

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
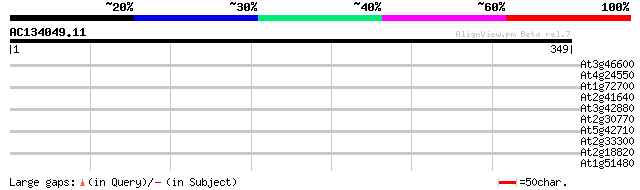
Query= AC134049.11 - phase: 0
(349 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g46600 scarecrow-like protein 30 1.5
At4g24550 clathrin coat assembly like protein 30 2.6
At1g72700 putative P-type transporting ATPase 30 2.6
At2g41640 unknown protein 29 3.3
At3g42880 receptor protein kinase -like protein 28 5.7
At2g30770 putative cytochrome P450 28 5.7
At5g42710 unknown protein 28 7.5
At2g33300 hypothetical protein 28 7.5
At2g18820 putative non-LTR retroelement reverse transcriptase 28 9.7
At1g51480 hypothetical protein 28 9.7
>At3g46600 scarecrow-like protein
Length = 583
Score = 30.4 bits (67), Expect = 1.5
Identities = 22/49 (44%), Positives = 27/49 (54%), Gaps = 1/49 (2%)
Query: 216 LEDEEFCRAAADGGFDIRLTFQPPNSPDLNVLDLGFFSAIQSLQQQEVT 264
LED +AA F++ L Q P S DL LG FS+I SL Q EV+
Sbjct: 82 LEDSLALQAAERSFFEV-LQDQTPISGDLEDGSLGNFSSITSLHQPEVS 129
>At4g24550 clathrin coat assembly like protein
Length = 451
Score = 29.6 bits (65), Expect = 2.6
Identities = 13/22 (59%), Positives = 15/22 (68%)
Query: 76 PMYNVVHIDEKWFYITKKSSSY 97
PMYNV + K+ I KKSSSY
Sbjct: 411 PMYNVSKLQVKYLQIAKKSSSY 432
>At1g72700 putative P-type transporting ATPase
Length = 1228
Score = 29.6 bits (65), Expect = 2.6
Identities = 23/86 (26%), Positives = 35/86 (39%), Gaps = 22/86 (25%)
Query: 284 QSSNNIFLTLQSCMTEIMKVKGSNNYKIPH-------MEKEKLLRRGML----PTQL--- 329
QS ++ T S T+ ++V+G NNY P E +L+ L P +
Sbjct: 477 QSQTKVYGTWDSSRTQEIEVEGDNNYNTPRAPIKGFGFEDNRLMNGNWLRESQPNDILQF 536
Query: 330 --------TCDPELFQETFEYLYNVD 347
T PEL +ET +Y Y +
Sbjct: 537 FRILAICHTAIPELNEETGKYTYEAE 562
>At2g41640 unknown protein
Length = 500
Score = 29.3 bits (64), Expect = 3.3
Identities = 20/63 (31%), Positives = 30/63 (46%), Gaps = 1/63 (1%)
Query: 245 NVLDLGFFSAIQSLQQQEVTNSVDALIEAVQKSFDAFSAQSSNNIFLTLQSCMTEIMKVK 304
NVLD G+ IQSL Q+E +V AL + S S+ L ++ + E+ +
Sbjct: 287 NVLDRGYSHRIQSLTQEETEANVTALDFKKKPKLVILSRNGSSRAILN-ENLLVELAEKT 345
Query: 305 GSN 307
G N
Sbjct: 346 GFN 348
>At3g42880 receptor protein kinase -like protein
Length = 633
Score = 28.5 bits (62), Expect = 5.7
Identities = 22/78 (28%), Positives = 32/78 (40%), Gaps = 3/78 (3%)
Query: 33 EGVLRRHSNVLKPYLNDSNKKARIEFCLDMLEESSIPHDPVFKPMYNVVHIDEKWF---Y 89
+ V+ +V+ + D NK AR F +M + H V P+ +EK Y
Sbjct: 374 KAVMANGLSVVVKRIRDMNKLAREAFDTEMQRFGKLRHPNVLTPLAYHYRREEKLVVSEY 433
Query: 90 ITKKSSSYYLLGDEDEPH 107
+ K S Y L GD H
Sbjct: 434 MPKSSLLYVLHGDRGVYH 451
>At2g30770 putative cytochrome P450
Length = 497
