

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133779.1 + phase: 0 /pseudo
(611 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
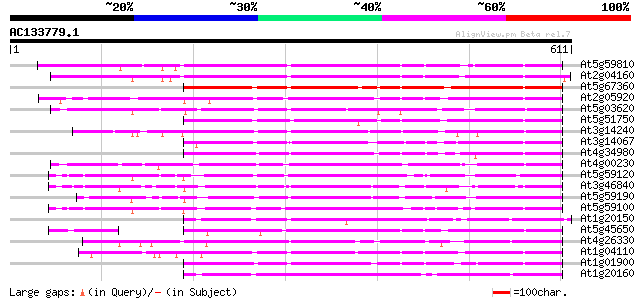
Score E
Sequences producing significant alignments: (bits) Value
At5g59810 subtilisin-like protease - like protein 444 e-125
At2g04160 subtilisin-like serine protease AIR3 411 e-115
At5g67360 cucumisin-like serine protease (gb|AAC18851.1) 319 2e-87
At2g05920 serine protease like protein 313 1e-85
At5g03620 cucumisin precursor -like protein 304 1e-82
At5g51750 serine protease-like protein 297 1e-80
At3g14240 unknown protein 289 3e-78
At3g14067 subtilisin-like serine proteinase, putative, 3' partial 288 6e-78
At4g34980 subtilisin proteinase - like 286 3e-77
At4g00230 subtilisin-type serine endopeptidase XSP1 285 5e-77
At5g59120 cucumisin precursor - like 285 7e-77
At3g46840 subtilisin-like proteinase 282 4e-76
At5g59190 cucumisin precursor - like 279 3e-75
At5g59100 cucumisin precursor - like 278 8e-75
At1g20150 unknown protein 278 8e-75
At5g45650 subtilisin-like protease 276 3e-74
At4g26330 subtilisin protease - like 275 5e-74
At1g04110 putative subtilisin protease 275 5e-74
At1g01900 putative subtilisin-like serine protease 272 3e-73
At1g20160 unknown protein 270 2e-72
>At5g59810 subtilisin-like protease - like protein
Length = 778
Score = 444 bits (1142), Expect = e-125
Identities = 265/592 (44%), Positives = 359/592 (59%), Gaps = 40/592 (6%)
Query: 31 ISFWFLVSYLFLQCYIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIY 90
++ +F ++ + YIVYLG+H H P SS L+ +SH L S +GSHE A+EAI Y
Sbjct: 28 VTLFFSPAFALKKSYIVYLGSHAHLPQISSAHLDGVAHSHRTFLASFVGSHENAKEAIFY 87
Query: 91 SYNKQINGFAAILEEEEAAQLANCTQHVL-----GNSLD*VQMMSTQLGKREGLVKIPL* 145
SY + INGFAAIL+E EAA++A V G L + L + G+V
Sbjct: 88 SYKRHINGFAAILDENEAAEIAKHPDVVSVFPNKGRKLHTTHSWNFMLLAKNGVVHKSS- 146
Query: 146 LTLIQVFGRNRRVLTTEE*V-----QFH*DGVGILLRKF----HVTVYLNGLPRKLIGAR 196
L +G + + + V F +G G + ++ H V N RKLIGAR
Sbjct: 147 LWNKAGYGEDTIIANLDTGVWPESKSFSDEGYGAVPARWKGRCHKDVPCN---RKLIGAR 203
Query: 197 FFNKAYEAFHGKLPS--SQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPR 254
+FNK Y A+ G LPS S +T RD G G+HTLSTA GNFV A +FGIGNGT GGSP+
Sbjct: 204 YFNKGYLAYTG-LPSNASYETCRDHDGHGSHTLSTAAGNFVPGANVFGIGNGTASGGSPK 262
Query: 255 SRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISI 314
+RVA YK CW D +CF AD+LAAI+ AI DG D++S S GG +D I+I
Sbjct: 263 ARVAAYKVCWPPVDGAECFDADILAAIEAAIEDGVDVLSASVGGDAGD----YMSDGIAI 318
Query: 315 GAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTI-NNKTLTGA 373
G+FHA+ + +V SAGN GP G+V+NVAPWV TV AS++DR+F + + + N ++ G
Sbjct: 319 GSFHAVKNGVTVVCSAGNSGPKSGTVSNVAPWVITVGASSMDREFQAFVELKNGQSFKGT 378
Query: 374 SLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIA 433
SL LP + + +I + DA AN DA C+ G+LDP KV GK++ C R G +
Sbjct: 379 SLSKPLPEEKMYSLISAADANVANGNVTDALLCKKGSLDPKKVKGKILVCLR-GDNARVD 437
Query: 434 EGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVV--STINYYDARSITTPKGSEITPED-I 490
+G +A +AGA G+++ N + G ++++ HV+ S I+Y D ++ + S P+ I
Sbjct: 438 KGMQAAAAGAAGMVLCND-KASGNEIISDAHVLPASQIDYKDGETLFSYLSSTKDPKGYI 496
Query: 491 KTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASV 550
K +P LN KPAP MASFSSRGPN + P ILKPD+TAPGVNI+AA++
Sbjct: 497 K-------APTATLN-TKPAPFMASFSSRGPNTITPGILKPDITAPGVNIIAAFTEATGP 548
Query: 551 SNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
++L +DNRR PFN + GTSMSCPH+ G GL+KTLHP+WSPAAI+SAIMTT
