

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134049.3 - phase: 0
(1113 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
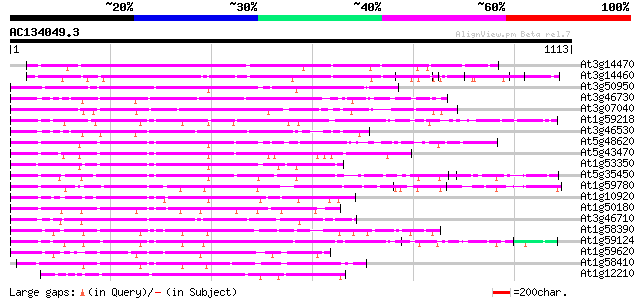
At3g14470 disease resistance protein, putative 465 e-131
At3g14460 disease resistance protein, putative 388 e-108
At3g50950 putative disease resistance protein 243 6e-64
At3g46730 disease resistance protein RPP8-like protein 232 1e-60
At3g07040 disease resistance gene (RPM1) 231 2e-60
At1g59218 resistance protein RPP13, putative 219 7e-57
At3g46530 disease resistance protein RPP13-like protein 219 9e-57
At5g48620 disease resistance protein 214 2e-55
At5g43470 disease resistance protein RPP8 214 3e-55
At1g53350 hypothetical protein 211 2e-54
At5g35450 disease resistance protein 207 2e-53
At1g59780 hypothetical protein 204 3e-52
At1g10920 disease resistance protein RPM1 isolog 199 1e-50
At1g50180 disease resistance protein, putative 198 1e-50
At3g46710 disease resistance protein RPP8-like protein 194 3e-49
At1g58390 disease resistance protein PRM1 192 7e-49
At1g59124 PRM1 homolog (RF45) 185 1e-46
At1g59620 PRM1 homolog (CW9) 184 2e-46
At1g58410 unknown protein 182 1e-45
At1g12210 NBS/LRR disease resistance protein 172 7e-43
>At3g14470 disease resistance protein, putative
Length = 1054
Score = 465 bits (1197), Expect = e-131
Identities = 327/997 (32%), Positives = 534/997 (52%), Gaps = 85/997 (8%)
Query: 33 RLASLLTTIKATLEDAEEKQFSDSEIGRDVKDWLLKLKDAAYTLDDIMDECATEALEMEY 92
RL++ L TI A L DAEEKQ ++ V+ W+ +L+D Y +D +D+ ATEAL +
Sbjct: 41 RLSTALLTITAVLIDAEEKQITNPV----VEKWVNELRDVVYHAEDALDDIATEALRLNI 96
Query: 93 KASKCGLSH-KHIAFRYKLAK-----------KMKRIGVWLDDIAAEKNKFHLTEIVRER 140
A + + + R L +++++ + L+ +A+++N L E+
Sbjct: 97 GAESSSSNRLRQLRGRMSLGDFLDGNSEHLETRLEKVTIRLERLASQRNILGLKEL---- 152
Query: 141 SGVVPDWR-QTTSIVTQPLVYGRNEDKDKIVDFLVGDASEQEDLSVYPIVGLGGLGKTTL 199
+ ++P R TTS+V + V+GR++DKD+I+ FL+ + + ++V IVG+GG+GKTTL
Sbjct: 153 TAMIPKQRLPTTSLVDESEVFGRDDDKDEIMRFLIPENGKDNGITVVAIVGIGGVGKTTL 212
Query: 200 AQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGATKKSCEDLDLELLQRKLQDLL 259
+QL++N + ++F K+W VSE+F + ++TK + E T + CE DL++LQ KL++ L
Sbjct: 213 SQLLYNDQHVRSYFGTKVWAHVSEEFDVFKITKKVYESVTSRPCEFTDLDVLQVKLKERL 272
Query: 260 RRK--RYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTTRLPKVAKIMGTIPHHELS 317
+LLVLDD+WN+ +W L+ +G+ ILVTTR +VA IM + H L
Sbjct: 273 TGTGLPFLLVLDDLWNENFADWDLLRQPFIHAAQGSQILVTTRSQRVASIMCAVHVHNLQ 332
Query: 318 RLSDEDCWELFKQRAFGPNE-VQQKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKE 376
LSD DCW LF + FG E +E+ + + I+ KC G PLA LG +LRF+ + E
Sbjct: 333 PLSDGDCWSLFMKTVFGNQEPCLNREIGDLAERIVHKCRGLPLAVKTLGGVLRFEGKVIE 392
Query: 377 WLYVKESKLWNLQGE-AYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWT 435
W V S++W+L + + ++P LR+SY +LP L++CF++C++FPK K ++ LW
Sbjct: 393 WERVLSSRIWDLPADKSNLLPVLRVSYYYLPAHLKRCFAYCSIFPKGHAFEKDKVVLLWM 452
Query: 436 ANGFISSNQMLE-ADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQ 494
A GF+ + + +++GNE ++EL RS + T+ T + MHD +++LA +
Sbjct: 453 AEGFLQQTRSSKNLEELGNEYFSELESRSLLQKTK-------TRYIMHDFINELAQFASG 505
Query: 495 DVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQ-LHHVKSLKTYMEFNFDVYEAGQL 553
+ +D +SE TR+ L Y R+++AE + L VK L+T++ +
Sbjct: 506 EFSSKFEDGCKLQVSERTRY-LSYLRDNYAEPMEFEALREVKFLRTFLPLSLTNSSRSCC 564
Query: 554 SPQVLNCYSLRVLLSHRLNNLSS-SIGRL--------KYLRYLDISEGRFKNLPNSLCKL 604
Q+++ L L R+ +LS I RL + R+LD+S + LP SLC +
Sbjct: 565 LDQMVSEKLLPTLTRLRVLSLSHYKIARLPPDFFKNISHARFLDLSRTELEKLPKSLCYM 624
Query: 605 CNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVG 664
NL+ L L C SL++LP ++ L L+ L L L +PR+ G+L SL TL+ + V
Sbjct: 625 YNLQTLLLSYCSSLKELPTDISNLINLRYLDLIG-TKLRQMPRRFGRLKSLQTLTTFFVS 683
Query: 665 EERGFLLEELGQL-NLKGQLHIKNLERLKSVTDAKKANM-SRKKLNQLWLSW-------E 715
G + ELG L +L G+L I L+R+ V DA +AN+ S+K L ++ W E
Sbjct: 684 ASDGSRISELGGLHDLHGKLKIVELQRVVDVADAAEANLNSKKHLREIDFVWRTGSSSSE 743
Query: 716 RNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKS 775
N +N ++ E L+P+ + + + Y G FP W+S PS + + + L +C+
Sbjct: 744 NNTNPHRTQNEAEVFEKLRPH-RHIEKLAIERYKGRRFPDWLSDPSFSRIVCIRLRECQY 802
Query: 776 CLNLPELWKLPSLKYLKLSNMIHVIYLFHESY---------DGEGLMALKTLFLEKLPNL 826
C +LP L +LP LK L +S M+ + + + Y D + +L+TL + LP+
Sbjct: 803 CTSLPSLGQLPCLKELHISGMVGLQSIGRKFYFSDQQLRDQDQQPFRSLETLRFDNLPDW 862
Query: 827 -----IGLSREERVMFPRLKALEITECPNLLG-LPC-LPSLSDLYIQ--GKYNQQLPSSI 877
+ ++R + +FP LK L I CP L G LP LPSL L+I G + Q
Sbjct: 863 QEWLDVRVTRGD--LFPSLKKLFILRCPELTGTLPTFLPSLISLHIYKCGLLDFQPDHHE 920
Query: 878 HKLGSLESLHF-SDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHI---HAL 933
+ +L++L S + L+ FP L + A+ L L + + L L H+ +AL
Sbjct: 921 YSYRNLQTLSIKSSCDTLVKFP---LNHFAN-LDKLEVDQCTSLYSLELSNEHLRGPNAL 976
Query: 934 QQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLK 970
+ L INDC+N++ LP + L ++ I C L+
Sbjct: 977 RNLRINDCQNLQLLPK--LNALPQNLQVTITNCRYLR 1011
Score = 41.2 bits (95), Expect = 0.003
Identities = 31/125 (24%), Positives = 65/125 (51%), Gaps = 7/125 (5%)
Query: 870 NQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIH 929
+++L ++ +L L H+ ++ P +N+ S + L R ++L+ LP + +
Sbjct: 570 SEKLLPTLTRLRVLSLSHY----KIARLPPDFFKNI-SHARFLDLSR-TELEKLPKSLCY 623
Query: 930 IHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSC 989
++ LQ L ++ C +++ELP ++ L +L+ LD++G ++ F L L+TL
Sbjct: 624 MYNLQTLLLSYCSSLKELPTDI-SNLINLRYLDLIGTKLRQMPRRFGRLKSLQTLTTFFV 682
Query: 990 SEVEG 994
S +G
Sbjct: 683 SASDG 687
Score = 32.0 bits (71), Expect = 2.0
Identities = 27/115 (23%), Positives = 51/115 (43%), Gaps = 6/115 (5%)
Query: 921 KMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQYLTC 980
K+LPT + L+ L ++ + I LP + + + + LD+ + KL Y+
Sbjct: 572 KLLPT----LTRLRVLSLSHYK-IARLPPDFFKNISHARFLDLSRTELEKLPKSLCYMYN 626
Query: 981 LETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIY 1035
L+TL + CS ++ + ++ L+ L L L +P G L L + +
Sbjct: 627 LQTLLLSYCSSLKELPTDISNLINLRYLDLIG-TKLRQMPRRFGRLKSLQTLTTF 680
>At3g14460 disease resistance protein, putative
Length = 1424
Score = 388 bits (997), Expect = e-108
Identities = 282/866 (32%), Positives = 443/866 (50%), Gaps = 70/866 (8%)
Query: 33 RLASLLTTIKATLEDAEEKQFSDSEIGRDVKDWLLKLKDAAYTLDDIMDECATEALEMEY 92
RL L T L DA+++ +E R+VK WL +KDA + +DI+DE TEAL
Sbjct: 38 RLKVALVTANPVLADADQR----AEHVREVKHWLTGIKDAFFQAEDILDELQTEALRRRV 93
Query: 93 KASKCGLSHK-------HIAFRYKLAKKMKRIGVWLDDIAAEKNKFHLTEIVRERSGVVP 145
A GL A + K+ KM+++ L+ L E R P
Sbjct: 94 VAEAGGLGGLFQNLMAGREAIQKKIEPKMEKVVRLLEHHVKHIEVIGLKEYSETRE---P 150
Query: 146 DWRQTTSI----VTQPLVYGRNEDKDKIVDFLVGDASEQEDLS-----VYPIVGLGGLGK 196
WRQ + + Q + GR EDK +V+ L+ D +++S V +VG+ G+GK
Sbjct: 151 QWRQASRSRPDDLPQGRLVGRVEDKLALVNLLLSD----DEISIGKPAVISVVGMPGVGK 206
Query: 197 TTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGATKKSCEDLDLELLQRKLQ 256
TTL ++VFN ++ HFE+K+W+ +F + +TKA+++ T + DL LQ +L+
Sbjct: 207 TTLTEIVFNDYRVTEHFEVKMWISAGINFNVFTVTKAVLQDITSSAVNTEDLPSLQIQLK 266
Query: 257 DLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTTRLPKVAKIMGTIPHHEL 316
L KR+LLVLDD W++ W+ + +G+ I++TTR V+ + +++
Sbjct: 267 KTLSGKRFLLVLDDFWSESDSEWESFQVAFTDAEEGSKIVLTTRSEIVSTVAKAEKIYQM 326
Query: 317 SRLSDEDCWELFKQRAFGPNEVQ--QKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREE 374
+++E+CWEL + AFG V +EL +GK I ++C G PLAA A+ S LR K
Sbjct: 327 KLMTNEECWELISRFAFGNISVGSINQELEGIGKRIAEQCKGLPLAARAIASHLRSKPNP 386
Query: 375 KEWLYVKESKLWNLQGEAYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLW 434
+W V SK ++ + ++P L+LSY LP +L++CF+ C++FPK + ++ L+ LW
Sbjct: 387 DDWYAV--SKNFSSYTNS-ILPVLKLSYDSLPPQLKRCFALCSIFPKGHVFDREELVLLW 443
Query: 435 TANGFI---SSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGS 491
A + S++ LE DIGN+ +L +SFF+ + +T F MHDL++DLA +
Sbjct: 444 MAIDLLYQPRSSRRLE--DIGNDYLGDLVAQSFFQRLDIT----MTSFVMHDLMNDLAKA 497
Query: 492 VTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDV-YEA 550
V+ D C +D+++ + TRH A + + L+T + FN E+
Sbjct: 498 VSGDFCFRLEDDNIPEIPSTTRHFSFSRSQCDASVAFRSICGAEFLRTILPFNSPTSLES 557
Query: 551 GQLSPQVLN-----CYSLRVL-LSH-RLNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCK 603
QL+ +VLN LR+L LSH ++ NL S+ LK LRYLD+S + K LP +C
Sbjct: 558 LQLTEKVLNPLLNALSGLRILSLSHYQITNLPKSLKGLKLLRYLDLSSTKIKELPEFVCT 617
Query: 604 LCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIV 663
LCNL+ L L C L LP + L L+ L L L +P I KL SL LS +++
Sbjct: 618 LCNLQTLLLSNCRDLTSLPKSIAELINLRLLDLVG-TPLVEMPPGIKKLRSLQKLSNFVI 676
Query: 664 GEERGFLLEELGQL-NLKGQLHIKNLERLKSVTDAKKANMSRKK-LNQLWLSWE------ 715
G G L EL +L +L+G L I L+ + ++AK A + RK L+ L L W
Sbjct: 677 GRLSGAGLHELKELSHLRGTLRISELQNVAFASEAKDAGLKRKPFLDGLILKWTVKGSGF 736
Query: 716 -RNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCK 774
+ L + +++L L+P+ L +F + Y G FP+W+ S + S+ L C
Sbjct: 737 VPGSFNALACDQKEVLRMLEPHPH-LKTFCIESYQGGAFPKWLGDSSFFGITSVTLSSCN 795
Query: 775 SCLNLPELWKLPSLKYLKLSNM-----IHVIYLFHESYDG----EGLMALKTLFLEKLPN 825
C++LP + +LPSLKYL + + + + F E+ + L LK + +
Sbjct: 796 LCISLPPVGQLPSLKYLSIEKFNILQKVGLDFFFGENNSRGVPFQSLQILKFYGMPRWDE 855
Query: 826 LIGLSREERVMFPRLKALEITECPNL 851
I E+ + FP L+ L I CP+L
Sbjct: 856 WICPELEDGI-FPCLQKLIIQRCPSL 880
Score = 55.8 bits (133), Expect = 1e-07
Identities = 67/257 (26%), Positives = 105/257 (40%), Gaps = 32/257 (12%)
Query: 840 LKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEELIYFPD 899
L++L I C L LP +L++ Y ++ L + H L S H + +Y D
Sbjct: 1093 LQSLHIDSCDGLTSLP--ENLTESY--PNLHELLIIACHSLESFPGSHPPTTLKTLYIRD 1148
Query: 900 GILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYI-NDCRNIEELPNEVMQRLHSL 958
N L+ PT L+ L+I + C N+ P + +L SL
Sbjct: 1149 CKKLNFTESLQ-------------PTRSYS--QLEYLFIGSSCSNLVNFPLSLFPKLRSL 1193
Query: 959 KELDIVGCDKLKLSSDFQYL----TCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLP 1014
