

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133779.5 + phase: 1 /pseudo
(547 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
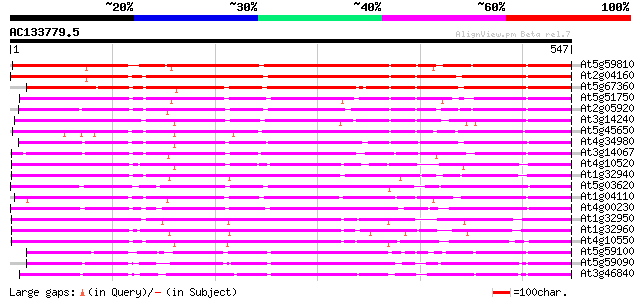
Score E
Sequences producing significant alignments: (bits) Value
At5g59810 subtilisin-like protease - like protein 539 e-153
At2g04160 subtilisin-like serine protease AIR3 522 e-148
At5g67360 cucumisin-like serine protease (gb|AAC18851.1) 412 e-115
At5g51750 serine protease-like protein 395 e-110
At2g05920 serine protease like protein 395 e-110
At3g14240 unknown protein 394 e-110
At5g45650 subtilisin-like protease 393 e-109
At4g34980 subtilisin proteinase - like 389 e-108
At3g14067 subtilisin-like serine proteinase, putative, 3' partial 389 e-108
At4g10520 putative subtilisin-like protease 382 e-106
At1g32940 unknown protein 381 e-106
At5g03620 cucumisin precursor -like protein 378 e-105
At1g04110 putative subtilisin protease 374 e-103
At4g00230 subtilisin-type serine endopeptidase XSP1 373 e-103
At1g32950 putative serine proteinase (At1g32950) 373 e-103
At1g32960 unknown protein 372 e-103
At4g10550 subtilisin-like protease -like protein 366 e-101
At5g59100 cucumisin precursor - like 363 e-100
At5g59090 cucumisin precursor - like 362 e-100
At3g46840 subtilisin-like proteinase 361 e-100
>At5g59810 subtilisin-like protease - like protein
Length = 778
Score = 539 bits (1388), Expect = e-153
Identities = 289/553 (52%), Positives = 372/553 (67%), Gaps = 32/553 (5%)
Query: 3 LGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRS 62
L S +GS ENAKEAI YSY + INGFAA+L+E EAA+IAK+P VVSVF +K KLHTT S
Sbjct: 71 LASFVGSHENAKEAIFYSYKRHINGFAAILDENEAAEIAKHPDVVSVFPNKGRKLHTTHS 130
Query: 63 WEFLGLRGNDI---NSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNI 119
W F+ L N + +S W K +GE+TII N+DTGVWPESKSFSD G G +PA+W+G
Sbjct: 131 WNFMLLAKNGVVHKSSLWNKAGYGEDTIIANLDTGVWPESKSFSDEGYGAVPARWKG--- 187
Query: 120 CQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPR--SQQTARDFVGHGTHTLSTAGG 177
K VPCNRKLIGAR+FNK Y G LP S +T RD GHG+HTLSTA G
Sbjct: 188 ------RCHKDVPCNRKLIGARYFNKGYLAYTG-LPSNASYETCRDHDGHGSHTLSTAAG 240
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDII 237
NFVPGA++F IGNGT GGSP+ARVA YKVCW D CF AD+L+AI+ AI+DGVD++
Sbjct: 241 NFVPGANVFGIGNGTASGGSPKARVAAYKVCWPPVDGAECFDADILAAIEAAIEDGVDVL 300
Query: 238 SVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAA 297
S S GG ++ + +D I+IG+FHA+ + +V SAGN GP G+V NVAPWV TV A
Sbjct: 301 SASVGG----DAGDYMSDGIAIGSFHAVKNGVTVVCSAGNSGPKSGTVSNVAPWVITVGA 356
Query: 298 STLDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTL 356
S++DR+F + + + N ++ G SL LP + ++++++ DA +AN DA C+ +L
Sbjct: 357 SSMDREFQAFVELKNGQSFKGTSLSKPLPEEKMYSLISAADANVANGNVTDALLCKKGSL 416
Query: 357 DPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVL--STI 414
DP KV GKI+ C R G V +G +A +AGA G++L N + +G ++S+ HVL S I
Sbjct: 417 DPKKVKGKILVCLR-GDNARVDKGMQAAAAGAAGMVLCND-KASGNEIISDAHVLPASQI 474
Query: 415 SYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSIL 474
Y G +L S K +P TLN KPAP MAS+SSRGPN + P IL
Sbjct: 475 DY------KDGETLFSYLSSTKDPKGYIKAPTATLN-TKPAPFMASFSSRGPNTITPGIL 527
Query: 475 KPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPN 534
KPD+TAPGVNI+AA++ ++L +D RR PFN GTSMSCPH++G GL+KTLHP+
Sbjct: 528 KPDITAPGVNIIAAFTEATGPTDLDSDNRR-TPFNTESGTSMSCPHISGVVGLLKTLHPH 586
Query: 535 WSPAAIKSAIRTT 547
WSPAAI+SAI TT
Sbjct: 587 WSPAAIRSAIMTT 599
>At2g04160 subtilisin-like serine protease AIR3
Length = 772
Score = 522 bits (1345), Expect = e-148
Identities = 286/552 (51%), Positives = 363/552 (64%), Gaps = 22/552 (3%)
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
D LGS GS+E A +AI YSY K INGFAA L+ + A +I+K+P+VVSVF +K KLHTT
Sbjct: 59 DFLGSFTGSRERATDAIFYSYTKHINGFAAHLDHDLAYEISKHPEVVSVFPNKALKLHTT 118
Query: 61 RSWEFLGLRGNDI---NSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGG 117
RSW+FLGL N +S W+K RFGE+TII N+DTGVWPESKSF D G+GPIP++W+G
Sbjct: 119 RSWDFLGLEHNSYVPSSSIWRKARFGEDTIIANLDTGVWPESKSFRDEGLGPIPSRWKG- 177
Query: 118 NICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGG 177
ICQ K T CNRKLIGAR+FNK Y G L S + RD GHG+HTLSTA G
Sbjct: 178 -ICQNQKDATFH---CNRKLIGARYFNKGYAAAVGHLNSSFDSPRDLDGHGSHTLSTAAG 233
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDII 237
+FVPG SIF GNGT KGGSPRARVA YKVCW C+ ADVL+A D AI DG D+I
Sbjct: 234 DFVPGVSIFGQGNGTAKGGSPRARVAAYKVCWPPVKGNECYDADVLAAFDAAIHDGADVI 293
Query: 238 SVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAA 297
SVS GG ++ F D ++IG+FHA + I++V SAGN GP +V NVAPW TV A
Sbjct: 294 SVSLGGEPTS----FFNDSVAIGSFHAAKKRIVVVCSAGNSGPADSTVSNVAPWQITVGA 349
Query: 298 STLDRDFSSVMTIGN-KTLTGASL-FVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRT 355
ST+DR+F+S + +GN K G SL LP + + I+ S +AK NA+ DA+ C+ +
Sbjct: 350 STMDREFASNLVLGNGKHYKGQSLSSTALPHAKFYPIMASVNAKAKNASALDAQLCKLGS 409
Query: 356 LDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTIS 415
LDP K GKI+ C R G+ V +G+ G G++L N + G LL++PHVL
Sbjct: 410 LDPIKTKGKILVCLR-GQNGRVEKGRAVALGGGIGMVLEN-TYVTGNDLLADPHVLPATQ 467
Query: 416 YPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILK 475
S R + I ++P++T KPAPVMAS+SS+GP+ V P ILK
Sbjct: 468 LTSKDSFAVSRYISQTKKPI-----AHITPSRTDLGLKPAPVMASFSSKGPSIVAPQILK 522
Query: 476 PDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNW 535
PD+TAPGV+++AAY+ S +N D RR FN + GTSMSCPH++G AGL+KT +P+W
