

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133780.8 - phase: 0 /pseudo
(287 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
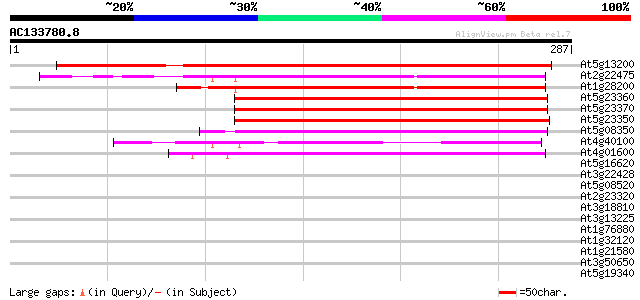
Score E
Sequences producing significant alignments: (bits) Value
At5g13200 ABA-responsive protein - like 313 9e-86
At2g22475 unknown protein 187 4e-48
At1g28200 FH protein interacting protein FIP1 187 7e-48
At5g23360 unknown protein 141 5e-34
At5g23370 putative protein 140 6e-34
At5g23350 unknown protein 138 3e-33
At5g08350 unknown protein 137 7e-33
At4g40100 putative protein 129 1e-30
At4g01600 putative ABA-repsonsive protein 127 5e-30
At5g16620 translocon Tic40-like protein 38 0.006
At3g22428 hypothetical protein 38 0.007
At5g08520 unknown protein 37 0.012
At2g23320 putative WRKY-type DNA-binding protein 37 0.012
At3g18810 protein kinase, putative 35 0.036
At3g13225 hypothetical protein, 5' partial 34 0.079
At1g76880 unknown protein 34 0.079
At1g32120 hypothetical protein 34 0.079
At1g21580 hypothetical protein 34 0.079
At3g50650 scarecrow-like 7 (SCL7) 34 0.10
At5g19340 putative protein 33 0.14
>At5g13200 ABA-responsive protein - like
Length = 272
Score = 313 bits (801), Expect = 9e-86
Identities = 148/254 (58%), Positives = 189/254 (74%), Gaps = 9/254 (3%)
Query: 25 PPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASADGQGQPQVHYY 84
P + + P P + + WGTH+MGAPA P +HPDN++AA A + Q Q Q
Sbjct: 18 PKTLETEHQPEPSSSSPDQKKWGTHVMGAPAAPVAHPDNQQAAAWVAGDNQQTQYQ---- 73
Query: 85 QQDHPYVQHSPVDKPSSS-PMESILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKM 143
PYV +SPV+ P+++ P+E ++ MF +WS+KAE A N+WHNLKTGPS+S A GK+
Sbjct: 74 ----PYVIYSPVEHPTTNNPLEPVIGMFHTWSRKAETVARNLWHNLKTGPSMSETAWGKV 129
Query: 144 NLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDR 203
NLT KAI++GGFESL++QIF T PNE LKKTFACYLSTTTGPVAGT+YLS+ +AFCSDR
Sbjct: 130 NLTAKAITKGGFESLFRQIFGTEPNETLKKTFACYLSTTTGPVAGTVYLSNARVAFCSDR 189
Query: 204 PLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNY 263
PL FTAPSGQ +WSYY+V+VPL + TVNPV+++E E+YIQ+ TVDGHDFWFMGFVNY
Sbjct: 190 PLYFTAPSGQESWSYYRVVVPLANVATVNPVVVKETPPEKYIQLTTVDGHDFWFMGFVNY 249
Query: 264 DKAVKNLSEGISHF 277
+KA +L +S F
Sbjct: 250 EKATHHLLTSVSDF 263
>At2g22475 unknown protein
Length = 299
Score = 187 bits (476), Expect = 4e-48
Identities = 100/269 (37%), Positives = 149/269 (55%), Gaps = 39/269 (14%)
Query: 16 DENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASADG 75
D P +P+ SS S + +W ++ S PD K A+ +A ++
Sbjct: 52 DSTPVKAPSRTSSGSKK----------SVHWSPELVSG----SQEPDQKAASSSSAGSN- 96
Query: 76 QGQPQVHYYQQDHPYVQHSPVDKPSSS---PMESILHMFDSW-------SKKAEATANNI 125
PY+ SP + +S ME++ + W +KK E+ A N
Sbjct: 97 -------------PYIARSPAETSDASLKDTMETVKGVLGRWGKRVAEAAKKTESLAGNT 143
