQSLLTRFW---WDTKPDVRKTS 732
Query: 240 WLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGGRLE 299
W++W + K GGLG R ++ N +LL K WR+L L R+L+ +Y +E
Sbjct: 733 WVAWSKLTNPKNAGGLGFRDIERCNDSLLAKLGWRLLNSPESLLSRILLGKYCHSSSFME 792
Query: 300 VGGRS-VSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
S S W + R+ + + G GW + +G S W+D
Sbjct: 793 CKLPSQPSHGWRSIIAGREILKE-GLGWL-------ITNGEKVSIWND 832
>At2g11240 pseudogene
Length = 1044
Score = 163 bits (413), Expect = 2e-40
Identities = 111/351 (31%), Positives = 173/351 (48%), Gaps = 20/351 (5%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLN-IVMKSLVE 59
W WI +C++T + S L+NGS +RGLRQGDPLSPFLF++ +E L+ + K+ ++
Sbjct: 428 WISWILQCITTVSYSFLLNGSAQGAVTPERGLRQGDPLSPFLFIICSEVLSGLCRKAQLD 487
Query: 60 AQLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKS 119
L G + NP V+HL FADDT+ + + L +E SG +N KS
Sbjct: 488 GSLL-GLRVSKGNP-RVNHLLFADDTIFFCRSDLKSCKTFLCILKKYEEASGQMINKSKS 545
Query: 120 MLVGVNIGSSWL-YEAASVLSCKV--GIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLS 176
+ + EA +L ++ G+ +L L G R L + IV+RI+ R
Sbjct: 546 AITFSRKTPDHIKTEAQQILGIQLVGGLGKYLGLPKMFGRKKRDL--FNQIVDRIRQRSL 603
Query: 177 GWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNR 236
W+SRFLS G+ +LKSVL S+P Y +S FK + I S +F+ S D +
Sbjct: 604 SWSSRFLSTAGKTTMLKSVLASMPTYTMSCFKLLVSLCKRIQSALTHFW---WDSSADKK 660
Query: 237 KISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGG 296
K+ W++W + K+ GGLG + + FN ALL K WR++ + R+L+ +Y
Sbjct: 661 KMCWIAWSKMAKNKKEGGLGFKDITNFNDALLAKLSWRIVQSPSCVLVRILLGKYCRTSS 720
Query: 297 RLEVGGRSVSC-WWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
L+ + S W + G Q+ + +G G+ T W++
Sbjct: 721 FLDCSVTAASSHGWRGICT--------GKDLIKSQLGKVIGSGLDTLVWNE 763
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 162 bits (410), Expect = 5e-40
Identities = 105/351 (29%), Positives = 172/351 (48%), Gaps = 20/351 (5%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W + I ECV + VL+NG+P E RGLRQGDPLSP+LF++ E L +++S +
Sbjct: 602 WIRLIMECVKSVRYQVLINGTPHGEIIPSRGLRQGDPLSPYLFVICTEMLVKMLQSAEQK 661
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
+G + P +SHL FADD++ + + + + + SG +VN+ KS
Sbjct: 662 NQITGLKVARGAP-PISHLLFADDSMFYCKVNDEALGQIIRIIEEYSLASGQRVNYLKS- 719
Query: 121 LVGVNIGSSWLYEAASVLSCKVGIV----PFLYLDMPIGGNPRRLCFWEPIVNRIKSRLS 176
+ G E ++ K+GI +YL +P ++ + +R+ ++
Sbjct: 720 --SIYFGKHISEERRCLVKRKLGIEREGGEGVYLGLPESFQGSKVATLSYLKDRLGKKVL 777
Query: 177 GWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNR 236
GW S FLS GG+ +LLK+V +LP Y +S FK P I I+S+ F+ ++ R
Sbjct: 778 GWQSNFLSPGGKEILLKAVAMALPTYTMSCFKIPKTICQQIESVMAEFW---WKNKKEGR 834
Query: 237 KISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGG 296
+ W +W + K GGLG ++++ FN+ALLGK WRM+ ++ L +V +RY
Sbjct: 835 GLHWKAWCHLSRPKAVGGLGFKEIEAFNIALLGKQLWRMITEKDSLMAKVFKSRYFSKSD 894
Query: 297 RLEVG-GRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
L G S W + + + G + +G+G + W+D
Sbjct: 895 PLNAPLGSRPSFAWKSIYEAQVLIKQG--------IRAVIGNGETINVWTD 937
>At2g16680 putative non-LTR retroelement reverse transcriptase
Length = 1319
Score = 161 bits (408), Expect = 9e-40
Identities = 108/352 (30%), Positives = 178/352 (49%), Gaps = 20/352 (5%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W WI CV++ T SVL+NG +RG+RQGDP+SPFLF+L E L +++ +
Sbjct: 554 WISWIMGCVTSVTYSVLINGQHFGHITPERGIRQGDPISPFLFVLCTEALIHILQQAENS 613
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