Sbjct: 523 PDITAPGVSVIAAYTGAVSPTNEQFDPRR-LLFNAISGTSMSCPHISGIAGLLKTRYPSW 581
Query: 536 SPAAIKSAIRTT 547
SPAAI+SAI TT
Sbjct: 582 SPAAIRSAIMTT 593
>At5g67360 cucumisin-like serine protease (gb|AAC18851.1)
Length = 757
Score = 412 bits (1060), Expect = e-115
Identities = 239/535 (44%), Positives = 333/535 (61%), Gaps = 29/535 (5%)
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
++Y+Y I+GF+ L +EEA + P V+SV ++LHTTR+ FLGL + +
Sbjct: 65 LLYTYENAIHGFSTRLTQEEADSLMTQPGVISVLPEHRYELHTTRTPLFLGLDEHTADLF 124
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
+ G + + ++G +DTGVWPESKS+SD G GPIP+ W+GG C+ T+ CNRK
Sbjct: 125 PEAGSYSD-VVVGVLDTGVWPESKSYSDEGFGPIPSSWKGG--CEAGTNFTASL--CNRK 179
Query: 137 LIGARFFNKAYQKRNGKLPRSQQTA--RDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIK 194
LIGARFF + Y+ G + S+++ RD GHGTHT STA G+ V GAS+ +GT +
Sbjct: 180 LIGARFFARGYESTMGPIDESKESRSPRDDDGHGTHTSSTAAGSVVEGASLLGYASGTAR 239
Query: 195 GGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFT 254
G +PRARVA YKVCW CF +D+L+AID+AI D V+++S+S GG S + +
Sbjct: 240 GMAPRARVAVYKVCW----LGGCFSSDILAAIDKAIADNVNVLSMSLGGGMS----DYYR 291
Query: 255 DEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-K 313
D ++IGAF A+ R IL+ SAGN GP+ S+ NVAPW+ TV A TLDRDF ++ +GN K
Sbjct: 292 DGVAIGAFAAMERGILVSCSAGNAGPSSSSLSNVAPWITTVGAGTLDRDFPALAILGNGK 351
Query: 314 TLTGASLFV-NLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREG 372
TG SLF P++ + + +A +NATN C TL P KV GKIV CDR G
Sbjct: 352 NFTGVSLFKGEALPDKLLPFIYAGNA--SNATN--GNLCMTGTLIPEKVKGKIVMCDR-G 406
Query: 373 KIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 432
V +G +AG G+IL N NG+ L+++ H+L + + P
Sbjct: 407 INARVQKGDVVKAAGGVGMILAN-TAANGEELVADAHLLPATTVGEKAGDIIRHYVTTDP 465
Query: 433 SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF 492
+ S +S T+ KP+PV+A++SSRGPN + P+ILKPD+ APGVNILAA++
Sbjct: 466 NPTAS-----ISILGTVVGVKPSPVVAAFSSRGPNSITPNILKPDLIAPGVNILAAWTGA 520
Query: 493 ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
A + L +D+RR FN++ GTSMSCPHV+G A L+K++HP WSPAAI+SA+ TT
Sbjct: 521 AGPTGLASDSRR-VEFNIISGTSMSCPHVSGLAALLKSVHPEWSPAAIRSALMTT 574
>At5g51750 serine protease-like protein
Length = 780
Score = 395 bits (1014), Expect = e-110
Identities = 231/555 (41%), Positives = 326/555 (58%), Gaps = 51/555 (9%)
Query: 10 KENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLR 69
+E I+Y+Y +G AA L +EEA ++ + VV+V ++LHTTRS FLGL
Sbjct: 72 EEGNNNRILYTYQTAFHGLAAQLTQEEAERLEEEDGVVAVIPETRYELHTTRSPTFLGLE 131
Query: 70 GNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSK 129
+ W + + ++G +DTG+WPES+SF+D G+ P+PA WRG C+ K +
Sbjct: 132 RQESERVWAERVTDHDVVVGVLDTGIWPESESFNDTGMSPVPATWRGA--CETGKRFLKR 189
Query: 130 KVPCNRKLIGARFFNKAYQKRNGKLPR--SQQTARDFVGHGTHTLSTAGGNFVPGASIFN 187
CNRK++GAR F + Y+ GK+ ++ RD GHGTHT +T G+ V GA++F
Sbjct: 190 N--CNRKIVGARVFYRGYEAATGKIDEELEYKSPRDRDGHGTHTAATVAGSPVKGANLFG 247
Query: 188 IGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSST 247
GT +G + +ARVA YKVCW CF +D+LSA+DQA+ DGV ++S+S GG ST
Sbjct: 248 FAYGTARGMAQKARVAAYKVCW----VGGCFSSDILSAVDQAVADGVQVLSISLGGGVST 303
Query: 248 NSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSV 307
S D +SI F A+ + + SAGN GP P S+ NV+PW+ TV AST+DRDF +
Sbjct: 304 YSR----DSLSIATFGAMEMGVFVSCSAGNGGPDPISLTNVSPWITTVGASTMDRDFPAT 359
Query: 308 MTIGN-KTLTGASLFVN---LPPNQDFTIVTSTDAKLANATNRD-ARFCRPRTLDPSKVN 362
+ IG +T G SL+ LP N+ + +V NA++ D FC LD V
Sbjct: 360 VKIGTMRTFKGVSLYKGRTVLPKNKQYPLVYLG----RNASSPDPTSFCLDGALDRRHVA 415
Query: 363 GKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNH-- 420
GKIV CDR G V +GQ AG G++L N NG+ L+++ H+L ++
Sbjct: 416 GKIVICDR-GVTPRVQKGQVVKRAGGIGMVLTN-TATNGEELVADSHMLPAVAVGEKEGK 473
Query: 421 --------SRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPS 472
S+ SL+I+ GT++ + KP+PV+A++SSRGPN +
Sbjct: 474 LIKQYAMTSKKATASLEIL------GTRIGI---------KPSPVVAAFSSRGPNFLSLE 518
Query: 473 ILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLH 532
ILKPD+ APGVNILAA++ + S+L +D RR FN++ GTSMSCPHV+G A LIK+ H
Sbjct: 519 ILKPDLLAPGVNILAAWTGDMAPSSLSSDPRR-VKFNILSGTSMSCPHVSGVAALIKSRH 577
Query: 533 PNWSPAAIKSAIRTT 547
P+WSPAAIKSA+ TT
Sbjct: 578 PDWSPAAIKSALMTT 592
>At2g05920 serine protease like protein
Length = 754
Score = 395 bits (1014), Expect = e-110
Identities = 240/545 (44%), Positives = 324/545 (59%), Gaps = 34/545 (6%)
Query: 9 SKENAKEAIIYSYNKQINGFAAMLEEEEA-AQIAKNPKVVSVFLSKEHKLHTTRSWEFLG 67
S+ N++ +++Y+Y +GF+A L+ EA + ++ + ++ +F + LHTTR+ EFLG
Sbjct: 52 SQLNSESSLLYTYTTSFHGFSAYLDSTEADSLLSSSNSILDIFEDPLYTLHTTRTPEFLG 111
Query: 68 LRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNT 127
L N G IIG +DTGVWPES+SF D + IP+KW+G C+
Sbjct: 112 L--NSEFGVHDLGSSSNGVIIGVLDTGVWPESRSFDDTDMPEIPSKWKGE--CESGSDFD 167
Query: 128 SKKVPCNRKLIGARFFNKAYQKRNG---KLPRSQQTARDFVGHGTHTLSTAGGNFVPGAS 184
SK CN+KLIGAR F+K +Q +G R + RD GHGTHT +TA G+ V AS
Sbjct: 168 SKL--CNKKLIGARSFSKGFQMASGGGFSSKRESVSPRDVDGHGTHTSTTAAGSAVRNAS 225
Query: 185 IFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGP 244
GT +G + RARVATYKVCWS T CFG+D+L+A+D+AI DGVD++S+S GG
Sbjct: 226 FLGYAAGTARGMATRARVATYKVCWS----TGCFGSDILAAMDRAILDGVDVLSLSLGGG 281
Query: 245 SSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDF 304
S+ + D I+IGAF A+ R + + SAGN GPT SV NVAPWV TV A TLDRDF
Sbjct: 282 SAP----YYRDTIAIGAFSAMERGVFVSCSAGNSGPTRASVANVAPWVMTVGAGTLDRDF 337
Query: 305 SSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNG 363