D C+ K S L LE+L I C +E F + L S+ LS+
Sbjct: 1194 SIRD---CESFKTFSIHAGLGDDRIALESLEIRDCPNLETFPQGGLPTPKLSSMLLSNCK 1250
Query: 1015 NLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGE 1074
L+ LPE + LT L + I CP++ +P S L L I C KL R + + +
Sbjct: 1251 KLQALPEKLFGLTSLLSLFIIKCPEIETIPGG-GFPSNLRTLCISLCDKLTPRIEWGLRD 1309
Query: 1075 DWPKIVHVQYIEIENDN 1091
+ +++ +EI+ N
Sbjct: 1310 ----LENLRNLEIDGGN 1322
Score = 49.7 bits (117), Expect = 9e-06
Identities = 76/271 (28%), Positives = 106/271 (39%), Gaps = 62/271 (22%)
Query: 765 LKSLELVDCKSCLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGLMALKTLFLEKLP 824
LK+L + DCK LN E + P+ Y +L YLF G L L P
Sbjct: 1141 LKTLYIRDCKK-LNFTESLQ-PTRSYSQLE------YLFI----GSSCSNLVNFPLSLFP 1188
Query: 825 NLIGLSREERVMFPR-------------LKALEITECPNLL-----GLPCLPSLSDLYIQ 866
L LS + F L++LEI +CPNL GLP P LS + +
Sbjct: 1189 KLRSLSIRDCESFKTFSIHAGLGDDRIALESLEIRDCPNLETFPQGGLPT-PKLSSMLLS 1247
Query: 867 G-KYNQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKL----- 920
K Q LP + L SL SL E+ P G S L+TL KL
Sbjct: 1248 NCKKLQALPEKLFGLTSLLSLFIIKCPEIETIPGG---GFPSNLRTLCISLCDKLTPRIE 1304
Query: 921 ---------------------KMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLK 959
+ P E + ++ L I+ N++ L + +++
Sbjct: 1305 WGLRDLENLRNLEIDGGNEDIESFPEEGLLPKSVFSLRISRFENLKTLNRKGFHDTKAIE 1364
Query: 960 ELDIVGCDKLKLSSDFQYLTCLETLAIGSCS 990
++I GCDKL++S D + L L L I SCS
Sbjct: 1365 TMEISGCDKLQISID-EDLPPLSCLRISSCS 1394
Score = 45.4 bits (106), Expect = 2e-04
Identities = 37/128 (28%), Positives = 65/128 (49%), Gaps = 10/128 (7%)
Query: 902 LRNLASPLKTLGFHRH-----SKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLH 956
+ NL LK L R+ +K+K LP + + LQ L +++CR++ LP + + L
Sbjct: 585 ITNLPKSLKGLKLLRYLDLSSTKIKELPEFVCTLCNLQTLLLSNCRDLTSLPKSIAE-LI 643
Query: 957 SLKELDIVGCDKLKLSSDFQYLTCLETLA---IGSCSEVEGFHEALQHMTTLKSLTLSDL 1013
+L+ LD+VG +++ + L L+ L+ IG S G HE + +L +S+L
Sbjct: 644 NLRLLDLVGTPLVEMPPGIKKLRSLQKLSNFVIGRLSGA-GLHELKELSHLRGTLRISEL 702
Query: 1014 PNLEYLPE 1021
N+ + E
Sbjct: 703 QNVAFASE 710
Score = 34.7 bits (78), Expect = 0.31
Identities = 20/57 (35%), Positives = 33/57 (57%), Gaps = 2/57 (3%)
Query: 607 LEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIV 663
LE L++ C +L+ P G +L ++ L +C L +LP ++ LTSL LS +I+
Sbjct: 1217 LESLEIRDCPNLETFPQGGLPTPKLSSMLLSNCKKLQALPEKLFGLTSL--LSLFII 1271
>At3g50950 putative disease resistance protein
Length = 852
Score = 243 bits (619), Expect = 6e-64
Identities = 223/794 (28%), Positives = 365/794 (45%), Gaps = 47/794 (5%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M +AV+ + L ++ ++ + ++ L S L +++ L+DAE ++ ++ +
Sbjct: 1 MVDAVVTVFLEKTLNILEEKGRTVSDYRKQLEDLQSELKYMQSFLKDAERQKRTNETLRT 60
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALE--MEYKASKCGLSHKH---IAFRYKLAKKMK 115
V D L++ Y +DI+ +C + E ++S LS H + +YK +K+++
Sbjct: 61 LVAD----LRELVYEAEDILVDCQLADGDDGNEQRSSNAWLSRLHPARVPLQYKKSKRLQ 116
Query: 116 RIGVWLDDIAAEKNKFH--LTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFL 173
I + I ++ + +T R W TQ V G DK KI ++L
Sbjct: 117 EINERITKIKSQVEPYFEFITPSNVGRDNGTDRWSSPVYDHTQ--VVGLEGDKRKIKEWL 174
Query: 174 VGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKA 233
S L + VG+GGLGKTT+AQ VFN +I + FE +IWV VS+ FT +++ ++
Sbjct: 175 F--RSNDSQLLIMAFVGMGGLGKTTIAQEVFNDKEIEHRFERRIWVSVSQTFTEEQIMRS 232
Query: 234 IIEGATKKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGA 293
I+ S D D+ L RK+Q L KRYL+V+DDVW+ W ++ L G+G
Sbjct: 233 ILRNLGDASVGD-DIGTLLRKIQQYLLGKRYLIVMDDVWDKNLSWWDKIYQGLP-RGQGG 290
Query: 294 SILVTTRLPKVAKIMGTIPH--HELSRLSDEDCWELFKQRAFGPNE--VQQKELVIVGKE 349
S++VTTR VAK + H LS ++ W LF AF N+ ++ EL VGKE
Sbjct: 291 SVIVTTRSESVAKRVQARDDKTHRPELLSPDNSWLLFCNVAFAANDGTCERPELEDVGKE 350
Query: 350 IIKKCGGFPLAAIALGSLLRFK-REEKEWLYVKESKLWNLQGEA----YVMPALRLSYLH 404
I+ KC G PL A+G LL K EW + E L+G VM +L+LSY
Sbjct: 351 IVTKCKGLPLTIKAVGGLLLCKDHVYHEWRRIAEHFQDELRGNTSETDNVMSSLQLSYDE 410
Query: 405 LPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSF 464
LP L+ C +L+P+D +I KQ L+ W GF+ A + G + ++ L R
Sbjct: 411 LPSHLKSCILTLSLYPEDCVIPKQQLVHGWIGEGFVMWRNGRSATESGEDCFSGLTNRCL 470
Query: 465 FENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFA 524
E + G I K+HD+V DL + + D+ RHL I +F
Sbjct: 471 IEVVDKTYSGTIITCKIHDMVRDLVIDIAK------KDSFSNPEGLNCRHLGI--SGNFD 522
Query: 525 EANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVL------LSHRLNNLSSSI 578
E H ++ + + + L+ + +C LRVL L+ + I
Sbjct: 523 EKQIKVNHKLRGVVSTTKTGEVNKLNSDLAKKFTDCKYLRVLDISKSIFDAPLSEILDEI 582
Query: 579 GRLKYLRYLDISEGR-FKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLR 637
L++L L +S P S+ L NL++L C +L++L + K+L L +
Sbjct: 583 ASLQHLACLSLSNTHPLIQFPRSMEDLHNLQILDASYCQNLKQLQPCIVLFKKLLVLDMT 642
Query: 638 DCDSLTSLPRQIGKLTSLNTLSKY-IVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTD 696
+C SL P+ IG L L L + G L E+ L +L + +L R + +
Sbjct: 643 NCGSLECFPKGIGSLVKLEVLLGFKPARSNNGCKLSEVKNLTNLRKLGL-SLTRGDQIEE 701
Query: 697 AKKANMSRKKLNQLWLSWERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQW 756
+ ++ L++L +S N +++ ++AL P +L+ + Y G P W
Sbjct: 702 EELDSLI--NLSKL-MSISINCYDSYGDDLITKIDALTP-PHQLHELSLQFYPGKSSPSW 757
Query: 757 ISIPSLNDLKSLEL 770
+S L L+ + +
Sbjct: 758 LSPHKLPMLRYMSI 771
Score = 35.0 bits (79), Expect = 0.24
Identities = 22/84 (26%), Positives = 40/84 (47%), Gaps = 3/84 (3%)
Query: 973 SDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEI 1032
+ Q+L CL ++ + + F +++ + L+ L S NL+ L CI L +
Sbjct: 583 ASLQHLACL---SLSNTHPLIQFPRSMEDLHNLQILDASYCQNLKQLQPCIVLFKKLLVL 639
Query: 1033 NIYSCPKLACLPTSIQQISGLEIL 1056
++ +C L C P I + LE+L
Sbjct: 640 DMTNCGSLECFPKGIGSLVKLEVL 663
>At3g46730 disease resistance protein RPP8-like protein
Length = 847
Score = 232 bits (591), Expect = 1e-60
Identities = 241/905 (26%), Positives = 408/905 (44%), Gaps = 114/905 (12%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M +AV VL + + E+ +G + L + LT I L+D E ++ D E+
Sbjct: 1 MVDAVTGFVLNKIGGYLINEVLALMGVKDDLEELKTELTCIHGYLKDVEARERED-EVS- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIAFRYKLAKKMKRIGVW 120
K+W + D AY ++D++D T L++E ++ + GL + K+ KK +
Sbjct: 59 --KEWTKLVLDIAYDIEDVLD---TYFLKLEERSLRRGL----LRLTNKIGKKRDAYNI- 108
Query: 121 LDDIAAEKNKFHLTEIVRERSGV---------------VPDWRQTTSIVTQPLVYGRNED 165
++DI K + RE G+ V R+ + + LV G +D
Sbjct: 109 VEDIRTLKRRILDITRKRETFGIGSFNEPRGENITNVRVRQLRRAPPVDQEELVVGLEDD 168
Query: 166 KDKIVDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDF 225
++ L+ D +E++ + I G+GGLGKT LA+ ++N + F+ + W VS+++
Sbjct: 169 VKILLVKLLSD-NEKDKSYIISIFGMGGLGKTALARKLYNSGDVKRRFDCRAWTYVSQEY 227
Query: 226 TLKRMTKAIIEGATKKSCEDLDL-------ELLQRKLQDLLRRKRYLLVLDDVWNDKQEN 278
+ + II S E+++ E L+ L LL K Y++V+DDVW+ +
Sbjct: 228 KTRDILIRIIRSLGIVSAEEMEKIKMFEEDEELEVYLYGLLEGKNYMVVVDDVWDP--DA 285
Query: 279 WQRLKSVLACGGKGASILVTTRLPKVAK-IMGTIPHHELSRLSDEDCWELFKQRAFGPNE 337
W+ LK L C +G+ +++TTR+ +A+ + GT+ H+L L+ E+ W LF+++AF E
Sbjct: 286 WESLKRALPCDHRGSKVIITTRIRAIAEGVEGTVYAHKLRFLTFEESWTLFERKAFSNIE 345
Query: 338 VQQKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWN--LQGEAYVM 395
++L GKE++KKCGG PLA + L LL KR EW V S LW ++
Sbjct: 346 KVDEDLQRTGKEMVKKCGGLPLAIVVLSGLLSRKR-TNEWHEVCAS-LWRRLKDNSIHIS 403
Query: 396 PALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEV 455
LS+ + +L+ CF + ++FP+D I + LI L A GFI ++ + +D+
Sbjct: 404 TVFDLSFKEMRHELKLCFLYFSVFPEDYEIKVEKLIHLLVAEGFIQEDEEMMMEDVARCY 463
Query: 456 WNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHL 515
+EL RS + E + G++ ++HDL+ DLA +++ + N + S+ R
Sbjct: 464 IDELVDRSLVK-AERIERGKVMSCRIHDLLRDLAIKKAKELNFVNVYNEKQHSSDICRRE 522
Query: 516 LIYN-RNSFAEANSIQLHHVKSLKTYME-FNFDVYEAGQLS---PQVLNCYSLRVLLSHR 570
++++ N + + ++S E F L +VLN L + +
Sbjct: 523 VVHHLMNDYYLCDRRVNKRMRSFLFIGERRGFGYVNTTNLKLKLLRVLNMEGLLFVSKNI 582
Query: 571 LNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKR 630
N L IG L +LRYL I++ LP S+ L L+ L G Q
Sbjct: 583 SNTLPDVIGELIHLRYLGIADTYVSILPASISNLRFLQTLDASGNDPFQ----------- 631
Query: 631 LQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLN------LKGQLH 684
D LTSL IGK + + ++GE G L+ L ++ L +L
Sbjct: 632 ----YTTDLSKLTSLRHVIGKF-----VGECLIGE--GVNLQTLRSISSYSWSKLNHEL- 679
Query: 685 IKNLERLKSVTDAKKANMSRKKLNQLWLSWERNEVSQLQENVEQILEALQPYAQKLYSFG 744
++NL+ L+ +K + R LN ++S+ + +N+ + ++ + S
Sbjct: 680 LRNLQDLEIYDHSKWVDQRRVPLN--FVSFSK------PKNLRVLKLEMRNFKLSSESRT 731
Query: 745 VGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLN-LPELWKLPSLKYLKLSNMIHVIYLF 803
G FP L+SL LV N +P L KLP L+ L L +
Sbjct: 732 TIGLVDVNFP---------SLESLTLVGTTLEENSMPALQKLPRLEDLVLKDC------- 775
Query: 804 HESYDGEGLMALKTLFLEKLPNLIGLSREERVMFPRLKALEITECPNLLGLPCLPSLSDL 863
+Y G +M++ +L NL +S E R L L I E +PSL L
Sbjct: 776 --NYSGVKIMSISAQGFGRLKNL-EMSMERR--GHGLDELRIEE-------EAMPSLIKL 823
Query: 864 YIQGK 868
++G+
Sbjct: 824 TVKGR 828
>At3g07040 disease resistance gene (RPM1)
Length = 926
Score = 231 bits (588), Expect = 2e-60
Identities = 234/950 (24%), Positives = 431/950 (44%), Gaps = 133/950 (14%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEK--QFSDSEI 58
MA A ++ +G + ++ E L G E +++ L +K+ LED + S +
Sbjct: 1 MASATVDFGIGRILSVLENETLLLSGVHGEIDKMKKELLIMKSFLEDTHKHGGNGSTTTT 60
Query: 59 GRDVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH--KHIAFRYKLAKKMKR 116
+ + ++ +D AY ++DI+DE A H +++ R+ +A+K+
Sbjct: 61 TQLFQTFVANTRDLAYQIEDILDEFGYHIHGYRSCAKIWRAFHFPRYMWARHSIAQKLGM 120
Query: 117 IGVWLDDIAAEKNKFHLTEIVRERSGVVP-----DWRQTTSIVTQPLVYGRNE------D 165
+ V + I+ +++ +E ++ ++P D + +I L + N
Sbjct: 121 VNVMIQSISDSMKRYYHSE--NYQAALLPPIDDGDAKWVNNISESSLFFSENSLVGIDAP 178
Query: 166 KDKIVDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDF 225
K K++ L+ S + V +VG+GG GKTTL+ +F + HFE WV +S+ +