Query: 126 WHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGP 185
W +L+T PS + AAMG++ + K +EGG+E +++Q F T P E+L +FACYLST+ GP
Sbjct: 144 WQHLRTAPSFADAAMGRIAQSTKVFAEGGYEKIFRQTFETDPEEQLLNSFACYLSTSAGP 203
Query: 186 VAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYI 245
V G LY+S LA+CSD PLS+ Q WSYYKV++PL ++ VNP N +E+YI
Sbjct: 204 VMGVLYISSAKLAYCSDNPLSY-KNGDQTEWSYYKVVIPLHQLKAVNPSASIVNPAEKYI 262
Query: 246 QIVTVDGHDFWFMGFVNYDKAVKNLSEGI 274
Q+++VD H+FWFMGF+NYD AV +L + +
Sbjct: 263 QVISVDNHEFWFMGFLNYDGAVTSLQDSL 291
>At1g28200 FH protein interacting protein FIP1
Length = 259
Score = 187 bits (474), Expect = 7e-48
Identities = 91/196 (46%), Positives = 127/196 (64%), Gaps = 11/196 (5%)
Query: 86 QDHPYVQHSPVDKPSSSPMESILHMFDSW-------SKKAEATANNIWHNLKTGPSVSSA 138
+ +PYV SP + + M+S+ W +KKAE A N W +LKTGPSV+ A
Sbjct: 63 ESNPYVSPSPAPR---NTMDSVKDTLGKWGKMAADATKKAEDLAGNFWQHLKTGPSVADA 119
Query: 139 AMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLA 198
A+ ++ K ++EGG+E ++KQ F P+EKL KT+ACYLST+ GPV G +YLS LA
Sbjct: 120 AVSRIAQGTKILAEGGYEKVFKQTFDCLPDEKLLKTYACYLSTSAGPVLGVMYLSTHKLA 179
Query: 199 FCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFM 258
F SD PLS+ Q WSYYKV++P ++ VNP R N S++YIQ++++D H+FWFM
Sbjct: 180 FSSDNPLSY-KEGEQTLWSYYKVVLPANQLKAVNPSTSRVNTSDKYIQVISIDNHEFWFM 238
Query: 259 GFVNYDKAVKNLSEGI 274
GFV Y+ AVK+L E +
Sbjct: 239 GFVTYESAVKSLQEAV 254
>At5g23360 unknown protein
Length = 210
Score = 141 bits (355), Expect = 5e-34
Identities = 67/160 (41%), Positives = 98/160 (60%)
Query: 116 KKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTF 175
KK ++ N K GP ++ K++L K + GG E +YK++F EKL K +
Sbjct: 47 KKTDSFTNGARDQDKLGPKLTETVKRKLSLGAKILQMGGLEKIYKRLFKVCDKEKLFKAY 106
Query: 176 ACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVI 235
CYLSTT G +AG L++S +AFCS+R + T+P G +T +YKV +PL KI VN
Sbjct: 107 QCYLSTTEGSIAGLLFISSKKIAFCSERSIKVTSPQGDLTRVHYKVSIPLCKINGVNQSQ 166
Query: 236 MRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
+ S+RY+++VTVD +DFWFMGFV+Y KA L + ++
Sbjct: 167 NTKKPSQRYLEVVTVDNYDFWFMGFVSYQKAFNCLEKALN 206
>At5g23370 putative protein
Length = 219
Score = 140 bits (354), Expect = 6e-34
Identities = 65/160 (40%), Positives = 98/160 (60%)
Query: 116 KKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTF 175
KK ++ N + K GP ++ K++L + + GG E +YK++F EKL K +
Sbjct: 55 KKNDSFTNGVRDQDKLGPKLTETVKRKLSLGARILQMGGLEKIYKRLFKVSDEEKLFKAY 114
Query: 176 ACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVI 235
CYLSTT GP+AG L++S +AFCS+R + +P G++ +YKV +PL KI VN
Sbjct: 115 QCYLSTTAGPIAGLLFISSKKIAFCSERSIKVASPQGELNRVHYKVSIPLCKINGVNQSQ 174
Query: 236 MRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
S++Y+++VTVDG DFWFMGF++Y KA L + +S