+ +GN K LTG SL+ + + + + C P +LD S V G
Sbjct: 338 PAFANLGNGKRLTGVSLYSGVGMG-----TKPLELVYNKGNSSSSNLCLPGSLDSSIVRG 392
Query: 364 KIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRT 423
KIV CDR G V +G AG G+I+ N +G+ L+++ H+L I+ +
Sbjct: 393 KIVVCDR-GVNARVEKGAVVRDAGGLGMIMAN-TAASGEELVADSHLLPAIAV----GKK 446
Query: 424 TGRSL-DIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPG 482
TG L + + SD K T L + L+ KP+PV+A++SSRGPN V P ILKPDV PG
Sbjct: 447 TGDLLREYVKSDSKP-TALLVFKGTVLD-VKPSPVVAAFSSRGPNTVTPEILKPDVIGPG 504
Query: 483 VNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKS 542
VNILA +S + L D+RR FN+M GTSMSCPH++G AGL+K HP WSP+AIKS
Sbjct: 505 VNILAGWSDAIGPTGLDKDSRR-TQFNIMSGTSMSCPHISGLAGLLKAAHPEWSPSAIKS 563
Query: 543 AIRTT 547
A+ TT
Sbjct: 564 ALMTT 568
>At3g14240 unknown protein
Length = 775
Score = 394 bits (1013), Expect = e-110
Identities = 234/556 (42%), Positives = 328/556 (58%), Gaps = 37/556 (6%)
Query: 5 SILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWE 64
S L S ++ +II++Y+ +GF+A L ++A+Q+ +P V+SV + LHTTRS E
Sbjct: 50 SSLASLTSSPPSIIHTYDTVFHGFSARLTSQDASQLLDHPHVISVIPEQVRHLHTTRSPE 109
Query: 65 FLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDK 124
FLGLR D ++ FG + +IG IDTGVWPE SF DRG+GP+P KW+G I D
Sbjct: 110 FLGLRSTDKAGLLEESDFGSDLVIGVIDTGVWPERPSFDDRGLGPVPIKWKGQCIASQDF 169
Query: 125 LNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQ--TARDFVGHGTHTLSTAGGNFVPG 182
++ CNRKL+GARFF Y+ NGK+ + + + RD GHGTHT S + G +V
Sbjct: 170 PESA----CNRKLVGARFFCGGYEATNGKMNETTEFRSPRDSDGHGTHTASISAGRYVFP 225
Query: 183 ASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAG 242
AS +G G +P+AR+A YKVCW+ + C+ +D+L+A D A+ DGVD+IS+S G
Sbjct: 226 ASTLGYAHGVAAGMAPKARLAAYKVCWN----SGCYDSDILAAFDTAVADGVDVISLSVG 281
Query: 243 GPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDR 302
G + D I+IGAF A+ R I + ASAGN GP +V NVAPW+ TV A T+DR
Sbjct: 282 GV----VVPYYLDAIAIGAFGAIDRGIFVSASAGNGGPGALTVTNVAPWMTTVGAGTIDR 337
Query: 303 DFSSVMTIGN-KTLTGASLF--VNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPS 359
DF + + +GN K ++G S++ L P + + +V L + C +LDP+
Sbjct: 338 DFPANVKLGNGKMISGVSVYGGPGLDPGRMYPLVYG--GSLLGGDGYSSSLCLEGSLDPN 395
Query: 360 KVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGN 419
V GKIV CDR G +G+ G G+I+ N +G+ L+++ HVL S +
Sbjct: 396 LVKGKIVLCDR-GINSRATKGEIVRKNGGLGMIIAN-GVFDGEGLVADCHVLPATSVGAS 453
Query: 420 HSRTTGRSLDIIPSDIKSGTKLRMS--PAKTLNRR------KPAPVMASYSSRGPNKVQP 471
D I I +K R S P T+ + +PAPV+AS+S+RGPN P
Sbjct: 454 GG-------DEIRRYISESSKSRSSKHPTATIVFKGTRLGIRPAPVVASFSARGPNPETP 506
Query: 472 SILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTL 531
ILKPDV APG+NILAA+ S + +D RR FN++ GTSM+CPHV+G A L+K
Sbjct: 507 EILKPDVIAPGLNILAAWPDRIGPSGVTSDNRR-TEFNILSGTSMACPHVSGLAALLKAA 565
Query: 532 HPNWSPAAIKSAIRTT 547
HP+WSPAAI+SA+ TT
Sbjct: 566 HPDWSPAAIRSALITT 581
>At5g45650 subtilisin-like protease
Length = 791
Score = 393 bits (1009), Expect = e-109
Identities = 236/575 (41%), Positives = 332/575 (57%), Gaps = 45/575 (7%)
Query: 3 LGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLS--KEHKLHTT 60
L S+ S+E+A+ +++YSY INGFAA L ++A+++ K +VVSVF S ++++ HTT
Sbjct: 51 LQSVKESEEDARASLLYSYKHSINGFAAELTPDQASKLEKLAEVVSVFKSHPRKYEAHTT 110
Query: 61 RSWEFLGL-----------RGNDINSAWQKGR-------FGENTIIGNIDTGVWPESKSF 102
RSWEF+GL R ND + ++ GR G+ I+G +D+GVWPESKSF
Sbjct: 111 RSWEFVGLEEEETDSDVPRRKNDADDRFRVGRNFLKKAKHGDGIIVGVLDSGVWPESKSF 170
Query: 103 SDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQ--- 159
+D+G+GP+P W+G ICQ S CNRK+IGAR++ K Y++ G +
Sbjct: 171 NDKGMGPVPKSWKG--ICQTGVAFNSSH--CNRKIIGARYYVKGYERYYGAFNATANKDF 226
Query: 160 -TARDFVGHGTHTLSTAGGNFVPGASIFN-IGNGTIKGGSPRARVATYKVCWSLTDATS- 216
+ RD GHG+HT STA G V GAS G+ GG+P AR+A YK CW+ +A
Sbjct: 227 LSPRDPDGHGSHTASTAVGRRVLGASALGGFAKGSASGGAPLARLAIYKACWAKPNAEKV 286
Query: 217 ----CFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLV 272
C D+L+AID AI DGV +IS+S G +T D I++GA HA+ RNI++
Sbjct: 287 EGNICLEEDMLAAIDDAIADGVHVISISIG---TTEPFPFTQDGIAMGALHAVKRNIVVA 343
Query: 273 ASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGNKTLTGASLFVNLPPNQDFTI 332
ASAGN GP PG++ N+APW+ TV ASTLDR F + +GN ++ +
Sbjct: 344 ASAGNSGPKPGTLSNLAPWIITVGASTLDRAFVGGLVLGNGYTIKTDSITAFKMDKFAPL 403
Query: 333 VTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVI 392
V +++ + + C P +L P V+GK+V C R G + +G E AG G+I
Sbjct: 404 VYASNVVVPGIALNETSQCLPNSLKPELVSGKVVLCLR-GAGSRIGKGMEVKRAGGAGMI 462
Query: 393 LRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRR 452
L N NG + S+ H + T G + L+ I +D K + P KT+ +
Sbjct: 463 LGN-IAANGNEVPSDSHFVPT---AGVTPTVVDKILEYIKTD--KNPKAFIKPGKTVYKY 516
Query: 453 KPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQ 512
+ AP M +SSRGPN V P+ILKPD+TAPG+ ILAA+S S S + D +R +N+
Sbjct: 517 QAAPSMTGFSSRGPNVVDPNILKPDITAPGLYILAAWSGADSPSKMSVD-QRVAGYNIYS 575
Query: 513 GTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
GTSMSCPHVAG L+K +HP WS AAI+SA+ TT
Sbjct: 576 GTSMSCPHVAGAIALLKAIHPKWSSAAIRSALMTT 610
>At4g34980 subtilisin proteinase - like
Length = 764
Score = 389 bits (999), Expect = e-108
Identities = 228/545 (41%), Positives = 328/545 (59%), Gaps = 29/545 (5%)
Query: 9 SKENAKEA-IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLG 67
S E A+E+ I++ Y+ +GF+A++ +EA + +P V++VF + +LHTTRS +FLG
Sbjct: 49 STEFAEESRIVHVYHTVFHGFSAVVTPDEADNLRNHPAVLAVFEDRRRELHTTRSPQFLG 108
