

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.24 + phase: 0
(456 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
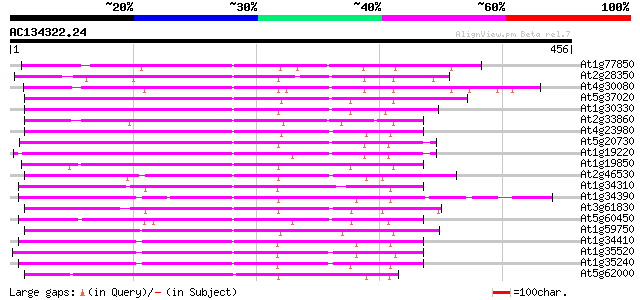
Score E
Sequences producing significant alignments: (bits) Value
At1g77850 putative auxin response factor protein (T32E8.16) 262 3e-70
At2g28350 unknown protein 239 3e-63
At4g30080 transcription factor like protein 235 3e-62
At5g37020 auxin response factor 8 (ARF8) 177 8e-45
At1g30330 putative protein 174 9e-44
At2g33860 auxin response transcription factor 3 (ETTIN/ARF3) 173 2e-43
At4g23980 auxin response factor 9 (ARF9) 171 6e-43
At5g20730 auxin response factor 7 (ARF7) 171 8e-43
At1g19220 auxin response factor (ARF) like protein 166 2e-41
At1g19850 transcription factor monopteros (MP) 162 3e-40
At2g46530 ARF1 family auxin responsive transcription factor like... 161 6e-40
At1g34310 hypothetical protein 159 4e-39
At1g34390 auxin response factor, putative 158 7e-39
At3g61830 auxin response factor-like protein 157 1e-38
At5g60450 auxin response factor 4 157 1e-38
At1g59750 auxin response factor 1 155 4e-38
At1g34410 auxin response factor, putative 153 2e-37
At1g35520 auxin response factor, putative 152 4e-37
At1g35240 hypothetical protein 152 5e-37
At5g62000 auxin response factor - like protein 149 2e-36
>At1g77850 putative auxin response factor protein (T32E8.16)
Length = 585
Score = 262 bits (670), Expect = 3e-70
Identities = 159/439 (36%), Positives = 231/439 (52%), Gaps = 74/439 (16%)
Query: 10 VRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVS 69
V P IW CA A+V+IP LHSRVYYFPQGH+E+ P S++ + S + CI++
Sbjct: 16 VDPTIWRACAGASVQIPVLHSRVYYFPQGHVEHCCPLLSTLPSSTSPVP-------CIIT 68
Query: 70 AVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVA----NLNDRNEVVSFVKTLTRSDV 125
++ LLADP TDEVF L+L P+T +++D N+V +F K LT SD
Sbjct: 69 SIQLLADPVTDEVFAHLILQPMTQQQFTPTNYSRFGRFDGDVDDNNKVTTFAKILTPSDA 128
Query: 126 NNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISE 185
NN F +PRFCAD+VFP L+ + + Q L+VTD+HG V F H+ RG P+R++L +
Sbjct: 129 NNGGGFSVPRFCADSVFPLLNFQIDPPVQKLYVTDIHGAVWDFRHIYRGTPRRHLL-TTG 187
Query: 186 WNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRRN-----------------TQFVAAAAEQ 228
W+ FV KKL+AGDSV+FM+ S ++F+G+RR + + ++
Sbjct: 188 WSKFVNSKKLIAGDSVVFMRKSADEMFIGVRRTPISSSDGGSSYYGGDEYNGYYSQSSVA 247
Query: 229 KKDE---------------LEKAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDE 273
K+D+ +AV +A+ A + FE+V+YP W +FVV V+
Sbjct: 248 KEDDGSPKKTFRRSGNGKLTAEAVTDAINRASQGLPFEVVFYPAA-GWSEFVVRAEDVES 306
Query: 274 SMKIQWNPRMRVK--MKTDKSSRIP-YQGTISIVSRTSNLWR-----MLQVNWDEFQVSQ 325
SM + W P RVK M+T+ SSRI +QG +S + + WR LQ+ WDE ++ Q
Sbjct: 307 SMSMYWTPGTRVKMAMETEDSSRITWFQGIVSSTYQETGPWRGSPWKQLQITWDEPEILQ 366
Query: 326 IPRRVNPWWVELISH-KPAPTPFPQTKKFRTTQ--------------------SSAQLSD 364
+RVNPW VE+ +H TPFP K+ + Q SSA D
Sbjct: 367 NVKRVNPWQVEIAAHATQLHTPFPPAKRLKYPQPGGGFLSGDDGEILYPQSGLSSAAAPD 426
Query: 365 KKETLLNGDGFPVDIQRVR 383
++ + FP +Q R
Sbjct: 427 PSPSMFSYSTFPAGMQGAR 445
>At2g28350 unknown protein
Length = 693
Score = 239 bits (609), Expect = 3e-63
Identities = 156/416 (37%), Positives = 217/416 (51%), Gaps = 76/416 (18%)
Query: 5 KQPSHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFR--P 62
+Q + P++WH CA + V+IP L+S V+YF QGH E+A H + R P
Sbjct: 2 EQEKSLDPQLWHACAGSMVQIPSLNSTVFYFAQGHTEHA--------HAPPDFHAPRVPP 53
Query: 63 FTLCIVSAVDLLADPHTDEVFVKLLLTPIT-NDVHLEN-------PKEEVANLNDRNEVV 114
LC V +V LAD TDEVF K+ L P+ ND+ LEN P N N + +
Sbjct: 54 LILCRVVSVKFLADAETDEVFAKITLLPLPGNDLDLENDAVLGLTPPSSDGNGNGKEKPA 113
Query: 115 SFVKTLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRG 174
SF KTLT+SD NN F +PR+CA+ +FP+LD AE Q + D+HGE KF H+ RG
Sbjct: 114 SFAKTLTQSDANNGGGFSVPRYCAETIFPRLDYSAEPPVQTVIAKDIHGETWKFRHIYRG 173
