

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.16 - phase: 0
(690 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
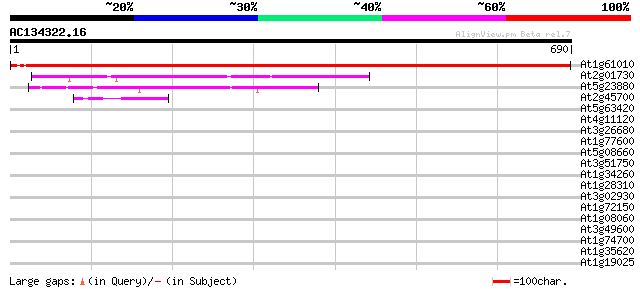
Score E
Sequences producing significant alignments: (bits) Value
At1g61010 putative cleavage and polyadenylation specificity factor 1136 0.0
At2g01730 putative cleavage and polyadenylation specifity factor 283 2e-76
At5g23880 polyadenylation cleavage/specificity factor 100 kDa su... 136 4e-32
At2g45700 unknown protein 48 2e-05
At5g63420 putative protein 41 0.002
At4g11120 translation elongation factor ts like protein 35 0.14
At3g26680 unknown protein 32 1.6
At1g77600 hypothetical protein 32 1.6
At5g08660 putative protein 31 2.6
At3g51750 unknown protein (At3g51750) 31 2.6
At1g34260 hypothetical protein 31 2.6
At1g28310 zinc finger like protein OBP3 30 3.5
At3g02930 unknown protein 30 4.5
At1g72150 putative cytosolic factor protein 30 4.5
At1g08060 storage protein, putative 30 5.9
At3g49600 putative protein 29 7.7
At1g74700 unknown protein 29 7.7
At1g35620 protein disulfide isomerase, putative (At1g35620) 29 7.7
At1g19025 unknown protein 29 7.7
>At1g61010 putative cleavage and polyadenylation specificity factor
Length = 693
Score = 1136 bits (2939), Expect = 0.0
Identities = 563/691 (81%), Positives = 635/691 (91%), Gaps = 7/691 (1%)
Query: 2 SSVKKRESNGGTINRETEDQLIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMA 61
+S+K+RE I+R+ DQLIVTPLGAG+EVGRSCVYM+++GK +LFDCGIHP YSGMA
Sbjct: 6 TSLKRREQ---PISRDG-DQLIVTPLGAGSEVGRSCVYMSFRGKNILFDCGIHPAYSGMA 61
Query: 62 ALPYFDEIDPSTVDVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYV 121
ALPYFDEIDPS++DVLLITHFH+DHAASLPYFLEKTTF GRVFMT+ATKAIYKLLL+DYV
Sbjct: 62 ALPYFDEIDPSSIDVLLITHFHIDHAASLPYFLEKTTFNGRVFMTHATKAIYKLLLTDYV 121
Query: 122 KVSKVSVDDMLYDEQDINRSMDKIEVIDFHQTVEVNGIRFWCYTAGHVLGAAMFMVDIAG 181
KVSKVSV+DML+DEQDIN+SMDKIEVIDFHQTVEVNGI+FWCYTAGHVLGAAMFMVDIAG
Sbjct: 122 KVSKVSVEDMLFDEQDINKSMDKIEVIDFHQTVEVNGIKFWCYTAGHVLGAAMFMVDIAG 181
Query: 182 VRVLYTGDYSREEDRHLRAAETPQFSPDVCIIESTYGVQHHQPRHTREKRFTDVIHSTIS 241
VR+LYTGDYSREEDRHLRAAE PQFSPD+CIIEST GVQ HQ RH REKRFTDVIHST++
Sbjct: 182 VRILYTGDYSREEDRHLRAAELPQFSPDICIIESTSGVQLHQSRHIREKRFTDVIHSTVA 241
Query: 242 QGGRVLIPAYALGRAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYETYTLSMND 301
QGGRVLIPA+ALGRAQELLLILDEYWANHP+L NIPIYYASPLAKKC+ VY+TY LSMND
Sbjct: 242 QGGRVLIPAFALGRAQELLLILDEYWANHPDLHNIPIYYASPLAKKCMAVYQTYILSMND 301
Query: 302 RIQN--AKSNPFAFKHISALSSIDIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNS 359
RI+N A SNPF FKHIS L+SID F DVGPSVVMA+PGGLQSGLSRQLFD WCSDKKN+