Query: 68 LRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNT 127
L+ W + +G + IIG DTG+WPE +SFSD +GPIP +WRG +C+ +
Sbjct: 109 LQNQ--KGLWSESDYGSDVIIGVFDTGIWPERRSFSDLNLGPIPKRWRG--VCESGARFS 164
Query: 128 SKKVPCNRKLIGARFFNKAYQKRN-GKLPRSQQ--TARDFVGHGTHTLSTAGGNFVPGAS 184
+ CNRK+IGARFF K Q G + ++ + + RD GHGTHT STA G AS
Sbjct: 165 PRN--CNRKIIGARFFAKGQQAAVIGGINKTVEFLSPRDADGHGTHTSSTAAGRHAFKAS 222
Query: 185 IFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGP 244
+ +G KG +P+AR+A YKVCW + C +D+L+A D A+ DGVD+IS+S GG
Sbjct: 223 MSGYASGVAKGVAPKARIAAYKVCWK---DSGCLDSDILAAFDAAVRDGVDVISISIGGG 279
Query: 245 SSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDF 304
S + D I+IG++ A ++ I + +SAGNEGP SV N+APWV TV AST+DR+F
Sbjct: 280 DGITSP-YYLDPIAIGSYGAASKGIFVSSSAGNEGPNGMSVTNLAPWVTTVGASTIDRNF 338
Query: 305 SSVMTIGN-KTLTGASLFVNLPPN-QDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVN 362
+ +G+ L G SL+ +P N + F +V + +++A+ C TLDP +V
Sbjct: 339 PADAILGDGHRLRGVSLYAGVPLNGRMFPVVYPGKSGMSSAS-----LCMENTLDPKQVR 393
Query: 363 GKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSR 422
GKIV CDR G VA+G AG G+IL N NG+ L+ + H++ + N
Sbjct: 394 GKIVICDR-GSSPRVAKGLVVKKAGGVGMILANGAS-NGEGLVGDAHLIPACAVGSNEGD 451
Query: 423 TTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPG 482
P+ I S + T+ KPAPV+AS+S RGPN + P ILKPD+ APG
Sbjct: 452 RIKAYASSHPNPIAS-----IDFRGTIVGIKPAPVIASFSGRGPNGLSPEILKPDLIAPG 506
Query: 483 VNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKS 542
VNILAA++ + L +D R+ FN++ GTSM+CPHV+G A L+K+ HP+WSPA I+S
Sbjct: 507 VNILAAWTDAVGPTGLPSDPRKT-EFNILSGTSMACPHVSGAAALLKSAHPDWSPAVIRS 565
Query: 543 AIRTT 547
A+ TT
Sbjct: 566 AMMTT 570
>At3g14067 subtilisin-like serine proteinase, putative, 3' partial
Length = 743
Score = 389 bits (998), Expect = e-108
Identities = 229/556 (41%), Positives = 331/556 (59%), Gaps = 43/556 (7%)
Query: 2 LLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTR 61
LL S+ S + A ++YSY++ ++GF+A L + A + ++P V+SV + ++HTT
Sbjct: 56 LLRSLPSSPQPA--TLLYSYSRAVHGFSARLSPIQTAALRRHPSVISVIPDQAREIHTTH 113
Query: 62 SWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQ 121
+ FLG N + W +GE+ I+G +DTG+WPE SFSD G+GPIP+ W+G C+
Sbjct: 114 TPAFLGFSQN--SGLWSNSNYGEDVIVGVLDTGIWPEHPSFSDSGLGPIPSTWKGE--CE 169
Query: 122 LDKLNTSKKVPCNRKLIGARFFNKAY-QKRNGK---LPRSQQTARDFVGHGTHTLSTAGG 177
+ + CNRKLIGAR F + Y +RNG + ++ RD GHGTHT STA G
Sbjct: 170 IGPDFPASS--CNRKLIGARAFYRGYLTQRNGTKKHAAKESRSPRDTEGHGTHTASTAAG 227
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDII 237
+ V AS++ GT G + +AR+A YK+CW+ C+ +D+L+A+DQA+ DGV +I
Sbjct: 228 SVVANASLYQYARGTATGMASKARIAAYKICWT----GGCYDSDILAAMDQAVADGVHVI 283
Query: 238 SVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAA 297
S+S G +S ++ E TD I+IGAF A I++ SAGN GP P + N+APW+ TV A
Sbjct: 284 SLSVG--ASGSAPEYHTDSIAIGAFGATRHGIVVSCSAGNSGPNPETATNIAPWILTVGA 341
Query: 298 STLDRDF-SSVMTIGNKTLTGASLFVNLP-PNQDFTIVTSTDAKLANATNRDARFCRPRT 355
ST+DR+F ++ +T K TG SL+ P+ ++V S D +R C P
Sbjct: 342 STVDREFAANAITGDGKVFTGTSLYAGESLPDSQLSLVYSGDC--------GSRLCYPGK 393
Query: 356 LDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTI- 414
L+ S V GKIV CDR G + V +G AG G+IL N E +G+ L ++ H++
Sbjct: 394 LNSSLVEGKIVLCDRGGNAR-VEKGSAVKLAGGAGMILANTAE-SGEELTADSHLVPATM 451
Query: 415 --SYPGNHSRTTGRSLDIIPSDIK-SGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQP 471
+ G+ R ++ D + I GT + SP P+P +A++SSRGPN + P
Sbjct: 452 VGAKAGDQIRDYIKTSDSPTAKISFLGTLIGPSP--------PSPRVAAFSSRGPNHLTP 503
Query: 472 SILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTL 531
ILKPDV APGVNILA ++ ++L D RR FN++ GTSMSCPHV+G A L++
Sbjct: 504 VILKPDVIAPGVNILAGWTGMVGPTDLDIDPRR-VQFNIISGTSMSCPHVSGLAALLRKA 562
Query: 532 HPNWSPAAIKSAIRTT 547
HP+WSPAAIKSA+ TT
Sbjct: 563 HPDWSPAAIKSALVTT 578
>At4g10520 putative subtilisin-like protease
Length = 756
Score = 382 bits (981), Expect = e-106
Identities = 230/564 (40%), Positives = 328/564 (57%), Gaps = 67/564 (11%)
Query: 2 LLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTR 61
+L S+LGSKE ++I+YSY +GFAA L E +A QI++ P+VV V + +++ TTR
Sbjct: 52 MLWSLLGSKEAVLDSIVYSYRHGFSGFAAKLTESQAQQISELPEVVQVIPNTLYEMTTTR 111
Query: 62 SWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQ 121
+W++LG+ + +S QK G N I+G ID+GVWPES+ F+D+G GPIP++W+GG C+
Sbjct: 112 TWDYLGVSPGNSDSLLQKANMGYNVIVGVIDSGVWPESEMFNDKGFGPIPSRWKGG--CE 169
Query: 122 LDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQ----TARDFVGHGTHTLSTAGG 177
+L + + CNRKLIGA++F G + R+Q + RDF GHGTH ST GG
Sbjct: 170 SGEL-FNASIHCNRKLIGAKYFVDGLVAEFGVVNRTQNPEYLSPRDFAGHGTHVASTIGG 228
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDII 237
+F+P S +G GT +GG+P +A YK CWS C GADVL A+D+AI DGVDI+
Sbjct: 229 SFLPNVSYVGLGRGTARGGAPGVHIAVYKACWS----GYCSGADVLKAMDEAIHDGVDIL 284
Query: 238 SVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAA 297
S+S GPS E T+ S+GAFHA+A+ I +V +AGN GPT ++ NVAPWV TVAA
Sbjct: 285 SLSL-GPSVPLFPE--TEHTSVGAFHAVAKGIPVVIAAGNAGPTAQTISNVAPWVLTVAA 341
Query: 298 STLDRDFSSVMTIGNK-TLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTL 356
+T DR F + +T+GN T+ G +++ P F +T ++ L+ C +
Sbjct: 342 TTQDRSFPTAITLGNNITILGQAIYGG--PELGFVGLTYPESPLSGD-------CEKLSA 392
Query: 357 DP-SKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTIS 415
+P S + GK+V C S A ++AG G+I+ P
Sbjct: 393 NPNSTMEGKVVLC-FAASTPSNAAIAAVINAGGLGLIMAKNP------------------ 433