+ SG + P V+HL F DDT L+ + ++ + L + +SG +N KS
Sbjct: 614 KKVSGIQFNGSGP-SVNHLLFVDDTQLVCRATKSDCEQMMLCLSQYGHISGQLINVEKSS 672
Query: 121 LV-GVNIGSS---WLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLS 176
+ GV + W+ + S + + G +L L + G+ + L + I +++S LS
Sbjct: 673 ITFGVKVDEDTKRWI-KNRSGIHLEGGTGKYLGLPENLSGSKQDLFGY--IKEKLQSHLS 729
Query: 177 GWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNR 236
GW + LS GG+ +LLKS+ +LPVY ++ F+ P G+ + + S+ ++F+ E +
Sbjct: 730 GWYDKTLSQGGKEILLKSIALALPVYIMTCFRLPKGLCTKLTSVMMDFW---WNSMEFSN 786
Query: 237 KISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGG 296
KI W+ + + L K GG G + L+ FN ALL K WR+ D + ++ +RY
Sbjct: 787 KIHWIGGKKLTLPKSLGGFGFKDLQCFNQALLAKQAWRLFSDSKSIVSQIFKSRYFMNTD 846
Query: 297 RLEV-GGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSDR 347
L G S W + R+ + G + R +G+G T+ W D+
Sbjct: 847 FLNARQGTRPSYTWRSILYGRELLNGG--------LKRLIGNGEQTNVWIDK 890
>At2g05980 putative non-LTR retroelement reverse transcriptase
Length = 1352
Score = 160 bits (405), Expect = 2e-39
Identities = 113/349 (32%), Positives = 177/349 (50%), Gaps = 23/349 (6%)
Query: 4 WIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVM-KSLVEAQL 62
WI+ C+ TA+ SV VNG + F +RGLRQG LSP+L+++ L+ ++ K+ VE ++
Sbjct: 775 WIELCIGTASFSVQVNGELSGFFRSERGLRQGCSLSPYLYVICMNVLSCMLDKAAVEKKI 834
Query: 63 FSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLV 122
Y N + ++HL FADD ++ + +++ A F AMS LK++ KS +
Sbjct: 835 --SYHPRCRN-MNLTHLCFADDIMVFSDGTSKSIQGTLAIFEKFAAMSWLKISLEKSTIF 891
Query: 123 GVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWNSRF 182
I + ++G +P YL +P+ + P+V +I++R++ W +RF
Sbjct: 892 MAGISPNAKTSILQQFPFELGTLPVKYLGLPLLTKRMTQSDYLPLVEKIRARITSWTNRF 951
Query: 183 LSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISWLS 242
LSF GRL L+KSVL+S+ + LS F+ P + I+ +F F G + N K + ++
Sbjct: 952 LSFAGRLQLIKSVLSSITNFWLSVFRLPKACLQEIEKMFSAFL---WSGPDLNTKKAKIA 1008
Query: 243 WQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVL---VARYGELGGRLE 299
W VC KE GGLG++ LKE N L K WR+L R LW + + + R E
Sbjct: 1009 WSEVCKLKEEGGLGLKPLKEANEVSLLKLIWRILSARDSLWVKWVNKHLIRKETFWSVKE 1068
Query: 300 VGGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRR--VGDGVVTSFWSD 346
G S W ++ + RD ++ R V G TSFW D
Sbjct: 1069 NTGLG-SWLWRKILKQRDKA----------RLFHRMEVRSGTFTSFWHD 1106
>At4g09710 RNA-directed DNA polymerase -like protein
Length = 1274
Score = 158 bits (400), Expect = 8e-39
Identities = 112/352 (31%), Positives = 169/352 (47%), Gaps = 28/352 (7%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W +W+ +CV T + S L+NGSP RGLRQGDPLSP+LF+L E L+ + + E
Sbjct: 543 WIRWVMQCVCTVSYSFLINGSPQGSVVPSRGLRQGDPLSPYLFILCTEVLSGLCRKAQEK 602
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
+ G + +P V+HL FADDT+ + AL L +E SG +N KS
Sbjct: 603 GVMVGIRVARGSP-QVNHLLFADDTMFFCKTNPTCCGALSNILKKYELASGQSINLAKSA 661
Query: 121 LVGVNIGSSWLYEAASVLSCKV----GIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLS 176
+ + + LS ++ GI YL +P R+ + IV+RI+ R
Sbjct: 662 ITFSSKTPQDIKRRVK-LSLRIDNEGGIGK--YLGLPEHFGRRKRDIFSSIVDRIRQRSH 718
Query: 177 GWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNR 236
W+ RFLS G+ +LLK+VL+S+P YA+ FK P + I S+ F+ D R
Sbjct: 719 SWSIRFLSSAGKQILLKAVLSSMPSYAMMCFKLPASLCKQIQSVLTRFW---WDSKPDKR 775
Query: 237 KISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGG 296
K++W+SW + L GGLG R+++ K WR+L + L RVL+ +Y