Sbjct: 179 KGKLIGRLL---SPEPQRIVVAVVGMGGSGKTTLSANIFKSQSVRRHFESYAWVTISKSY 235
Query: 226 TLKRMTKAIIEGATKKSCEDLDLEL-------LQRKLQDLLRRKRYLLVLDDVWNDKQEN 278
++ + + +I+ K++ + EL L KL + L+ KRY++VLDDVW
Sbjct: 236 VIEDVFRTMIKEFYKEADTQIPAELYSLGYRELVEKLVEYLQSKRYIVVLDDVWTTGL-- 293
Query: 279 WQRLKSVLACGGKGASILVTTRLPKVAKIMGTI--PHHELSRLSDEDCWELFKQRAFGPN 336
W+ + L G G+ +++TTR VA I HE+ L +++ W LF +AF P
Sbjct: 294 WREISIALPDGIYGSRVMMTTRDMNVASFPYGIGSTKHEIELLKEDEAWVLFSNKAF-PA 352
Query: 337 EVQQ---KELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGE-- 391
++Q + L + ++++++C G PLA +LGS++ K+ E EW V + W L
Sbjct: 353 SLEQCRTQNLEPIARKLVERCQGLPLAIASLGSMMSTKKFESEWKKVYSTLNWELNNNHE 412
Query: 392 -AYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADD 450
V + LS+ LP L++CF +C+LFP + + ++ LI +W A F+ + ++A++
Sbjct: 413 LKIVRSIMFLSFNDLPYPLKRCFLYCSLFPVNYRMKRKRLIRMWMAQRFVEPIRGVKAEE 472
Query: 451 IGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVT--QDVCCITDDNSMRTM 508
+ + NEL +R+ + FG+ FKMHD++ ++A SV+ + C + +D+S
Sbjct: 473 VADSYLNELVYRNMLQVILWNPFGRPKAFKMHDVIWEIALSVSKLERFCDVYNDDSDGDD 532
Query: 509 SEET------RHLLIYNRNSFAEANSIQLHHV---KSLKTYMEFNFDVYEAGQLSPQVLN 559
+ ET RHL I + + LH + S K ME L P LN
Sbjct: 533 AAETMENYGSRHLCIQKEMTPDSIRATNLHSLLVCSSAKHKME----------LLPS-LN 581
Query: 560 CYSLRVLLSHRLNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQ 619
L ++ L + + L+YL++S+ + K LP + KL NLE L ++
Sbjct: 582 LLRALDLEDSSISKLPDCLVTMFNLKYLNLSKTQVKELPKNFHKLVNLETLNTKHS-KIE 640
Query: 620 KLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQL-- 677
+LP G+ +LK+L+ L + R G ++ N Y++G + +L L
Sbjct: 641 ELPLGMWKLKKLR--------YLITFRRNEGHDSNWN----YVLGTRVVPKIWQLKDLQV 688
Query: 678 ----NLKGQLHIKNLERLKSVTDAKKANMSRKKLNQLWLSWERNEVSQLQENVEQILEAL 733
N + +L IKNL + +T + R+ L S N++ ++
Sbjct: 689 MDCFNAEDEL-IKNLGCMTQLTRISLVMVRREHGRDLCDS--LNKIKRI----------- 734
Query: 734 QPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLNLPELWKLPSLKYLK- 792
+++S+ S+++ + LE+ D + ++ +L+ L+ +
Sbjct: 735 ---------------------RFLSLTSIDEEEPLEIDDLIATASIEKLFLAGKLERVPS 773
Query: 793 -LSNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSREERVMFPR---------LKA 842
+ + ++ YL G L L ++ LP L+ LS M PR LK
Sbjct: 774 WFNTLQNLTYL---GLRGSQLQENAILSIQTLPRLVWLSFYNAYMGPRLRFAQGFQNLKI 830
Query: 843 LEITECPNLLGL----PCLPSLSDLYIQG-KYNQQLPSSIHKLGSLESLH 887
LEI + +L + + L LY++ + + +P I L +L+ LH
Sbjct: 831 LEIVQMKHLTEVVIEDGAMFELQKLYVRACRGLEYVPRGIENLINLQELH 880
>At1g59218 resistance protein RPP13, putative
Length = 1049
Score = 219 bits (558), Expect = 7e-57
Identities = 286/1146 (24%), Positives = 496/1146 (42%), Gaps = 159/1146 (13%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MA ++ + +L L+ +E LF G + + L L + + L+DA+ K+ + +
Sbjct: 1 MAGELISFGIQNLWNLLSQECELFQGVEDQVTELKRDLNLLSSFLKDADAKKHTSAV--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEY-KASKCGLSHKHIAF----RYKLAKKMK 115
VK+ + ++K+ Y +D ++ T LE K S S + +A R + A +
Sbjct: 58 -VKNCVEEIKEIIYDGEDTIE---TFVLEQNLGKTSGIKKSIRRLACIIPDRRRYALGIG 113
Query: 116 RIGVWLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKI-----V 170
+ + + + F + + + + P + + + + +++D D + V
Sbjct: 114 GLSNRISKVIRDMQSFGVQQAIVDGGYKQPQGDKQREMRPR---FSKDDDSDFVGLEANV 170
Query: 171 DFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRM 230
LVG ++ ++ V I G+GGLGKTTLA+ VFNH+ + + F+ WVCVS+DFT +
Sbjct: 171 KKLVGYLVDEANVQVVSITGMGGLGKTTLAKQVFNHEDVKHQFDGLSWVCVSQDFTRMNV 230
Query: 231 TKAIIEGATKKSCEDLDLELLQRKLQD----LLRRKRYLLVLDDVWNDKQENWQRLKSVL 286
+ I+ K E +E+ Q LQ LL + L+VLDD+W ++E+W+ +K +
Sbjct: 231 WQKILRDLKPKEEEKKIMEMTQDTLQGELIRLLETSKSLIVLDDIW--EKEDWELIKPIF 288
Query: 287 ACGGKGASILVTTRLPKVAKIMGT-IPHHELSRLSDEDCWELFKQRAFGPNEVQQ----K 341
KG +L+T+R VA T + + L+ ED W LF++ A + + +
Sbjct: 289 P-PTKGWKVLLTSRNESVAMRRNTSYINFKPECLTTEDSWTLFQRIALPMKDAAEFKIDE 347
Query: 342 ELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA--------- 392
E +GK +IK CGG PLA LG +L K +W + E+ +L G
Sbjct: 348 EKEELGKLMIKHCGGLPLAIRVLGGMLAEKYTSHDWRRLSENIGSHLVGGRTNFNDDNNN 407
Query: 393 ---YVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFIS----SNQM 445
YV L LS+ LP L+ CF + A FP D I+ + L W A G ++
Sbjct: 408 TCNYV---LSLSFEELPSYLKHCFLYLAHFPDDYEINVKNLSYYWAAEGIFQPRHYDGEI 464
Query: 446 LEADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHD--LAGSVTQDVCCITDDN 503
+ D+G+ EL R+ + +V + +HD++ + L + ++ IT
Sbjct: 465 IR--DVGDVYIEELVRRNMVISERDVKTSRFETCHLHDMMREVCLLKAKEENFLQITSSR 522
Query: 504 SM--RTMSEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAG----QLSPQV 557
+ ++S T L+Y + ++ K + N ++ G L
Sbjct: 523 TSTGNSLSIVTSRRLVYQYPITLDVEK-DINDPKLRSLVVVANTYMFWGGWSWMLLGSSF 581
Query: 558 LNCYSLRVLLSHRL----NNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLD 613
+ LRVL HR L+SSIG+L +LRYL++ ++P SL L L L L
Sbjct: 582 IRLELLRVLDIHRAKLKGGKLASSIGQLIHLRYLNLKHAEVTHIPYSLGNLKLLIYLNLV 641
Query: 614 GCVSLQKL-PGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLE 672
VS L P L +++L+ L +LP+ +G+ T L + + + F +
Sbjct: 642 ILVSGSTLVPNVLKEMQQLRYL---------ALPKDMGRKTKLELSNLVKLETLKNFSTK 692
Query: 673 ELGQLNLKGQLHIKNLE-RLKSVTDAKKANMS---RKKLNQLWLSWERNEVSQLQENVEQ 728
+L+G + ++ L L+ T + S K L L ++ +E+ + +
Sbjct: 693 NCSLEDLRGMVRLRTLTIELRKETSLETLAASIGGLKYLESLTITDLGSEMRTKEAGIVF 752
Query: 729 ILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLN-LPELWKLPS 787
L+ KLY + +FP + L +L L C+ + +P L KL
Sbjct: 753 DFVYLKTLTLKLYMPRLS--KEQHFP--------SHLTTLYLQHCRLEEDPMPILEKLHQ 802
Query: 788 LKYLKLSNMIHVIYLFHESYDGEGLMALKTLF--LEKLPNLIGLS-----REERVMFPRL 840
LK L+L +S+ G+ ++ F L+KL ++ GL + E P L
Sbjct: 803 LKELELR---------RKSFSGKEMVCSSGGFPQLQKL-SIKGLEEWEDWKVEESSMPVL 852
Query: 841 KALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEELIYFPDG 900
L+I +C L LP ++ LPS + + SL F EE
Sbjct: 853 HTLDIRDCRKLKQLP--------------DEHLPSHLTSI----SLFFCCLEE------- 887
Query: 901 ILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKE 960
P+ TL ++H+ LQ L+ + I +LH LK
Sbjct: 888 ------DPMPTL------------ERLVHLKELQLLFRSFSGRIMVCAGSGFPQLHKLKL 929
Query: 961 LDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLP 1020
++ G ++ + + L TL I C +++ L++L L++L E
Sbjct: 930 SELDGLEEWIVEDG--SMPQLHTLEIRRCPKLKKLPNGFPQ---LQNLELNELEEWEEWI 984
Query: 1021 ECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPKIV 1080
G++ LLH + I++CPKL LP ++ I L+ L++ + +KR K GED+ K+
Sbjct: 985 VEDGSMPLLHTLRIWNCPKLKQLPDGLRFIYSLKNLTVP--KRWKKRLSKG-GEDYYKVQ 1041
Query: 1081 HVQYIE 1086
H+ +E
Sbjct: 1042 HIPSVE 1047
>At3g46530 disease resistance protein RPP13-like protein
Length = 835
Score = 219 bits (557), Expect = 9e-57
Identities = 199/748 (26%), Positives = 359/748 (47%), Gaps = 86/748 (11%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M +A+ E V+G + + +E S+F+ ++ L + LT I L+D E ++ D E+
Sbjct: 1 MVDAITEFVVGKIGNYLIEEASMFMAVKEDLEELKTELTCIHGYLKDVEARERED-EVS- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIAFRYKLAKKMKRIGVW 120
K+W + D AY ++D++D T L++E ++ + GL K+ +KM +
Sbjct: 59 --KEWSKLVLDFAYDVEDVLD---TYHLKLEERSQRRGLRR----LTNKIGRKMDAYSI- 108
Query: 121 LDDIAAEKNKFHLTEIVRERSGV-----VPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVG 175
+DDI K + RE G+ T+S+ + L R+ D++++V L
Sbjct: 109 VDDIRILKRRILDITRKRETYGIGGLKEPQGGGNTSSLRVRQLRRARSVDQEEVVVGLED 168
Query: 176 DAS---------EQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFT 226
DA E+++ + I G+GGLGKT LA+ ++N + FE + W VS+++
Sbjct: 169 DAKILLEKLLDYEEKNRFIISIFGMGGLGKTALARKLYNSRDVKERFEYRAWTYVSQEYK 228
Query: 227 LKRMTKAIIEGATKKSCEDLDL------ELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQ 280
+ II S E+L+ E L+ L LL K+YL+V+DD+W ++E W
Sbjct: 229 TGDILMRIIRSLGMTSGEELEKIRKFAEEELEVYLYGLLEGKKYLVVVDDIW--EREAWD 286
Query: 281 RLKSVLACGGKGASILVTTRLPKVAK-IMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQ 339
LK L C +G+ +++TTR+ VA+ + G H+L L+ E+ WELF+QRAF + +
Sbjct: 287 SLKRALPCNHEGSRVIITTRIKAVAEGVDGRFYAHKLRFLTFEESWELFEQRAFRNIQRK 346
Query: 340 QKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA-YVMP-A 397
++L+ GKE+++KC G PL + L LL ++ EW V S L+ ++ +V P
Sbjct: 347 DEDLLKTGKEMVQKCRGLPLCIVVLAGLLS-RKTPSEWNDVCNSLWRRLKDDSIHVAPIV 405
Query: 398 LRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWN 457
LS+ L + + CF + ++FP+D I + LI L A GFI ++ + +D+
Sbjct: 406 FDLSFKELRHESKLCFLYLSIFPEDYEIDLEKLIHLLVAEGFIQGDEEMMMEDVARYYIE 465
Query: 458 ELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCIT--DDNSMRTMSEETRHL 515
EL RS E G++ ++HDL+ D+A ++++ + +D+ + S R
Sbjct: 466 ELIDRSLLEAVRRER-GKVMSCRIHDLLRDVAIKKSKELNFVNVYNDHVAQHSSTTCRRE 524
Query: 516 LIYNRNSFAEANSIQLHHVKSLKTYMEFNFDV---YEAGQLSPQVLNCYSLRVLLSHRLN 572
+++++ + + ++S + EF+ V +E +L +VL+ SL L ++N
Sbjct: 525 VVHHQFKRYSSEKRKNKRMRSFLYFGEFDHLVGLDFETLKLL-RVLDFGSL--WLPFKIN 581
Query: 573 NLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQ 632
G L +LRYL I + + +++L+ LQ
Sbjct: 582 ------GDLIHLRYLGIDGNSINDF----------------------DIAAIISKLRFLQ 613
Query: 633 NLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGE-ERGFLLEELGQLNLKGQLHIKNLERL 691
L + D + + KLTSL ++++G G L+ ++ L + + +L
Sbjct: 614 TLFVSD-NYFIEETIDLRKLTSL----RHVIGNFFGGLLIGDVANLQTLTSISFDSWNKL 668
Query: 692 K-----SVTDAKKANMSRKKLNQLWLSW 714
K ++ D + MSR K ++ +SW
Sbjct: 669 KPELLINLRDLGISEMSRSKERRVHVSW 696
>At5g48620 disease resistance protein
Length = 908
Score = 214 bits (545), Expect = 2e-55
Identities = 267/1008 (26%), Positives = 441/1008 (43%), Gaps = 142/1008 (14%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAE + L L +L+ +E G D++ + L L ++++ L+DA+ K+
Sbjct: 1 MAEGFVSFGLEKLWDLLSRESERLQGIDEQLDGLKRQLRSLQSLLKDADAKKHGSDR--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH--KHIAFRYKLAKKMKRIG 118
V+++L +KD + +DI++ L E K K + + + R+K+A ++ I