Query: 175 FPKRNMLYISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRR----------------- 217
P+R++L + W++FV +KKL+AGDS++F++ +G + VGIRR
Sbjct: 174 TPRRHLL-TTGWSTFVNQKKLIAGDSIVFLRSESGDLCVGIRRAKRGGLGSNAGSDNPYP 232
Query: 218 -------------------------NTQFVAAAAEQKKDELEKAVMEALKLAEENKAFEI 252
N AAA + + E AV EA+ A +AFE+
Sbjct: 233 GFSGFLRDDESTTTTSKLMMMKRNGNNDGNAAATGRVRVE---AVAEAVARAACGQAFEV 289
Query: 253 VYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKSSRIP-YQGTISIVSRTSN 309
VYYP+ +F V V +M+I+W MR KM +T+ SSRI + GT+S V
Sbjct: 290 VYYPRAST-PEFCVKAADVRSAMRIRWCSGMRFKMAFETEDSSRISWFMGTVSAVQVADP 348
Query: 310 L------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFRTTQ 357
+ WR+LQV WDE + Q +RV+PW VEL+S+ P +PF KK R Q
Sbjct: 349 IRWPNSPWRLLQVAWDEPDLLQNVKRVSPWLVELVSNMPTIHLSPFSPRKKIRIPQ 404
>At4g30080 transcription factor like protein
Length = 670
Score = 235 bits (600), Expect = 3e-62
Identities = 173/519 (33%), Positives = 245/519 (46%), Gaps = 107/519 (20%)
Query: 12 PEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAV 71
P++WH CA V++P ++S+V+YFPQGH ENA P LC V A+
Sbjct: 18 PQLWHACAGGMVRMPPMNSKVFYFPQGHAENAYDCVDFGN------LPIPPMVLCRVLAI 71
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLN----DRNEVVSFVKTLTRSDVNN 127
+AD +DEVF KL L P+ +D ++++ + + N + + SF KTLT+SD NN
Sbjct: 72 KYMADAESDEVFAKLRLIPLKDDEYVDHEYGDGEDSNGFESNSEKTPSFAKTLTQSDANN 131
Query: 128 ARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWN 187
F +PR+CA+ +FP+LD AE Q + DVHG+V KF H+ RG P+R++L + W+
Sbjct: 132 GGGFSVPRYCAETIFPRLDYNAEPPVQTILAKDVHGDVWKFRHIYRGTPRRHLL-TTGWS 190
Query: 188 SFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--------NTQFVA---------------- 223
+FV +KKLVAGDS++FM+ G + VGIRR ++ A
Sbjct: 191 NFVNQKKLVAGDSIVFMRAENGDLCVGIRRAKRGGIGNGPEYSAGWNPIGGSCGYSSLLR 250
Query: 224 ------------AAAEQKKDELEKAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVV 271
+ A++K ++V+EA LA + FE+VYYP+ +F V
Sbjct: 251 EDESNSLRRSNCSLADRKGKVTAESVIEAATLAISGRPFEVVYYPRAST-SEFCVKALDA 309
Query: 272 DESMKIQWNPRMRVKM--KTDKSSRIP-YQGTISIVSRTSNL------WRMLQVNWDEFQ 322
+M+I W MR KM +T+ SSRI + GT+S V+ + + WR+LQV WDE
Sbjct: 310 RAAMRIPWCSGMRFKMAFETEDSSRISWFMGTVSAVNVSDPIRWPNSPWRLLQVAWDEPD 369
Query: 323 VSQIPRRVNPWWVELISH-KPAP-TPF-PQTKKFRTTQ---------SSAQLSDKKETLL 370
+ Q +RVNPW VEL+S+ P P T F P KK R Q S S L+
Sbjct: 370 LLQNVKRVNPWLVELVSNVHPIPLTSFSPPRKKMRLPQHPDYNNLINSIPVPSFPSNPLI 429
Query: 371 NG-------DGFPVDIQRVRHDLVSISGPIHS---HIILNSSETKFP------------- 407
D PV +Q RH+ G S H LN P
Sbjct: 430 RSSPLSSVLDNVPVGLQGARHNAHQYYGLSSSDLHHYYLNRPPPPPPPSSLQLSPSLGLR 489
Query: 408 ---------------ATHNCNTKNDSDGSIMLFGKIIQP 431
T CN I+LFGK+I P
Sbjct: 490 NIDTKNEKGFCFLTMGTTPCNDTKSKKSHIVLFGKLILP 528
>At5g37020 auxin response factor 8 (ARF8)
Length = 811
Score = 177 bits (450), Expect = 8e-45
Identities = 118/374 (31%), Positives = 189/374 (49%), Gaps = 16/374 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTH-SFLQSFRPFTLCIVSAV 71
E+WH CA V +P SRV YFPQGH E + +++ H S P +C + V
Sbjct: 22 ELWHACAGPLVSLPSSGSRVVYFPQGHSEQVAATTNKEVDGHIPNYPSLPPQLICQLHNV 81
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSF 131
+ AD TDEV+ ++ L P+T + E + + F KTLT SD + F
Sbjct: 82 TMHADVETDEVYAQMTLQPLTPEEQKETFVPIELGIPSKQPSNYFCKTLTASDTSTHGGF 141
Query: 132 HIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVK 191
+PR A+ VFP LD + +Q L D+H KF H+ RG PKR++L + W+ FV
Sbjct: 142 SVPRRAAEKVFPPLDYTLQPPAQELIARDLHDVEWKFRHIFRGQPKRHLL-TTGWSVFVS 200
Query: 192 RKKLVAGDSVIFMKDSTGKIFVGIRRNT--QFVAAAAEQKKDELEKAVMEALKLAE-ENK 248
K+LVAGDSVIF+++ ++F+GIR T Q + ++ D + ++ A A N
Sbjct: 201 AKRLVAGDSVIFIRNEKNQLFLGIRHATRPQTIVPSSVLSSDSMHIGLLAAAAHASATNS 260
Query: 249 AFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTISIVS 305
F + ++P+ +FV+ + +++ +I R R+ +T++SS Y GTI+ +S