Sbjct: 302 RIRNQFANSNPFVFKHISPLNSIDDFNDVGPSVVMATPGGLQSGLSRQLFDSWCSDKKNA 361
Query: 360 CVIPGYVVEGTLAKTILNEPKEVTLMNGLSAPLHMQVHYISFSAHADSAQTSAFLEELNP 419
C+IPGY+VEGTLAKTI+NEPKEVTLMNGL+APL+MQVHYISFSAHAD AQTS FL+EL P
Sbjct: 362 CIIPGYMVEGTLAKTIINEPKEVTLMNGLTAPLNMQVHYISFSAHADYAQTSTFLKELMP 421
Query: 420 PNIILVHGAANEMGRLKQKLMTQFADRNTKILTPKNCQSVEMYFNSQKMAKTIGKLAEKT 479
PNIILVHG ANEM RLKQKL+T+F D NTKI+TPKNC+SVEMYFNS+K+AKTIG+LAEKT
Sbjct: 422 PNIILVHGEANEMMRLKQKLLTEFPDGNTKIMTPKNCESVEMYFNSEKLAKTIGRLAEKT 481
Query: 480 PEVGETVSGLLVKKGFTYQIMAPDDLHVFSQLSTANVTQRITIPYSGAFCVIQSRLKQIY 539
P+VG+TVSG+LVKKGFTYQIMAPD+LHVFSQLSTA VTQRITIP+ GAF VI+ RL++I+
Sbjct: 482 PDVGDTVSGILVKKGFTYQIMAPDELHVFSQLSTATVTQRITIPFVGAFGVIKHRLEKIF 541
Query: 540 ESVEPSVDEESGVPMLLVHDRVTVKHESEKHVSLHWASDPINDMVSDSVVALVLNINRDL 599
ESVE S DEESG+P L VH+RVTVK ESEKH+SL W+SDPI+DMVSDS+VAL+LNI+R++
Sbjct: 542 ESVEFSTDEESGLPALKVHERVTVKQESEKHISLQWSSDPISDMVSDSIVALILNISREV 601
Query: 600 PKIV-AESDATKIEEENEKKTEKVMQALLNSLFGNVKVGENGKLIINIDGNVAELNKESG 658
PKIV E DA K EEEN KK EKV+ ALL SLFG+VK+GENGKL+I +DGNVA+L+KESG
Sbjct: 602 PKIVMEEEDAVKSEEENGKKVEKVIYALLVSLFGDVKLGENGKLVIRVDGNVAQLDKESG 661
Query: 659 EVESENEGLKERVRTAFRRIQSSVKPIPLSA 689
EVESE+ GLKERVR AF RIQS+VKPIPLSA
Sbjct: 662 EVESEHSGLKERVRVAFERIQSAVKPIPLSA 692
>At2g01730 putative cleavage and polyadenylation specifity factor
Length = 837
Score = 283 bits (724), Expect = 2e-76
Identities = 160/431 (37%), Positives = 237/431 (54%), Gaps = 23/431 (5%)
Query: 27 LGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPS-----TVDVLLITH 81
LGAG E+G+SCV +T GK ++FDCG+H G P F I S + ++ITH
Sbjct: 8 LGAGQEIGKSCVVVTINGKKIMFDCGMHMGCDDHNRYPNFSLISKSGDFDNAISCIIITH 67
Query: 82 FHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVD----DMLYDEQD 137
FH+DH +LPYF E + G ++M+Y TKA+ L+L DY +V VD + L+
Sbjct: 68 FHMDHVGALPYFTEVCGYNGPIYMSYPTKALSPLMLEDY---RRVMVDRRGEEELFTTTH 124
Query: 138 INRSMDKIEVIDFHQTVEVN-GIRFWCYTAGHVLGAAMFMVDIAGVRVLYTGDYSREEDR 196
I M K+ ID QT++V+ ++ Y AGHVLGA M + ++YTGDY+ DR
Sbjct: 125 IANCMKKVIAIDLKQTIQVDEDLQIRAYYAGHVLGAVMVYAKMGDAAIVYTGDYNMTTDR 184
Query: 197 HLRAAETPQFSPDVCIIESTYGVQHHQPRHTREKRFTDVIHSTISQGGRVLIPAYALGRA 256
HL AA+ + D+ I ESTY ++ RE+ F +H ++ GG+ LIP++ALGRA
Sbjct: 185 HLGAAKIDRLQLDLLISESTYATTIRGSKYPREREFLQAVHKCVAGGGKALIPSFALGRA 244
Query: 257 QELLLILDEYWANHPELQNI--PIYYASPLAKKCLTVYETYT--LSMNDRIQNAKSNPFA 312
QEL ++LD+YW E NI PIY++S L + Y+ S N + ++ NPF
Sbjct: 245 QELCMLLDDYW----ERMNIKVPIYFSSGLTIQANMYYKMLISWTSQNVKEKHNTHNPFD 300
Query: 313 FKHISALSSIDIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVVEGTLA 372