Sbjct: 549 TDLDSDNRR-TPFNTESGTSMSCPHISGVVGLLKTLHPHWSPAAIRSAIMTT 599
>At2g04160 subtilisin-like serine protease AIR3
Length = 772
Score = 411 bits (1056), Expect = e-115
Identities = 252/590 (42%), Positives = 341/590 (57%), Gaps = 39/590 (6%)
Query: 45 YIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAILE 104
Y+VY GAH H + ++ +HYD LGS GS E A +AI YSY K INGFAA L+
Sbjct: 32 YVVYFGAHSHVGEITEDAMDRVKETHYDFLGSFTGSRERATDAIFYSYTKHINGFAAHLD 91
Query: 105 EEEAAQLANCTQHVLGNSLD*VQMMSTQ----LGKREGLVKIPL*LTLIQVFGRNRRVLT 160
+ A +++ + V +++ +T+ LG + FG + +
Sbjct: 92 HDLAYEISKHPEVVSVFPNKALKLHTTRSWDFLGLEHNSYVPSSSIWRKARFGEDTIIAN 151
Query: 161 TEE*V-----QFH*DGVG--------ILLRKFHVTVYLNGLPRKLIGARFFNKAYEAFHG 207
+ V F +G+G I + T + N RKLIGAR+FNK Y A G
Sbjct: 152 LDTGVWPESKSFRDEGLGPIPSRWKGICQNQKDATFHCN---RKLIGARYFNKGYAAAVG 208
Query: 208 KLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLT 267
L SS + RD G G+HTLSTA G+FV +IFG GNGT KGGSPR+RVA YK CW
Sbjct: 209 HLNSSFDSPRDLDGHGSHTLSTAAGDFVPGVSIFGQGNGTAKGGSPRARVAAYKVCWPPV 268
Query: 268 DVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLV 327
+C+ ADVLAA D AI+DGAD+ISVS GG+P + F D ++IG+FHA + I++V
Sbjct: 269 KGNECYDADVLAAFDAAIHDGADVISVSLGGEPTS----FFNDSVAIGSFHAAKKRIVVV 324
Query: 328 ASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTI-NNKTLTGASLFVNLPPNQDFL 386
SAGN GP +V+NVAPW TV AST+DR+F+S + + N K G SL P+ F
Sbjct: 325 CSAGNSGPADSTVSNVAPWQITVGASTMDREFASNLVLGNGKHYKGQSLSSTALPHAKFY 384
Query: 387 -IIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVG 445
I+ S +AK N + +DAQ C+ G+LDP K GK++ C R G+ + +G+ G +G
Sbjct: 385 PIMASVNAKAKNASALDAQLCKLGSLDPIKTKGKILVCLR-GQNGRVEKGRAVALGGGIG 443
Query: 446 VIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPEDIKTNATIRMSPANALN 505
+++ N V G LLA+PHV+ S + T + I ++P+
Sbjct: 444 MVLEN-TYVTGNDLLADPHVLPATQLTSKDSFAVSRYISQTKKPI-----AHITPSRTDL 497
Query: 506 GRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNI 565
G KPAPVMASFSS+GP+ V P ILKPD+TAPGV+++AAY+ S +N D RR FN
Sbjct: 498 GLKPAPVMASFSSKGPSIVAPQILKPDITAPGVSVIAAYTGAVSPTNEQFDPRR-LLFNA 556
Query: 566 QQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT-----DCNGQMEN 610
GTSMSCPH+ G AGL+KT +P+WSPAAI+SAIMTT D G ++N
Sbjct: 557 ISGTSMSCPHISGIAGLLKTRYPSWSPAAIRSAIMTTATIMDDIPGPIQN 606
>At5g67360 cucumisin-like serine protease (gb|AAC18851.1)
Length = 757
Score = 319 bits (818), Expect = 2e-87
Identities = 190/420 (45%), Positives = 260/420 (61%), Gaps = 30/420 (7%)
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQTA--RDFVGPGTHTLSTAGGNFVQNATIFGIGNGT 247
RKLIGARFF + YE+ G + S+++ RD G GTHT STA G+ V+ A++ G +GT
Sbjct: 178 RKLIGARFFARGYESTMGPIDESKESRSPRDDDGHGTHTSSTAAGSVVEGASLLGYASGT 237
Query: 248 IKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVI 307
+G +PR+RVA YK CW + CF +D+LAAID+AI D +++S+S GG +
Sbjct: 238 ARGMAPRARVAVYKVCW----LGGCFSSDILAAIDKAIADNVNVLSMSLGGGMSD----Y 289
Query: 308 FTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTINN 367
+ D ++IGAF A+ R IL+ SAGN GP+ S++NVAPW+ TV A TLDRDF ++ + N
Sbjct: 290 YRDGVAIGAFAAMERGILVSCSAGNAGPSSSSLSNVAPWITTVGAGTLDRDFPALAILGN 349
Query: 368 -KTLTGASLFVN--LPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACD 424
K TG SLF LP D L+ +N T+ C GTL P KV GK+V CD
Sbjct: 350 GKNFTGVSLFKGEALP---DKLLPFIYAGNASNATN--GNLCMTGTLIPEKVKGKIVMCD 404
Query: 425 REGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSE 484
R G + +G +AG VG+I+ N +G+ L+A+ H++ + K +
Sbjct: 405 R-GINARVQKGDVVKAAGGVGMILANTA-ANGEELVADAHLLPATTVGE-------KAGD 455