Sbjct: 261 CFTVFFHPRASQ-SEFVIQLSKYIKAVFHTRISVGMRFRMLFETEESSVRRYMGTITGIS 319
Query: 306 RTSNL------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFPQTKKFRTTQS 358
++ WR ++V WDE + RV+ W +E ++ P P+ FP K
Sbjct: 320 DLDSVRWPNSHWRSVKVGWDESTAGERQPRVSLWEIEPLTTFPMYPSLFPLRLKRPWHAG 379
Query: 359 SAQLSDKKETLLNG 372
++ L D + L +G
Sbjct: 380 TSSLPDGRGDLGSG 393
>At1g30330 putative protein
Length = 933
Score = 174 bits (441), Expect = 9e-44
Identities = 115/350 (32%), Positives = 177/350 (49%), Gaps = 16/350 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTH-SFLQSFRPFTLCIVSAV 71
E+WH CA V +P + SRV YFPQGH E + S++ H S P +C + V
Sbjct: 23 ELWHACAGPLVSLPPVGSRVVYFPQGHSEQVAASTNKEVDAHIPNYPSLHPQLICQLHNV 82
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSF 131
+ AD TDEV+ ++ L P+ + + R F KTLT SD + F
Sbjct: 83 TMHADVETDEVYAQMTLQPLNAQEQKDPYLPAELGVPSRQPTNYFCKTLTASDTSTHGGF 142
Query: 132 HIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVK 191
+PR A+ VFP LD + +Q L D+H KF H+ RG PKR++L + W+ FV
Sbjct: 143 SVPRRAAEKVFPPLDYSQQPPAQELMARDLHDNEWKFRHIFRGQPKRHLL-TTGWSVFVS 201
Query: 192 RKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVM-EALKLAEENK 248
K+LVAGDSV+F+ + ++ +GIRR Q V ++ D + ++ A A N
Sbjct: 202 AKRLVAGDSVLFIWNDKNQLLLGIRRANRPQTVMPSSVLSSDSMHLGLLAAAAHAAATNS 261
Query: 249 AFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTISIV- 304
F I Y P+ +FV+ +++ ++ R R+ +T++SS Y GTI+ +
Sbjct: 262 RFTIFYNPRASP-SEFVIPLAKYVKAVYHTRVSVGMRFRMLFETEESSVRRYMGTITGIC 320
Query: 305 ----SRTSNL-WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFP 348
+R +N WR ++V WDE + RV+ W +E ++ P P+PFP
Sbjct: 321 DLDPTRWANSHWRSVKVGWDESTAGERQPRVSLWEIEPLTTFPMYPSPFP 370
>At2g33860 auxin response transcription factor 3 (ETTIN/ARF3)
Length = 608
Score = 173 bits (439), Expect = 2e-43
Identities = 112/346 (32%), Positives = 170/346 (48%), Gaps = 33/346 (9%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA + +PK S V YFPQGHLE A S++I P C + V
Sbjct: 54 ELWHACAGPLISLPKRGSLVLYFPQGHLEQAPDFSAAI-------YGLPPHVFCRILDVK 106
Query: 73 LLADPHTDEVFVKLLLTPITNDVH---------LENPKEEVANLNDRNEVVSFVKTLTRS 123
L A+ TDEV+ ++ L P + D+ ++ +E+ L N F KTLT S
Sbjct: 107 LHAETTTDEVYAQVSLLPESEDIERKVREGIIDVDGGEEDYEVLKRSNTPHMFCKTLTAS 166
Query: 124 DVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYI 183
D + F +PR A++ FP LD SQ L D+HG +F H+ RG P+R++L
Sbjct: 167 DTSTHGGFSVPRRAAEDCFPPLDYSQPRPSQELLARDLHGLEWRFRHIYRGQPRRHLL-T 225
Query: 184 SEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRRNTQF---VAAAAEQKKDELEKAVMEA 240
+ W++FV +KKLV+GD+V+F++ GK+ +G+RR +Q A +A+ ++ E
Sbjct: 226 TGWSAFVNKKKLVSGDAVLFLRGDDGKLRLGVRRASQIEGTAALSAQYNQNMNHNNFSEV 285
Query: 241 LKLAEENKAFEIVYYPQGDDWCDFVVDG----NVVDESMKIQWNPRMRVKMKTDKSSRIP 296
+ F I Y P+ W +F++ VVD I + RV+ + R P
Sbjct: 286 AHAISTHSVFSISYNPKA-SWSNFIIPAPKFLKVVDYPFCIGMRFKARVESEDASERRSP 344
Query: 297 YQGTISIVSRTSNL------WRMLQVNWDEFQVSQIPRRVNPWWVE 336
G IS +S + WR L V WD+ + +RV+PW +E
Sbjct: 345 --GIISGISDLDPIRWPGSKWRCLLVRWDDIVANGHQQRVSPWEIE 388
>At4g23980 auxin response factor 9 (ARF9)
Length = 638
Score = 171 bits (434), Expect = 6e-43
Identities = 111/334 (33%), Positives = 168/334 (50%), Gaps = 13/334 (3%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSIT-HTHSFLQSFRPFTLCIVSAV 71
E+W CA V +P+ RVYYFPQGH+E S+ + +T L P LC V V
Sbjct: 12 ELWKLCAGPLVDVPQAQERVYYFPQGHMEQLEASTQQVDLNTMKPLFVLPPKILCNVMNV 71
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSF 131
L A+ TDEV+ ++ L P+ +V + R +V SF K LT SD + F
Sbjct: 72 SLQAEKDTDEVYAQITLIPVGTEVDEPMSPDPSPPELQRPKVHSFSKVLTASDTSTHGGF 131
Query: 132 HIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVK 191
+ R A P LD+ ++ +Q L DVHG KF H+ RG P+R++L + W++FV
Sbjct: 132 SVLRKHATECLPPLDMTQQTPTQELVAEDVHGYQWKFKHIFRGQPRRHLL-TTGWSTFVT 190