FK++ + GP V+ A+PG L +G S ++F W N +PGY V GT+
Sbjct: 301 FKNVKDFDR-SLIHAPGPCVLFATPGMLCAGFSLEVFKHWAPSPLNLVALPGYSVAGTVG 359
Query: 373 -KTILNEPKEVTLMNGLSAPLHMQVHYISFSAHADSAQTSAFLEELNPPNIILVHGAANE 431
K + +P V L NG + +VH ++FS H D+ + L+P N++LVHG
Sbjct: 360 HKLMAGKPTTVDLYNGTKVDVRCKVHQVAFSPHTDAKGIMDLTKFLSPKNVVLVHGEKPS 419
Query: 432 MGRLKQKLMTQ 442
M LK+K+ ++
Sbjct: 420 MMILKEKITSE 430
>At5g23880 polyadenylation cleavage/specificity factor 100 kDa
subunit
Length = 739
Score = 136 bits (342), Expect = 4e-32
Identities = 106/372 (28%), Positives = 187/372 (49%), Gaps = 24/372 (6%)
Query: 24 VTPL-GAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPSTVDVLLITHF 82
VTPL G NE S + ++ G L DCG + + P ST+D +L++H
Sbjct: 7 VTPLCGVYNENPLSYL-VSIDGFNFLIDCGWNDLFDTSLLEPLSRVA--STIDAVLLSHP 63
Query: 83 HLDHAASLPYFLEKTTFKGRVFMTYATKAIYKL-LLSDYVK-VSKVSVDDM-LYDEQDIN 139
H +LPY +++ V YAT+ +++L LL+ Y + +S+ V D L+ DI+
Sbjct: 64 DTLHIGALPYAMKQLGLSAPV---YATEPVHRLGLLTMYDQFLSRKQVSDFDLFTLDDID 120
Query: 140 RSMDKIEVIDFHQTVEVNG----IRFWCYTAGHVLGAAMFMVDIAGVRVLYTGDYSREED 195
+ + + + Q ++G I + AGH+LG +++ + G V+Y DY+ ++
Sbjct: 121 SAFQNVIRLTYSQNYHLSGKGEGIVIAPHVAGHMLGGSIWRITKDGEDVIYAVDYNHRKE 180
Query: 196 RHLRAAETPQF-SPDVCIIESTYGVQHHQP-RHTREKRFTDVIHSTISQGGRVLIPAYAL 253
RHL F P V I ++ + + +Q R R+K F D I + GG VL+P
Sbjct: 181 RHLNGTVLQSFVRPAVLITDAYHALYTNQTARQQRDKEFLDTISKHLEVGGNVLLPVDTA 240
Query: 254 GRAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYETYTLSMNDRI----QNAKSN 309
GR ELLLIL+++W+ + PIY+ + ++ + +++ M+D I + ++ N
Sbjct: 241 GRVLELLLILEQHWSQRG--FSFPIYFLTYVSSSTIDYVKSFLEWMSDSISKSFETSRDN 298
Query: 310 PFAFKHISALSSIDIFKDV--GPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVV 367
F +H++ L + + GP VV+AS L++G +R++F W +D +N +
Sbjct: 299 AFLLRHVTLLINKTDLDNAPPGPKVVLASMASLEAGFAREIFVEWANDPRNLVLFTETGQ 358
Query: 368 EGTLAKTILNEP 379
GTLA+ + + P
Sbjct: 359 FGTLARMLQSAP 370
>At2g45700 unknown protein
Length = 723
Score = 47.8 bits (112), Expect = 2e-05
Identities = 31/118 (26%), Positives = 53/118 (44%), Gaps = 26/118 (22%)
Query: 79 ITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDI 138
+THFHLDH L K+ G+++ + T + + I
Sbjct: 420 LTHFHLDHYQGLT----KSFSHGKIYCSLVTAKLVNM---------------------KI 454
Query: 139 NRSMDKIEVIDFHQTVEVNGIRFWCYTAGHVLGAAMFMVDIA-GVRVLYTGDYSREED 195
++++V+D Q V ++GI C+ A H G+ M + + A G VL+TGD+ E+
Sbjct: 455 GIPWERLQVLDLGQKVNISGIDVTCFDANHCPGSIMILFEPANGKAVLHTGDFRYSEE 512
>At5g63420 putative protein
Length = 528
Score = 41.2 bits (95), Expect = 0.002
Identities = 28/109 (25%), Positives = 49/109 (44%), Gaps = 14/109 (12%)