Sbjct: 58 -VRNFLEDVKDLVFDAEDIIESYVLNKLRGEGKGVKKHVRRLARFLTDRHKVASDIEGIT 116
Query: 119 VWLDDIAAEKNKFHLTEIV--------RERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIV 170
+ D+ E F + +I+ +ER V + RQT ++ + G + +++V
Sbjct: 117 KRISDVIGEMQSFGIQQIIDGVRSLSLQERQRVQREIRQTYPDSSESDLVGVEQSVEELV 176
Query: 171 DFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRM 230
LV E + V I G+GG+GKTTLA+ VF+HD + HF+ WVCVS+ FTLK +
Sbjct: 177 GHLV----ENDIYQVVSIAGMGGIGKTTLARQVFHHDLVRRHFDGFAWVCVSQQFTLKHV 232
Query: 231 TKAIIEGATKK--SCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLAC 288
+ I++ + +D LQ KL LL RYLLVLDDVW K+E+W R+K+V
Sbjct: 233 WQRILQELQPHDGNILQMDESALQPKLFQLLETGRYLLVLDDVW--KKEDWDRIKAVFP- 289
Query: 289 GGKGASILVTTRLPKVA-KIMGTIPHHELSRLSDEDCWELFKQRAF---GPNEVQ-QKEL 343
+G +L+T+R V T S L+ E+ W+L ++ F EV+ +E+
Sbjct: 290 RKRGWKMLLTSRNEGVGIHADPTCLTFRASILNPEESWKLCERIVFPRRDETEVRLDEEM 349
Query: 344 VIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA--------YVM 395
+GKE++ CGG PLA ALG LL K EW V ++ + G + V
Sbjct: 350 EAMGKEMVTHCGGLPLAVKALGGLLANKHTVPEWKRVSDNIGSQIVGGSCLDDNSLNSVN 409
Query: 396 PALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEV 455
L LSY LP L+ F + A FP+D I Q L + W A G + + D G
Sbjct: 410 RILSLSYEDLPTHLKHRFLYLAHFPEDSKIYTQDLFNYWAAEGIYDGSTI---QDSGEYY 466
Query: 456 WNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHL 515
EL R+ + +MHD++ ++ S + ++N ++ + + T
Sbjct: 467 LEELVRRNLVIADNRYLSLEFNFCQMHDMMREVCLSKAK------EENFLQIIKDPTSTS 520
Query: 516 LIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLNNLS 575
I N S + + +H K+ N +P+V + R + + +
Sbjct: 521 TI-NAQSPSRSRRFSIHSGKAFHILGHRN---------NPKVRSLIVSRFEEDFWIRS-A 569
Query: 576 SSIGRLKYLRYLDISEGRFK--NLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQN 633
S L LR LD+S +F+ LP+S+ L +L L L G V + LP + LK L
Sbjct: 570 SVFHNLTLLRVLDLSRVKFEGGKLPSSIGGLIHLRYLSLYGAV-VSHLPSTMRNLKLLLF 628
Query: 634 LSLR-DCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLK 692
L+LR D +P + ++ L LS +++ L ELG L NLE L
Sbjct: 629 LNLRVDNKEPIHVPNVLKEMLELRYLSLPQEMDDKTKL--ELGDL--------VNLEYL- 677
Query: 693 SVTDAKKANMSRKKLNQLWLSWERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAY 752
+ S + + V+ L KL + GV
Sbjct: 678 -----------------WYFSTQHSSVTDLLR------------MTKLRNLGVSLSERCN 708
Query: 753 FPQWISIPSLNDLKSLELVDCKSCLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGL 812
F S SL +L++LE++ + L PE+ + + L + IH+ L GL
Sbjct: 709 FETLSS--SLRELRNLEML---NVLFSPEIVMVDHMGEFVLDHFIHLKQL--------GL 755
Query: 813 MALKTLFLEKLPNLIGLSREERVMFPRLKALEITEC-------PNLLGLPCLPSLSDLYI 865
+ + K+P ++ P L + + C P L L L S++ Y
Sbjct: 756 ----AVRMSKIP-------DQHQFPPHLAHIHLVHCVMKEDPMPILEKLLHLKSVALSY- 803
Query: 866 QGKY-NQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLASP-LKTLGFHRHSKLKML 923
G + +++ S L +L S EL + I+ + P L+TL H KLK L
Sbjct: 804 -GAFIGRRVVCSKGGFPQLCALGISGESEL---EEWIVEEGSMPCLRTLTIHDCEKLKEL 859
Query: 924 PTEMIHIHALQQLYINDCRN--IEEL--PNEVMQRLHSLKELDIVGCD 967
P + +I +L++L I + + E+L E ++ + ++ + CD
Sbjct: 860 PDGLKYITSLKELKIREMKREWKEKLVPGGEDYYKVQHIPDVQFINCD 907
Score = 40.0 bits (92), Expect = 0.007
Identities = 74/323 (22%), Positives = 135/323 (40%), Gaps = 75/323 (23%)
Query: 777 LNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSREERVM 836
L+LP+ ++ L+L +++++ YL++ S + L L + KL NL G+S ER
Sbjct: 654 LSLPQ--EMDDKTKLELGDLVNLEYLWYFSTQHSSVTDL--LRMTKLRNL-GVSLSERCN 708
Query: 837 FPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEELI- 895
F + L SS+ +L +LE L+ + E++
Sbjct: 709 F---------------------------------ETLSSSLRELRNLEMLNVLFSPEIVM 735
Query: 896 --YFPDGILRNLASPLKTLGFH-RHSKLK---MLPTEMIHIHALQQLYINDCRNIEELPN 949
+ + +L + LK LG R SK+ P + HIH + + ++E P
Sbjct: 736 VDHMGEFVLDHFIH-LKQLGLAVRMSKIPDQHQFPPHLAHIHLVHCV-------MKEDPM 787
Query: 950 EVMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLT 1009
++++L LK + + Y + + CS+ GF + L +L
Sbjct: 788 PILEKLLHLKSVAL------------SYGAFIGRRVV--CSK-GGFPQ-------LCALG 825
Query: 1010 LSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQ 1069
+S LE G++ L + I+ C KL LP ++ I+ L+ L I + + K
Sbjct: 826 ISGESELEEWIVEEGSMPCLRTLTIHDCEKLKELPDGLKYITSLKELKIREMKREWKEKL 885
Query: 1070 KEIGEDWPKIVHVQYIEIENDNL 1092
GED+ K+ H+ ++ N +L
Sbjct: 886 VPGGEDYYKVQHIPDVQFINCDL 908
>At5g43470 disease resistance protein RPP8
Length = 908
Score = 214 bits (544), Expect = 3e-55
Identities = 232/901 (25%), Positives = 406/901 (44%), Gaps = 129/901 (14%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA + L L +L+ +E G D + + L L ++++ L+DA+ K+
Sbjct: 1 MAEAFVSFGLEKLWDLLSRESERLQGIDGQLDGLKRQLRSLQSLLKDADAKKHGSDR--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHI-------AFRYKLAKK 113
V+++L +KD + +DI++ L + K K KH+ R+K+A
Sbjct: 58 -VRNFLEDVKDLVFDAEDIIESYVLNKLSGKGKGVK-----KHVRRLACFLTDRHKVASD 111
Query: 114 MKRIGVWLDDIAAEKNKFHLTEIV--------RERSGVVPDWRQTTSIVTQPLVYGRNED 165
++ I + ++ E F + +I+ +ER V + RQT ++ + G +
Sbjct: 112 IEGITKRISEVIGEMQSFGIQQIIDGGRSLSLQERQRVQREIRQTYPDSSESDLVGVEQS 171
Query: 166 KDKIVDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDF 225
++V LV E + V I G+GG+GKTTLA+ VF+HD + HF+ WVCVS+ F
Sbjct: 172 VKELVGHLV----ENDVHQVVSIAGMGGIGKTTLARQVFHHDLVRRHFDGFAWVCVSQQF 227
Query: 226 TLKRMTKAIIEGATKKSCEDLDLE--LLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLK 283
T K + + I++ + L ++ LQRKL LL RYL+VLDDVW K+E+W +K
Sbjct: 228 TQKHVWQRILQELQPHDGDILQMDEYALQRKLFQLLEAGRYLVVLDDVW--KKEDWDVIK 285
Query: 284 SVLACGGKGASILVTTRLPKVA-KIMGTIPHHELSRLSDEDCWELFKQRAF---GPNEVQ 339
+V +G +L+T+R V T S L+ E+ W+L ++ F EV+
Sbjct: 286 AVFP-RKRGWKMLLTSRNEGVGIHADPTCLTFRASILNPEESWKLCERIVFPRRDETEVR 344
Query: 340 -QKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEAY----- 393
+E+ +GKE++ CGG PLA ALG LL K EW V ++ + G ++
Sbjct: 345 LDEEMEAMGKEMVTHCGGLPLAVKALGGLLANKHTVPEWKRVFDNIGSQIVGGSWLDDNS 404
Query: 394 ---VMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADD 450
V L LSY LP L+ CF A FP+D IS L W A G + + +D
Sbjct: 405 LNSVYRILSLSYEDLPTHLKHCFLNLAHFPEDSEISTYSLFYYWAAEGIYDGSTI---ED 461
Query: 451 IGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQD---VCCITDDNSMRT 507
G EL R+ +N Q +MHD++ ++ S ++ + I D T
Sbjct: 462 SGEYYLEELVRRNLVIADDNYLSWQSKYCQMHDMMREVCLSKAKEENFLQIIIDPTCTST 521
Query: 508 MSEE----TRHLLIYNRNSF----------------------AEANSIQLHHVKSLKTYM 541
++ + +R L I++ +F S + H +L +
Sbjct: 522 INAQSPSRSRRLSIHSGKAFHILGHKNKTKVRSLIVPRFEEDYWIRSASVFHNLTLLRVL 581
Query: 542 EFNFDVYEAGQLSPQVLNCYSLRVLLSH--RLNNLSSSIGRLKYLRYLDISEGRFK--NL 597
+ ++ +E G+L + LR L + ++++L S++ LK L YL++ + ++
Sbjct: 582 DLSWVKFEGGKLPCSIGGLIHLRYLSLYEAKVSHLPSTMRNLKLLLYLNLRVDTEEPIHV 641
Query: 598 PNSLCKLCNLEV------------LKLDGCVSLQKLPG---------GLTRLKRLQNLSL 636
PN L ++ L L+L V+L+ L G L R+ +L+ L++
Sbjct: 642 PNVLKEMIQLRYLSLPLKMDDKTKLELGDLVNLEYLYGFSTQHSSVTDLLRMTKLRYLAV 701
Query: 637 R-----DCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKNL--- 688
+ ++L+S R++ L +LN L ++++ +G+ L +H+K L
Sbjct: 702 SLSERCNFETLSSSLRELRNLETLNFLFSL-----ETYMVDYMGEFVLDHFIHLKQLGLA 756
Query: 689 ERLKSVTDAKKANMSRKKLNQLWLSWERNEVSQLQE--NVEQILEALQPYAQKLYSFGVG 746
R+ + D + L ++ E + + L++ +++ + A + + G
Sbjct: 757 VRMSKIPDQHQFPPHLVHLFLIYCGMEEDPMPILEKLLHLKSVRLARKAFLGSRMVCSKG 816
Query: 747 GY---------TGAYFPQWI-SIPSLNDLKSLELVDCKSCLNLPE-LWKLPSLKYLKLSN 795
G+ + +WI S+ L++L + DCK LP+ L + SLK LK+
Sbjct: 817 GFPQLCVIEISKESELEEWIVEEGSMPCLRTLTIDDCKKLKELPDGLKYITSLKELKIEG 876
Query: 796 M 796
M
Sbjct: 877 M 877
Score = 39.3 bits (90), Expect = 0.013
Identities = 57/231 (24%), Positives = 97/231 (41%), Gaps = 47/231 (20%)
Query: 871 QQLPSSIHKLGSLESLHFSDNEELI---YFPDGILRNLASPLKTLGFH-RHSKLK---ML 923
+ L SS+ +L +LE+L+F + E Y + +L + LK LG R SK+
Sbjct: 710 ETLSSSLRELRNLETLNFLFSLETYMVDYMGEFVLDHFIH-LKQLGLAVRMSKIPDQHQF 768
Query: 924 PTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDI-----VGCDKLKLSSDFQYL 978
P ++H L++ C +EE P ++++L LK + + +G + F L
Sbjct: 769 PPHLVH------LFLIYC-GMEEDPMPILEKLLHLKSVRLARKAFLGSRMVCSKGGFPQL 821
Query: 979 TCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCP 1038
+E I SE+E + M L++LT+ D C
Sbjct: 822 CVIE---ISKESELEEWIVEEGSMPCLRTLTIDD------------------------CK 854
Query: 1039 KLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYIEIEN 1089
KL LP ++ I+ L+ L I + K GED+ K+ H+ ++ N
Sbjct: 855 KLKELPDGLKYITSLKELKIEGMKREWKEKLVPGGEDYYKVQHIPDVQFIN 905
>At1g53350 hypothetical protein
Length = 906
Score = 211 bits (537), Expect = 2e-54
Identities = 204/702 (29%), Positives = 332/702 (47%), Gaps = 65/702 (9%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEAV+ + L EL+ +E + G D++ + L L +++ L+DA+ K+ +E R
Sbjct: 1 MAEAVVSFGVEKLWELLSRESARLNGIDEQVDGLKRQLGRLQSLLKDADAKK---NETER 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHK--HIAFRYKLAKKMKRIG 118
V+++L +KD Y DDI++ L + K K + + R K A ++ I
Sbjct: 58 -VRNFLEDVKDIVYDADDIIESFLLNELRGKEKGIKKQVRTLACFLVDRRKFASDIEGIT 116
Query: 119 VWLDDIAAEKNKFHLTEI---------VRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKI 169
+ ++ + I ++ER + RQT S ++ + G ++ +++
Sbjct: 117 KRISEVIVGMQSLGIQHIADGGGRSLSLQERQREI---RQTFSRNSESDLVGLDQSVEEL 173
Query: 170 VDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKR 229
VD LV E + + V + G+GG+GKTTLA+ VF+HD + HF+ WVCVS+ FT K
Sbjct: 174 VDHLV----ENDSVQVVSVSGMGGIGKTTLARQVFHHDIVRRHFDGFSWVCVSQQFTRKD 229
Query: 230 MTKAIIEGAT--KKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLA 287
+ + I++ + +D LQ +L +LL RYLLVLDDVW K+E+W R+K+V
Sbjct: 230 VWQRILQDLRPYDEGIIQMDEYTLQGELFELLESGRYLLVLDDVW--KEEDWDRIKAVFP 287
Query: 288 CGGKGASILVTTRLPKVAKIMGTIPHHELSR-LSDEDCWELFKQRAFG-PNEVQQKELVI 345
+G +L+T+R + R L+ E W+LF++ ++ + K
Sbjct: 288 -HKRGWKMLLTSRNEGLGLHADPTCFAFRPRILTPEQSWKLFERIVSSRRDKTEFKVDEA 346
Query: 346 VGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA--------YVMPA 397