Query: 192 RKKLVAGDSVIFMKDSTGKIFVGIRRNT--QFVAAAAEQKKDELEKAVMEALKLAEENKA 249
K+LVAGD+ +F++ G++ VG+RR Q ++ + V+ + A + K
Sbjct: 191 SKRLVAGDTFVFLRGENGELRVGVRRANLQQSSMPSSVISSHSMHLGVLATARHATQTKT 250
Query: 250 FEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMK--TDKSSRIPYQGTISIVSRT 307
IVYY F++ N E+M +++ MR KM+ + S Y GT+ V
Sbjct: 251 MFIVYYKPRTS--QFIISLNKYLEAMSNKFSVGMRFKMRFEGEDSPERRYSGTVIGVKDC 308
Query: 308 S-----NLWRMLQVNWDEFQVSQIPRRVNPWWVE 336
S + WR L+V+WDE P +V+PW +E
Sbjct: 309 SPHWKDSKWRCLEVHWDEPASISRPNKVSPWEIE 342
>At5g20730 auxin response factor 7 (ARF7)
Length = 1164
Score = 171 bits (433), Expect = 8e-43
Identities = 112/350 (32%), Positives = 170/350 (48%), Gaps = 18/350 (5%)
Query: 9 HVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIV 68
++ E+WH CA + +P S V YFPQGH E + S T + +C++
Sbjct: 20 NINSELWHACAGPLISLPPAGSLVVYFPQGHSEQVAASMQKQTDFIPSYPNLPSKLICML 79
Query: 69 SAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNA 128
V L ADP TDEV+ ++ L P+ ++ +R F KTLT SD +
Sbjct: 80 HNVTLNADPETDEVYAQMTLQPVNKYDRDALLASDMGLKLNRQPNEFFCKTLTASDTSTH 139
Query: 129 RSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNS 188
F +PR A+ +FP LD + Q L D+H F H+ RG PKR++L + W+
Sbjct: 140 GGFSVPRRAAEKIFPALDFSMQPPCQELVAKDIHDNTWTFRHIYRGQPKRHLL-TTGWSV 198
Query: 189 FVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAEE 246
FV K+L AGDSV+F++D ++ +GIRR Q +++ D + V+ A A
Sbjct: 199 FVSTKRLFAGDSVLFIRDGKAQLLLGIRRANRQQPALSSSVISSDSMHIGVLAAAAHANA 258
Query: 247 NKA-FEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKSSRIPYQGTISI 303
N + F I Y P+ +FVV ++M Q + MR +M +T++ Y GT++
Sbjct: 259 NNSPFTIFYNPRAAP-AEFVVPLAKYTKAMYAQVSLGMRFRMIFETEECGVRRYMGTVTG 317
Query: 304 VSR------TSNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPAPTPF 347
+S ++ WR LQ+ WDE P RV+ W +E P TPF
Sbjct: 318 ISDLDPVRWKNSQWRNLQIGWDESAAGDRPSRVSVWDIE-----PVLTPF 362
>At1g19220 auxin response factor (ARF) like protein
Length = 1086
Score = 166 bits (421), Expect = 2e-41
Identities = 109/355 (30%), Positives = 172/355 (47%), Gaps = 20/355 (5%)
Query: 4 QKQPSHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPF 63
+K+P + ++WH CA V +P + S V YFPQGH E + S T +
Sbjct: 16 EKKP--INSQLWHACAGPLVSLPPVGSLVVYFPQGHSEQVAASMQKQTDFIPNYPNLPSK 73
Query: 64 TLCIVSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRS 123
+C++ +V L AD TDEV+ ++ L P+ ++ +R F KTLT S
Sbjct: 74 LICLLHSVTLHADTETDEVYAQMTLQPVNKYDREALLASDMGLKLNRQPTEFFCKTLTAS 133
Query: 124 DVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYI 183
D + F +PR A+ +FP LD + +Q + D+H F H+ RG PKR++L
Sbjct: 134 DTSTHGGFSVPRRAAEKIFPPLDFSMQPPAQEIVAKDLHDTTWTFRHIYRGQPKRHLL-T 192
Query: 184 SEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRRNTQFVAAAAEQ--KKDELEKAVMEAL 241
+ W+ FV K+L AGDSV+F++D ++ +GIRR + + D + ++ A
Sbjct: 193 TGWSVFVSTKRLFAGDSVLFVRDEKSQLMLGIRRANRQTPTLSSSVISSDSMHIGILAAA 252
Query: 242 KLAEENKA-FEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKSSRIPYQ 298
A N + F I + P+ +FVV ++++ Q + MR +M +T+ Y
Sbjct: 253 AHANANSSPFTIFFNPRASP-SEFVVPLAKYNKALYAQVSLGMRFRMMFETEDCGVRRYM 311
Query: 299 GTISIVSR------TSNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPAPTPF 347
GT++ +S + WR LQV WDE P RV+ W +E P TPF
Sbjct: 312 GTVTGISDLDPVRWKGSQWRNLQVGWDESTAGDRPSRVSIWEIE-----PVITPF 361
>At1g19850 transcription factor monopteros (MP)
Length = 902
Score = 162 bits (411), Expect = 3e-40
Identities = 106/340 (31%), Positives = 172/340 (50%), Gaps = 15/340 (4%)
Query: 10 VRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSS--SSITHTHSFLQSFRPFTLCI 67
+ E+WH CA V +P++ S VYYF QGH E + S+ S+ T ++ + +C
Sbjct: 51 INSELWHACAGPLVCLPQVGSLVYYFSQGHSEQVAVSTRRSATTQVPNY-PNLPSQLMCQ 109
Query: 68 VSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNN 127
V V L AD +DE++ ++ L P+ ++ + + ++ F KTLT SD +
