

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134049.2 - phase: 0
(170 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
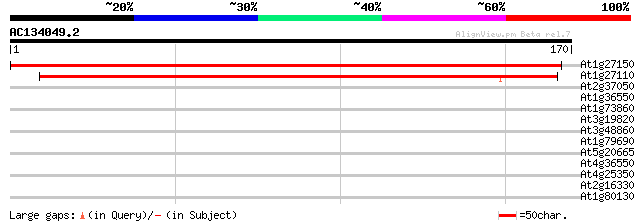
At1g27150 unknown protein 225 1e-59
At1g27110 hypothetical protein 192 6e-50
At2g37050 putative receptor-like protein kinase 28 2.4
At1g36550 hypothetical protein 28 2.4
At1g73860 kinesin-related protein 27 4.1
At3g19820 cell elongation protein, Dwarf1 27 5.3
At3g48860 putative protein 27 7.0
At1g79690 unknown protein 27 7.0
At5g20665 unknown protein 26 9.1
At4g36550 putative protein 26 9.1
At4g25350 putative protein 26 9.1
At2g16330 putative replication protein A1 26 9.1
At1g80130 unknown protein 26 9.1
>At1g27150 unknown protein
Length = 468
Score = 225 bits (573), Expect = 1e-59
Identities = 102/167 (61%), Positives = 133/167 (79%)
Query: 1 MGINHHVLPQNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQ 60
+G+ VLP N+ E++I+G+LAFPLLELG+M+EA A+++G+EIN +D W+ H CHVLQ
Sbjct: 144 LGLVQQVLPANQEESYIHGLLAFPLLELGRMEEAAAASRKGYEINKEDAWAHHCLCHVLQ 203
Query: 61 YECRFREAVEFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKEL 120
+ECRF+EAVEFME + +W S SFM THNWWHVALCYLEG +PM +V E+YD++IWKEL
Sbjct: 204 HECRFKEAVEFMEALAGTWPSCSSFMYTHNWWHVALCYLEGGSPMSKVEEIYDHHIWKEL 263
Query: 121 DKTDATVPEVYLNAVALLLRLCVRDELEFFGDRLKMLADRLADQVSY 167
+K DA PEVYLNA+ LL+RL VRD L+ F DRLK LA RL +Q ++
Sbjct: 264 EKDDAVPPEVYLNALGLLIRLDVRDALDGFEDRLKNLAVRLTNQANW 310
>At1g27110 hypothetical protein
Length = 519
Score = 192 bits (489), Expect = 6e-50
Identities = 92/158 (58%), Positives = 120/158 (75%), Gaps = 1/158 (0%)
Query: 10 QNEGENFIYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAV 69
+NEG+ ++ GMLAF L+ELG ++EAEEAA++G EIN D W+ HA CHVLQ ECRF+EAV
Sbjct: 132 KNEGQVYVNGMLAFCLIELGHLREAEEAARKGCEINENDSWAHHALCHVLQTECRFKEAV 191
Query: 70 EFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRVLEVYDNYIWKELDKTDATVPE 129
+FMEE S SW+S S +HNWWHVA+CYLEG + + +V EVYD+ +WKEL+K DA +
Sbjct: 192 KFMEEHSDSWDSCSSLRFSHNWWHVAVCYLEGGSHISKVEEVYDHQMWKELEKDDAVARD 251
Query: 130 VYLNAVALLLRLCVRDEL-EFFGDRLKMLADRLADQVS 166
VY +A+ LLLRL R +L + F DRL+ LAD L D+VS
Sbjct: 252 VYTDALGLLLRLDTRGKLDDGFQDRLEKLADSLTDKVS 289
>At2g37050 putative receptor-like protein kinase
Length = 933
Score = 28.1 bits (61), Expect = 2.4
Identities = 17/72 (23%), Positives = 29/72 (39%)
Query: 19 GMLAFPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEECSPS 78
G F ++ G+ +E +E A + N+ G + A L R V+F+ C
Sbjct: 610 GSGGFGIVYYGKTREGKEIAVKVLANNSYQGKREFANEVTLLSRIHHRNLVQFLGYCQEE 669
Query: 79 WNSFLSFMLTHN 90
+ L + HN
Sbjct: 670 GKNMLVYEFMHN 681
>At1g36550 hypothetical protein
Length = 353
Score = 28.1 bits (61), Expect = 2.4
Identities = 21/72 (29%), Positives = 31/72 (42%), Gaps = 5/72 (6%)
Query: 98 YLEGNAPMQRVLEVYDNYIWKELDKTDATVPEVYLNAVALLLRLCVRDELE-----FFGD 152
Y+E N E YD + L + + V E Y L+LR + ++ E F GD
Sbjct: 120 YIEINRVRVAATEFYDYALSWNLTQGNRYVEEYYKEMETLMLRADISEDREATMSRFLGD 179
Query: 153 RLKMLADRLADQ 164
+ + DRL Q
Sbjct: 180 LNRDIQDRLETQ 191
>At1g73860 kinesin-related protein
Length = 1050
Score = 27.3 bits (59), Expect = 4.1
Identities = 15/49 (30%), Positives = 29/49 (58%), Gaps = 2/49 (4%)
Query: 94 VALCYLEGNAPMQRVLEVYDNYIWKELDKTDATVPEVYLNAVALLLRLC 142