+GKE++ CGG PLA LG LL K EW V + + ++ G++ V
Sbjct: 347 MGKEMVTYCGGLPLAVKVLGGLLAKKHTVLEWKRVHSNIVTHIVGKSGLSDDNSNSVYRV 406
Query: 398 LRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISS-NQMLEADDIGNEVW 456
L LSY LP++L+ CF + A FP+D I ++L + W A G I+ + D G
Sbjct: 407 LSLSYEDLPMQLKHCFFYLAHFPEDYKIDVKILFNYWVAEGIITPFHDGSTIQDTGESYL 466
Query: 457 NELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLL 516
EL R+ E+ +I +MHD++ ++ S + ++N +R + T
Sbjct: 467 EELVRRNMVVVEESYLTSRIEYCQMHDMMREVCLSKAK------EENFIRVVKVPTTTST 520
Query: 517 IYNRNSFAEANSIQLH---------HVKSLKTYMEFNFDVYEAGQLSPQVLNCYS-LRVL 566
N S + + LH H + K F V E P+ C LRVL
Sbjct: 521 TINAQSPCRSRRLVLHSGNALHMLGHKDNKKARSVLIFGV-EEKFWKPRGFQCLPLLRVL 579
Query: 567 ----LSHRLNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVS--LQK 620
+ L SSIG L +LR+L + E +LP+SL L L L L G L
Sbjct: 580 DLSYVQFEGGKLPSSIGDLIHLRFLSLYEAGVSHLPSSLGNLKLLLCLNL-GVADRLLVH 638
Query: 621 LPGGLTRLKRLQNLSL-RDCDSLTSLPRQIGKLTSLNTLSKY 661
+P L ++ L+ L L R + T L ++G L +L +L+ +
Sbjct: 639 VPNVLKEMQELRYLRLPRSMPAKTKL--ELGDLVNLESLTNF 678
Score = 30.0 bits (66), Expect = 7.7
Identities = 39/168 (23%), Positives = 74/168 (43%), Gaps = 17/168 (10%)
Query: 876 SIHKLGSLESLHFSDNEELIYFPDG----ILRNLASPLKTLGFHRHSKLKMLPTEMIHIH 931
S+ +L +LE+L F D +++ G +L + TL H L P +
Sbjct: 713 SLRELRNLETLSFHDFQKVSVANHGGELLVLDFIHLKDLTLSMH----LPRFPDQYRFPP 768
Query: 932 ALQQLYINDCRNIEELPNEVMQRLHSLKELDI-----VGCDKLKLSSDFQYLTCLETLAI 986
L +++ CR +EE P ++++L LK + + +G + F L L+ +
Sbjct: 769 HLAHIWLIGCR-MEEDPMPILEKLLHLKSVYLSSGAFLGRRMVCSKGGFPQLLALK---M 824
Query: 987 GSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINI 1034
E+ + M L++LT+ + L+ LP+ + +T L E+ I
Sbjct: 825 SYKKELVEWRVEEGSMPCLRTLTIDNCKKLKQLPDGLKYVTCLKELKI 872
>At5g35450 disease resistance protein
Length = 901
Score = 207 bits (528), Expect = 2e-53
Identities = 243/937 (25%), Positives = 413/937 (43%), Gaps = 125/937 (13%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAE V+ + L L+ +E G D++ + L L +++ L+DA+ K+
Sbjct: 1 MAEGVVSFGVQKLWALLNRESERLNGIDEQVDGLKRQLRGLQSLLKDADAKKHGSDR--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASK-------CGLSHKHIAFRYKLAKK 113
V+++L +KD + +DI++ L E K K C L+ +H K+A
Sbjct: 58 -VRNFLEDVKDLVFDAEDIIESYVLNKLRGEGKGVKNHVRRLACFLTDRH-----KVASD 111
Query: 114 MKRIGVWLDDIAAEKNKFHLTEIVRERS------GVVPDWRQTTSIVTQPLVYGRNEDKD 167
++ I + + E + + + + + + RQT ++ + G +
Sbjct: 112 IEGITKRISKVIGEMQSLGIQQQIIDGGRSLSLQDIQREIRQTFPNSSESDLVGVEQS-- 169
Query: 168 KIVDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTL 227
V+ LVG E +++ V I G+GG+GKTTLA+ +F+HD + HF+ WVCVS+ FT
Sbjct: 170 --VEELVGPMVEIDNIQVVSISGMGGIGKTTLARQIFHHDLVRRHFDGFAWVCVSQQFTQ 227
Query: 228 KRMTKAIIEGATKKSCEDLDLE--LLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSV 285
K + + I++ E L ++ +Q KL LL RYL+VLDDVW K+E+W R+K V
Sbjct: 228 KHVWQRILQELRPHDGEILQMDEYTIQGKLFQLLETGRYLVVLDDVW--KEEDWDRIKEV 285
Query: 286 LACGGKGASILVTTRLPKVA-KIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELV 344
+G +L+T+R V T L+ ++ W+LF++ NE + +E+
Sbjct: 286 FP-RKRGWKMLLTSRNEGVGLHADPTCLSFRARILNPKESWKLFERIVPRRNETEYEEME 344
Query: 345 IVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA--------YVMP 396
+GKE++ CGG PLA LG LL K EW V E+ + G++ V
Sbjct: 345 AIGKEMVTYCGGLPLAVKVLGGLLANKHTASEWKRVSENIGAQIVGKSCLDDNSLNSVYR 404
Query: 397 ALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVW 456
L LSY LP L+ CF + A FP+D I + L W A G +L D G +
Sbjct: 405 ILSLSYEDLPTDLKHCFLYLAHFPEDYKIKTRTLYSYWAAEGIYDGLTIL---DSGEDYL 461
Query: 457 NELYWRSFFENTENVGFGQITIFKMHDLVHDLAGS------VTQDVCCITDDNSMRTMS- 509
EL R+ ++ ++ + +MHD++ ++ S Q + T +++ S
Sbjct: 462 EELVRRNLVIAEKSNLSWRLKLCQMHDMMREVCISKAKVENFLQIIKVPTSTSTIIAQSP 521
Query: 510 EETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVL--- 566
+R L +++ +F L H K +++ + Q + + + LRVL
Sbjct: 522 SRSRRLTVHSGKAFH-----ILGHKKKVRSLLVLGLKEDLWIQSASRFQSLPLLRVLDLS 576
Query: 567 -LSHRLNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQ-KLPGG 624
+ L SSIG L +LR+L + + +LP+++ L + L L + + +P
Sbjct: 577 SVKFEGGKLPSSIGGLIHLRFLSLHQAVVSHLPSTIRNLKLMLYLNLHVAIGVPVHVPNV 636
Query: 625 LTRLKRLQNLSL-RDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQL 683
L + L+ LSL D T L ++G L +L L +
Sbjct: 637 LKEMLELRYLSLPLDMHDKTKL--ELGDLVNLEYLWCFST-------------------- 674
Query: 684 HIKNLERLKSVTDAKKANMSRKKLNQLWLSW-ERNEVSQLQENVEQI--LEALQ-PYAQK 739
+ SVTD + KL +S+ ER L ++ Q LE L Y++K
Sbjct: 675 ------QHSSVTDL----LRMTKLRFFGVSFSERCTFENLSSSLRQFRKLETLSFIYSRK 724
Query: 740 LYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLNLPELWKLPSLKYLKLSNMIHV 799
Y + Y G + +I + L+ L +P+ +LP ++ H+
Sbjct: 725 TY---MVDYVGEFVLDFIHLKKLSLGVHLS--------KIPDQHQLP-------PHIAHI 766
Query: 800 IYLF-HESYDG----EGLMALKTLFLEKLPNLIGLSREERVMFPRLKALEITECPNL--- 851
LF H D E L+ LK++ L + + + FP+L+AL+I+E L
Sbjct: 767 YLLFCHMEEDPMPILEKLLHLKSVELRRKAFIGRRMVCSKGGFPQLRALQISEQSELEEW 826
Query: 852 -LGLPCLPSLSDLYIQG-KYNQQLPSSIHKLGSLESL 886
+ +P L DL I + ++LP + + SL+ L
Sbjct: 827 IVEEGSMPCLRDLIIHSCEKLEELPDGLKYVTSLKEL 863
Score = 43.9 bits (102), Expect = 5e-04
Identities = 56/230 (24%), Positives = 102/230 (44%), Gaps = 48/230 (20%)
Query: 871 QQLPSSIHKLGSLESLHFSDNEELI---YFPDGILRNLASPLKTLGFHRHSKLK---MLP 924
+ L SS+ + LE+L F + + Y + +L + +LG H SK+ LP
Sbjct: 702 ENLSSSLRQFRKLETLSFIYSRKTYMVDYVGEFVLDFIHLKKLSLGVHL-SKIPDQHQLP 760
Query: 925 TEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDI-----VGCDKLKLSSDFQYLT 979
+ HI Y+ C ++EE P ++++L LK +++ +G + F L
Sbjct: 761 PHIAHI------YLLFC-HMEEDPMPILEKLLHLKSVELRRKAFIGRRMVCSKGGFPQLR 813
Query: 980 CLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPK 1039
L+ I SE+E E++ E G++ L ++ I+SC K
Sbjct: 814 ALQ---ISEQSELE-----------------------EWIVE-EGSMPCLRDLIIHSCEK 846
Query: 1040 LACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYIEIEN 1089
L LP ++ ++ L+ L I + K +K +GED+ K+ H+ ++ N
Sbjct: 847 LEELPDGLKYVTSLKELKIEGMKREWK--EKLVGEDYYKVQHIPDVQFFN 894
>At1g59780 hypothetical protein
Length = 906
Score = 204 bits (518), Expect = 3e-52
Identities = 240/924 (25%), Positives = 414/924 (43%), Gaps = 129/924 (13%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M ++++ + L +L+ +E F G +++ L L + A L DA+ K+ + +
Sbjct: 6 MVDSIVSFGVEKLWKLLSQEYERFQGVEEQITELRDDLKMLMAFLSDADAKKQTRAL--- 62
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATE-ALEMEYKASKCGLSHKHIAFRY--------KLA 111
++ L ++K+ Y +DI++ + ++ M A G + IA + K+
Sbjct: 63 -ARNCLEEIKEITYDAEDIIEIFLLKGSVNMRSLACFPG-GRREIALQITSISKRISKVI 120
Query: 112 KKMKRIGVWLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVD 171
+ M+ +G+ D + + ++ R+R + R T S ++ + G ++ +K+V+
Sbjct: 121 QVMQNLGIKSDIMDGVDSH---AQLERKR-----ELRHTFSSESESNLVGLEKNVEKLVE 172
Query: 172 FLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMT 231
LVG+ S I GLGGLGKTTLA+ +F+HDK+ +HF+ WVCVS++FT K +
Sbjct: 173 ELVGNDSSHG----VSITGLGGLGKTTLARQIFDHDKVKSHFDGLAWVCVSQEFTRKDVW 228
Query: 232 KAIIEGATKK-SCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGG 290
K I+ + K DL + +Q+KL LL K+ L+V DD+W K+E+W R+ +
Sbjct: 229 KTILGNLSPKYKDSDLPEDDIQKKLFQLLETKKALIVFDDLW--KREDWYRIAPMFPERK 286
Query: 291 KGASILVTTRLPKVAKIMGTIPH---HELSRLSDEDCWELFKQRAFGPNE-----VQQKE 342
G +L+T+R + PH + L+ ++CW+L ++ AF + + KE
Sbjct: 287 AGWKVLLTSRNDAIH------PHCVTFKPELLTHDECWKLLQRIAFSKQKTITGYIIDKE 340
Query: 343 LVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKL---------WNLQGEAY 393
+V + KE+ K C PLA LG LL K ++W + E+ + N +
Sbjct: 341 MVKMAKEMTKHCKRLPLAVKLLGGLLDAKHTLRQWKLISENIISHIVVGGTSSNENDSSS 400
Query: 394 VMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANG--FISSNQMLEADDI 451
V L LS+ LP L+ C + A +P+D I + L +W A G + + + D+
Sbjct: 401 VNHVLSLSFEGLPGYLKHCLLYLASYPEDHEIEIERLSYVWAAEGITYPGNYEGATIRDV 460
Query: 452 GNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLA------GSVTQDVCCITDDNSM 505
+ EL R+ + + + ++HDL+ ++ + Q V T +S+
Sbjct: 461 ADLYIEELVKRNMVISERDALTSRFEKCQLHDLMREICLLKAKEENFLQIVTDPTSSSSV 520
Query: 506 RTM-SEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLR 564
++ S +R L++YN + F+ N ++ ++SL L
Sbjct: 521 HSLASSRSRRLVVYNTSIFSGENDMKNSKLRSL-------------------------LF 555
Query: 565 VLLSHRLNNLSSSIGRLKYLRYLDISEGRFK--NLPNSLCKLCNLEVLKLDGCVSLQKLP 622
+ + + ++ S+ L LR LD+ +FK LP+S+ KL +L+ L L S+ LP
Sbjct: 556 IPVGYSRFSMGSNFIELPLLRVLDLDGAKFKGGKLPSSIGKLIHLKYLSLYQ-ASVTYLP 614
Query: 623 GGLTRLKRLQNLSLR-DCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKG 681
L LK L L+LR + L ++P ++ L LS + E ELG L LK
Sbjct: 615 SSLRNLKSLLYLNLRINSGQLINVPNVFKEMLELRYLS--LPWERSSLTKLELGNL-LKL 671
Query: 682 QLHIKNLERLKSVTDAKKANMSRKKLNQLWLSWERNEVSQLQENVEQILEALQPYAQKLY 741
+ I + SVTD + M++ + Q+ +S E + L + + + + L
Sbjct: 672 ETLINFSTKDSSVTDLHR--MTKLRTLQILISGEGLHMETLSSALSML-----GHLEDLT 724
Query: 742 SFGVGGYTGAYFPQWISIPSLND-------LKSLELVDCKSCLN-LPELWKLPSLKYLKL 793
P+ I P L D L ++ LV C + +P L KL LK
Sbjct: 725 VTPSENSVQFKHPKLIYRPMLPDVQHFPSHLTTISLVYCFLEEDPMPTLEKLLQLK---- 780
Query: 794 SNMIHVIYLFHESY-------DGEGLMALKTLFLEKLPNLIGLSREERVMFPRLKALEIT 846
V+ L++ +Y G G L L + L L EE M P L L I
Sbjct: 781 -----VVSLWYNAYVGRRMVCTGGGFPPLHRLEIWGLDALEEWIVEEGSM-PLLHTLHIV 834
Query: 847 ECPNLL----GLPCLPSLSDLYIQ 866
+C L GL + SL +L I+
Sbjct: 835 DCKKLKEIPDGLRFISSLKELAIR 858
Score = 47.8 bits (112), Expect = 4e-05
Identities = 85/353 (24%), Positives = 145/353 (40%), Gaps = 88/353 (24%)
Query: 761 SLNDLKSLELVDCK----SCLNLPELWK-LPSLKYLKLSNMIHVIYLFHESYDGEGLMAL 815
SL +LKSL ++ + +N+P ++K + L+YL L ++ L L
Sbjct: 616 SLRNLKSLLYLNLRINSGQLINVPNVFKEMLELRYLSLP------------WERSSLTKL 663
Query: 816 KTLFLEKLPNLIGLSREERVMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPS 875
+ L KL LI S ++ +T+ + L L L + +G + + L S
Sbjct: 664 ELGNLLKLETLINFSTKDS---------SVTDLHRMTKLRTLQIL--ISGEGLHMETLSS 712