Sbjct: 110 VHNVTLHADKDSDEIYAQMSLQPVHSERDVFPVPDFGMLRGSKHPTEFFCKTLTASDTST 169
Query: 128 ARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWN 187
F +PR A+ +FP LD A+ +Q L V D+H F H+ RG PKR++L + W+
Sbjct: 170 HGGFSVPRRAAEKLFPPLDYSAQPPTQELVVRDLHENTWTFRHIYRGQPKRHLL-TTGWS 228
Query: 188 SFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAE 245
FV K+L AGDSV+F++D ++ VG+RR Q ++ D + V+ A A
Sbjct: 229 LFVGSKRLRAGDSVLFIRDEKSQLMVGVRRANRQQTALPSSVLSADSMHIGVLAAAAHAT 288
Query: 246 ENKAFEIVYYPQGDDWCDFVVDGNVVDESM---KIQWNPRMRVKMKTDKSSRIPYQGTIS 302
N+ +++Y +FV+ +++ ++ R + +T+ S + Y GTI
Sbjct: 289 ANRTPFLIFYNPRACPAEFVIPLAKYRKAICGSQLSVGMRFGMMFETEDSGKRRYMGTIV 348
Query: 303 IVSRTSNL------WRMLQVNWDEFQVSQIPRRVNPWWVE 336
+S L WR LQV WDE + P RV+PW +E
Sbjct: 349 GISDLDPLRWPGSKWRNLQVEWDEPGCNDKPTRVSPWDIE 388
>At2g46530 ARF1 family auxin responsive transcription factor like
protein
Length = 622
Score = 161 bits (408), Expect = 6e-40
Identities = 113/365 (30%), Positives = 183/365 (49%), Gaps = 21/365 (5%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSS-ITHTHSFLQSFRPFTLCIVSAV 71
E+W CA V++P+ RV+YFPQGH+E S++ + + + P LC V +V
Sbjct: 42 ELWKACAGPLVEVPRYGERVFYFPQGHMEQLVASTNQGVVDQEIPVFNLPPKILCRVLSV 101
Query: 72 DLLADPHTDEVFVKLLLTPITND---VHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNA 128
L A+ TDEV+ ++ L P + L+ P E A + V SFVK LT SD +
Sbjct: 102 TLKAEHETDEVYAQITLQPEEDQSEPTSLDPPLVEPA----KPTVDSFVKILTASDTSTH 157
Query: 129 RSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNS 188
F + R A P LD+ + +Q L D+HG +F H+ RG P+R++L + W++
Sbjct: 158 GGFSVLRKHATECLPSLDMTQPTPTQELVARDLHGYEWRFKHIFRGQPRRHLL-TTGWST 216
Query: 189 FVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAEE 246
FV K+LVAGD+ +F++ TG + VG+RR Q A+ + V+ A
Sbjct: 217 FVTSKRLVAGDAFVFLRGETGDLRVGVRRLAKQQSTMPASVISSQSMRLGVLATASHAVT 276
Query: 247 NKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMK--TDKSSRIPYQGTI--- 301
+V+Y F++ N +MK ++ MR +M+ ++S + GTI
Sbjct: 277 TTTIFVVFYKPRIS--QFIISVNKYMMAMKNGFSLGMRYRMRFEGEESPERIFTGTIIGS 334
Query: 302 -SIVSR-TSNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKP-APTPFPQTKKFRTTQS 358
+ S+ ++ WR LQ+ WDE Q P +V+PW +E S PTP Q K + ++
Sbjct: 335 GDLSSQWPASKWRSLQIQWDEPSSIQRPNKVSPWEIEPFSPSALTPTPTQQQSKSKRSRP 394
Query: 359 SAQLS 363
++++
Sbjct: 395 ISEIT 399
>At1g34310 hypothetical protein
Length = 591
Score = 159 bits (401), Expect = 4e-39
Identities = 112/340 (32%), Positives = 169/340 (48%), Gaps = 22/340 (6%)
Query: 8 SHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCI 67
S+V ++W CA IPKL +VYYFPQGH+E S+ + + C
Sbjct: 22 SYVYEQLWKLCAGPLCDIPKLGEKVYYFPQGHIELVETSTREELNELQPICDLPSKLQCR 81
Query: 68 VSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLND---RNEVVSFVKTLTRSD 124
V A+ L + ++DE + ++ L P T EN + + N+ R V SF K LT SD
Sbjct: 82 VIAIHLKVENNSDETYAEITLMPDTTVS--ENLQVVIPTQNENQFRPLVNSFTKVLTASD 139
Query: 125 VNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYIS 184
+ F +P+ A P LD+ +Q L D+HG +F H RG P+R++L +
Sbjct: 140 TSAHGGFFVPKKHAIECLPSLDMSQPLPAQELLAIDLHGNQWRFNHNYRGTPQRHLL-TT 198
Query: 185 EWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIR--RNTQFVAAAAEQKKDELEKAVMEALK 242
WN+F KKLVAGD ++F++ TG++ VGIR R+ Q ++ D + V+ + K
Sbjct: 199 GWNAFTTSKKLVAGDVIVFVRGETGELRVGIRRARHQQGNIPSSIVSIDCMRHGVVASAK 258
Query: 243 LAEENKA-FEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMKTDKSSRIPYQGTI 301
A +N+ F +VY P D + D V + K R ++++ D S GTI
Sbjct: 259 HAFDNQCMFTVVYKPSYDKFLDAV--------NNKFNVGSRFTMRLEGDDFSERRCFGTI 310
Query: 302 SIVSRTS-----NLWRMLQVNWDEFQVSQIPRRVNPWWVE 336
VS S + WR L+V WDEF P++V+PW +E
Sbjct: 311 IGVSDFSPHWKCSEWRSLEVQWDEFTSFPGPKKVSPWDIE 350
>At1g34390 auxin response factor, putative
Length = 600
Score = 158 bits (399), Expect = 7e-39
Identities = 126/445 (28%), Positives = 208/445 (46%), Gaps = 35/445 (7%)