Query: 485 ITPEDIKT--NATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILA 542
I + T N T +S + G KP+PV+A+FSSRGPN + P ILKPD+ APGVNILA
Sbjct: 456 IIRHYVTTDPNPTASISILGTVVGVKPSPVVAAFSSRGPNSITPNILKPDLIAPGVNILA 515
Query: 543 AYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
A++ A + L +D+RR FNI GTSMSCPHV G A L+K++HP WSPAAI+SA+MTT
Sbjct: 516 AWTGAAGPTGLASDSRR-VEFNIISGTSMSCPHVSGLAALLKSVHPEWSPAAIRSALMTT 574
Score = 30.4 bits (67), Expect = 3.0
Identities = 23/67 (34%), Positives = 34/67 (50%), Gaps = 10/67 (14%)
Query: 45 YIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAILE 104
YIV++ PSS DL + Y S L S ++ E ++Y+Y I+GF+ L
Sbjct: 32 YIVHMAK---SQMPSSFDLHSNWYD------SSLRSISDSAE-LLYTYENAIHGFSTRLT 81
Query: 105 EEEAAQL 111
+EEA L
Sbjct: 82 QEEADSL 88
>At2g05920 serine protease like protein
Length = 754
Score = 313 bits (803), Expect = 1e-85
Identities = 223/601 (37%), Positives = 314/601 (52%), Gaps = 72/601 (11%)
Query: 32 SFWFLVSYLFLQCYIVYLGAHV----HGPTPSSVDLETATYSHYDLLGSILGSHEEAEEA 87
S + ++LFL + ++ H P S +H+D S L S E +
Sbjct: 10 SITIITTFLFLLLHTTAKKTYIIRVNHSDKPESF------LTHHDWYTSQLNS----ESS 59
Query: 88 IIYSYNKQINGFAAILEEEEAAQLANCTQHVLGNSLD*VQMMSTQLGKREGLVKIPL*LT 147
++Y+Y +GF+A L+ EA L + + +L D + + T + P L
Sbjct: 60 LLYTYTTSFHGFSAYLDSTEADSLLSSSNSILDIFEDPLYTLHT--------TRTPEFLG 111
Query: 148 LIQVFGRNRRVLTTEE*VQFH*D-GVGILLRKFHVTVYLNGLP----------------- 189
L FG + ++ + D GV R F T + +P
Sbjct: 112 LNSEFGVHDLGSSSNGVIIGVLDTGVWPESRSFDDTD-MPEIPSKWKGECESGSDFDSKL 170
Query: 190 --RKLIGARFFNKAYEAFHGKLPSSQQTA---RDFVGPGTHTLSTAGGNFVQNATIFGIG 244
+KLIGAR F+K ++ G SS++ + RD G GTHT +TA G+ V+NA+ G
Sbjct: 171 CNKKLIGARSFSKGFQMASGGGFSSKRESVSPRDVDGHGTHTSTTAAGSAVRNASFLGYA 230
Query: 245 NGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNP 304
GT +G + R+RVATYK CWS CFG+D+LAA+D+AI DG D++S+S GG
Sbjct: 231 AGTARGMATRARVATYKVCWS----TGCFGSDILAAMDRAILDGVDVLSLSLGG----GS 282
Query: 305 EVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMT 364
+ D I+IGAF A+ R + + SAGN GPT SV NVAPWV TV A TLDRDF +
Sbjct: 283 APYYRDTIAIGAFSAMERGVFVSCSAGNSGPTRASVANVAPWVMTVGAGTLDRDFPAFAN 342
Query: 365 I-NNKTLTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVAC 423
+ N K LTG SL+ + L ++ ++ + C PG+LD S V GK+V C
Sbjct: 343 LGNGKRLTGVSLYSGVGMGTKPLELVYNKGNSSS-----SNLCLPGSLDSSIVRGKIVVC 397
Query: 424 DREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGS 483
DR G + +G AG +G+IM N G+ L+A+ H++ I K
Sbjct: 398 DR-GVNARVEKGAVVRDAGGLGMIMAN-TAASGEELVADSHLLPAI-------AVGKKTG 448
Query: 484 EITPEDIKTNA--TIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNIL 541
++ E +K+++ T + + KP+PV+A+FSSRGPN V P ILKPDV PGVNIL
Sbjct: 449 DLLREYVKSDSKPTALLVFKGTVLDVKPSPVVAAFSSRGPNTVTPEILKPDVIGPGVNIL 508
Query: 542 AAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMT 601
A +S + L D+RR FNI GTSMSCPH+ G AGL+K HP WSP+AIKSA+MT
Sbjct: 509 AGWSDAIGPTGLDKDSRR-TQFNIMSGTSMSCPHISGLAGLLKAAHPEWSPSAIKSALMT 567
Query: 602 T 602
T
Sbjct: 568 T 568
>At5g03620 cucumisin precursor -like protein
Length = 766
Score = 304 bits (778), Expect = 1e-82
Identities = 221/577 (38%), Positives = 305/577 (52%), Gaps = 55/577 (9%)
Query: 45 YIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAILE 104
YIVY+G L A +H++LL +++G +A E IYSY K INGF A L
Sbjct: 35 YIVYMGEATEN------SLVEAAENHHNLLMTVIGDESKARELKIYSYGKNINGFVARLF 88
Query: 105 EEEAAQLANCTQHVLGNSLD*VQMMSTQ----LGKREGLVKIPL*LTLIQVFGRNRRVLT 160
EA +L+ V Q+ +T+ LG E K + + + G +
Sbjct: 89 PHEAEKLSREEGVVSVFKNTQRQLHTTRSWDFLGLVESKYKRSVGIESNIIVGVLDTGID 148