Query: 876 SIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKL---KMLPTEMIHIHA 932
++ LG LE L + +E + F +H KL MLP
Sbjct: 713 ALSMLGHLEDLTVTPSENSVQF------------------KHPKLIYRPMLPDVQHFPSH 754
Query: 933 LQQLYINDCRNIEELPNEVMQRLHSLKELDI-----VGCDKLKLSSDFQYLTCLETLAIG 987
L + + C +EE P +++L LK + + VG + F L LE +
Sbjct: 755 LTTISLVYCF-LEEDPMPTLEKLLQLKVVSLWYNAYVGRRMVCTGGGFPPLHRLEIWGL- 812
Query: 988 SCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSI 1047
+AL+ E++ E G++ LLH ++I C KL +P +
Sbjct: 813 ---------DALE----------------EWIVE-EGSMPLLHTLHIVDCKKLKEIPDGL 846
Query: 1048 QQISGLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYI------EIENDNLIH 1094
+ IS L+ L+I K+ ++ + GED+ K+ HV I E EN+ +I+
Sbjct: 847 RFISSLKELAIRTNEKVFQKKVSKGGEDYYKMQHVPLIRYNWPQEPENNEVIY 899
>At1g10920 disease resistance protein RPM1 isolog
Length = 821
Score = 199 bits (505), Expect = 1e-50
Identities = 206/735 (28%), Positives = 337/735 (45%), Gaps = 83/735 (11%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAE V+ + L EL+ +E + G ++ + L L +++ L+DA+ K+
Sbjct: 1 MAEGVVLFGVHKLWELLNRESARLNGIGEQVDGLKRQLGRLQSLLKDADAKKHESER--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIAFRYKLAKKMKR-IGV 119
V+++L ++D Y +DI++ L E++ + G+ K A+++ + +
Sbjct: 58 -VRNFLEDVRDIVYDAEDIIESF----LLNEFRTKEKGIK--------KHARRLACFLSL 104
Query: 120 WLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
+ +I + L E RE+ + RQT + ++ + G + V+ L G E
Sbjct: 105 GIQEIIDGASSMSLQERQREQKEI----RQTFANSSESDLVGVEQS----VEALAGHLVE 156
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
+++ V I G+GG+GKTTLA+ VF+HD + HF+ WV VS+ FT K + + I +
Sbjct: 157 NDNIQVVSISGMGGIGKTTLARQVFHHDMVQRHFDGFAWVFVSQQFTQKHVWQRIWQELQ 216
Query: 240 KKS--CEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILV 297
++ +D +LQ KL LL RYL+VLDDVW K+E+W R+K+V +G +L+
Sbjct: 217 PQNGDISHMDEHILQGKLFKLLETGRYLVVLDDVW--KEEDWDRIKAVFP-RKRGWKMLL 273
Query: 298 TTRLPKVA-----KIMGTIPHHELSRLSDEDCWELFKQRAFGPNEV--QQKELVIVGKEI 350
T+R V K G + L+ E+ W+L ++ F + ++ +GKE+
Sbjct: 274 TSRNEGVGIHADPKSFG----FKTRILTPEESWKLCEKIVFHRRDETGTLSDMEAMGKEM 329
Query: 351 IKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEA-------YVMPALRLSYL 403
+ CGG PLA LG LL K EW V ++ +L G + + L LSY
Sbjct: 330 VTCCGGLPLAVKVLGGLLATKHTVPEWKRVYDNIGPHLAGRSSLDDNLNSIYRVLSLSYE 389
Query: 404 HLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFI-SSNQMLEADDIGNEVWNELYWR 462
+LP+ L+ CF + A FP+ I + L + A G I SS+ D G + EL R
Sbjct: 390 NLPMCLKHCFLYLAHFPEYYEIHVKRLFNYLAAEGIITSSDDGTTIQDKGEDYLEELARR 449
Query: 463 SFFENTENVGFGQITIFKMHDLVHDLAGSVTQD--------VCCITDDNSMRTMSEETRH 514
+ +N F + +MHD++ ++ S ++ V T + R++S ++R
Sbjct: 450 NMITIDKNYMFLRKKHCQMHDMMREVCLSKAKEENFLEIFKVSTATSAINARSLS-KSRR 508
Query: 515 LLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRL--- 571
L ++ N+ V+SL Y F + +P + LRVL R+
Sbjct: 509 LSVHGGNALPSLGQTINKKVRSL-LYFAFEDEFCILESTTPCFRSLPLLRVLDLSRVKFE 567
Query: 572 -NNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKL--DGCVSLQKLPGGLTRL 628
L SSIG L +LR+L + +LP+SL L L L L +G V + + + L
Sbjct: 568 GGKLPSSIGDLIHLRFLSLHRAWISHLPSSLRNLKLLLYLNLGFNGMVHVPNVLKEMQEL 627
Query: 629 KRLQ---------NLSLRDCDSLTSLPRQIGK---------LTSLNTLSKYIVGEERGFL 670
+ LQ L L D +L SL K +T L LS +I L
Sbjct: 628 RYLQLPMSMHDKTKLELSDLVNLESLMNFSTKYASVMDLLHMTKLRELSLFITDGSSDTL 687
Query: 671 LEELGQLNLKGQLHI 685
LGQL LH+
Sbjct: 688 SSSLGQLRSLEVLHL 702
Score = 35.8 bits (81), Expect = 0.14
Identities = 52/184 (28%), Positives = 81/184 (43%), Gaps = 12/184 (6%)
Query: 782 LWKLPSLKYLKL--SNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSREERVMFPR 839
L L L YL L + M+HV + E + L ++ + L L E +M
Sbjct: 598 LRNLKLLLYLNLGFNGMVHVPNVLKEMQELRYLQLPMSMHDKTKLELSDLVNLESLMNFS 657
Query: 840 LKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEE--LIYF 897
K + + LL + L LS L+I + L SS+ +L SLE LH D +E + Y
Sbjct: 658 TKYASVMD---LLHMTKLRELS-LFITDGSSDTLSSSLGQLRSLEVLHLYDRQEPRVAYH 713
Query: 898 PDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHS 957
I+ N LK L H + P + + L +Y+ C ++EE P +++RL
Sbjct: 714 GGEIVLNCIH-LKELELAIH--MPRFPDQYLFHPHLSHIYL-WCCSMEEDPIPILERLLH 769
Query: 958 LKEL 961
LK +
Sbjct: 770 LKSV 773
>At1g50180 disease resistance protein, putative
Length = 857
Score = 198 bits (504), Expect = 1e-50
Identities = 199/714 (27%), Positives = 317/714 (43%), Gaps = 87/714 (12%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA++ + + L +L+ +E G + +L L + L+DA+EKQ
Sbjct: 1 MAEAIVSVTVQKLGQLLLEEPLFLFGIGDQVKQLQDELKRLNCFLKDADEKQHESER--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSH-----KHIAFRYKLAKKMK 115
V++W+ +++A+Y +DI++ +A + K K L + + +++
Sbjct: 58 -VRNWVAGIREASYDAEDILEAFFLKAESRKQKGMKRVLRRLACILNEAVSLHSVGSEIR 116
Query: 116 RIGVWLDDIAAEKNKFHLTEIVRER----SGVVPDWRQTTSIVTQPLVYGRNEDKDKIVD 171
I L IAA F + E + S + + RQ+ V + + G + +K+V+
Sbjct: 117 EITSRLSKIAASMLDFGIKESMGREGLSLSDSLREQRQSFPYVVEHNLVGLEQSLEKLVN 176
Query: 172 FLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMT 231
LV S E L V I G+GGLGKTTLA+ +F+H K+ HF+ WV VS+D + +
Sbjct: 177 DLV---SGGEKLRVTSICGMGGLGKTTLAKQIFHHHKVRRHFDRFAWVYVSQDCRRRHVW 233
Query: 232 KAIIEGATKKSCEDLDLEL----LQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLA 287
+ I + K L L L +L L+R + L+VLDD+W ++ W LK V
Sbjct: 234 QDIFLNLSYKDENQRILSLRDEQLGEELHRFLKRNKCLIVLDDIWG--KDAWDCLKHVFP 291
Query: 288 CGGKGASILVTTRLPKVAKI---MGTIPHHELSRLSDEDCWELFKQRAFGPNE----VQQ 340
G+ I++TTR +VA G + HE L+ E+ WEL ++ + E +
Sbjct: 292 -HETGSEIILTTRNKEVALYADPRGVL--HEPQLLTCEESWELLEKISLSGRENIEPMLV 348
Query: 341 KELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEW--------LYVKESKLWNLQGEA 392
K++ +GK+I+ +CGG PLA LG LL K EW YV N
Sbjct: 349 KKMEEIGKQIVVRCGGLPLAITVLGGLLATKSTWNEWQRVCENIKSYVSNGGSSNGSKNM 408
Query: 393 YVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEA---- 448
V L LSY +LP ++QCF + A +P+D + L+ A G + + EA
Sbjct: 409 LVADVLCLSYEYLPPHVKQCFLYFAHYPEDYEVHVGTLVSYCIAEGMVMPVKHTEAGTTV 468
Query: 449 DDIGNEVWNELYWRSF-FENTENVGFGQITIFKMHDLVHDLA------GSVTQDVCCITD 501
+D+G + EL RS ++ ++ +MHDL+ ++ S Q +
Sbjct: 469 EDVGQDYLEELVKRSMVMVGRRDIVTSEVMTCRMHDLMREVCLQKAKQESFVQVIDSRDQ 528
Query: 502 DNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLK----------TYMEFNFDVYEAG 551
D + +S T R S + HH+KSL ++ E G
Sbjct: 529 DEAEAFISLSTN---TSRRISVQLHGGAEEHHIKSLSQVSFRKMKLLRVLDLEGAQIEGG 585
Query: 552 QLSPQVLNCYSLRVLLSHRLNN---LSSSIGRLKYLRYLDISEGRFKNLPNSLCKL---- 604
+L V + LR LS RL N L+SSIG LK + LD+ +PN L
Sbjct: 586 KLPDDVGDLIHLR-NLSVRLTNVKELTSSIGNLKLMITLDLFVKGQLYIPNQLWDFPVGK 644
Query: 605 CNLEVLKLDGCVSLQKLPGGLTRLKRLQ-NLSLRDCD--SLTSLPRQIGKLTSL 655
CN L +T L+RL NLS ++ D ++SL + + +L L
Sbjct: 645 CNPRDLL------------AMTSLRRLSINLSSQNTDFVVVSSLSKVLKRLRGL 686
Score = 35.8 bits (81), Expect = 0.14
Identities = 34/136 (25%), Positives = 55/136 (40%), Gaps = 13/136 (9%)
Query: 970 KLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLTL-------------SDLPNL 1016
KL + + + L L + C V+ L+ + LK L L +L NL
Sbjct: 722 KLPGEQSFSSDLGALRLWQCGLVDDPFMVLEKLPNLKILQLFEGSFVGSKLCCSKNLENL 781
Query: 1017 EYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDW 1076
E G + L + + C KL +P + + L+ + I + +K K GED+
Sbjct: 782 EEWTVEDGAMMRLVTVELKCCNKLKSVPEGTRFLKNLQEVEIGNRTKAFKDKLISGGEDF 841
Query: 1077 PKIVHVQYIEIENDNL 1092
K+ HV + EN L
Sbjct: 842 YKVQHVPCVVFENCEL 857
>At3g46710 disease resistance protein RPP8-like protein
Length = 847
Score = 194 bits (492), Expect = 3e-49
Identities = 184/715 (25%), Positives = 335/715 (46%), Gaps = 55/715 (7%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M +A+ E V+G + + +E + +G + L + LT I+ L++ E D E+
Sbjct: 1 MVDAITEFVVGKIDNYLIEEAPMLIGVKDDLEELKTELTCIQVYLKNVEVCDKED-EVS- 58
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGL-------SHKHIAFRYKLAKK 113
K+W + D AY ++D++D T L++E + + GL S K A Y +
Sbjct: 59 --KEWTKLVLDIAYDVEDVLD---TYFLKLEKRLHRLGLMRLTNIISDKKDA--YNILDD 111
Query: 114 MKRIGVWLDDIAAEKNKFHLTEIVRER----SGVVPDWRQTTSIVTQPLVYGRNEDKDKI 169
+K + D+ + + + R + V + R+ S + V G +D +
Sbjct: 112 IKTLKRRTLDVTRKLEMYGIGNFNEHRVVASTSRVREVRRARSDDQEERVVGLTDDAKVL 171
Query: 170 VDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKR 229
+ L+ D + + + + I G+ GLGKT+LA+ +FN + FE ++W VS + +
Sbjct: 172 LTKLLDDDGDNK-IYMISIFGMEGLGKTSLARKLFNSSDVKESFEYRVWTNVSGECNTRD 230
Query: 230 MTKAII---EGATKKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVL 286
+ II E ++ E + + L+ L D+L+ KRYL+V+DD+W + E + LK L
Sbjct: 231 ILMRIISSLEETSEGELEKMAQQELEVYLHDILQEKRYLVVVDDIW--ESEALESLKRAL 288
Query: 287 ACGGKGASILVTTRLPKVAKIMGT-IPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVI 345
C +G+ +++TT + VA+ + H + L+ ++ W LF+++AF +EL
Sbjct: 289 PCSYQGSRVIITTSIRVVAEGRDKRVYTHNIRFLTFKESWNLFEKKAFRYILKVDQELQK 348
Query: 346 VGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEAYVMPALRLSYLHL 405
+GKE+++KCGG P + L L+ +++ EW V S L +V LS+ +
Sbjct: 349 IGKEMVQKCGGLPRTTVVLAGLMS-RKKPNEWNDV-WSSLRVKDDNIHVSSLFDLSFKDM 406
Query: 406 PVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFF 465
+L+ CF + ++FP+D + + LI L A GFI ++ + +D+ +L + S
Sbjct: 407 GHELKLCFLYLSVFPEDYEVDVEKLIQLLVAEGFIQEDEEMTMEDVARYYIEDLVYISLV 466
Query: 466 ENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCIT--DDNSMRTMS--EETRHLLIYNRN 521
E + G++ F++HDLV + ++++ + D+ T S E HL+ N
Sbjct: 467 EVVKRKK-GKLMSFRIHDLVREFTIKKSKELNFVNVYDEQHSSTTSRREVVHHLMDDNYL 525
Query: 522 SFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLN--------- 572
N+ +++++ F + + L LRVL L+
Sbjct: 526 CDRRVNT-------QMRSFLFFGKRRNDITYVETITLKLKLLRVLNLGGLHFICQGYSPW 578
Query: 573 NLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQ 632
+L IG L +LRYL I++ NLP+ + L L+ L G S +++ L+ L L+
Sbjct: 579 SLPDVIGGLVHLRYLGIADTVVNNLPDFISNLRFLQTLDASG-NSFERMT-DLSNLTSLR 636