Query: 416 YPGNHSRTTGRSLDIIPSDIKSGTKL------------RMSPAKTLNRRKPAPVMASYSS 463
HS T R + D + GT + ++ +KTL + + +A++SS
Sbjct: 434 ---THSLTPTRKFPWVSIDFELGTDILFYIRSTRSPIVKIQASKTLFGQSVSTKVATFSS 490
Query: 464 RGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAG 523
RGPN V P+ILKPD+ APGVNILAA S +S ++ F +M GTSM+ P V+G
Sbjct: 491 RGPNSVSPAILKPDIAAPGVNILAAISPNSSIND--------GGFAMMSGTSMATPVVSG 542
Query: 524 TAGLIKTLHPNWSPAAIKSAIRTT 547
L+K+LHP+WSP+AIKSAI TT
Sbjct: 543 VVVLLKSLHPDWSPSAIKSAIVTT 566
>At1g32940 unknown protein
Length = 774
Score = 381 bits (978), Expect = e-106
Identities = 221/557 (39%), Positives = 325/557 (57%), Gaps = 35/557 (6%)
Query: 2 LLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTR 61
+L S+LGSK +A E+++YSY +GFAA L E +A ++A +P+VV V ++L TTR
Sbjct: 52 MLSSLLGSKVDAHESMVYSYRHGFSGFAAKLTESQAKKLADSPEVVHVMADSFYELATTR 111
Query: 62 SWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQ 121
+W++LGL + N+ G+ IIG IDTGVWPES+SF+D G+GPIP+ W+GG C+
Sbjct: 112 TWDYLGLSVANPNNLLNDTNMGDQVIIGFIDTGVWPESESFNDNGVGPIPSHWKGG--CE 169
Query: 122 LDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKL----PRSQQTARDFVGHGTHTLSTAGG 177
+ S CNRKLIGA++F + N R +ARDF+GHGTHT S AGG
Sbjct: 170 SGEKFISTN--CNRKLIGAKYFINGFLAENEGFNTTESRDYISARDFIGHGTHTASIAGG 227
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTD--ATSCFGADVLSAIDQAIDDGVD 235
+FVP S + G ++GG+PRAR+A YK CW + A +C +D+L A+D+++ DGVD
Sbjct: 228 SFVPNISYKGLAGGNLRGGAPRARIAIYKACWYVDQLGAVACSSSDILKAMDESMHDGVD 287
Query: 236 IISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTV 295
++S+S G E D I+ GAFHA+A+ I++V + GN GP +V+N APW+ TV
Sbjct: 288 VLSLSLGAQIPLYPETDLRDRIATGAFHAVAKGIIVVCAGGNSGPAAQTVLNTAPWIITV 347
Query: 296 AASTLDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRD-ARFCRP 353
AA+TLDR F + +T+GN K + G +L+ Q+ + + A TN + C
Sbjct: 348 AATTLDRSFPTPITLGNRKVILGQALYT----GQELGFTSLVYPENAGFTNETFSGVCER 403
Query: 354 RTLDPSK-VNGKIVACDREGKIKSVAE--GQEALSAGAKGVILRNQPEINGKTLLSEPHV 410
L+P++ + GK+V C + + +AG GVI+ P N T +
Sbjct: 404 LNLNPNRTMAGKVVLCFTTNTLFTAVSRAASYVKAAGGLGVIIARNPGYN-LTPCRDDFP 462
Query: 411 LSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQ 470
I Y G + + +S +++ P++TL + +A++SSRGPN +
Sbjct: 463 CVAIDY------ELGTDVLLYIRSTRSPV-VKIQPSRTLVGQPVGTKVATFSSRGPNSIS 515
Query: 471 PSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKT 530
P+ILKPD+ APGV+ILAA S +++S F+++ GTSM+ P VAG L+K
Sbjct: 516 PAILKPDIGAPGVSILAATSPDSNSS--------VGGFDILAGTSMAAPVVAGVVALLKA 567
Query: 531 LHPNWSPAAIKSAIRTT 547
LHPNWSPAA +SAI TT
Sbjct: 568 LHPNWSPAAFRSAIVTT 584
>At5g03620 cucumisin precursor -like protein
Length = 766
Score = 378 bits (971), Expect = e-105
Identities = 232/553 (41%), Positives = 317/553 (56%), Gaps = 39/553 (7%)
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
+LL +++G + A+E IYSY K INGF A L EA ++++ VVSVF + + +LHTT
Sbjct: 56 NLLMTVIGDESKARELKIYSYGKNINGFVARLFPHEAEKLSREEGVVSVFKNTQRQLHTT 115
Query: 61 RSWEFLGLRGNDINSAWQKG-RFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNI 119
RSW+FLGL + S +++ N I+G +DTG+ ES SF+D+G+GP PAKW+G +
Sbjct: 116 RSWDFLGL----VESKYKRSVGIESNIIVGVLDTGIDVESPSFNDKGVGPPPAKWKGKCV 171
Query: 120 CQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNF 179
+ CN K+IGA++F + + G TA D GHGTHT ST G
Sbjct: 172 ------TGNNFTRCNNKVIGAKYF---HIQSEGLPDGEGDTAADHDGHGTHTSSTIAGVS 222
Query: 180 VPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISV 239
V AS+F I NGT +GG P AR+A YKVCW + C D+L+A D+AI DGVDIIS+
Sbjct: 223 VSSASLFGIANGTARGGVPSARIAAYKVCWD----SGCTDMDMLAAFDEAISDGVDIISI 278
Query: 240 SAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAAST 299
S GG S F D I+IGAFHA+ R IL SAGN GP +V N+APWV TVAA++
Sbjct: 279 SIGGASLP----FFEDPIAIGAFHAMKRGILTTCSAGNNGPGLFTVSNLAPWVMTVAANS 334
Query: 300 LDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIVT-STDAKLANATNRDARFCRPRTLD 357
LDR F +V+ +GN T +G SL P + + + + S + L+ + C P TL
Sbjct: 335 LDRKFETVVKLGNGLTASGISLNGFNPRKKMYPLTSGSLASNLSAGGYGEPSTCEPGTLG 394
Query: 358 PSKVNGKIVACD---REGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTI 414
KV GK+V C+ EG + S GVI++ L EP ++T
Sbjct: 395 EDKVMGKVVYCEAGREEGGNGGQGQDHVVRSLKGAGVIVQ----------LLEPTDMATS 444
Query: 415 SYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSIL 474
+ S I I S + KT + AP ++S+S+RGP ++ P+IL
Sbjct: 445 TLIAG-SYVFFEDGTKITEYINSTKNPQAVIFKTKTTKMLAPSISSFSARGPQRISPNIL 503
Query: 475 KPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPN 534
KPD++APG+NILAAYS AS + D RR F++M GTSM+CPH A A +K+ HP+
Sbjct: 504 KPDISAPGLNILAAYSKLASVTGYPDDNRRTL-FSIMSGTSMACPHAAAAAAYVKSFHPD 562
Query: 535 WSPAAIKSAIRTT 547
WSPAAIKSA+ TT
Sbjct: 563 WSPAAIKSALMTT 575
>At1g04110 putative subtilisin protease
Length = 775
Score = 374 bits (959), Expect = e-103
Identities = 230/554 (41%), Positives = 316/554 (56%), Gaps = 36/554 (6%)
Query: 5 SILGSKENAKEA---IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTR 61
++LG +E +E ++YSY I GFAA L E EA + +P+VV+V ++ TT
Sbjct: 56 AVLGVEEEEEEPSSRLLYSYGSAIEGFAAQLTESEAEILRYSPEVVAVRPDHVLQVQTTY 115
Query: 62 SWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQ 121
S++FLGL G + W K RFG+ TIIG +DTGVWPES SF D G+ IP KW+G ICQ
Sbjct: 116 SYKFLGLDGFGNSGVWSKSRFGQGTIIGVLDTGVWPESPSFDDTGMPSIPRKWKG--ICQ 173
Query: 122 LDKLNTSKKVPCNRKLIGARFFNKAYQKRNG-----KLPRSQQTARDFVGHGTHTLSTAG 176
+ +S CNRKLIGARFF + ++ N +PR +ARD GHGTHT ST G
Sbjct: 174 EGESFSSSS--CNRKLIGARFFIRGHRVANSPEESPNMPREYISARDSTGHGTHTASTVG 231