Query: 161 TEE*VQFH*DGVGILLRKFH-VTVYLNGLPR---KLIGARFFNKAYEAFHGKLPSSQ-QT 215
E F+ GVG K+ V N R K+IGA++F+ E LP + T
Sbjct: 149 VES-PSFNDKGVGPPPAKWKGKCVTGNNFTRCNNKVIGAKYFHIQSEG----LPDGEGDT 203
Query: 216 ARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGA 275
A D G GTHT ST G V +A++FGI NGT +GG P +R+A YK CW C
Sbjct: 204 AADHDGHGTHTSSTIAGVSVSSASLFGIANGTARGGVPSARIAAYKVCWDS----GCTDM 259
Query: 276 DVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGP 335
D+LAA D+AI DG D+IS+S GG F D I+IGAFHA+ R IL SAGN GP
Sbjct: 260 DMLAAFDEAISDGVDIISISIGGASLP----FFEDPIAIGAFHAMKRGILTTCSAGNNGP 315
Query: 336 TPGSVTNVAPWVFTVAASTLDRDFSSVMTINNKTLTGASLFVNLPPNQDFLIIISTDAKF 395
+V+N+APWV TVAA++LDR F +V+ + N LT + + +N + + +++ +
Sbjct: 316 GLFTVSNLAPWVMTVAANSLDRKFETVVKLGN-GLTASGISLNGFNPRKKMYPLTSGSLA 374
Query: 396 ANVTD---VDAQFCRPGTLDPSKVNGKVVACD---REGKINSIAEGQEALSAGAVGVIMR 449
+N++ + C PGTL KV GKVV C+ EG + S GVI++
Sbjct: 375 SNLSAGGYGEPSTCEPGTLGEDKVMGKVVYCEAGREEGGNGGQGQDHVVRSLKGAGVIVQ 434
Query: 450 --NQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPEDI--KTNATIRMSPANALN 505
++ TL+A +V + D IT S P+ + KT T
Sbjct: 435 LLEPTDMATSTLIAGSYVF----FEDGTKITEYINSTKNPQAVIFKTKTT---------- 480
Query: 506 GRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNI 565
+ AP ++SFS+RGP ++ P ILKPD++APG+NILAAYS LASV+ DNRR F+I
Sbjct: 481 -KMLAPSISSFSARGPQRISPNILKPDISAPGLNILAAYSKLASVTGYPDDNRRTL-FSI 538
Query: 566 QQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
GTSM+CPH A +K+ HP+WSPAAIKSA+MTT
Sbjct: 539 MSGTSMACPHAAAAAAYVKSFHPDWSPAAIKSALMTT 575
>At5g51750 serine protease-like protein
Length = 780
Score = 297 bits (760), Expect = 1e-80
Identities = 178/421 (42%), Positives = 252/421 (59%), Gaps = 29/421 (6%)
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQ--TARDFVGPGTHTLSTAGGNFVQNATIFGIGNGT 247
RK++GAR F + YEA GK+ + + RD G GTHT +T G+ V+ A +FG GT
Sbjct: 193 RKIVGARVFYRGYEAATGKIDEELEYKSPRDRDGHGTHTAATVAGSPVKGANLFGFAYGT 252
Query: 248 IKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVI 307
+G + ++RVA YK CW V CF +D+L+A+DQA+ DG ++S+S GG +T
Sbjct: 253 ARGMAQKARVAAYKVCW----VGGCFSSDILSAVDQAVADGVQVLSISLGGGVSTYSR-- 306
Query: 308 FTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTINN 367
D +SI F A+ + + SAGN GP P S+TNV+PW+ TV AST+DRDF + + I
Sbjct: 307 --DSLSIATFGAMEMGVFVSCSAGNGGPDPISLTNVSPWITTVGASTMDRDFPATVKIGT 364
Query: 368 -KTLTGASLFVN---LPPNQDF-LIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVA 422
+T G SL+ LP N+ + L+ + +A + T FC G LD V GK+V
Sbjct: 365 MRTFKGVSLYKGRTVLPKNKQYPLVYLGRNASSPDPTS----FCLDGALDRRHVAGKIVI 420
Query: 423 CDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKG 482
CDR G + +GQ AG +G+++ N +G+ L+A+ H++ + ++ +G
Sbjct: 421 CDR-GVTPRVQKGQVVKRAGGIGMVLTNTA-TNGEELVADSHMLPAV------AVGEKEG 472
Query: 483 SEITPEDIKTN-ATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNIL 541
I + + AT + G KP+PV+A+FSSRGPN + ILKPD+ APGVNIL
Sbjct: 473 KLIKQYAMTSKKATASLEILGTRIGIKPSPVVAAFSSRGPNFLSLEILKPDLLAPGVNIL 532
Query: 542 AAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMT 601
AA++ + S+L +D RR FNI GTSMSCPHV G A LIK+ HP+WSPAAIKSA+MT
Sbjct: 533 AAWTGDMAPSSLSSDPRR-VKFNILSGTSMSCPHVSGVAALIKSRHPDWSPAAIKSALMT 591
Query: 602 T 602
T
Sbjct: 592 T 592
>At3g14240 unknown protein
Length = 775
Score = 289 bits (740), Expect = 3e-78
Identities = 211/567 (37%), Positives = 306/567 (53%), Gaps = 61/567 (10%)
Query: 69 SHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAILEEEEAAQLANCTQHVLGNSLD*VQM 128
