

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.2 + phase: 0 /pseudo
(840 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
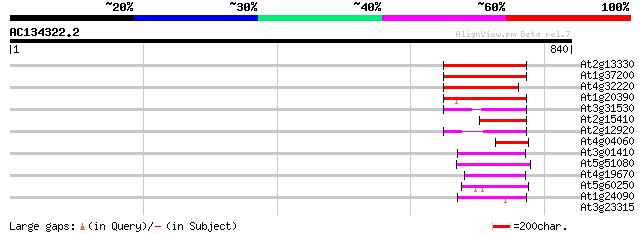
At2g13330 F14O4.9 116 6e-26
At1g37200 hypothetical protein 114 2e-25
At4g32220 hypothetical protein 99 1e-20
At1g20390 hypothetical protein 99 1e-20
At3g31530 hypothetical protein 79 8e-15
At2g15410 putative retroelement pol polyprotein 65 1e-10
At2g12920 pseudogene 48 3e-05
At4g04060 putative transposon protein 46 1e-04
At3g01410 putative RNase H 45 1e-04
At5g51080 unknown protein 45 2e-04
At4g19670 putative protein 45 2e-04
At5g60250 unknown protein 44 3e-04
At1g24090 unknown protein 43 6e-04
At3g23315 unknown protein 29 9.6
>At2g13330 F14O4.9
Length = 889
Score = 116 bits (290), Expect = 6e-26
Identities = 61/126 (48%), Positives = 85/126 (67%), Gaps = 2/126 (1%)
Query: 650 LADFLVEMTAEK--GSMEAAKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSS 707
LADF++++ G+ KW+L VDGSSN +GSG G+ L P G +IEQSL+ F +S
Sbjct: 622 LADFVIKLPLADLDGTNSNKKWLLHVDGSSNRQGSGVGIQLTSPTGEVIEQSLQLGFNAS 681
Query: 708 NNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQ 767
NN++EYEAL AG++LA + ++E+ A SDSQLV Q G ++ KD ++ YLE V L Q
Sbjct: 682 NNESEYEALIAGIKLAQEKGIREIHAYSDSQLVTSQFHGEYEAKDERMEAYLELVKTLAQ 741
Query: 768 KFVSFK 773
+F SFK
Sbjct: 742 QFESFK 747
>At1g37200 hypothetical protein
Length = 1564
Score = 114 bits (286), Expect = 2e-25
Identities = 60/126 (47%), Positives = 84/126 (66%), Gaps = 2/126 (1%)
Query: 650 LADFLVEMTAEK--GSMEAAKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSS 707
LADF++E+ G+ KW+L VDGSSN +GSG G+ L P G +IEQS R F +S
Sbjct: 1020 LADFVIELPLADLDGTNSNKKWLLHVDGSSNRQGSGEGIQLTSPTGEVIEQSFRLGFNAS 1079
Query: 708 NNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQ 767
NN++EYEAL G++LA + ++++ A SDSQLV Q G ++ KD ++ YLE V LTQ
Sbjct: 1080 NNESEYEALIDGIKLAQGMRIRDIHAHSDSQLVTSQFHGEYEAKDERMEAYLELVKTLTQ 1139
Query: 768 KFVSFK 773
+F SF+
Sbjct: 1140 QFESFE 1145
>At4g32220 hypothetical protein
Length = 452
Score = 99.0 bits (245), Expect = 1e-20
Identities = 51/114 (44%), Positives = 77/114 (66%), Gaps = 1/114 (0%)
Query: 650 LADFLVEMTAEK-GSMEAAKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSSN 708
LADF+ E++ + + W+L DGSS+L+GS G+ L+ P ++EQSLR FK+SN
Sbjct: 189 LADFMTELSPDVIDVIPDENWILYSDGSSSLQGSSLGILLQSPTREILEQSLRLQFKASN 248
Query: 709 NQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESV 762
N+AEYEAL AG+RLA + K+++A SDSQLV + +G F+ K+ ++ +L V
Sbjct: 249 NEAEYEALLAGLRLAKGLGAKQIKAFSDSQLVVSRFSGEFEAKNERMRAHLSMV 302
>At1g20390 hypothetical protein
Length = 1791
Score = 99.0 bits (245), Expect = 1e-20
Identities = 51/129 (39%), Positives = 80/129 (61%), Gaps = 5/129 (3%)
Query: 650 LADFLVEMTAEKGSMEAA-----KWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDF 704
LADFL+E+ + + +W L VDGSS+ +GSG G+ L P ++EQS R F
Sbjct: 1186 LADFLIELPLQSAERAVSGNRGEEWSLYVDGSSSARGSGIGIRLVSPTAEVLEQSFRLRF 1245
Query: 705 KSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMY 764