Query: 177 GNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDI 236
G+ V A++ G G +G +P A +A YKVCW C+ +D+L+AID AI D VD+
Sbjct: 232 GSSVSMANVLGNGAGVARGMAPGAHIAVYKVCW----FNGCYSSDILAAIDVAIQDKVDV 287
Query: 237 ISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVA 296
+S+S GG ++ D I+IG F A+ R I ++ +AGN GP SV N APWV T+
Sbjct: 288 LSLSLGG----FPIPLYDDTIAIGTFRAMERGISVICAAGNNGPIESSVANTAPWVSTIG 343
Query: 297 ASTLDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRT 355
A TLDR F +V+ + N K L G SL+ P + ++ + FC +
Sbjct: 344 AGTLDRRFPAVVRLANGKLLYGESLY---PGKGIKNAGREVEVIYVTGGDKGSEFCLRGS 400
Query: 356 LDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVL--ST 413
L ++ GK+V CDR +S +G+ AG +IL N EIN + + H+L +
Sbjct: 401 LPREEIRGKMVICDRGVNGRS-EKGEAVKEAGGVAMILAN-TEINQEEDSIDVHLLPATL 458
Query: 414 ISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSI 473
I Y + + + P K R+ T+ R AP +A +S+RGP+ PSI
Sbjct: 459 IGYTESVLLKAYVNATVKP-------KARIIFGGTVIGRSRAPEVAQFSARGPSLANPSI 511
Query: 474 LKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHP 533
LKPD+ APGVNI+AA+ + L D+RR F VM GTSMSCPHV+G LI++ +P
Sbjct: 512 LKPDMIAPGVNIIAAWPQNLGPTGLPYDSRR-VNFTVMSGTSMSCPHVSGITALIRSAYP 570
Query: 534 NWSPAAIKSAIRTT 547
NWSPAAIKSA+ TT
Sbjct: 571 NWSPAAIKSALMTT 584
>At4g00230 subtilisin-type serine endopeptidase XSP1
Length = 749
Score = 373 bits (957), Expect = e-103
Identities = 227/550 (41%), Positives = 320/550 (57%), Gaps = 42/550 (7%)
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
+LL S+ S+E AKE +YSY K N FAA L EA ++ + +VVSV ++ KLHTT
Sbjct: 58 NLLSSLNISQEEAKERKVYSYTKAFNAFAAKLSPHEAKKMMEMEEVVSVSRNQYRKLHTT 117
Query: 61 RSWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNIC 120
+SW+F+GL +A + + + IIG +DTG+ P+S+SF D G+GP PAKW+G C
Sbjct: 118 KSWDFVGLP----LTAKRHLKAERDVIIGVLDTGITPDSESFLDHGLGPPPAKWKGS--C 171
Query: 121 QLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQ-QTARDFVGHGTHTLSTAGGNF 179
K T CN K+IGA++F K +G +P + ++ D GHGTHT ST G
Sbjct: 172 GPYKNFTG----CNNKIIGAKYF-----KHDGNVPAGEVRSPIDIDGHGTHTSSTVAGVL 222
Query: 180 VPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISV 239
V AS++ I NGT +G P AR+A YKVCW+ + C D+L+ + AI DGV+IIS+
Sbjct: 223 VANASLYGIANGTARGAVPSARLAMYKVCWA---RSGCADMDILAGFEAAIHDGVEIISI 279
Query: 240 SAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAAST 299
S GGP + S +D IS+G+FHA+ + IL VASAGN+GP+ G+V N PW+ TVAAS
Sbjct: 280 SIGGPIADYS----SDSISVGSFHAMRKGILTVASAGNDGPSSGTVTNHEPWILTVAASG 335
Query: 300 LDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDP 358
+DR F S + +GN K+ +G + + P + + +V+ DA AR+C +LD
Sbjct: 336 IDRTFKSKIDLGNGKSFSGMGISMFSPKAKSYPLVSGVDAAKNTDDKYLARYCFSDSLDR 395
Query: 359 SKVNGKIVACDR-EGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYP 417
KV GK++ C G ++S + S G G I+ + ++ + P S S
Sbjct: 396 KKVKGKVMVCRMGGGGVESTIK-----SYGGAGAIIVSDQYLDNAQIFMAP-ATSVNSSV 449
Query: 418 GNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPD 477
G DII I S KT PAP +AS+SSRGPN +LKPD
Sbjct: 450 G----------DIIYRYINSTRSASAVIQKTRQVTIPAPFVASFSSRGPNPGSIRLLKPD 499
Query: 478 VTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSP 537
+ APG++ILAA++L S + L DT+ F ++ GTSM+CPHVAG A +K+ HP+W+P
Sbjct: 500 IAAPGIDILAAFTLKRSLTGLDGDTQFS-KFTILSGTSMACPHVAGVAAYVKSFHPDWTP 558
Query: 538 AAIKSAIRTT 547
AAIKSAI T+
Sbjct: 559 AAIKSAIITS 568
>At1g32950 putative serine proteinase (At1g32950)
Length = 763
Score = 373 bits (957), Expect = e-103
Identities = 216/558 (38%), Positives = 322/558 (56%), Gaps = 48/558 (8%)
Query: 2 LLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTR 61
+L S+LGSK++A E+++YSY +GFAA L + +A +IA +P+V+ V ++L TTR
Sbjct: 52 MLSSLLGSKDDAHESMVYSYRHGFSGFAAKLTKSQAKKIADSPEVIHVIPDSYYELATTR 111
Query: 62 SWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQ 121
W++LG ++ + G+ TIIG IDTGVWPES+SF+D G+GP+P+ W+GG
Sbjct: 112 IWDYLGPSADNSKNLVSDTNMGDQTIIGVIDTGVWPESESFNDYGVGPVPSHWKGGCEPG 171
Query: 122 LDKLNTSKKVPCNRKLIGARFFNKAY---QKRNGKLPRSQQTARDFVGHGTHTLSTAGGN 178
+ ++T+ CNRKLIGA++F + + N +ARDF GHGTH S AGG+
Sbjct: 172 ENFISTN----CNRKLIGAKYFINGFLAENQFNATESPDYISARDFDGHGTHVASIAGGS 227
Query: 179 FVPGASIFNIGNGTIKGGSPRARVATYKVCWSLT--DATSCFGADVLSAIDQAIDDGVDI 236
FVP S +G GT++GG+PRAR+A YK CW + D +C +D++ AID+AI DGVD+
Sbjct: 228 FVPNVSYKGLGRGTLRGGAPRARIAMYKACWYINELDGVTCSFSDIMKAIDEAIHDGVDV 287
Query: 237 ISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVA 296
+S+S GG NSE D I+ GAFHA+A+ I++V + GN GP+ +VVN APW+ TVA
Sbjct: 288 LSISLGGRVPLNSETDLRDGIATGAFHAVAKGIVVVCAGGNAGPSSQTVVNTAPWILTVA 347
Query: 297 ASTLDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRT 355
A+TLDR F++ + +GN + + G ++++ P FT + + N+ + + C
Sbjct: 348 ATTLDRSFATPIILGNNQVILGQAMYIG--PELGFTSLVYPEDP-GNSIDTFSGVCESLN 404
Query: 356 LDPSK-VNGKIVAC---DREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVL 411
L+ ++ + GK+V C R+ + S A + G +I RN
Sbjct: 405 LNSNRTMAGKVVLCFTTARDFTVVSTAASIVKAAGGLGLIIARN---------------- 448
Query: 412 STISYPGNHSRTTGRSLDIIPSDIKSGTKLR--MSPAKTLNRRKPAPVMASYSSRGPNKV 469
PG + + D + GT + + TL +A++SSRGPN +
Sbjct: 449 -----PGYNLAPCSDDFPCVAIDNELGTDILFYIRYTGTLVGEPVGTKVATFSSRGPNSI 503
Query: 470 QPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIK 529
P+ILKPD+ APGV+ILAA S DT F + GTSM+ P ++G L+K
Sbjct: 504 SPAILKPDIAAPGVSILAATSP--------NDTLNAGGFVMRSGTSMAAPVISGVIALLK 555
Query: 530 TLHPNWSPAAIKSAIRTT 547
+LHP+WSPAA +SAI TT