+H+ S L S + +II++Y+ +GF+A L ++A+QL + HV+ + V+
Sbjct: 43 THFHWYTSSLASLTSSPPSIIHTYDTVFHGFSARLTSQDASQLLD-HPHVISVIPEQVRH 101
Query: 129 MSTQ-----LGKRE----GLVKIPL*LTLIQVFGRNRRVLTTEE*V-----QFH*DGVGI 174
+ T LG R GL++ FG + + + V F G+G
Sbjct: 102 LHTTRSPEFLGLRSTDKAGLLEE-------SDFGSDLVIGVIDTGVWPERPSFDDRGLGP 154
Query: 175 LLRKFHVTVYLN------GLPRKLIGARFFNKAYEAFHGKLPSSQQ--TARDFVGPGTHT 226
+ K+ + RKL+GARFF YEA +GK+ + + + RD G GTHT
Sbjct: 155 VPIKWKGQCIASQDFPESACNRKLVGARFFCGGYEATNGKMNETTEFRSPRDSDGHGTHT 214
Query: 227 LSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIY 286
S + G +V A+ G +G G +P++R+A YK CW+ C+ +D+LAA D A+
Sbjct: 215 ASISAGRYVFPASTLGYAHGVAAGMAPKARLAAYKVCWNS----GCYDSDILAAFDTAVA 270
Query: 287 DGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPW 346
DG D+IS+S GG + D I+IGAF A+ R I + ASAGN GP +VTNVAPW
Sbjct: 271 DGVDVISLSVGGVVVP----YYLDAIAIGAFGAIDRGIFVSASAGNGGPGALTVTNVAPW 326
Query: 347 VFTVAASTLDRDF-SSVMTINNKTLTGASLF--VNLPPNQDFLIIISTDAKFANVTDVDA 403
+ TV A T+DRDF ++V N K ++G S++ L P + + ++ +
Sbjct: 327 MTTVGAGTIDRDFPANVKLGNGKMISGVSVYGGPGLDPGRMYPLVYG--GSLLGGDGYSS 384
Query: 404 QFCRPGTLDPSKVNGKVVACDREGKINSIA-EGQEALSAGAVGVIMRNQPEVDGKTLLAE 462
C G+LDP+ V GK+V CDR INS A +G+ G +G+I+ N DG+ L+A+
Sbjct: 385 SLCLEGSLDPNLVKGKIVLCDRG--INSRATKGEIVRKNGGLGMIIANGV-FDGEGLVAD 441
Query: 463 PHVVSTINYYDARSITTPKGSEIT---PEDIKTNATIRMSPANALNGRK----PAPVMAS 515
HV+ A S+ G EI E K+ ++ + G + PAPV+AS
Sbjct: 442 CHVLP------ATSVGASGGDEIRRYISESSKSRSSKHPTATIVFKGTRLGIRPAPVVAS 495
Query: 516 FSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPH 575
FS+RGPN P ILKPDV APG+NILAA+ S + +DNRR FNI GTSM+CPH
Sbjct: 496 FSARGPNPETPEILKPDVIAPGLNILAAWPDRIGPSGVTSDNRRT-EFNILSGTSMACPH 554
Query: 576 VVGTAGLIKTLHPNWSPAAIKSAIMTT 602
V G A L+K HP+WSPAAI+SA++TT
Sbjct: 555 VSGLAALLKAAHPDWSPAAIRSALITT 581
>At3g14067 subtilisin-like serine proteinase, putative, 3' partial
Length = 743
Score = 288 bits (737), Expect = 6e-78
Identities = 180/423 (42%), Positives = 247/423 (57%), Gaps = 35/423 (8%)
Query: 190 RKLIGARFFNKAY----EAFHGKLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGN 245
RKLIGAR F + Y ++ RD G GTHT STA G+ V NA+++
Sbjct: 181 RKLIGARAFYRGYLTQRNGTKKHAAKESRSPRDTEGHGTHTASTAAGSVVANASLYQYAR 240
Query: 246 GTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPE 305
GT G + ++R+A YK CW+ C+ +D+LAA+DQA+ DG +IS+S G + PE
Sbjct: 241 GTATGMASKARIAAYKICWT----GGCYDSDILAAMDQAVADGVHVISLSVGASGSA-PE 295
Query: 306 VIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDF-SSVMT 364
TD I+IGAF A I++ SAGN GP P + TN+APW+ TV AST+DR+F ++ +T
Sbjct: 296 Y-HTDSIAIGAFGATRHGIVVSCSAGNSGPNPETATNIAPWILTVGASTVDREFAANAIT 354
Query: 365 INNKTLTGASLFV--NLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVA 422
+ K TG SL+ +LP +Q L+ D ++ C PG L+ S V GK+V
Sbjct: 355 GDGKVFTGTSLYAGESLPDSQLSLVYSG---------DCGSRLCYPGKLNSSLVEGKIVL 405
Query: 423 CDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKG 482
CDR G + +G AG G+I+ N E G+ L A+ H+V A + G
Sbjct: 406 CDRGGNAR-VEKGSAVKLAGGAGMILANTAE-SGEELTADSHLV------PATMVGAKAG 457
Query: 483 SEITPEDIKT--NATIRMSPANALNG-RKPAPVMASFSSRGPNKVQPYILKPDVTAPGVN 539
+I + IKT + T ++S L G P+P +A+FSSRGPN + P ILKPDV APGVN
Sbjct: 458 DQIR-DYIKTSDSPTAKISFLGTLIGPSPPSPRVAAFSSRGPNHLTPVILKPDVIAPGVN 516
Query: 540 ILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAI 599
ILA ++ + ++L D RR FNI GTSMSCPHV G A L++ HP+WSPAAIKSA+
Sbjct: 517 ILAGWTGMVGPTDLDIDPRR-VQFNIISGTSMSCPHVSGLAALLRKAHPDWSPAAIKSAL 575