Sbjct: 776 KMAWVSWDKLTLPINEGGLGFREIE-------AKLSWRILKEPHSLLSRVLLGKYCNTSS 828
Query: 297 RLEVGGRS--VSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
++ S W + RD + G GW +G G + W++
Sbjct: 829 FMDCSASPSFASHGWRGILAGRDLLRK-GLGW-------SIGQGDSINVWTE 872
>At1g31030 F17F8.5
Length = 872
Score = 158 bits (399), Expect = 1e-38
Identities = 115/355 (32%), Positives = 163/355 (45%), Gaps = 40/355 (11%)
Query: 4 WIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLF 63
WI C++TA+ SV VNG F KRGLRQG LSP+LF++ + L+ ++ + F
Sbjct: 275 WINLCITTASFSVQVNGDLVGYFQSKRGLRQGCSLSPYLFVICMDVLSKMLDKAAGVRKF 334
Query: 64 SGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLVG 123
+ + ++HL FADD ++L ++ + F SGL+++ KS L
Sbjct: 335 GFHP--KCQRLGLTHLSFADDLMVLSDGKTRSIEGILEVFDEFCKRSGLRISLEKSTLYM 392
Query: 124 VNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWNSRFL 183
+ E A+ VG +P YL +P+ + P++ +IK R++ W RF
Sbjct: 393 AGVSPIIKQEIAAKFLFDVGQLPVRYLGLPLVTKRLTSADYSPLLEQIKKRIATWTFRFF 452
Query: 184 SFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISWLSW 243
SF GR L+KSVL S+ + L+ F+ P I ID L +F G S KI SW
Sbjct: 453 SFAGRFNLIKSVLWSICNFWLAAFRLPRQCIREIDKLCSSFLWSGSEMSSHKAKI---SW 509
Query: 244 QTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARY------------ 291
VC K GGLG+R LKE N K WR++ + LW + VA Y
Sbjct: 510 DIVCKPKAEGGLGLRNLKEANDVSCLKLVWRIISNSNSLWTK-WVAEYLIRKKSIWSLKQ 568
Query: 292 ------------------GELGGRLEVG-GRSVSCW---WSEVRRLRDGVGDGGS 324
+ R+EVG G S S W WS RL D VGD G+
Sbjct: 569 STSMGSWIWRKILKIRDVAKSFSRVEVGNGESASFWYDHWSAHGRLIDTVGDKGT 623
>At2g41580 putative non-LTR retroelement reverse transcriptase
Length = 1094
Score = 156 bits (394), Expect = 4e-38
Identities = 100/295 (33%), Positives = 153/295 (50%), Gaps = 11/295 (3%)
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W W+ CVS+ + SVL+NG P RGLRQGDPLSPFLF+L E L ++ +
Sbjct: 331 WINWMMACVSSVSYSVLINGQPFGHITPHRGLRQGDPLSPFLFVLCTEALIHILNQAEKI 390
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
SG P V+HL FADDTLL+ S + L + +SG +N KS
Sbjct: 391 GKISGIQFNGTGP-SVNHLLFADDTLLICKASQLECAEIMHCLSQYGHISGQMINSEKSA 449
Query: 121 LV-GVNIG---SSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLS 176
+ G + W+ + + + G +L L G+ + L + I +++SRLS
Sbjct: 450 ITFGAKVNEETKQWIMNRSGI-QTEGGTGKYLGLPECFQGSKQVLFGF--IKEKLQSRLS 506
Query: 177 GWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNR 236
GW ++ LS GG+ +LLKS+ + PVYA++ F+ + + + S+ ++F+ +D +
Sbjct: 507 GWYAKTLSQGGKDILLKSIAMAFPVYAMTCFRLSKTLCTKLTSVMMDFW---WNSVQDKK 563
Query: 237 KISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARY 291
KI W+ Q + L K GG G + L+ FN ALL K R+ D L ++L +RY
Sbjct: 564 KIHWIGAQKLMLPKFLGGFGFKDLQCFNQALLAKQASRLHTDSDSLLSQILKSRY 618
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.348 0.155 0.570
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,300,949
Number of Sequences: 26719
Number of extensions: 553767
Number of successful extensions: 2418
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 63
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 2167
Number of HSP's gapped (non-prelim): 101
length of query: 624
length of database: 11,318,596
effective HSP length: 105
effective length of query: 519
effective length of database: 8,513,101
effective search space: 4418299419
effective search space used: 4418299419
T: 11
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC134967.10