Sbjct: 556 SLHPDWSPAAFRSAIVTT 573
>At1g32960 unknown protein
Length = 777
Score = 372 bits (954), Expect = e-103
Identities = 219/565 (38%), Positives = 318/565 (55%), Gaps = 51/565 (9%)
Query: 2 LLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTR 61
+L S+LGSK++A ++++YSY +GFAA L + +A +IA P+VV V H+L TTR
Sbjct: 55 MLASLLGSKKDADDSMVYSYRHGFSGFAAKLTKSQAKKIADLPEVVHVIPDGFHELATTR 114
Query: 62 SWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQ 121
+WE+LGL + + G+ IIG IDTGVWPES+SF+D G+GPIP KW+GG C+
Sbjct: 115 TWEYLGLSSANPKNLLNDTNMGDQVIIGVIDTGVWPESESFNDNGVGPIPRKWKGG--CE 172
Query: 122 LDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKL----PRSQQTARDFVGHGTHTLSTAGG 177
+ + CNRKLIGA++F + N R +ARDF GHGTH S AGG
Sbjct: 173 SGE--NFRSTDCNRKLIGAKYFINGFLAENKGFNTTESRDYISARDFDGHGTHVASIAGG 230
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTD--ATSCFGADVLSAIDQAIDDGVD 235
+FVP S + GT++GG+PRAR+A YK CW + +C +D++ AID+AI DGVD
Sbjct: 231 SFVPNVSYKGLAGGTLRGGAPRARIAMYKACWFHEELKGVTCSDSDIMKAIDEAIHDGVD 290
Query: 236 IISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTV 295
++S+S G NSE DE + G FHA+A+ I++V + GN+GP +VVN+APW+ TV
Sbjct: 291 VLSISLVGQIPLNSETDIRDEFATGLFHAVAKGIVVVCAGGNDGPAAQTVVNIAPWILTV 350
Query: 296 AASTLDRDFSSVMTIG-NKTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARF---C 351
AA+TLDR F + +T+G NK + G + + P T + + NA N + F C
Sbjct: 351 AATTLDRSFPTPITLGNNKVILGQATYTG--PELGLTSLVYPE----NARNNNETFSGVC 404
Query: 352 RPRTLDPSKVNG-KIVACDREGKIKSVAEGQEAL--SAGAKGVILRNQPEINGKTLLSEP 408
L+P+ K+V C + + + +AG G+I+ P + + ++
Sbjct: 405 ESLNLNPNYTMAMKVVLCFTASRTNAAISRAASFVKAAGGLGLIISRNP-VYTLSPCNDD 463
Query: 409 HVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSP------AKTLNRRKPAPVMASYS 462
+ Y + +DI S + SP ++TL+ + + ++S
Sbjct: 464 FPCVAVDYE-------------LGTDILSYIRSTRSPVVKIQRSRTLSGQPVGTKVVNFS 510
Query: 463 SRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVA 522
SRGPN + P+ILKPD+ APGV ILAA S DT F ++ GTSM+ P ++
Sbjct: 511 SRGPNSMSPAILKPDIAAPGVRILAATS--------PNDTLNVGGFAMLSGTSMATPVIS 562
Query: 523 GTAGLIKTLHPNWSPAAIKSAIRTT 547
G L+K LHP WSPAA +SAI TT
Sbjct: 563 GVIALLKALHPEWSPAAFRSAIVTT 587
>At4g10550 subtilisin-like protease -like protein
Length = 778
Score = 366 bits (939), Expect = e-101
Identities = 219/560 (39%), Positives = 320/560 (57%), Gaps = 42/560 (7%)
Query: 2 LLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTR 61
+L S+LGSKE+A ++++YSY +GFAA L E +A +IA P VV V +KL TTR
Sbjct: 57 MLWSLLGSKEDANDSMVYSYRHGFSGFAAKLTESQAKKIADLPDVVHVIPDSFYKLATTR 116
Query: 62 SWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQ 121
+W++LGL + S + GE IIG IDTGVWPES+ F+D G GP+P+ W+GG C+
Sbjct: 117 TWDYLGLSAANPKSLLHETNMGEQIIIGVIDTGVWPESEVFNDSGFGPVPSHWKGG--CE 174
Query: 122 L-DKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQ----TARDFVGHGTHTLSTAG 176
+ N+S CN+KLIGA++F + N + + RD GHGTH + AG
Sbjct: 175 TGENFNSSN---CNKKLIGAKYFINGFLAENESFNSTNSLDFISPRDLDGHGTHVSTIAG 231
Query: 177 GNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSL--TDATSCFGADVLSAIDQAIDDGV 234
G+FVP S + GT++GG+PRA +A YK CW L D T+C AD+L A+D+A+ DGV
Sbjct: 232 GSFVPNISYKGLAGGTVRGGAPRAHIAMYKACWYLDDDDTTTCSSADILKAMDEAMHDGV 291
Query: 235 DIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFT 294
D++S+S G E D I+ GAFHA+ + I +V S GN GP +V N APW+ T
Sbjct: 292 DVLSISLGSSVPLYGETDIRDGITTGAFHAVLKGITVVCSGGNSGPDSLTVTNTAPWIIT 351
Query: 295 VAASTLDRDFSSVMTIG-NKTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRP 353
VAA+TLDR F++ +T+G NK + G +++ P FT + + N+ + C
Sbjct: 352 VAATTLDRSFATPLTLGNNKVILGQAMYTG--PGLGFTSLVYPE-NPGNSNESFSGTCEE 408
Query: 354 RTLDPSK-VNGKIVAC----DREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEP 408
+ ++ + GK+V C G + S A + AG GVI+ P + L +
Sbjct: 409 LLFNSNRTMEGKVVLCFTTSPYGGAVLSAA--RYVKRAGGLGVIIARHPGYAIQPCLDDF 466
Query: 409 HVLSTISYPGNHSRTTGRSLDIIPSDIKSGTK-LRMSPAKTLNRRKPAPVMASYSSRGPN 467
++ G DI+ SG+ +++ P+KTL + +A++SSRGPN
Sbjct: 467 PCVAVDWELGT---------DILLYTRSSGSPVVKIQPSKTLVGQPVGTKVATFSSRGPN 517
Query: 468 KVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGL 527
+ P+ILKPD+ APGV+ILA A+ + +D F ++ GTSM+ P ++G A L
Sbjct: 518 SIAPAILKPDIAAPGVSILA-----ATTNTTFSDQ----GFIMLSGTSMAAPAISGVAAL 568
Query: 528 IKTLHPNWSPAAIKSAIRTT 547
+K LH +WSPAAI+SAI TT
Sbjct: 569 LKALHRDWSPAAIRSAIVTT 588
>At5g59100 cucumisin precursor - like
Length = 741
Score = 363 bits (933), Expect = e-100
Identities = 229/532 (43%), Positives = 307/532 (57%), Gaps = 43/532 (8%)
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
++ SY K NGFAA L E E ++A +VVSVF S++ KL TT SW F+GL+
Sbjct: 71 LVRSYKKSFNGFAARLTESERKRLAGMERVVSVFPSRKLKLQTTSSWNFMGLKEGIKTKR 130
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
+ +TIIG ID+G++PES SFSD+G GP P KW+G K CN K
Sbjct: 131 TRS--IESDTIIGVIDSGIYPESDSFSDQGFGPPPKKWKG-------TCAGGKNFTCNNK 181
Query: 137 LIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGG 196
+IGAR + A K N QTARD+ GHGTHT S A GN V ++ + +GNGT +GG
Sbjct: 182 VIGARDYT-AKSKAN-------QTARDYSGHGTHTASIAAGNAVANSNFYGLGNGTARGG 233
Query: 197 SPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDE 256
P AR+A YKVC D C G ++SA D AI DGVD+IS+S + EE D
Sbjct: 234 VPAARIAVYKVC----DNEGCDGEAMMSAFDDAIADGVDVISISIVLDNIPPFEE---DP 286
Query: 257 ISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-KTL 315
I+IGAFHA+A +L V +AGN GP +V + APWVF+VAAS +R F + + +G+ K L