++NN AEYE L AG+RLA + + + A +DSQL+ Q++G ++ K+ ++ YL+ V
Sbjct: 1246 VATNNVAEYEVLIAGLRLAAGMQITTIHAFTDSQLIAGQLSGEYEAKNEKMDAYLKIVQL 1305
Query: 765 LTQKFVSFK 773
+T+ F +FK
Sbjct: 1306 MTKDFENFK 1314
>At3g31530 hypothetical protein
Length = 831
Score = 79.3 bits (194), Expect = 8e-15
Identities = 47/126 (37%), Positives = 69/126 (54%), Gaps = 13/126 (10%)
Query: 650 LADFLVEMTAEKGSMEA--AKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSS 707
LADF+VE+ ++ W+L VDGSS+ +GSG G+ L P G +++
Sbjct: 393 LADFIVELPTKEARENPLDTTWLLHVDGSSSKQGSGVGIRLTSPTG-----------EAT 441
Query: 708 NNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQ 767
NN AEYEAL AG+ LA + + + A DSQL+ Q G + T+D ++ YL V L +
Sbjct: 442 NNVAEYEALVAGLNLAWGLKIGKTRAFCDSQLIANQFNGEYTTQDKKMEAYLIHVQNLAK 501
Query: 768 KFVSFK 773
F F+
Sbjct: 502 NFDEFE 507
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 65.5 bits (158), Expect = 1e-10
Identities = 33/70 (47%), Positives = 49/70 (69%)
Query: 704 FKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVM 763
F +SNN+AEYEAL AG+RLA + VK+++A DSQLV Q +GN++ K+ ++ YL+ V
Sbjct: 1190 FPASNNEAEYEALIAGLRLAHGIEVKKIQAYCDSQLVASQFSGNYEAKNERMDAYLKVVR 1249
Query: 764 YLTQKFVSFK 773
L+ F F+
Sbjct: 1250 ELSYNFEVFE 1259
>At2g12920 pseudogene
Length = 863
Score = 47.8 bits (112), Expect = 3e-05
Identities = 36/126 (28%), Positives = 54/126 (42%), Gaps = 32/126 (25%)
Query: 650 LADFLVEMTAEKGSME--AAKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSS 707
LADF VE+ ++ W+L VDGS +
Sbjct: 526 LADFNVELPTKEARKNWLDTTWLLHVDGSFS----------------------------- 556
Query: 708 NNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQ 767
QAEYEAL AG++LA + + ++ A DS LV Q G + +D ++ YL L +
Sbjct: 557 -KQAEYEALVAGLKLAQGLKIAKIRAFCDSSLVANQFNGEYTARDERMGAYLTHSQDLAK 615
Query: 768 KFVSFK 773
+F F+
Sbjct: 616 QFDEFE 621
>At4g04060 putative transposon protein
Length = 375
Score = 45.8 bits (107), Expect = 1e-04
Identities = 22/49 (44%), Positives = 32/49 (64%)
Query: 728 VKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQKFVSFKYYM 776
++++ A SDSQLV Q G ++ KD ++ YLE V LTQ+F SF+ M
Sbjct: 3 IRDIHAHSDSQLVISQFHGEYEAKDERMEAYLELVKTLTQQFESFELTM 51
>At3g01410 putative RNase H
Length = 290
Score = 45.4 bits (106), Expect = 1e-04
Identities = 32/104 (30%), Positives = 47/104 (44%), Gaps = 2/104 (1%)
Query: 671 LSVDGSS--NLKGSGAGVTLEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKVVV 728
+ DG+S N +GAG L ++ ++NN AEY AL G+R AL
Sbjct: 153 IEFDGASKGNPGKAGAGAVLRASDNSVLFYLREGVGNATNNVAEYRALLLGLRSALDKGF 212
Query: 729 KELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQKFVSF 772
K + DS LV QV G ++T P++ + L F +F
Sbjct: 213 KNVHVLGDSMLVCMQVQGAWKTNHPKMAELCKQAKELMNSFKTF 256
>At5g51080 unknown protein
Length = 322
Score = 45.1 bits (105), Expect = 2e-04
Identities = 33/113 (29%), Positives = 57/113 (50%), Gaps = 2/113 (1%)
Query: 670 VLSVDGSS--NLKGSGAGVTLEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKVV 727
++ DG+S N SGA L+ G LI + + ++NN AEY L G++ A++