Query: 600 MTT 602
+TT
Sbjct: 576 VTT 578
>At4g34980 subtilisin proteinase - like
Length = 764
Score = 286 bits (731), Expect = 3e-77
Identities = 176/424 (41%), Positives = 250/424 (58%), Gaps = 34/424 (8%)
Query: 190 RKLIGARFFNKAYEA-FHGKLPSSQQ--TARDFVGPGTHTLSTAGGNFVQNATIFGIGNG 246
RK+IGARFF K +A G + + + + RD G GTHT STA G A++ G +G
Sbjct: 170 RKIIGARFFAKGQQAAVIGGINKTVEFLSPRDADGHGTHTSSTAAGRHAFKASMSGYASG 229
Query: 247 TIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPN-TNPE 305
KG +P++R+A YK CW + C +D+LAA D A+ DG D+IS+S GG T+P
Sbjct: 230 VAKGVAPKARIAAYKVCWKDSG---CLDSDILAAFDAAVRDGVDVISISIGGGDGITSP- 285
Query: 306 VIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTI 365
+ D I+IG++ A ++ I + +SAGNEGP SVTN+APWV TV AST+DR+F + +
Sbjct: 286 -YYLDPIAIGSYGAASKGIFVSSSAGNEGPNGMSVTNLAPWVTTVGASTIDRNFPADAIL 344
Query: 366 NN-KTLTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACD 424
+ L G SL+ +P N ++ + A C TLDP +V GK+V CD
Sbjct: 345 GDGHRLRGVSLYAGVPLNGRMFPVVYPGKSGMS----SASLCMENTLDPKQVRGKIVICD 400
Query: 425 REGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSE 484
R G +A+G AG VG+I+ N +G+ L+ + H++ A ++ + +G
Sbjct: 401 R-GSSPRVAKGLVVKKAGGVGMILANGAS-NGEGLVGDAHLIP------ACAVGSNEGDR 452
Query: 485 ITPEDIKTNATIRMSPANALN------GRKPAPVMASFSSRGPNKVQPYILKPDVTAPGV 538
I K A+ +P +++ G KPAPV+ASFS RGPN + P ILKPD+ APGV
Sbjct: 453 I-----KAYASSHPNPIASIDFRGTIVGIKPAPVIASFSGRGPNGLSPEILKPDLIAPGV 507
Query: 539 NILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSA 598
NILAA++ + L +D R+ FNI GTSM+CPHV G A L+K+ HP+WSPA I+SA
Sbjct: 508 NILAAWTDAVGPTGLPSDPRK-TEFNILSGTSMACPHVSGAAALLKSAHPDWSPAVIRSA 566
Query: 599 IMTT 602
+MTT
Sbjct: 567 MMTT 570
>At4g00230 subtilisin-type serine endopeptidase XSP1
Length = 749
Score = 285 bits (729), Expect = 5e-77
Identities = 205/576 (35%), Positives = 299/576 (51%), Gaps = 63/576 (10%)
Query: 45 YIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAILE 104
YI+YLG + E +H +LL S+ S EEA+E +YSY K N FAA L
Sbjct: 38 YIIYLGDRPD-------NTEETIKTHINLLSSLNISQEEAKERKVYSYTKAFNAFAAKLS 90
Query: 105 EEEAAQLANCTQHVLGNSLD*VQMMSTQLGKREGLVKIPL*LTLIQVFGRNRRVLT---- 160
EA ++ + V S+ Q K V +PL T + R V+
Sbjct: 91 PHEAKKMMEMEEVV---SVSRNQYRKLHTTKSWDFVGLPL--TAKRHLKAERDVIIGVLD 145
Query: 161 ---TEE*VQFH*DGVGILLRKFHVTV--YLN--GLPRKLIGARFFNKAYEAFHGKLPSSQ 213
T + F G+G K+ + Y N G K+IGA++F G +P+ +
Sbjct: 146 TGITPDSESFLDHGLGPPPAKWKGSCGPYKNFTGCNNKIIGAKYFKH-----DGNVPAGE 200
Query: 214 -QTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDC 272
++ D G GTHT ST G V NA+++GI NGT +G P +R+A YK CW+ + C
Sbjct: 201 VRSPIDIDGHGTHTSSTVAGVLVANASLYGIANGTARGAVPSARLAMYKVCWARSG---C 257
Query: 273 FGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFT-DEISIGAFHALARNILLVASAG 331
D+LA + AI+DG ++IS+S GG P ++ D IS+G+FHA+ + IL VASAG
Sbjct: 258 ADMDILAGFEAAIHDGVEIISISIGG-----PIADYSSDSISVGSFHAMRKGILTVASAG 312
Query: 332 NEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTI-NNKTLTGASLFVNLPPNQDFLIIIS 390
N+GP+ G+VTN PW+ TVAAS +DR F S + + N K+ +G + + P + + ++
Sbjct: 313 NDGPSSGTVTNHEPWILTVAASGIDRTFKSKIDLGNGKSFSGMGISMFSPKAKSYPLVSG 372
Query: 391 TDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEAL--SAGAVGVIM 448
DA A++C +LD KV GKV+ C G G E+ S G G I+
Sbjct: 373 VDAAKNTDDKYLARYCFSDSLDRKKVKGKVMVCRMGG------GGVESTIKSYGGAGAII 426
Query: 449 RNQPEVDGKTLLAEP--HVVSTINYYDARSITTPKGSEITPEDIKTNATIRMSPANALNG 506
+ +D + P V S++ R I + + + + + TI
Sbjct: 427 VSDQYLDNAQIFMAPATSVNSSVGDIIYRYINSTRSASAVIQKTR-QVTI---------- 475