Sbjct: 287 IAIGAFHAMAVGVLTVNAAGNNGPKISTVTSTAPWVFSVAASVTNRAFMAKVVLGDGKIL 346
Query: 316 TGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIK 375
G S+ ++ +V A L+ + AR C P+ LD V GKIV CD K
Sbjct: 347 IGRSVNTYDMNGTNYPLVYGKSAALSTCSVDKARLCEPKCLDGKLVKGKIVLCD---STK 403
Query: 376 SVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDI 435
+ E Q+ GA G I++N PE + + S P +S+ N +SL +
Sbjct: 404 GLIEAQK---LGAVGSIVKN-PEPDRAFIRSFP-----VSFLSNDDY---KSLVSYMNST 451
Query: 436 KSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASA 495
K+ + + N+R AP++AS+SSRGP+ + ILKPD+TAPGV ILAAYS +S
Sbjct: 452 KNPKATVLKSEEISNQR--APLVASFSSRGPSSIVSDILKPDITAPGVEILAAYSPDSSP 509
Query: 496 SNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
+ DTRR ++V+ GTSM+CPHVAG A +KT HP WSP+ I+SAI TT
Sbjct: 510 TESEFDTRR-VKYSVLSGTSMACPHVAGVAAYVKTFHPQWSPSMIQSAIMTT 560
>At5g59090 cucumisin precursor - like
Length = 736
Score = 362 bits (929), Expect = e-100
Identities = 228/533 (42%), Positives = 303/533 (56%), Gaps = 52/533 (9%)
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
++ SY + NGFAA L E E IA+ VVSVF +K +LHTT SW+F+G++ +
Sbjct: 69 LVRSYKRSFNGFAARLTESERTLIAEIEGVVSVFPNKILQLHTTTSWDFMGVKEG--KNT 126
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
+ +TIIG IDTG+WPESKSFSD+G GP P KW+G +C + K CN K
Sbjct: 127 KRNLAIESDTIIGVIDTGIWPESKSFSDKGFGPPPKKWKG--VC-----SGGKNFTCNNK 179
Query: 137 LIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGG 196
LIGAR + + + RD GHGTHT STA GN V S F IGNGT++GG
Sbjct: 180 LIGARDY-------------TSEGTRDTSGHGTHTASTAAGNAVKDTSFFGIGNGTVRGG 226
Query: 197 SPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDE 256
P +R+A YKVC TD+ C +LS+ D AI DGVD+I++S G + E+ D
Sbjct: 227 VPASRIAAYKVC---TDS-GCSSEALLSSFDDAIADGVDLITISIGFQFPSIFED---DP 279
Query: 257 ISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-KTL 315
I+IGAFHA+A+ IL V+SAGN GP P +V +VAPW+FTVAAST +R F + + +GN KTL
Sbjct: 280 IAIGAFHAMAKGILTVSSAGNSGPKPTTVSHVAPWIFTVAASTTNRGFITKVVLGNGKTL 339
Query: 316 TGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIK 375
G S+ + + +V A + + A C P L+ S+V GKI+ C K
Sbjct: 340 AGRSVNAFDMKGKKYPLVYGKSAASSACDAKTAALCAPACLNKSRVKGKILVCGGPSGYK 399
Query: 376 SVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDI 435
A S GA +I ++ P V T P S + + S I
Sbjct: 400 I------AKSVGAIAIIDKS----------PRPDVAFTHHLPA--SGLKAKDFKSLVSYI 441
Query: 436 KSGTKLRMSPAKTLN-RRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFAS 494
+S + + KT + +PV+AS+SSRGPN + ILKPD+TAPGV ILAA+S
Sbjct: 442 ESQDSPQAAVLKTETIFNRTSPVIASFSSRGPNTIAVDILKPDITAPGVEILAAFSPNGE 501
Query: 495 ASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
S DTRR ++V GTSM+CPHVAG A +KT +P WSP+ I+SAI TT
Sbjct: 502 PSE--DDTRR-VKYSVFSGTSMACPHVAGVAAYVKTFYPRWSPSMIQSAIMTT 551
>At3g46840 subtilisin-like proteinase
Length = 739
Score = 361 bits (927), Expect = e-100
Identities = 225/539 (41%), Positives = 313/539 (57%), Gaps = 43/539 (7%)
Query: 10 KENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLR 69
+ + ++ ++ +Y + NGFAA L + E +A +VVSVF +K+ KL TT SW F+GL+
Sbjct: 64 ESSIEDRLVRNYKRSFNGFAARLTKSEREILASMDEVVSVFPNKKLKLQTTTSWNFMGLK 123
Query: 70 GNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSK 129
+ + +TIIG ID+G++PES SFS +G GP P KW+G +C+ K
Sbjct: 124 --ESKRTKRNTIIESDTIIGVIDSGIYPESDSFSGKGFGPPPKKWKG--VCK-----GGK 174
Query: 130 KVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIG 189
N KLIGAR++ KL ++ARD++GHG+HT STA GN V S + +G
Sbjct: 175 NFTWNNKLIGARYYTP-------KLEGFPESARDYMGHGSHTASTAAGNAVKHVSFYGLG 227
Query: 190 NGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNS 249
NGT +GG P AR+A YKVC D + G +L+A D AI D VDII++S GG +S+
Sbjct: 228 NGTARGGVPAARIAVYKVCDPGVDGCTTDG--ILAAFDDAIADKVDIITISIGGDNSSPF 285
Query: 250 EEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMT 309
EE D I+IGAFHA+A+ IL+V SAGN GP P +V ++APW+FTVAAS +R F + +
Sbjct: 286 EE---DPIAIGAFHAMAKGILIVNSAGNSGPEPSTVASIAPWMFTVAASNTNRAFVTKVV 342
Query: 310 IGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVAC 368
+GN KT+ G S+ + + +V A ++ A FC P LD +V GKIV C
Sbjct: 343 LGNGKTVVGRSVNSFDLNGKKYPLVYGKSAS-SSCGAASAGFCSPGCLDSKRVKGKIVLC 401
Query: 369 DREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSL 428
D S EA + GA I+R+ V S S+P + +
Sbjct: 402 D------SPQNPDEAQAMGAIASIVRSH----------RTDVASIFSFPVSVLLEDDYNT 445
Query: 429 DIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAA 488
+ + K + ++T+ ++ APV+ASY SRGPN + P ILKPD+TAPG I+AA
Sbjct: 446 VLSYMNSTKNPKAAVLKSETIFNQR-APVVASYFSRGPNTIIPDILKPDITAPGSEIVAA 504
Query: 489 YSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
YS A S I+DTRR ++V GTSMSCPHVAG A +K+ HP WSP+ I+SAI TT
Sbjct: 505 YSPDAPPS--ISDTRR-VKYSVDTGTSMSCPHVAGVAAYLKSFHPRWSPSMIQSAIMTT 560
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.132 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,262,386
Number of Sequences: 26719
Number of extensions: 539334
Number of successful extensions: 1683
Number of sequences better than 10.0: 69
Number of HSP's better than 10.0 without gapping: 59
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1216
Number of HSP's gapped (non-prelim): 96
length of query: 547
length of database: 11,318,596
effective HSP length: 104
effective length of query: 443
effective length of database: 8,539,820
effective search space: 3783140260
effective search space used: 3783140260
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC133779.5