Query: 633 NLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKN 687
+L+ R L L L +L ++S Y + + LL L L + + HI N
Sbjct: 637 HLTGRFIGEL--LIGDAVNLQTLRSISSYSWSKLKHELLINLRDLEIY-EFHILN 688
>At1g58390 disease resistance protein PRM1
Length = 907
Score = 192 bits (489), Expect = 7e-49
Identities = 227/926 (24%), Positives = 410/926 (43%), Gaps = 136/926 (14%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MA ++ + L +L+ +E F G + + L L + + L+DA+ K+ + + +
Sbjct: 1 MAGELVSFGIKKLWDLLSQECEQFQGVEDQVTGLKRDLNLLSSFLKDADAKKHTTAVVRN 60
Query: 61 DVKDWLLKLKDAAYTLDDIMDECA-------TEALEMEYKASKCGLSHKHIAFRYKLAKK 113
V++ +K+ Y +DI++ T ++M + C +S R + A
Sbjct: 61 VVEE----IKEIVYDAEDIIETYLLKEKLWKTSGIKMRIRRHACIISD-----RRRNALD 111
Query: 114 MKRIGVWLDDIAAEKNKFHLTEIVRERSGVVP------DWRQTTSIVTQPLVYGRNEDKD 167
+ I + D+ + F + + + + + P + RQT S + G +
Sbjct: 112 VGGIRTRISDVIRDMQSFGVQQAIVDGGYMQPQGDRQREMRQTFSKDYESDFVGLEVNVK 171
Query: 168 KIVDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTL 227
K+V +LV ++E++ V I G+GGLGKTTLA+ VFNH+ + + F+ WVCVS++FT
Sbjct: 172 KLVGYLV----DEENVQVVSITGMGGLGKTTLARQVFNHEDVKHQFDRLAWVCVSQEFTR 227
Query: 228 KRMTKAIIEGATKKSCEDLDLELLQRKLQD----LLRRKRYLLVLDDVWNDKQENWQRLK 283
K + + I++ T + +D L++ + +L D LL + L+V DD+W D E+W +K
Sbjct: 228 KNVWQMILQNLTSREKKDEILQMEEAELHDKLFQLLETSKSLIVFDDIWKD--EDWDLIK 285
Query: 284 SVLACGGKGASILVTTRLPKVAKIMGTIPHHELSR--LSDEDCWELFKQRAFGPNEVQQ- 340
+ KG +L+T++ VA + G I + L+ ED W LF++ AF + +
Sbjct: 286 PIFP-PNKGWKVLLTSQNESVA-VRGDIKYLNFKPECLAIEDSWTLFQRIAFPKKDASES 343
Query: 341 ---KELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNLQG-----EA 392
+E+ +GK+++K CGG PLA LG LL K +W + + ++ G +
Sbjct: 344 KVDEEMEDMGKQMLKHCGGLPLAIKVLGGLLAAKYTMHDWERLSVNIGSDIVGRTSSNNS 403
Query: 393 YVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEAD--- 449
+ L +S+ LP L+ CF + A FP+D I+ + L W A G ++ +
Sbjct: 404 SIYHVLSMSFEELPSYLKHCFLYLAHFPEDHKINVEKLSYCWAAEGISTAEDYHNGETIQ 463
Query: 450 DIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCI--TDDNSMRT 507
D+G EL R+ + + +HD++ ++VC ++N ++
Sbjct: 464 DVGQSYLEELVRRNMIIWERDATASRFGTCHLHDMM--------REVCLFKAKEENFLQI 515
Query: 508 MSEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVLL 567
+ NS + S +L V T + D+ +L V+ + L V
Sbjct: 516 AVKSVGVTSSSTGNSQSPCRSRRL--VYQCPTTLHVERDINNP-KLRSLVVLWHDLWV-- 570
Query: 568 SHRLNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTR 627
L +S RLK LR LD+ F+ + KLP G+
Sbjct: 571 -ENWKLLGTSFTRLKLLRVLDLFYVDFEGM----------------------KLPFGIGN 607
Query: 628 LKRLQNLSLRDCDSLTSLPRQIGKL-----TSLNTLSKYIVGEERGFLLEELGQLNL--- 679
L L+ LSL+D ++ LP +G L +L+ +++I + + EL L L
Sbjct: 608 LIHLRYLSLQDA-KVSHLPSSLGNLMLLIYLNLDVDTEFIFVPDVFMRMHELRYLKLPLH 666
Query: 680 ---KGQLHIKNLERLKSVTDAKKANMSRKKL----NQLWLSWERNEVSQLQENVEQILEA 732
K +L ++NL +L+++ + S K L + L+ V+ + E + +
Sbjct: 667 MHKKTRLSLRNLVKLETLVYFSTWHSSSKDLCGMTRLMTLAIRLTRVT----STETLSAS 722
Query: 733 LQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLNLPELWKLPS-LKYL 791
+ Y + VG ++ + I + ++ LK L L+D L +P PS L ++
Sbjct: 723 ISGLRNLEYLYIVGTHSKKMREEGIVLDFIH-LKHL-LLD----LYMPRQQHFPSRLTFV 776
Query: 792 KLS-------------NMIHV--IYLFHESYDGEGLMALKTLF--LEKLPNLIGLSREER 834
KLS ++H+ + L SY G ++ F L+KL ++GL++ E
Sbjct: 777 KLSECGLEEDPMPILEKLLHLKGVILLKGSYCGRRMVCSGGGFPQLKKL-EIVGLNKWEE 835
Query: 835 VM-----FPRLKALEITECPNLLGLP 855
+ P L+ L I +C L +P
Sbjct: 836 WLVEEGSMPLLETLSILDCEELKEIP 861
Score = 31.2 bits (69), Expect = 3.4
Identities = 71/328 (21%), Positives = 116/328 (34%), Gaps = 85/328 (25%)
Query: 837 FPRLKALEITECPNL--------LGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHF 888
F RLK L + + + G+ L L L +Q LPSS+ L L L+
Sbjct: 580 FTRLKLLRVLDLFYVDFEGMKLPFGIGNLIHLRYLSLQDAKVSHLPSSLGNLMLLIYLNL 639
Query: 889 SDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHA---------------- 932
+ E I+ PD +R L H H K ++ ++ +
Sbjct: 640 DVDTEFIFVPDVFMRMHELRYLKLPLHMHKKTRLSLRNLVKLETLVYFSTWHSSSKDLCG 699
Query: 933 ---LQQLYINDCR-NIEELPNEVMQRLHSLKELDIVGCDKLKLSS-----DFQYL----- 978
L L I R E + + L +L+ L IVG K+ DF +L
Sbjct: 700 MTRLMTLAIRLTRVTSTETLSASISGLRNLEYLYIVGTHSKKMREEGIVLDFIHLKHLLL 759
Query: 979 -----------TCLETLAIGSCSEVEGFHEALQHMTTLKSLTL--------------SDL 1013
+ L + + C E L+ + LK + L
Sbjct: 760 DLYMPRQQHFPSRLTFVKLSECGLEEDPMPILEKLLHLKGVILLKGSYCGRRMVCSGGGF 819
Query: 1014 PNLEYLPECIG------------NLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDC 1061
P L+ L E +G ++ LL ++I C +L +P ++ I LE++
Sbjct: 820 PQLKKL-EIVGLNKWEEWLVEEGSMPLLETLSILDCEELKEIPDGLRFIYSLELV----- 873
Query: 1062 SKLEKRCQKEI---GEDWPKIVHVQYIE 1086
L R +K+ GED+ K+ H+ +E
Sbjct: 874 -MLGTRWKKKFSVGGEDYYKVQHIPSVE 900
>At1g59124 PRM1 homolog (RF45)
Length = 1155
Score = 185 bits (469), Expect = 1e-46
Identities = 262/1044 (25%), Positives = 435/1044 (41%), Gaps = 117/1044 (11%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MA ++ + +L L+ +E LF G + + L L + + L+DA K+ + +
Sbjct: 1 MAGELISFGIQNLWNLLSQECELFQGVEDQVTELKRDLNMLSSFLKDANAKKHTSAV--- 57
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEY-KASKCGLSHKHIAF----RYKLAKKMK 115
VK+ + ++K+ Y +D ++ T LE K S S + +A R + A +
Sbjct: 58 -VKNCVEEIKEIIYDGEDTIE---TFVLEQNLGKTSGIKKSIRRLACIIPDRRRYALGIG 113
Query: 116 RIGVWLDDIAAEKNKFHLTEIVRERSGVVP------DWRQTTSIVTQPLVYGRNEDKDKI 169
+ + + + F + + + + P + RQ S G + K+
Sbjct: 114 GLSNRISKVIRDMQSFGVQQAIVDGGYKQPQGDKQREMRQKFSKDDDSDFVGLEANVKKL 173
Query: 170 VDFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKR 229
V +LV +A+ Q V I G+GGLGKTTLA+ VFNH+ + + F+ WVCVS+DFT
Sbjct: 174 VGYLVDEANVQ----VVSITGMGGLGKTTLAKQVFNHEDVKHQFDGLSWVCVSQDFTRMN 229
Query: 230 MTKAIIEGATKKSCEDLDLELLQRKLQD----LLRRKRYLLVLDDVWNDKQENWQRLKSV 285
+ + I+ K E +E+ Q LQ LL + L+VLDD+W ++E+W+ +K +
Sbjct: 230 VWQKILRDLKPKEEEKKIMEMTQDTLQGELIRLLETSKSLIVLDDIW--EKEDWELIKPI 287
Query: 286 LACGGKGASILVTTRLPKVAKIMGT-IPHHELSRLSDEDCWELFKQRAFGPNEVQQ---- 340
KG +L+T+R VA T + + L+ ED W LF++ A + +
Sbjct: 288 FP-PTKGWKVLLTSRNESVAMRRNTSYINFKPECLTTEDSWTLFQRIALPMKDAAEFKID 346
Query: 341 KELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKES---------KLWNLQGE 391
+E +GK +IK CGG PLA LG +L K +W + E+ +N
Sbjct: 347 EEKEELGKLMIKHCGGLPLAIRVLGGMLAEKYTSHDWRRLSENIGSHLVGGRTNFNDDNN 406
Query: 392 AYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQM--LEAD 449
L LS+ LP L+ CF + A FP+D I + L W A G
Sbjct: 407 NTCNNVLSLSFEELPSYLKHCFLYLAHFPEDYEIKVENLSYYWAAEGIFQPRHYDGETIR 466
Query: 450 DIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHD--LAGSVTQDVCCITDDNSMRT 507
D+G+ EL R+ + +V + +HD++ + L + ++ IT
Sbjct: 467 DVGDVYIEELVRRNMVISERDVKTSRFETCHLHDMMREVCLLKAKEENFLQITSSRPSTA 526
Query: 508 MSEET---RHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLR 564
+ T R + + I +++L ++++ + ++L L
Sbjct: 527 NLQSTVTSRRFVYQYPTTLHVEKDINNPKLRALVVVTLGSWNLAGSSFTRLELLRVLDL- 585
Query: 565 VLLSHRLNNLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGG 624
+ + + L+S IG+L +LRYL + ++P SL L L L L +P
Sbjct: 586 IEVKIKGGKLASCIGKLIHLRYLSLEYAEVTHIPYSLGNLKLLIYLNLASFGRSTFVPNV 645
Query: 625 LTRLKRLQNLSL-RDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQL 683
L ++ L+ L+L D T L ++ L L TL + E L + G + L L
Sbjct: 646 LMGMQELRYLALPSDMGRKTKL--ELSNLVKLETLENF--STENSSLEDLCGMVRL-STL 700
Query: 684 HIKNLERLKSVTDAKKANMSRKKLNQLWLSWERNEVSQLQENVEQILEALQPYAQKLYSF 743
+IK +E T A K L +L + +E+ + + L+ KLY
Sbjct: 701 NIKLIEETSLETLAASIG-GLKYLEKLEIYDHGSEMRTKEAGIVFDFVHLKRLWLKLYMP 759
Query: 744 GVGGYTGAYFPQWISIPSLNDLKSLELVDCKSCLN-LPELWKLPSLKYLKLSNMIHVIYL 802
+ T +FP + L +L L C+ + +P L KL LK L+L
Sbjct: 760 RLS--TEQHFP--------SHLTTLYLESCRLEEDPMPILEKLLQLKELELG-------- 801
Query: 803 FHESYDGE-------GLMALKTLFLEKLPNLIGLSREERVMFPRLKALEITECPNLLGLP 855
ES+ G+ G L+ L L KL EE M P L+ L+I + P
Sbjct: 802 -FESFSGKKMVCSSGGFPQLQRLSLLKLEEWEDWKVEESSM-PLLRTLDIQKDP------ 853
Query: 856 CLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFH 915
LP+L L Y ++L + FP L+ L +
Sbjct: 854 -LPTLGRLV----YLKELQLGFRTFSGRIMVCSGGG-----FPQ---------LQKLSIY 894
Query: 916 RHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDF 975
R + + E + L LYI+DC +++LP + +Q ++SLK L I K +LS
Sbjct: 895 RLEEWEEWIVEQGSMPFLHTLYIDDCPKLKKLP-DGLQFIYSLKNLKISERWKERLSEGG 953
Query: 976 QYLTCLETLAIGSCSEVEGFHEAL 999
+ E + VE +H L
Sbjct: 954 E-----EYYKVQHIPSVEFYHRVL 972
Score = 50.8 bits (120), Expect = 4e-06
Identities = 92/366 (25%), Positives = 143/366 (38%), Gaps = 65/366 (17%)
Query: 777 LNLPELWKLP-SLKYLKLSNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSREERV 835
L E+ +P SL LKL +++ ++ LM ++ L LP+ +G R+ ++
Sbjct: 610 LEYAEVTHIPYSLGNLKLLIYLNLASFGRSTFVPNVLMGMQELRYLALPSDMG--RKTKL 667
Query: 836 MFPRLKALEITE--------CPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLH 887
L LE E +L G+ L +L+ I+ + L +SI L LE L
Sbjct: 668 ELSNLVKLETLENFSTENSSLEDLCGMVRLSTLNIKLIEETSLETLAASIGGLKYLEKLE 727
Query: 888 FSDN-EELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEE 946
D+ E+ GI+ + LK L + + L TE L LY+ CR +EE
Sbjct: 728 IYDHGSEMRTKEAGIVFDFVH-LKRLWLKLY--MPRLSTEQHFPSHLTTLYLESCR-LEE 783
Query: 947 LPNEVMQRLHSLKELDI-----VGCDKLKLSSDFQYLTCLETLAIGSCSE--VEGFHEAL 999
P ++++L LKEL++ G + S F L L L + + VE L
Sbjct: 784 DPMPILEKLLQLKELELGFESFSGKKMVCSSGGFPQLQRLSLLKLEEWEDWKVEESSMPL 843
Query: 1000 QHMTTLKSLTLSDLPNLEYLPEC------------------------------------- 1022
++ L L L YL E
Sbjct: 844 LRTLDIQKDPLPTLGRLVYLKELQLGFRTFSGRIMVCSGGGFPQLQKLSIYRLEEWEEWI 903
Query: 1023 --IGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPKIV 1080
G++ LH + I CPKL LP +Q I L+ L I S+ K E GE++ K+
Sbjct: 904 VEQGSMPFLHTLYIDDCPKLKKLPDGLQFIYSLKNLKI---SERWKERLSEGGEEYYKVQ 960
Query: 1081 HVQYIE 1086
H+ +E
Sbjct: 961 HIPSVE 966
>At1g59620 PRM1 homolog (CW9)
Length = 628
Score = 184 bits (468), Expect = 2e-46