Sbjct: 175 NTTKPSQKYLEVVTVDGFDFWFMGFLSYQKAFNCLEQALS 214
>At5g23350 unknown protein
Length = 218
Score = 138 bits (348), Expect = 3e-33
Identities = 65/161 (40%), Positives = 98/161 (60%)
Query: 116 KKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTF 175
KK ++ N K GP ++ K++L K + GG E +YK++F EKL K +
Sbjct: 55 KKTDSFTNGARDQDKLGPKLTETVKRKLSLGAKILQMGGLEKIYKRLFKVCDQEKLFKAY 114
Query: 176 ACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVI 235
CYLSTT GP+AG L++S +AFCS+R + +P G ++ +YKV +PL KI VN
Sbjct: 115 QCYLSTTAGPIAGLLFISSKKIAFCSERSIKVASPQGVLSRVHYKVSIPLCKINGVNQSQ 174
Query: 236 MRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGISH 276
+ S++Y++IVT+D DFWFMGFV+Y KA L + +++
Sbjct: 175 NTKKPSQKYLEIVTIDNFDFWFMGFVSYQKAFNCLEKALNN 215
>At5g08350 unknown protein
Length = 222
Score = 137 bits (345), Expect = 7e-33
Identities = 68/178 (38%), Positives = 103/178 (57%), Gaps = 5/178 (2%)
Query: 98 KPSSSPMESILHMFDSWSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFES 157
K S ++SIL KK + N + K P ++ K++L + + GG E
Sbjct: 41 KSEQSNVKSILKR-----KKTDGFTNGVRDQSKIRPKLTETVKRKLSLGARILQVGGLEK 95
Query: 158 LYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWS 217
++K++F EKL K + CYLSTT GP+AG L++S +AFCS+R + +P G +
Sbjct: 96 IFKRLFRVSEGEKLFKMYQCYLSTTAGPIAGLLFISSKKMAFCSERSIKVDSPQGDIIRV 155
Query: 218 YYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
+YKV +PL KI VN + S++Y+++VTVDG DFWFMGF++Y KA L + +S
Sbjct: 156 HYKVSIPLCKIDRVNQSQNTKKPSQKYLEVVTVDGFDFWFMGFLSYQKAFNCLEKALS 213
>At4g40100 putative protein
Length = 225
Score = 129 bits (325), Expect = 1e-30
Identities = 78/230 (33%), Positives = 120/230 (51%), Gaps = 58/230 (25%)
Query: 54 PAVPSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSS---PMESILHM 110
P + S P + AL ++SA + +PYV +P + +S MES+ +
Sbjct: 32 PELVSESPAPDEKALSSSSA-----------ARSNPYVARAPTETSDASLKETMESVKGV 80
Query: 111 FDSWSK-------KAEATANNIW-HNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQI 162
W + KAE+ A N W H L+ AAMG++ + K ++EGG+E +++Q
Sbjct: 81 LGRWGRRVGEAAMKAESLAGNTWQHPLR-------AAMGRIAQSTKVLAEGGYEKIFRQT 133
Query: 163 FTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVM 222
F T P E+L+ +FACYLST+ GPV G LY V+
Sbjct: 134 FETVPEEQLQNSFACYLSTSAGPVMGVLY-----------------------------VV 164
Query: 223 VPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSE 272
+PL ++ +VNP I N +E+YIQ+++VD H+FWFMGF+NY+ AV +L +
Sbjct: 165 IPLHQLKSVNPSISTVNPAEKYIQVISVDDHEFWFMGFLNYEGAVTSLQD 214
>At4g01600 putative ABA-repsonsive protein
Length = 233
Score = 127 bits (320), Expect = 5e-30
Identities = 74/202 (36%), Positives = 115/202 (56%), Gaps = 9/202 (4%)
Query: 82 HYYQQDHPYVQ----HSPVDKPSSSPMESILHM----FDSWSKKAEATANNIWHNLKTGP 133
H + D+PYV S DK S + +L+ + ++KAEA + +LK P