Query: 22 LIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGI-HPGYSGMAALPYFDEIDPST------- 73
L + P+G E+G +C+ + + +L D GI P Y P +I P T
Sbjct: 110 LRILPIGGLGEIGMNCMLVGNYDRYILIDAGIMFPDYDE----PGIQKIMPDTGFIRRWK 165
Query: 74 --VDVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDY 120
++ ++ITH H DH +LP+ + +F + T + K L ++
Sbjct: 166 HKIEAVVITHGHEDHIGALPWVIPALDPNTPIFASSFTMELIKKRLKEH 214
>At4g11120 translation elongation factor ts like protein
Length = 395
Score = 35.0 bits (79), Expect = 0.14
Identities = 35/158 (22%), Positives = 69/158 (43%), Gaps = 11/158 (6%)
Query: 538 IYESVEPSVDEESGVPMLLVHDRVTVKHESEK---HVSLHWASDPINDMVSDSVVALVLN 594
++ S +P + +G+ L V T ++ +++H + + D V + +
Sbjct: 243 LHTSPQPGLGRIAGIVSLEVEGENTQLEAIQRVGSELAMHVVAAKPLFLSKDLVSSEAMA 302
Query: 595 INRDLPKIVAESDATKIEEENEKKTEKVMQALLNSLFGNVKVGENGKLI---INIDGNVA 651
R++ K AES +N+ EK+++ L F V + E ++ INI V
Sbjct: 303 NEREILKSQAESTG-----KNQMAIEKIVEGRLRKYFEEVALMEQKFIVNDAINIKTLVD 357
Query: 652 ELNKESGEVESENEGLKERVRTAFRRIQSSVKPIPLSA 689
L+KE G + L+ V R+++S +P+ +A
Sbjct: 358 NLSKEVGSPVKVTDFLRVEVGEGIERLEASDEPVAQTA 395
>At3g26680 unknown protein
Length = 484
Score = 31.6 bits (70), Expect = 1.6
Identities = 29/113 (25%), Positives = 44/113 (38%), Gaps = 26/113 (23%)
Query: 79 ITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDI 138
+THFH DH L K G ++ + T + +L LS +
Sbjct: 174 LTHFHADHYIG----LTKAWSHGPIYCSSLTSRLLRLSLS-------------------V 210
Query: 139 NRSMDKIEVIDFHQTVEVNGIRFWCYTAGHVLGAAMFMVDIA-GVRVLYTGDY 190
N S I ++ +NGI+ A H GAA+ + G L+TGD+
Sbjct: 211 NPS--SIHPLELDVEYTINGIKVTLIEANHCPGAALIHFRLLDGTCYLHTGDF 261
>At1g77600 hypothetical protein
Length = 1303
Score = 31.6 bits (70), Expect = 1.6
Identities = 21/80 (26%), Positives = 37/80 (46%), Gaps = 9/80 (11%)
Query: 592 VLNINRDLPKIVAESDATKIEEENEKKTEKVMQALLNSLFGNVKVGENGKLIINIDGNVA 651
V I+ L K+ A+ T +++N+K K+ S GN GE G+V+
Sbjct: 1215 VPEISHTLAKVTAQKQTTTTKQQNKKVPAKLNPPAAKSKKGNSDSGE---------GSVS 1265
Query: 652 ELNKESGEVESENEGLKERV 671
E+ S ++SE +K+ +
Sbjct: 1266 EVTDTSDNIKSECRLIKKSI 1285
>At5g08660 putative protein
Length = 649
Score = 30.8 bits (68), Expect = 2.6
Identities = 23/91 (25%), Positives = 40/91 (43%), Gaps = 19/91 (20%)
Query: 294 TYTLSMNDRIQNAKSNPFAFKHISALSSI----------DIFKDVGPSVVMASPGGLQSG 343
TYT++ + +I++AKS A ++ S + D+ +G S+ S GG SG
Sbjct: 82 TYTMAPSQKIRSAKSTQTAVSKVTEASKLLGKAGLGRAKDVLDTLGSSMTDLSSGGFTSG 141
Query: 344 LSRQLFDMWCSDKKNSCVIPGYVVEGTLAKT 374
+ + K N I + V T+ K+
Sbjct: 142 V---------ATKGNELGILAFEVANTIVKS 163
>At3g51750 unknown protein (At3g51750)