Identities = 194/677 (28%), Positives = 311/677 (45%), Gaps = 106/677 (15%)
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAE +L + L +L+ +E F G ++FN L S L ++ LEDA+ K+ + +
Sbjct: 1 MAETLLSFGVEKLWDLLVRESDRFQGVKKQFNELRSDLNKLRCFLEDADAKKHQSAMVSN 60
Query: 61 DVKDWLLKLKDAAYTLDDIMDECA-------TEALEMEYKASKCGLSHKHIAFRYKLAKK 113
VK+ +K+ Y +DI++ T ++ K C L R K+A
Sbjct: 61 TVKE----VKEIVYDTEDIIETFLRKKQLGRTRGMKKRIKEFACVLPD-----RRKIAID 111
Query: 114 MKRIGVWLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFL 173
M+ + + A+K+K ++ RQT S + ++ G E+ K+V L
Sbjct: 112 MEGLSKRI----AKKDKRNM--------------RQTFSNNNESVLVGLEENVKKLVGHL 153
Query: 174 VGDASEQEDLS-VYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTK 232
V E ED S V I G+GG+GKTTLA+ VFNH+ + +HF WVCVS+ FT K + +
Sbjct: 154 V----EVEDSSQVVSITGMGGIGKTTLARQVFNHETVKSHFAQLAWVCVSQQFTRKYVWQ 209
Query: 233 AIIEGATKKSCEDLDLEL----LQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLAC 288
I+ K E + LE+ LQ KL LL ++ L+VLDD+W ++E+W ++ +
Sbjct: 210 TILR---KVGPEYIKLEMTEDELQEKLFRLLGTRKALIVLDDIW--REEDWDMIEPIFPL 264
Query: 289 GGKGASILVTTRLPKVAKIMGTIPHHELSR---LSDEDCWELFKQRAF-GPNEVQQK--- 341
GKG +L+T+R VA + P+ + + L+ E+ W +F++ F G N + K
Sbjct: 265 -GKGWKVLLTSRNEGVA--LRANPNGFIFKPDCLTPEESWTIFRRIVFPGENTTEYKVDE 321
Query: 342 ELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEW--------LYVKESKLWNLQGEAY 393
++ +GK++IK CGG PLA LG LL EW ++ +N + +
Sbjct: 322 KMEELGKQMIKHCGGLPLALKVLGGLLVVHFTLDEWKRIYGNIKSHIVGGTSFNDKNMSS 381
Query: 394 VMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEAD--DI 451
V L LS+ LP+ L+ CF + A FP+D I + L W A G A +
Sbjct: 382 VYHILHLSFEELPIYLKHCFLYLAQFPEDFTIDLEKLSYYWAAEGMPRPRYYDGATIRKV 441
Query: 452 GNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEE 511
G+ EL R+ + + + + K D D+ G + + +
Sbjct: 442 GDGYIEELVKRNMVISERDASKPRRLVVKGGDKT-DMEG---------------KLKNPK 485
Query: 512 TRHLLIYN-----RNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVL 566
R LL R +QL V L +EF G+L + LR L
Sbjct: 486 LRSLLFIEELGGYRGFEVWFTRLQLMRVLDLHG-VEF------GGELPSSIGLLIHLRYL 538
Query: 567 LSHR--LNNLSSSIGRLKYLRYLD--ISEGRFKNLPNSLCKLCNLEV----LKLDGCVSL 618
+R ++L SS+ LK L YL+ + E + +PN L ++ L+ L++D V L
Sbjct: 539 SLYRAKASHLPSSMQNLKMLLYLNLCVQESCYIYIPNFLKEMLELKYLSLPLRMDDKVKL 598
Query: 619 QKLPGGLTRLKRLQNLS 635
+ G L L++L+N S
Sbjct: 599 EL--GNLVNLEKLENFS 613
Score = 33.5 bits (75), Expect = 0.69
Identities = 35/106 (33%), Positives = 50/106 (47%), Gaps = 11/106 (10%)
Query: 563 LRVLLSHRLN---NLSSSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQ 619
+RVL H + L SSIG L +LRYL + + +LP+S+ L L L L CV
Sbjct: 511 MRVLDLHGVEFGGELPSSIGLLIHLRYLSLYRAKASHLPSSMQNLKMLLYLNL--CVQES 568
Query: 620 ---KLPGGLTRLKRLQNLSL-RDCDSLTSLPRQIGKLTSLNTLSKY 661
+P L + L+ LSL D L ++G L +L L +
Sbjct: 569 CYIYIPNFLKEMLELKYLSLPLRMDDKVKL--ELGNLVNLEKLENF 612
Score = 30.8 bits (68), Expect = 4.5
Identities = 33/134 (24%), Positives = 61/134 (44%), Gaps = 29/134 (21%)
Query: 991 EVEGFHEALQHMTTLKSLTLSDLPNLEY---LPECIGNLTLLHEINIYSCPKLACLPTSI 1047
E+ G+ T L+ + + DL +E+ LP IG L L +++Y K + LP+S+
Sbjct: 494 ELGGYRGFEVWFTRLQLMRVLDLHGVEFGGELPSSIGLLIHLRYLSLYRA-KASHLPSSM 552
Query: 1048 QQISGLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYI---------------------E 1086
Q + L L++ C +++ C I +++ ++Y+ +
Sbjct: 553 QNLKMLLYLNL--C--VQESCYIYIPNFLKEMLELKYLSLPLRMDDKVKLELGNLVNLEK 608
Query: 1087 IENDNLIHGGHGGS 1100
+EN + HGG GGS
Sbjct: 609 LENFSTEHGGVGGS 622
>At1g58410 unknown protein
Length = 899
Score = 182 bits (461), Expect = 1e-45
Identities = 187/724 (25%), Positives = 324/724 (43%), Gaps = 73/724 (10%)
Query: 13 LSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGRDVKDWLLKLKDA 72
L + + +E F G + + L S L +K+ L+DA+ K+ I V+ + ++KD
Sbjct: 11 LWDRLSQEYDQFKGVEDQVTELKSNLNLLKSFLKDADAKK----HISEMVRHCVEEIKDI 66
Query: 73 AYTLDDIMDE-CATEALEMEYKASK-CGLSHKHIAFRYKLAKKMKRIGVWLDDIAAEKNK 130
Y +DI++ E +EM+ K I R +LA + I + + +
Sbjct: 67 VYDTEDIIETFILKEKVEMKRGIMKRIKRFASTIMDRRELASDIGGISKRISKVIQDMQS 126
Query: 131 FHLTEIVRERSGVVP-------DWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASEQEDL 183
F + +I+ + S + R T S ++ G + K+V +LV E++D
Sbjct: 127 FGVQQIITDGSRSSHPLQERQREMRHTFSRDSENDFVGMEANVKKLVGYLV----EKDDY 182
Query: 184 SVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGATKKSC 243
+ + G+GGLGKTTLA+ VFNHD + + F+ WV VS++FT + + I++ T K
Sbjct: 183 QIVSLTGMGGLGKTTLARQVFNHDVVKDRFDGFAWVSVSQEFTRISVWQTILQNLTSKER 242
Query: 244 EDLDLELLQRKLQD----LLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTT 299
+D + + L D LL + L+VLDD+W K+E+W +K + KG +L+T+
Sbjct: 243 KDEIQNMKEADLHDDLFRLLESSKTLIVLDDIW--KEEDWDLIKPIFP-PKKGWKVLLTS 299
Query: 300 RLPKVA-KIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQ----KELVIVGKEIIKKC 354
R +A + T + LS D W LF+ A + + +E+ +GK++IK C
Sbjct: 300 RTESIAMRGDTTYISFKPKCLSIPDSWTLFQSIAMPRKDTSEFKVDEEMENMGKKMIKHC 359
Query: 355 GGFPLAAIALGSLLRFKREEKEWLYVKESKLWNL-----QGEAYVMPALRLSYLHLPVKL 409
GG LA LG LL K +W + E+ ++ + + L +S+ LP L
Sbjct: 360 GGLSLAVKVLGGLLAAKYTLHDWKRLSENIGSHIVERTSGNNSSIDHVLSVSFEELPNYL 419
Query: 410 RQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEAD---DIGNEVWNELYWRSFFE 466
+ CF + A FP+D I + L W A G IS + + + D G+ EL R+
Sbjct: 420 KHCFLYLAHFPEDHEIDVEKLHYYWAAEG-ISERRRYDGETIRDTGDSYIEELVRRNMVI 478
Query: 467 NTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEA 526
+ +V + ++HD++ ++ C+ + + H N + +
Sbjct: 479 SERDVMTSRFETCRLHDMMREI---------CLFKAKEENFLQIVSNHSPTSNPQTLGAS 529
Query: 527 NSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVLNCYSLRVLLSHRLNNLSSSI-GRLKYLR 585
LH+ +L E + +P++ + + + +R LS SI R+K LR
Sbjct: 530 RRFVLHNPTTLHV---------ERYKNNPKLRSLVVVYDDIGNRRWMLSGSIFTRVKLLR 580
Query: 586 YLDISEGRFK--NLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLT 643
LD+ + +FK LP+ + KL +L L L + LP L L L L +R +
Sbjct: 581 VLDLVQAKFKGGKLPSDIGKLIHLRYLSLKD-AKVSHLPSSLRNLVLLIYLDIRTDFTDI 639
Query: 644 SLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANMS 703
+P + L L E F+ E K +L + NLE+L+++ + + S
Sbjct: 640 FVPNVFMGMRELRYL------ELPRFMHE-------KTKLELSNLEKLEALENFSTKSSS 686
Query: 704 RKKL 707
+ L
Sbjct: 687 LEDL 690
>At1g12210 NBS/LRR disease resistance protein
Length = 885
Score = 172 bits (437), Expect = 7e-43
Identities = 166/634 (26%), Positives = 292/634 (45%), Gaps = 60/634 (9%)
Query: 62 VKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIAFRYKLAKKMKRIGVWL 121
V+ WL +++ +D++ C E + CG K++ Y K R+ V L
Sbjct: 72 VQVWLTRIQTIENQFNDLLSTCNAEIQRL----CLCGFCSKNVKMSYLYGK---RVIVLL 124
Query: 122 DDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASEQE 181
++ ++ + +IV E + + + + Q + G++ DK+ + L+ D
Sbjct: 125 REVEGLSSQ-GVFDIVTEAAPIA----EVEELPIQSTIVGQDSMLDKVWNCLMEDK---- 175
Query: 182 DLSVYPIVGLGGLGKTTLAQLVFNH-DKIVNHFELKIWVCVSEDFTLKRMTKAIIE--GA 238
+ + + G+GG+GKTTL + N K+ F++ IWV VS++ T+ ++ K+I E G
Sbjct: 176 -VWIVGLYGMGGVGKTTLLTQINNKFSKLGGGFDVVIWVVVSKNATVHKIQKSIGEKLGL 234
Query: 239 TKKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSV---LACGGKGASI 295
K+ ++ + + ++LRRK+++L+LDD+W + LK + G G +
Sbjct: 235 VGKNWDEKNKNQRALDIHNVLRRKKFVLLLDDIWEKVE-----LKVIGVPYPSGENGCKV 289
Query: 296 LVTTRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCG 355
TT +V MG E+S L + W+L K++ ++ + +++ +KC
Sbjct: 290 AFTTHSKEVCGRMGVDNPMEISCLDTGNAWDLLKKKVGENTLGSHPDIPQLARKVSEKCC 349
Query: 356 GFPLAAIALGSLLRFKREEKEWLYVKE--SKLWNLQG-EAYVMPALRLSYLHLPVK-LRQ 411
G PLA +G + FKR +EW + E + + G E ++P L+ SY L + +
Sbjct: 350 GLPLALNVIGETMSFKRTIQEWRHATEVLTSATDFSGMEDEILPILKYSYDSLNGEDAKS 409
Query: 412 CFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLE-ADDIGNEVWNELYWRSFFENTEN 470
CF +C+LFP+D I K++LI+ W GFI Q E A + G ++ L S
Sbjct: 410 CFLYCSLFPEDFEIRKEMLIEYWICEGFIKEKQGREKAFNQGYDILGTLVRSSLLLE--- 466
Query: 471 VGFGQITIFKMHDLVHDLAGSVTQDV-----CCITDDNSMRTMSEETRHLLIYNRNSFAE 525
G + MHD+V ++A + D+ CI E + R S
Sbjct: 467 -GAKDKDVVSMHDMVREMALWIFSDLGKHKERCIVQAGIGLDELPEVENWRAVKRMSLMN 525
Query: 526 ANSIQL----HHVKSLKTYMEFNFDVYEAGQLSPQVLNCY-SLRVL---LSHRLNNLSSS 577
N ++ V+ + +++ N Y+ +S + C SL VL +H L+ L
Sbjct: 526 NNFEKILGSPECVELITLFLQNN---YKLVDISMEFFRCMPSLAVLDLSENHSLSELPEE 582
Query: 578 IGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLR 637
I L L+YLD+S + LP+ L +L L LKL+ L+ + G++ L L+ L LR
Sbjct: 583 ISELVSLQYLDLSGTYIERLPHGLHELRKLVHLKLERTRRLESI-SGISYLSSLRTLRLR 641
Query: 638 DCDSL--TSLPRQIGKLTSL----NTLSKYIVGE 665
D + T L +++ L L +S +VGE
Sbjct: 642 DSKTTLDTGLMKELQLLEHLELITTDISSGLVGE 675
Score = 38.5 bits (88), Expect = 0.022
Identities = 35/120 (29%), Positives = 53/120 (44%), Gaps = 9/120 (7%)
Query: 779 LPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLIGLSREERVMFP 838
LPE+ ++K + L N F + + L TLFL+ L+ +S E P
Sbjct: 509 LPEVENWRAVKRMSLMNNN-----FEKILGSPECVELITLFLQNNYKLVDISMEFFRCMP 563
Query: 839 RLKALEITECPNLLGLPC----LPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSDNEEL 894
L L+++E +L LP L SL L + G Y ++LP +H+L L L L
Sbjct: 564 SLAVLDLSENHSLSELPEEISELVSLQYLDLSGTYIERLPHGLHELRKLVHLKLERTRRL 623
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.138 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,607,097
Number of Sequences: 26719
Number of extensions: 1154396
Number of successful extensions: 7323
Number of sequences better than 10.0: 401
Number of HSP's better than 10.0 without gapping: 248
Number of HSP's successfully gapped in prelim test: 155
Number of HSP's that attempted gapping in prelim test: 3299
Number of HSP's gapped (non-prelim): 2199
length of query: 1113
length of database: 11,318,596
effective HSP length: 110
effective length of query: 1003
effective length of database: 8,379,506
effective search space: 8404644518
effective search space used: 8404644518
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC134049.3