Sbjct: 187 IIEFDGASKGNPGLSGAAAVLKTEDGSLIFKMRQGLGIATNNAAEYHGLILGLKHAIEKG 246
Query: 728 VKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQKFVSFKYYMCLES 780
+++ K+DS+LV Q+ G ++ L + L+ K +SF+ L S
Sbjct: 247 YTKIKVKTDSKLVCMQMKGQWKVNHEVLSKLHKEAKQLSDKCLSFEISHVLRS 299
>At4g19670 putative protein
Length = 532
Score = 44.7 bits (104), Expect = 2e-04
Identities = 30/92 (32%), Positives = 47/92 (50%), Gaps = 1/92 (1%)
Query: 682 SGAGVTLEGPYGV-LIEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLV 740
SG GV LE + LI+ + DF + A Y AL G+ +AL+ + + A +DS+L+
Sbjct: 79 SGIGVVLERSGDLELIQVQKKLDFYVEESVANYLALMDGLEVALQNNLSSVVAVTDSELL 138
Query: 741 FRQVTGNFQTKDPQLLMYLESVMYLTQKFVSF 772
+ Q+T Q + P L+ E V+ T F
Sbjct: 139 YNQITREEQLEIPLLVALRERVLEKTSNLNGF 170
>At5g60250 unknown protein
Length = 655
Score = 44.3 bits (103), Expect = 3e-04
Identities = 30/110 (27%), Positives = 53/110 (47%), Gaps = 10/110 (9%)
Query: 677 SNLKGSGAGVTLEGPYGVLI---EQSLRFDFKS-------SNNQAEYEALFAGMRLALKV 726
S+ G G + +GV I +L F+ K S AE +AL G+ ALK+
Sbjct: 154 SDENGKGKMTDVVSGFGVAICDQRDNLLFEMKGPLIDNGMSRQGAELKALIRGLTEALKL 213
Query: 727 VVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQKFVSFKYYM 776
+K + DS +F+ VTG + K ++ + L+ + + Q F S+++ +
Sbjct: 214 GIKHIVFFCDSYPIFQYVTGKWMAKQKKISLLLDDLQSIMQHFSSYQHVL 263
>At1g24090 unknown protein
Length = 535
Score = 43.1 bits (100), Expect = 6e-04
Identities = 34/108 (31%), Positives = 54/108 (49%), Gaps = 5/108 (4%)
Query: 671 LSVDGSS--NLKGSGAGVTLEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKVVV 728
+ DG+S N SGA L+ G LI + + ++NN AEY AL G++ A++
Sbjct: 219 IEFDGASKGNPGLSGAAAVLKTEDGSLICRVRQGLGIATNNAAEYHALILGLKYAIEKGY 278
Query: 729 KELEAKSDSQLVF---RQVTGNFQTKDPQLLMYLESVMYLTQKFVSFK 773
K ++ K DS+LV +Q+ G ++ L + L K VSF+
Sbjct: 279 KNIKVKGDSKLVCMQKQQIKGQWKVNHEVLAKLHKEAKLLCNKCVSFE 326
>At3g23315 unknown protein
Length = 191
Score = 29.3 bits (64), Expect = 9.6
Identities = 25/119 (21%), Positives = 52/119 (43%), Gaps = 6/119 (5%)
Query: 660 EKGSMEAAKWV---LSVDGSSNLKGSGAGVTLEGPYGVLIEQSL-RFDFKSSNNQAEYEA 715
+K A+WV V + + SG + G ++ +F + + +AE A
Sbjct: 28 DKWQKPGAEWVKCNYDVSNHAGRQDSGLRWIIRNSQGTCLDCGCGKFQGRQTIKEAECTA 87
Query: 716 LFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQKFVSFKY 774
L ++ A + + +E + D+ V R + + +P+L YLE + ++ F + K+
Sbjct: 88 LIWAIQCAWDLGYRRVEFEGDNITVNRLIRN--KETNPRLRYYLEFIQQWSKAFTTVKF 144
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.358 0.156 0.572
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,378,564
Number of Sequences: 26719
Number of extensions: 539586
Number of successful extensions: 1955
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 1941
Number of HSP's gapped (non-prelim): 14
length of query: 840
length of database: 11,318,596
effective HSP length: 108
effective length of query: 732
effective length of database: 8,432,944
effective search space: 6172915008
effective search space used: 6172915008
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC134322.2