Sbjct: 23 HGGRYDNPYVHITSPTSASDKRSKDKVLEVLNRCGKKVEDATRKAEALVGGLKDHLKFSP 82
Query: 134 SVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLS 193
S+S AAM +++ K I EGG E ++++ F EKL +F CY+STT+GPV G +Y+S
Sbjct: 83 SISDAAMARLSQGTKMIVEGGPERVFQREFGVLAVEKLLDSFVCYISTTSGPVTGVIYIS 142
Query: 194 DIHLAFCSDRPLSF-TAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDG 252
+ +AFCSD + ++ G +YYKV++ KI +++ SERY+ +VT DG
Sbjct: 143 NRRIAFCSDYAIRLPSSAGGNGVAAYYKVVMEWEKISSISSSTNVLKPSERYVHMVTRDG 202
Query: 253 HDFWFMGFVNYDKAVKNLSEGI 274
+FWFMGFV+Y A L++ +
Sbjct: 203 FEFWFMGFVSYIDAFNCLNKAL 224
>At5g16620 translocon Tic40-like protein
Length = 447
Score = 38.1 bits (87), Expect = 0.006
Identities = 39/167 (23%), Positives = 66/167 (39%), Gaps = 25/167 (14%)
Query: 1 MNNNNNNNNQ--NQSSPDENPFASPTPPSSSSSSVPLPPQNH---ATTENWGTHMMGAPA 55
MN N N+Q N P +PF P PP +S +S P Q+ AT + T + P+
Sbjct: 142 MNQMNTQNSQFNNSGFPSGSPFPFPFPPQTSPASSPFQSQSQSSGATVDVTATKVETPPS 201
Query: 56 VPSSHPDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMESILHMFDSWS 115
K + S + + ++++ + SP + SP + + ++ S
Sbjct: 202 TKPKPTPAKDIEVDKPSVVLEASKE-KKEEKNYAFEDISPEETTKESPFSNYAEVSETNS 260
Query: 116 KKAE----------------ATANNIWHNL---KTGPSVSSAAMGKM 143
K ATA+ ++ +L K GP +S A+ KM
Sbjct: 261 PKETRLFEDVLQNGAGPANGATASEVFQSLGGGKGGPGLSVEALEKM 307
>At3g22428 hypothetical protein
Length = 342
Score = 37.7 bits (86), Expect = 0.007
Identities = 20/69 (28%), Positives = 30/69 (42%), Gaps = 5/69 (7%)
Query: 3 NNNNNNNQNQSSP-----DENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVP 57
+NNNNNN+ +S P + P P+ S P+P Q++ +N P P
Sbjct: 45 SNNNNNNKKESKPMMIGVQKTADKKPLKPNPSPKLAPIPKQSNPNHQNQTGPSSSRPMAP 104
Query: 58 SSHPDNKKA 66
PD+ A
Sbjct: 105 FPFPDSSPA 113
Score = 29.6 bits (65), Expect = 2.0
Identities = 16/47 (34%), Positives = 25/47 (53%), Gaps = 5/47 (10%)
Query: 5 NNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMM 51
+N N+QNQ+ P + +P P SS ++ PPQ +G HM+
Sbjct: 86 SNPNHQNQTGPSSSRPMAPFPFPDSSPALGPPPQ-----PTYGFHML 127
>At5g08520 unknown protein
Length = 298
Score = 37.0 bits (84), Expect = 0.012
Identities = 21/55 (38%), Positives = 28/55 (50%), Gaps = 7/55 (12%)
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAV 56
+NNNNNNN N SSP + + + S + P PP +GT +G PAV
Sbjct: 203 SNNNNNNNNNNSSPAVAGGGNKSAKQAVSQAPPGPPM-------YGTPAIGQPAV 250
>At2g23320 putative WRKY-type DNA-binding protein
Length = 317
Score = 37.0 bits (84), Expect = 0.012
Identities = 32/137 (23%), Positives = 56/137 (40%), Gaps = 29/137 (21%)
Query: 12 QSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTA 71
Q P PF SP PP PPQ M+ + SS ++L +
Sbjct: 110 QEEPKTTPFQSPLPP---------PPQ-----------MIRKGSFSSSMKTIDFSSLSSV 149
Query: 72 SADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMESILHMFDSWSKKAEATANNIWHNLKT 131