Score = 28.5 bits (62), Expect = 5.7
Identities = 15/55 (27%), Positives = 34/55 (61%), Gaps = 4/55 (7%)
Query: 244 LNVLDLGFFSAIQSLQQQEVTNSVDALIEAVQKSFDAFSAQSSNNIFLTLQSCMT 298
LN+L + + +++ EV +A+IE ++K+ + S+++ + +F+TL S +T
Sbjct: 134 LNLLTNKMVESFEKVREDEV----NAMIEKLEKASSSSSSENLSELFITLPSDVT 184
>At5g42710 unknown protein
Length = 807
Score = 28.1 bits (61), Expect = 7.5
Identities = 11/38 (28%), Positives = 20/38 (51%)
Query: 38 RHSNVLKPYLNDSNKKARIEFCLDMLEESSIPHDPVFK 75
+H +LK Y + + KKA ++ C+ + S + H K
Sbjct: 516 KHDLLLKSYNDSNEKKAEVDTCIKSSQVSGVEHKKEIK 553
>At2g33300 hypothetical protein
Length = 265
Score = 28.1 bits (61), Expect = 7.5
Identities = 21/79 (26%), Positives = 37/79 (46%), Gaps = 8/79 (10%)
Query: 168 KPIISINREVTRMMYINEVLPAL----KEKWPQEHAHETIYIQQDNAPCHVPLEDEEFCR 223
K + N+ +T + N V+ K + HAHET ++Q + HVP ++ +
Sbjct: 10 KDAYTYNKALTLNLKKNNVVKVCSGLKKREHQSSHAHETGEVKQIDEQNHVP--EKVSMK 67
Query: 224 AAADGG--FDIRLTFQPPN 240
+ D + +RLT Q P+
Sbjct: 68 SVRDNSTCYKVRLTLQKPD 86
>At2g18820 putative non-LTR retroelement reverse transcriptase
Length = 1412
Score = 27.7 bits (60), Expect = 9.7
Identities = 22/68 (32%), Positives = 32/68 (46%), Gaps = 6/68 (8%)
Query: 237 QPPNSPDLNVLDLGFFSAIQSLQQQEVTN-SVDALIEAVQKSFDAFSAQSSNNIFLTLQS 295
+PPN D+ + + FFS + S Q + T SVD L +Q +S N L +
Sbjct: 665 RPPNQDDIKIEAVRFFSDLLSSQPSDFTGISVDELKGILQY---RYSLHEQN--LLVAEI 719
Query: 296 CMTEIMKV 303
E+MKV
Sbjct: 720 TEAEVMKV 727
>At1g51480 hypothetical protein
Length = 941
Score = 27.7 bits (60), Expect = 9.7
Identities = 27/131 (20%), Positives = 53/131 (39%), Gaps = 26/131 (19%)
Query: 37 RRHSNVLKPYLNDS-NKKARIEFCLDMLEESSIPHDPVFKPMYNVVHIDEKW---FYITK 92
+ H +L P S N K + ++ S DPVF P++ + E++ F I +
Sbjct: 14 KSHIEMLHPACEMSWNLKQHDDTAASLMVIQSPTSDPVFNPVHTIKQFGERYTQDFVIRE 73
Query: 93 KSSSYYLLGDEDEPHRMCRNKNYIGKVMFLVVVARPRFDDDGNETFSGKIGCF----PIV 148
+ S+ +L H C+ +++ + + N+ F+ GCF +
Sbjct: 74 RDKSFGVL-----IHCFCKMADWLLLIPW-------------NKIFTAACGCFFSDRNYI 115
Query: 149 HKVAAQRSSIN 159
HK+ A ++
Sbjct: 116 HKMEANLDDLH 126
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,850,427
Number of Sequences: 26719
Number of extensions: 344524
Number of successful extensions: 757
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 753
Number of HSP's gapped (non-prelim): 10
length of query: 349
length of database: 11,318,596
effective HSP length: 100
effective length of query: 249
effective length of database: 8,646,696
effective search space: 2153027304
effective search space used: 2153027304
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC134049.11