Query: 507 RKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQ 566
PAP +ASFSSRGPN +LKPD+ APG++ILAA++L S++ L D + F I
Sbjct: 476 --PAPFVASFSSRGPNPGSIRLLKPDIAAPGIDILAAFTLKRSLTGLDGDTQFS-KFTIL 532
Query: 567 QGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
GTSM+CPHV G A +K+ HP+W+PAAIKSAI+T+
Sbjct: 533 SGTSMACPHVAGVAAYVKSFHPDWTPAAIKSAIITS 568
>At5g59120 cucumisin precursor - like
Length = 732
Score = 285 bits (728), Expect = 7e-77
Identities = 214/572 (37%), Positives = 289/572 (50%), Gaps = 63/572 (11%)
Query: 43 QCYIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAI 102
Q YIVY+G+ S D T T H ++L + G E ++ SY + NGFAA
Sbjct: 30 QVYIVYMGS-----LSSRADY-TPTSDHMNILQEVTGE-SSIEGRLVRSYKRSFNGFAAR 82
Query: 103 LEEEEAAQLANCTQHVLGNSLD*VQMMSTQ----LGKREGL-VKIPL*LTLIQVFGRNRR 157
L E E ++A V +Q+ +T +G +EG+ K + + G
Sbjct: 83 LTESERERVAKMVGVVSVFPNKKLQLQTTTSWDFMGLKEGIKTKRNPTVESDTIIGVIDS 142
Query: 158 VLTTEE*VQFH*DGVGILLRKFHVTVYLNG----LPRKLIGARFFNKAYEAFHGKLPSSQ 213
+T E F G G +K+ V G KLIGAR + +
Sbjct: 143 GITPES-QSFSDKGFGPPPQKWK-GVCSGGKNFTCNNKLIGARDY-------------TS 187
Query: 214 QTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCF 273
+ RD G GTHT STA GN V +A+ FGIGNGT++GG P SRVA YK C C
Sbjct: 188 EGTRDMDGHGTHTASTAAGNAVVDASFFGIGNGTVRGGVPASRVAAYKVCTP----TGCS 243
Query: 274 GADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNE 333
+L+A D AI DG DLI++S G K + D I+IGAFHA+A+ +L V SAGN
Sbjct: 244 SEALLSAFDDAIADGVDLITISIGDK---TASMFQNDPIAIGAFHAMAKGVLTVNSAGNS 300
Query: 334 GPTPGSVTNVAPWVFTVAASTLDRDF-SSVMTINNKTLTGASLFVNLPPNQDFLIIISTD 392
GP P SV+ VAPW+ TVAAST +R F + V+ N KTL G S+ +D+ ++
Sbjct: 301 GPKPISVSGVAPWILTVAASTTNRGFVTKVVLGNGKTLVGKSVNAYEMKGKDYPLVYGKS 360
Query: 393 AKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQP 452
A + A C +D S+V GK++ C G + + S GAVG+I R P
Sbjct: 361 AASSACDAESAGLCELSCVDKSRVKGKILVCGGPGGLKIVE------SVGAVGLIYRT-P 413
Query: 453 EVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPEDI--KTNATIRMSPANALNGRKPA 510
+ D P + + D S+ + S +P+ I KT A + +
Sbjct: 414 KPD--VAFIHPLPAAGLLTEDFESLVSYLESTDSPQAIVLKTEAIF----------NRTS 461
Query: 511 PVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTS 570
PV+ASFSSRGPN + ILKPD+TAPGV ILAAYS S D+ R +++ GTS
Sbjct: 462 PVIASFSSRGPNTIAVDILKPDITAPGVEILAAYSPAGEPSQ---DDTRHVKYSVLSGTS 518
Query: 571 MSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
MSCPHV G A +KT +P WSP+ I+SAIMTT
Sbjct: 519 MSCPHVAGVAAYVKTFNPKWSPSMIQSAIMTT 550
>At3g46840 subtilisin-like proteinase
Length = 739
Score = 282 bits (721), Expect = 4e-76
Identities = 211/581 (36%), Positives = 303/581 (51%), Gaps = 74/581 (12%)
Query: 43 QCYIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAI 102
Q YIVY+GA P+ VD ++ H +L + G E+ ++ +Y + NGFAA
Sbjct: 33 QEYIVYMGA-----LPARVDYMPMSH-HTSILQDVTGE-SSIEDRLVRNYKRSFNGFAAR 85
Query: 103 LEEEEAAQLANCTQHV-----------LGNSLD*VQMMSTQLGKREGLVKIPL*LTLIQV 151
L + E LA+ + V S + + + ++ KR +++ T+I V
Sbjct: 86 LTKSEREILASMDEVVSVFPNKKLKLQTTTSWNFMGLKESKRTKRNTIIESD---TIIGV 142
Query: 152 FGRNRRVLTTEE*VQFH*DGVGILLRKFHVTVYLNGLP----RKLIGARFFNKAYEAFHG 207
E F G G +K+ V G KLIGAR++ E F
Sbjct: 143 IDSG----IYPESDSFSGKGFGPPPKKWK-GVCKGGKNFTWNNKLIGARYYTPKLEGF-- 195
Query: 208 KLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLT 267
++ARD++G G+HT STA GN V++ + +G+GNGT +GG P +R+A YK C
Sbjct: 196 -----PESARDYMGHGSHTASTAAGNAVKHVSFYGLGNGTARGGVPAARIAVYKVCDPGV 250
Query: 268 DVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLV 327