+ + Q ++H++Q+ P +S +S+ S+SK + N+ NL T
Sbjct: 150 TTESDNQKKIHHHQRPSE-------TAPFASQTQSLSTTVSSFSKSTKRKCNS--ENLLT 200
Query: 132 GPSVSSAAMGKMNLTVK 148
G S+++ G+ + + K
Sbjct: 201 GKCASASSSGRCHCSKK 217
>At3g18810 protein kinase, putative
Length = 700
Score = 35.4 bits (80), Expect = 0.036
Identities = 19/47 (40%), Positives = 27/47 (57%), Gaps = 1/47 (2%)
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPS-SSSSSVPLPPQNHATTENWG 47
NN NNN+N NQ++ + SP PPS +S + P PP+ A + G
Sbjct: 126 NNGNNNDNNNQNNGGGSNNRSPPPPSRNSDRNSPSPPRALAPPRSSG 172
Score = 32.0 bits (71), Expect = 0.39
Identities = 20/61 (32%), Positives = 28/61 (45%), Gaps = 16/61 (26%)
Query: 5 NNNNNQNQSSP----------------DENPFASPTPPSSSSSSVPLPPQNHATTENWGT 48
++++NQ QSSP D++ SP PPSS S S P P N+ N G
Sbjct: 20 SSSDNQQQSSPPPSDSSSPSPPAPPPPDDSSNGSPQPPSSDSQSPPSPQGNNNNDGNNGN 79
Query: 49 H 49
+
Sbjct: 80 N 80
Score = 31.2 bits (69), Expect = 0.67
Identities = 22/69 (31%), Positives = 32/69 (45%), Gaps = 12/69 (17%)
Query: 14 SPDENPFASPTPPSSSSS---SVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQT 70
SP+ +P SPTPPS SSS PP + +++ +P P D+ + Q
Sbjct: 6 SPENSP-PSPTPPSPSSSDNQQQSSPPPSDSSSP--------SPPAPPPPDDSSNGSPQP 56
Query: 71 ASADGQGQP 79
S+D Q P
Sbjct: 57 PSSDSQSPP 65
Score = 28.1 bits (61), Expect = 5.7
Identities = 12/40 (30%), Positives = 21/40 (52%)
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHA 41
NNN N+NN N ++ + N + S++ P PP ++
Sbjct: 115 NNNGNDNNGNNNNGNNNDNNNQNNGGGSNNRSPPPPSRNS 154
>At3g13225 hypothetical protein, 5' partial
Length = 366
Score = 34.3 bits (77), Expect = 0.079
Identities = 22/77 (28%), Positives = 32/77 (40%), Gaps = 9/77 (11%)
Query: 15 PDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAP-AVPSSHPDNKKAALQTASA 73
P E+ P PP S S +P PP N + + +G P VP S+ SA
Sbjct: 23 PSESEDVPPPPPDSYSEPIPPPPDNGHVASSLSSDSLGVPYTVPQSY--------MHQSA 74
Query: 74 DGQGQPQVHYYQQDHPY 90
D Q + Y + ++ Y
Sbjct: 75 DYATQYNLSYPESNYQY 91
>At1g76880 unknown protein
Length = 603
Score = 34.3 bits (77), Expect = 0.079
Identities = 16/33 (48%), Positives = 19/33 (57%)
Query: 2 NNNNNNNNQNQSSPDENPFASPTPPSSSSSSVP 34
NNNNNNNN N S P + P+ SSS+P
Sbjct: 169 NNNNNNNNNNSSIFSTPPPVTTVMPTLPSSSIP 201
Score = 30.8 bits (68), Expect = 0.88
Identities = 22/63 (34%), Positives = 28/63 (43%), Gaps = 16/63 (25%)
Query: 2 NNNNNNNNQNQ--SSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSS 59
NNNNNNNN + S+P PT PSSS +PP T + P+ P+
Sbjct: 170 NNNNNNNNNSSIFSTPPPVTTVMPTLPSSS-----IPPY---------TQQINVPSFPNI 215
Query: 60 HPD 62
D
Sbjct: 216 SGD 218
>At1g32120 hypothetical protein
Length = 1206
Score = 34.3 bits (77), Expect = 0.079
Identities = 30/121 (24%), Positives = 52/121 (42%), Gaps = 10/121 (8%)