V LC ++ NAP Q +L V + + + +++ + +P++ + V LLL C
Sbjct: 125 VGLCLMQ-NAPTQSLLSVLNGILDESIERKNGEIPQI-PSFVLLLLGKC 171
>At3g19820 cell elongation protein, Dwarf1
Length = 561
Score = 26.9 bits (58), Expect = 5.3
Identities = 14/39 (35%), Positives = 22/39 (55%), Gaps = 3/39 (7%)
Query: 15 NFIYGMLAFP---LLELGQMKEAEEAAKRGFEINNQDGW 50
+F+ GM+ P ++ +G EEA K+G +INN W
Sbjct: 270 DFVEGMVYNPTEGVMMVGTYASKEEAKKKGNKINNVGWW 308
>At3g48860 putative protein
Length = 609
Score = 26.6 bits (57), Expect = 7.0
Identities = 18/82 (21%), Positives = 38/82 (45%), Gaps = 9/82 (10%)
Query: 27 ELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEECSPSWNSFLSFM 86
ELG+ ++ + A K+ + ++++ S+H Q++ R+R S+N +
Sbjct: 475 ELGEAEQEDVAFKQIYVLDDETYESKHCRTVTYQFKSRYR---------LDSYNYIMQAW 525
Query: 87 LTHNWWHVALCYLEGNAPMQRV 108
L + W L +E + +RV
Sbjct: 526 LMYFWGRAKLHGVEDDIAEERV 547
>At1g79690 unknown protein
Length = 772
Score = 26.6 bits (57), Expect = 7.0
Identities = 23/74 (31%), Positives = 35/74 (47%), Gaps = 14/74 (18%)
Query: 110 EVYDNYI-WKELD-KTDATV-------PEVYLNAVALLLRLCVRDE-----LEFFGDRLK 155
E Y++ I W +LD K D T+ E++ + +RD+ L+ FGD LK
Sbjct: 433 EYYESDIAWMDLDSKLDITIGPYETYEDEIFGYKATFETFIGIRDDKATADLKLFGDNLK 492
Query: 156 MLADRLADQVSYDS 169
+L D L + Y S
Sbjct: 493 LLEDNLPLESVYKS 506
>At5g20665 unknown protein
Length = 301
Score = 26.2 bits (56), Expect = 9.1
Identities = 9/27 (33%), Positives = 19/27 (70%)
Query: 26 LELGQMKEAEEAAKRGFEINNQDGWSQ 52
+++G+ +E E +RG + +N++GW Q
Sbjct: 195 MQVGEEEEEELMIERGEKSSNEEGWHQ 221
>At4g36550 putative protein
Length = 680
Score = 26.2 bits (56), Expect = 9.1
Identities = 10/21 (47%), Positives = 17/21 (80%)
Query: 124 DATVPEVYLNAVALLLRLCVR 144
++ VPE NA+++LL+LCV+
Sbjct: 556 ESNVPEEQENAISILLQLCVQ 576
>At4g25350 putative protein
Length = 710
Score = 26.2 bits (56), Expect = 9.1
Identities = 15/42 (35%), Positives = 25/42 (58%), Gaps = 3/42 (7%)
Query: 120 LDKTDATVPEVYLNAVALLLRLCVRDELEFFGDRLKMLADRL 161
++KT P L+A+ +L++ +DEL+F D LK + RL
Sbjct: 206 MNKTREITP---LSAIKTILKVHKQDELKFTRDNLKEVEKRL 244
>At2g16330 putative replication protein A1
Length = 233
Score = 26.2 bits (56), Expect = 9.1
Identities = 17/60 (28%), Positives = 29/60 (48%), Gaps = 7/60 (11%)
Query: 25 LLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEECSPSWNSFLS 84
L+E+G + E K F+I ++ VLQ E + +A EF +EC +++S
Sbjct: 143 LIEIGNLTEDYRGRKLPFKIMDKHK-------EVLQCEAQNDQAFEFKDECFTYTTNYVS 195
>At1g80130 unknown protein
Length = 305
Score = 26.2 bits (56), Expect = 9.1
Identities = 16/44 (36%), Positives = 23/44 (51%), Gaps = 1/44 (2%)
Query: 7 VLPQNEGENFIYGMLAFPLLEL-GQMKEAEEAAKRGFEINNQDG 49
++ N G + + G A L E+ G MK+AEE +R N DG
Sbjct: 172 MIDSNPGNSLLTGNYAKFLKEVKGDMKKAEEYCERAILGNTNDG 215
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,989,950
Number of Sequences: 26719
Number of extensions: 163718
Number of successful extensions: 347
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 341
Number of HSP's gapped (non-prelim): 13
length of query: 170
length of database: 11,318,596
effective HSP length: 92
effective length of query: 78
effective length of database: 8,860,448
effective search space: 691114944
effective search space used: 691114944
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC134049.2