Query: 8 SHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCI 67
S++ ++W CA IPKL ++YYFPQG++E S+ + + C
Sbjct: 22 SYMYEQLWKLCAGPLCDIPKLGEKIYYFPQGNIELVEASTREELNELKPICDLPSKLQCR 81
Query: 68 VSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNN 127
V A+ L + ++DE + ++ L P T V + E R V SF K LT SD +
Sbjct: 82 VIAIQLKVENNSDETYAEITLMPDTTQVVIPTQNEN----QFRPLVNSFTKVLTASDTSG 137
Query: 128 ARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWN 187
F +P+ A P LD+ +Q L TD+HG +F H RG P+R++L + WN
Sbjct: 138 G--FFVPKKHAIECLPPLDMSQPLPTQELLATDLHGNQWRFNHNYRGTPQRHLL-TTGWN 194
Query: 188 SFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAE 245
+F KKLVAGD ++F++ TG++ VGIRR + Q ++ + + V+ + K A
Sbjct: 195 AFTTSKKLVAGDVIVFVRGETGELRVGIRRAGHQQGNIPSSIISIESMRHGVIASAKHAF 254
Query: 246 ENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWN--PRMRVKMKTDKSSRIPYQGTISI 303
+N+ IV Y F+V + +++ ++N R ++ + D S Y GTI
Sbjct: 255 DNQCMFIVVYKPSIRSSQFIVSYDKFLDAVNNKFNVGSRFTMRFEGDDFSERRYFGTIIG 314
Query: 304 VSRTS-----NLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFPQTKKFRTTQ 357
VS S + WR L+V WDEF P +V+PW +E + PA P P K + +
Sbjct: 315 VSDFSPHWKCSEWRNLEVQWDEFASFSRPNKVSPWEIEHL--MPALNVPRPSLLKNKRLR 372
Query: 358 SSAQLSDKKETLLNGDGFPVDIQRVRHDLVSISGPIHSHIILNSSETKFPATHNCNTKND 417
++ LL P+ Q +S++ P++ + T+ T++
Sbjct: 373 EVNEIGSSSSHLLP----PILTQGQEIGQLSVASPMNISL-----------TYRDTTEDV 417
Query: 418 SDGSIMLFGKIIQPHVS-NFHNSHI 441
+ S +L +QP N++N +
Sbjct: 418 MNPSRLLMSYPVQPMPKLNYNNQMV 442
>At3g61830 auxin response factor-like protein
Length = 602
Score = 157 bits (397), Expect = 1e-38
Identities = 111/358 (31%), Positives = 175/358 (48%), Gaps = 29/358 (8%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSF-LQSFRPFTLCIVSAV 71
E+W CA V++P+ RV+YFPQGH+E S++ ++ + P LC V V
Sbjct: 25 ELWKVCAGPLVEVPRAQERVFYFPQGHMEQLVASTNQGINSEEIPVFDLPPKILCRVLDV 84
Query: 72 DLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLND------RNEVVSFVKTLTRSDV 125
L A+ TDEV+ ++ L P E + E +L+ + E SFVK LT SD
Sbjct: 85 TLKAEHETDEVYAQITLQP-------EEDQSEPTSLDPPIVGPTKQEFHSFVKILTASDT 137
Query: 126 NNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISE 185
+ F + R A P LD+ + +Q L D+HG +F H+ RG P+R++L +
Sbjct: 138 STHGGFSVLRKHATECLPSLDMTQATPTQELVTRDLHGFEWRFKHIFRGQPRRHLL-TTG 196
Query: 186 WNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKL 243
W++FV K+LVAGD+ +F++ G + VG+RR Q + + V+
Sbjct: 197 WSTFVSSKRLVAGDAFVFLRGENGDLRVGVRRLARHQSTMPTSVISSQSMHLGVLATASH 256
Query: 244 AEENKAFEIVYYPQGDDWCDFVVDGNVVDESMK--IQWNPRMRVKMKTDKSSRIPYQGTI 301
A +V+Y F+V N E++K R R++ + ++S + GTI
Sbjct: 257 AVRTTTIFVVFYKPRIS--QFIVGVNKYMEAIKHGFSLGTRFRMRFEGEESPERIFTGTI 314
Query: 302 ----SIVSR-TSNLWRMLQVNWDEFQVSQIPRRVNPWWVE-LISHKPAPTPF--PQTK 351
+ S+ ++ WR LQV WDE Q P +V+PW +E ++ P TP PQ+K
Sbjct: 315 VGSGDLSSQWPASKWRSLQVQWDEPTTVQRPDKVSPWEIEPFLATSPISTPAQQPQSK 372
>At5g60450 auxin response factor 4
Length = 788
Score = 157 bits (396), Expect = 1e-38
Identities = 112/350 (32%), Positives = 169/350 (48%), Gaps = 25/350 (7%)
Query: 8 SHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLE-NASPSSSSITHTHSFLQSFRPFTLC 66
S + E+WH CA +PK + V YFPQGHLE +A S SS F P +C
Sbjct: 60 SSIYSELWHACAGPLTCLPKKGNVVVYFPQGHLEQDAMVSYSSPLEIPKF--DLNPQIVC 117
Query: 67 IVSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLN---DRNEVVS-------F 116
V V LLA+ TDEV+ ++ L P+ L +EV L +RN S F
Sbjct: 118 RVVNVQLLANKDTDEVYTQVTLLPLQEFSMLNGEGKEVKELGGEEERNGSSSVKRTPHMF 177
Query: 117 VKTLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFP 176
KTLT SD + F +PR A++ F LD + + SQ L D+HG KF H+ RG P
Sbjct: 178 CKTLTASDTSTHGGFSVPRRAAEDCFAPLDYKQQRPSQELIAKDLHGVEWKFRHIYRGQP 237
Query: 177 KRNMLYISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRRNTQFVAAAAEQ--KKDELE 234