Query: 5 NNNNNQNQSSP--DENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPD 62
N+ +N SP + P+ S + SSSS+ PP++H + G + P++ +S
Sbjct: 759 NSRGQKNPPSPRVSKEPYMSSSLSVSSSSTRKKPPRSHEAANSRGYN--HTPSLRASKEP 816
Query: 63 NKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMESILHMFDSWSKKAEATA 122
K ++L +S+ + P+ H Y + P S + L+ S S AT
Sbjct: 817 YKSSSLSGSSSTRKKPPRSHEASSSRGY------NHPPSPRVSKELNKTPSISGSPSATR 870
Query: 123 N 123
N
Sbjct: 871 N 871
>At1g21580 hypothetical protein
Length = 1696
Score = 34.3 bits (77), Expect = 0.079
Identities = 25/92 (27%), Positives = 43/92 (46%), Gaps = 14/92 (15%)
Query: 14 SPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASA 73
+P +P+ SP P S PLPP + P SHP++ ++ +
Sbjct: 7 NPTYDPWNSPYSPHLHPPSAPLPPP---------PPLPPPPPPRQSHPESPNLYGRSTQS 57
Query: 74 DGQGQPQVHYY---QQDHPYVQHSPVDKPSSS 102
+GQ Q +H Y +QD P ++ V++P+S+
Sbjct: 58 NGQRQDYLHQYSHHRQDLP--PNTVVNQPTSN 87
>At3g50650 scarecrow-like 7 (SCL7)
Length = 542
Score = 33.9 bits (76), Expect = 0.10
Identities = 29/147 (19%), Positives = 55/147 (36%), Gaps = 2/147 (1%)
Query: 1 MNNNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSH 60
+N +++ + Q P SP+PPS P P + + + P +
Sbjct: 132 VNKDDSASQQLPPPPASTAIWSPSPPSPQHPPPPPPQPDFDLNQPIFKAIHDYARKPETK 191
Query: 61 PDNKKAALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMESILHMFDSWSKKAEA 120
PD ++ S G +V YY + + H + PSSS S+ S+ +A
Sbjct: 192 PDTLIRIKESVSESGDPIQRVGYYFAE--ALSHKETESPSSSSSSSLEDFILSYKTLNDA 249
Query: 121 TANNIWHNLKTGPSVSSAAMGKMNLTV 147
+ + +L ++ A N+ +
Sbjct: 250 CPYSKFAHLTANQAILEATNQSNNIHI 276
>At5g19340 putative protein
Length = 255
Score = 33.5 bits (75), Expect = 0.14
Identities = 25/76 (32%), Positives = 37/76 (47%), Gaps = 8/76 (10%)
Query: 2 NNNNNNNNQNQSS--PDENPFASPTPPSSS---SSSVPLPPQNHATTENWGTHMMG-APA 55
NNNNNNNN N+ S D++P SP PP + + L Q TT T + +P+
Sbjct: 122 NNNNNNNNNNRGSWFLDDDP--SPRPPKCTVLWKELLRLKKQRTTTTTTASTRVSSLSPS 179
Query: 56 VPSSHPDNKKAALQTA 71
SS + +++ A
Sbjct: 180 SSSSSTSSSSSSIGDA 195
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.312 0.127 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,231,099
Number of Sequences: 26719
Number of extensions: 332704
Number of successful extensions: 2619
Number of sequences better than 10.0: 154
Number of HSP's better than 10.0 without gapping: 64
Number of HSP's successfully gapped in prelim test: 92
Number of HSP's that attempted gapping in prelim test: 2055
Number of HSP's gapped (non-prelim): 421
length of query: 287
length of database: 11,318,596
effective HSP length: 98
effective length of query: 189
effective length of database: 8,700,134
effective search space: 1644325326
effective search space used: 1644325326
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC133780.8