Length = 110
Score = 30.8 bits (68), Expect = 2.6
Identities = 21/74 (28%), Positives = 38/74 (50%), Gaps = 3/74 (4%)
Query: 600 PKIVAESDATKIEEENEKKTEK--VMQALLNSLFGNVKVGENGKLIINIDGNVAELNKES 657
PK V E + K EEE ++K K ++ L+++ GK NI ++ E +++
Sbjct: 14 PKEVVEKNKDKQEEEEDEKGAKGGILNNLISNFMDCTTTTVEGKSSENIQKSLCE-DEDE 72
Query: 658 GEVESENEGLKERV 671
E ES + G+ E++
Sbjct: 73 KEQESPSVGILEKI 86
>At1g34260 hypothetical protein
Length = 1456
Score = 30.8 bits (68), Expect = 2.6
Identities = 17/64 (26%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Query: 407 SAQTSAFLEELNPPNIILVHGAANEMGRLKQKLMTQFADRNTKILTPKNCQSVEMYFNSQ 466
S++++ LE L PP +++ G+ +G+ K +++ +AD + + L + C S Y S
Sbjct: 1152 SSESTNRLETLPPPEVLVTFGSVKSVGKPKYSIVSLYAD-DFRDLRKRCCSSELDYIASL 1210
Query: 467 KMAK 470
K
Sbjct: 1211 SRCK 1214
>At1g28310 zinc finger like protein OBP3
Length = 311
Score = 30.4 bits (67), Expect = 3.5
Identities = 16/78 (20%), Positives = 34/78 (43%), Gaps = 3/78 (3%)
Query: 251 YALGRAQELLLILDEYWANHPELQNIPI---YYASPLAKKCLTVYETYTLSMNDRIQNAK 307
Y+L + + YW L+N+P+ Y + K+ T T +++ ++
Sbjct: 45 YSLSQPRHFCKACKRYWTRGGTLRNVPVGGSYRKNKRVKRPSTATTTTASTVSTTNSSSP 104
Query: 308 SNPFAFKHISALSSIDIF 325
+NP H S+++ +F
Sbjct: 105 NNPHQISHFSSMNHHPLF 122
>At3g02930 unknown protein
Length = 806
Score = 30.0 bits (66), Expect = 4.5
Identities = 19/82 (23%), Positives = 39/82 (47%), Gaps = 5/82 (6%)
Query: 592 VLNINRDLPKIVAESDATKIEEENEKKTEKVMQALLNSLFGNVKVGENGKLI-----INI 646
+L+ + IV E+D ++++++ K K + LL + ENG+L ++
Sbjct: 615 MLDKETEFQSIVHENDELRVKQDDSLKKIKELSELLEEALAKKHIEENGELSESEKDYDL 674
Query: 647 DGNVAELNKESGEVESENEGLK 668
V E ++E+G +E + K
Sbjct: 675 LPKVVEFSEENGYRSAEEKSSK 696
>At1g72150 putative cytosolic factor protein
Length = 573
Score = 30.0 bits (66), Expect = 4.5
Identities = 29/129 (22%), Positives = 59/129 (45%), Gaps = 21/129 (16%)
Query: 454 KNCQSVEMYFNSQKMAKTIGKLAEKTPEVGETVSGLLVKKGFTYQIMAPDDLHVFSQLST 513
K+ ++ Y +++ G L++ TP ET++ +VK Y I P S+ T
Sbjct: 442 KSADTIFKYIAPEQVPVKYGGLSKDTPLTEETITEAIVKPAANYTIELP-----ASEACT 496
Query: 514 ANVTQRI---TIPY--------SGAFCVIQSRLKQIYESVEPSVDE--ESGVPMLLVHDR 560
+ R+ + Y G++ VI S+ ++I + EP + + + G P +V
Sbjct: 497 LSWELRVLGADVSYGAQFEPTTEGSYAVIVSKTRKIGSTDEPVITDSFKVGEPGKIV--- 553
Query: 561 VTVKHESEK 569
+T+ +++ K
Sbjct: 554 ITIDNQTSK 562
>At1g08060 storage protein, putative
Length = 1892
Score = 29.6 bits (65), Expect = 5.9
Identities = 27/113 (23%), Positives = 46/113 (39%), Gaps = 7/113 (6%)
Query: 435 LKQKLMTQFADRNTKILTPKNCQSVEMYFNSQKMAKTIGKLAEKTPEVGETVSGLLVKKG 494
L+++ +F D NT L P N +S + + K + T E G+ V