+R++L + W+ FV +K LV+GD+V+F++D G++ +GIRR + + +K+
Sbjct: 238 RRHLL-TTGWSIFVSQKNLVSGDAVLFLRDEGGELRLGIRRAARPRNGLPDSIIEKNSCS 296
Query: 235 KAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDESMK--IQWNPRMRVKMKTDKS 292
+ F + Y P+ +FV+ S++ + R R++ + D S
Sbjct: 297 NILSLVANAVSTKSMFHVFYSPRATH-AEFVIPYEKYITSIRSPVCIGTRFRMRFEMDDS 355
Query: 293 SRIPYQGTISIVSR------TSNLWRMLQVNWDEFQVSQIPRRVNPWWVE 336
G ++ V ++ WR L V WDE VS RV+PW ++
Sbjct: 356 PERRCAGVVTGVCDLDPYRWPNSKWRCLLVRWDESFVSDHQERVSPWEID 405
>At1g59750 auxin response factor 1
Length = 665
Score = 155 bits (392), Expect = 4e-38
Identities = 104/347 (29%), Positives = 163/347 (46%), Gaps = 12/347 (3%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RVYYFP+GH+E S + LC V +
Sbjct: 22 ELWHACAGPLVTLPREGERVYYFPEGHMEQLEASMHQGLEQQMPSFNLPSKILCKVINIQ 81
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFH 132
A+P TDEV+ ++ L P + +P V ++ V SF KTLT SD + F
Sbjct: 82 RRAEPETDEVYAQITLLPELDQSEPTSPDAPVQE-PEKCTVHSFCKTLTASDTSTHGGFS 140
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+ R AD+ P LD+ + Q L TD+H F H+ RG P+R++L + W+ FV
Sbjct: 141 VLRRHADDCLPPLDMSQQPPWQELVATDLHNSEWHFRHIFRGQPRRHLL-TTGWSVFVSS 199
Query: 193 KKLVAGDSVIFMKDSTGKIFVGIRRN--TQFVAAAAEQKKDELEKAVMEALKLAEENKAF 250
KKLVAGD+ IF++ ++ VG+RR+ Q ++ + V+ A
Sbjct: 200 KKLVAGDAFIFLRGENEELRVGVRRHMRQQTNIPSSVISSHSMHIGVLATAAHAITTGTI 259
Query: 251 EIVYYPQGDDWCDFVVDGN--VVDESMKIQWNPRMRVKMKTDKSSRIPYQGTISIVSRTS 308
V+Y +F+V N + ++ K+ R +++ + +++ + GTI V
Sbjct: 260 FSVFYKPRTSRSEFIVSVNRYLEAKTQKLSVGMRFKMRFEGEEAPEKRFSGTIVGVQENK 319
Query: 309 NL------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKPAPTPFPQ 349
+ WR L+V WDE P RV+PW +E + P+ PQ
Sbjct: 320 SSVWHDSEWRSLKVQWDEPSSVFRPERVSPWELEPLVANSTPSSQPQ 366
>At1g34410 auxin response factor, putative
Length = 620
Score = 153 bits (387), Expect = 2e-37
Identities = 105/338 (31%), Positives = 164/338 (48%), Gaps = 14/338 (4%)
Query: 8 SHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCI 67
S++ ++W CA IPKL VYYFPQG++E S+ + + C
Sbjct: 34 SYMYEQLWKLCAGPLCDIPKLGENVYYFPQGNIELVQASTREELNELQPICDLPSKLQCR 93
Query: 68 VSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNN 127
V A+ L + ++DE++ ++ L P T V + E R V SF K LT SD +
Sbjct: 94 VIAIHLKVENNSDEIYAEITLMPDTTQVVIPTQSEN----RFRPLVNSFTKVLTASDTSA 149
Query: 128 ARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWN 187
F +P+ A P LD+ +Q + D+H +F H RG P+R+ L + WN
Sbjct: 150 YGGFSVPKKHAIECLPPLDMSQPLPAQEILAIDLHDNQWRFRHNYRGTPQRHSL-TTGWN 208
Query: 188 SFVKRKKLVAGDSVIFMKDSTGKIFVGIR--RNTQFVAAAAEQKKDELEKAVMEALKLAE 245
F+ KKLV GD ++F++ TG++ VGIR R+ Q ++ D + V+ + K A
Sbjct: 209 EFITSKKLVKGDVIVFVRGETGELRVGIRRARHQQGNIPSSIVSIDCMRHGVIASAKHAF 268
Query: 246 ENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWN--PRMRVKMKTDKSSRIPYQGTISI 303
+N+ IV Y F+V + +++ ++N R ++ + D S Y GTI
Sbjct: 269 DNQCIFIVVYKPSIRSSQFIVSYDKFLDAVNNKFNVGSRFTMRFEGDDFSERRYFGTIIG 328
Query: 304 VSRTS-----NLWRMLQVNWDEFQVSQIPRRVNPWWVE 336
VS S + WR L+V WDEF P +V+PW +E
Sbjct: 329 VSDFSPHWKCSEWRSLEVQWDEFASFSRPNKVSPWEIE 366
>At1g35520 auxin response factor, putative
Length = 576
Score = 152 bits (384), Expect = 4e-37
Identities = 106/342 (30%), Positives = 167/342 (47%), Gaps = 16/342 (4%)
Query: 3 LQKQPSHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRP 62
+ + S++ ++W CA IPKL +VYYFPQG++E S+ + +
Sbjct: 17 IDRSKSYMYEQLWKLCAGPLCDIPKLGEKVYYFPQGNIELVEASTREELNELQPICDLPS 76
Query: 63 FTLCIVSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTR 122
C V A+ L + ++DE + K+ L P T V + E R V SF K LT
Sbjct: 77 KLQCRVIAIHLKVENNSDETYAKITLMPDTTQVVIPTQNEN----QFRPLVNSFTKVLTA 132
Query: 123 SDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLY 182
SD++ F +P+ A P LD+ +Q L D+HG F H RG P+R++L
Sbjct: 133 SDISANGVFSVPKKHAIECLPPLDMSQPLPAQELLAIDLHGNQWSFRHSYRGTPQRHLL- 191