Sbjct: 465 LEKEYPQKFQDDNTDCLPPANAESKRLPVGETSLEKDTDFPLKSTTETGKMVL------- 517
Query: 495 FTYQIMAPDDLHVFSQLSTANVTQRITIPYSGAFCVIQSRLKQIYESVEPSVD 547
+ I+ D V ST TQ++ + +G I LK+ ++ E +D
Sbjct: 518 YASPIVETRDDSVICSPSTNLETQKLLVSKTGLETDIVLPLKRKRDTAEIELD 570
>At3g49600 putative protein
Length = 1672
Score = 29.3 bits (64), Expect = 7.7
Identities = 21/85 (24%), Positives = 41/85 (47%), Gaps = 4/85 (4%)
Query: 597 RDLPKIVAESDATKIEEENEKKTEKVMQALLNSLFGNVK----VGENGKLIINIDGNVAE 652
RD K +SD ++IE +N+K+ ++ + K V ++GK D E
Sbjct: 386 RDDKKAARDSDDSEIEYQNKKQLRSKVEVYSAGMSQKRKEEEDVTKHGKDKYRSDSRGKE 445
Query: 653 LNKESGEVESENEGLKERVRTAFRR 677
+ ++S + E+E E K+ +++R
Sbjct: 446 VARDSDDSEAEYENRKKLKNESYQR 470
>At1g74700 unknown protein
Length = 281
Score = 29.3 bits (64), Expect = 7.7
Identities = 19/87 (21%), Positives = 38/87 (42%), Gaps = 15/87 (17%)
Query: 75 DVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYD 134
D L I+H H+DH LP ++ AT+ +YK + + S+ + +
Sbjct: 48 DFLFISHSHMDHIGGLPMYV-------------ATRGLYK--MKPPTIIVPASIKETVES 92
Query: 135 EQDINRSMDKIEVIDFHQTVEVNGIRF 161
+++R +D E+ +++ G F
Sbjct: 93 LFEVHRKLDSSELKHNLVGLDIGGEEF 119
>At1g35620 protein disulfide isomerase, putative (At1g35620)
Length = 440
Score = 29.3 bits (64), Expect = 7.7
Identities = 16/58 (27%), Positives = 30/58 (51%), Gaps = 4/58 (6%)
Query: 580 INDMVSDSVVALVLNINRDLPKIVAESD----ATKIEEENEKKTEKVMQALLNSLFGN 633
+ + V S + L+L IN D K++ + + T +E+E + EK+ +AL + N
Sbjct: 232 LEEFVKQSFLPLILPINHDTLKLLKDDERKIVLTIVEDETHESLEKLYKALRAAAHAN 289
>At1g19025 unknown protein
Length = 549
Score = 29.3 bits (64), Expect = 7.7
Identities = 27/112 (24%), Positives = 50/112 (44%), Gaps = 21/112 (18%)
Query: 79 ITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDI 138
+TH H DH + + VF Y+T LLL + ++ D+ + +I
Sbjct: 31 LTHAHKDHTVGV------SPSNIVVFPIYSTSLTISLLLQRFPQL-----DESYFVRVEI 79
Query: 139 NRSMDKIEVIDFHQTVEVNG-IRFWCYTAGHVLGAAMFMVDIAGVRVLYTGD 189
+S+ ++D + +G + + A H GA MF+ + + +L+TGD
Sbjct: 80 GQSV----IVD-----DPDGEFKVTAFDANHCPGAVMFLFEGSFGNILHTGD 122
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,975,003
Number of Sequences: 26719
Number of extensions: 644881
Number of successful extensions: 1897
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 1876
Number of HSP's gapped (non-prelim): 23
length of query: 690
length of database: 11,318,596
effective HSP length: 106
effective length of query: 584
effective length of database: 8,486,382
effective search space: 4956047088
effective search space used: 4956047088
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC134322.16