Query: 183 ISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIR--RNTQFVAAAAEQKKDELEKAVMEA 240
+ WN F KKLV GD ++F++ TG++ VGIR R+ Q ++ D + V+ +
Sbjct: 192 TTGWNEFTTSKKLVKGDVIVFVRGETGELRVGIRRARHQQGNIPSSIVSIDCMRHGVIAS 251
Query: 241 LKLAEENKA-FEIVYYPQGDDWCDFVVDGNVVDESMKIQWN--PRMRVKMKTDKSSRIPY 297
K A +N+ F +VY P+ F+V + +++ ++N R ++ + D S Y
Sbjct: 252 AKHAFDNQCMFIVVYKPRSS---QFIVSYDKFLDAVNNKFNVGSRFTMRFEGDDLSERRY 308
Query: 298 QGTISIVSRTSNLWR---MLQVNWDEFQVSQIPRRVNPWWVE 336
GTI VS S W+ + WDEF P +V+PW +E
Sbjct: 309 FGTIIGVSNFSPHWKCSDWRSLEWDEFASFLRPNKVSPWEIE 350
>At1g35240 hypothetical protein
Length = 615
Score = 152 bits (383), Expect = 5e-37
Identities = 108/339 (31%), Positives = 164/339 (47%), Gaps = 18/339 (5%)
Query: 8 SHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCI 67
S++ ++W CA IPKL VYYFPQG++E S+ + + C
Sbjct: 22 SYMYEQLWKLCAGPLCDIPKLGENVYYFPQGNIELVDASTREELNELQPICDLPSKLQCR 81
Query: 68 VSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNN 127
V A+ L + ++DE + ++ L P T V + E R V SF K LT SD +
Sbjct: 82 VIAIHLKVENNSDETYAEITLMPDTTQVVIPTQSEN----QFRPLVNSFTKVLTASDTSA 137
Query: 128 ARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWN 187
F +P+ A P LD+ +Q L D+HG +F H RG P+R+ L + WN
Sbjct: 138 YGGFFVPKKHAIECLPPLDMSQPLPAQELLAKDLHGNQWRFRHSYRGTPQRHSL-TTGWN 196
Query: 188 SFVKRKKLVAGDSVIFMKDSTGKIFVGIR--RNTQFVAAAAEQKKDELEKAVMEALKLAE 245
F KKLV GD ++F++ TG++ VGIR R+ Q ++ D + V+ + K A
Sbjct: 197 EFTTSKKLVKGDVIVFVRGETGELRVGIRRARHQQGNIPSSIVSIDCMRHGVIASAKHAL 256
Query: 246 ENKA-FEIVYYPQGDDWCDFVVDGNVVDESM--KIQWNPRMRVKMKTDKSSRIPYQGTIS 302
+N+ F +VY P+ F+V + ++M K R ++ + D S Y GTI
Sbjct: 257 DNQCIFIVVYKPRSS---QFIVSYDKFLDAMNNKFIVGSRFTMRFEGDDFSERRYFGTII 313
Query: 303 IVSRTS-----NLWRMLQVNWDEFQVSQIPRRVNPWWVE 336
V+ S + WR L+V WDEF P +V+PW +E
Sbjct: 314 GVNDFSPHWKCSEWRSLEVQWDEFASFSRPNKVSPWEIE 352
>At5g62000 auxin response factor - like protein
Length = 371
Score = 149 bits (377), Expect = 2e-36
Identities = 102/314 (32%), Positives = 153/314 (48%), Gaps = 13/314 (4%)
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RV+YFPQGH+E AS + ++ L LC V VD
Sbjct: 61 ELWHACAGPLVTVPRQDDRVFYFPQGHIEQASTNQAA--EQQMPLYDLPSKLLCRVINVD 118
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFH 132
L A+ TDEV+ ++ L P N KE R +V SF KTLT SD + F
Sbjct: 119 LKAEADTDEVYAQITLLPEANQDENAIEKEAPLPPPPRFQVHSFCKTLTASDTSTHGGFS 178
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+ R AD P LD+ + +Q L D+H +F H+ RG P+R++L S W+ FV
Sbjct: 179 VLRRHADECLPPLDMSRQPPTQELVAKDLHANEWRFRHIFRGQPRRHLLQ-SGWSVFVSS 237
Query: 193 KKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKDELEKAVMEALKLAEENKAF 250
K+LVAGD+ IF++ G++ VG+RR Q ++ + V+ A
Sbjct: 238 KRLVAGDAFIFLRGENGELRVGVRRAMRQQGNVPSSVISSHSMHLGVLATAWHAISTGTM 297
Query: 251 EIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMK--TDKSSRIPYQGTISIVSRT- 307
VYY +F+V + ES+K ++ MR KM+ +++ + GTI + +
Sbjct: 298 FTVYYKPRTSPSEFIVPFDQYMESVKNNYSIGMRFKMRFEGEEAPEQRFTGTIVGIEESD 357
Query: 308 -----SNLWRMLQV 316
+ WR L+V
Sbjct: 358 PTRWPKSKWRSLKV 371
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.133 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,564,536
Number of Sequences: 26719
Number of extensions: 451998
Number of successful extensions: 1432
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 31
Number of HSP's successfully gapped in prelim test: 28
Number of HSP's that attempted gapping in prelim test: 1299
Number of HSP's gapped (non-prelim): 64
length of query: 456
length of database: 11,318,596
effective HSP length: 103
effective length of query: 353
effective length of database: 8,566,539
effective search space: 3023988267
effective search space used: 3023988267
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC134322.24