D C +LAA D AI D D+I++S GG N++P D I+IGAFHA+A+ IL+V
Sbjct: 251 D--GCTTDGILAAFDDAIADKVDIITISIGGD-NSSP--FEEDPIAIGAFHAMAKGILIV 305
Query: 328 ASAGNEGPTPGSVTNVAPWVFTVAASTLDRDF-SSVMTINNKTLTGASLFVNLPPNQDFL 386
SAGN GP P +V ++APW+FTVAAS +R F + V+ N KT+ G S+ + +
Sbjct: 306 NSAGNSGPEPSTVASIAPWMFTVAASNTNRAFVTKVVLGNGKTVVGRSVNSFDLNGKKYP 365
Query: 387 IIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGV 446
++ A ++ A FC PG LD +V GK+V CD S EA + GA+
Sbjct: 366 LVYGKSAS-SSCGAASAGFCSPGCLDSKRVKGKIVLCD------SPQNPDEAQAMGAIAS 418
Query: 447 IMRNQPEVDGKTLLAEPHVVSTINYYDA-----RSITTPKGSEITPEDIKTNATIRMSPA 501
I+R+ D ++ + P V + Y+ S PK + + E I
Sbjct: 419 IVRSH-RTDVASIFSFPVSVLLEDDYNTVLSYMNSTKNPKAAVLKSETI----------- 466
Query: 502 NALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGF 561
N R APV+AS+ SRGPN + P ILKPD+TAPG I+AAYS A S ++D RR
Sbjct: 467 --FNQR--APVVASYFSRGPNTIIPDILKPDITAPGSEIVAAYSPDAPPS--ISDTRR-V 519
Query: 562 PFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
+++ GTSMSCPHV G A +K+ HP WSP+ I+SAIMTT
Sbjct: 520 KYSVDTGTSMSCPHVAGVAAYLKSFHPRWSPSMIQSAIMTT 560
>At5g59190 cucumisin precursor - like
Length = 693
Score = 279 bits (714), Expect = 3e-75
Identities = 198/547 (36%), Positives = 289/547 (52%), Gaps = 68/547 (12%)
Query: 73 LLGSILGSHEEAEEAIIYSYNKQINGFAAILEEEEAAQLANCTQHVLGNSLD*VQMMST- 131
L+G+I SH ++ SY + NGFAA L + E+ +L N + V ++ +T
Sbjct: 22 LVGTIAASH-----LLVRSYKRSFNGFAANLSQAESQKLQNMKEVVSVFPSKSHELTTTR 76
Query: 132 --------QLGKREGLVKIPL*LTLIQVFGRNRRVLTTEE*VQFH*DGVGILLRKFHVTV 183
+ +RE + + + + +I + + E F +G G +K+ +
Sbjct: 77 SWDFVGFGEKARRESVKESDVIVGVI-----DSGIWPESE--SFDDEGFGPPPKKWKGSC 129
Query: 184 YLNGLP----RKLIGARFFNKAYEAFHGKLPSSQQTARDFVGPGTHTLSTAGGNFVQNAT 239
GL KLIGARF+NK ++ ARD G GTHT STA GN VQ A+
Sbjct: 130 K-GGLKFACNNKLIGARFYNKFADS-----------ARDEEGHGTHTASTAAGNAVQAAS 177
Query: 240 IFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGK 299
+G+ GT +GG P +R+A YK C++ C D+LAA D AI DG D+IS+S
Sbjct: 178 FYGLAQGTARGGVPSARIAAYKVCFNR-----CNDVDILAAFDDAIADGVDVISISISAD 232
Query: 300 PNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDF 359
+N + ++IG+FHA+ R I+ SAGN GP GSV NV+PW+ TVAAS DR F
Sbjct: 233 YVSN---LLNASVAIGSFHAMMRGIITAGSAGNNGPDQGSVANVSPWMITVAASGTDRQF 289
Query: 360 -SSVMTINNKTLTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNG 418
V+ N K LTG S+ F I+ + N + A +C G +D V G
Sbjct: 290 IDRVVLGNGKALTGISVNTFNLNGTKFPIVYGQNVS-RNCSQAQAGYCSSGCVDSELVKG 348
Query: 419 KVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSIT 478
K+V CD +EA AGA+GVI++N D ++ P S++ + D +SI
Sbjct: 349 KIVLCD------DFLGYREAYLAGAIGVIVQNTLLPDSAFVV--PFPASSLGFEDYKSIK 400
Query: 479 TPKGSEITP--EDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAP 536
+ S P E ++T + + AP + SFSSRGP+ V +LKPDV+AP
Sbjct: 401 SYIESAEPPQAEILRTEEIVD----------REAPYVPSFSSRGPSFVIQNLLKPDVSAP 450
Query: 537 GVNILAAYSLLASVSNLVT-DNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAI 595
G+ ILAA+S +AS S+ + +++R +++ GTSM+CPHV G A +K+ HP+WSP+AI
Sbjct: 451 GLEILAAFSPVASPSSFLNPEDKRSVRYSVMSGTSMACPHVAGVAAYVKSFHPDWSPSAI 510
Query: 596 KSAIMTT 602
KSAIMTT
Sbjct: 511 KSAIMTT 517
>At5g59100 cucumisin precursor - like
Length = 741
Score = 278 bits (710), Expect = 8e-75
Identities = 213/570 (37%), Positives = 295/570 (51%), Gaps = 52/570 (9%)
Query: 43 QCYIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAI 102
Q YIVYLG+ PS + T H +L I G E ++ SY K NGFAA
Sbjct: 33 QVYIVYLGS-----LPSREEY-TPMSDHMSILQEITGE-SLIENRLVRSYKKSFNGFAAR 85
Query: 103 LEEEEAAQLANCTQHVLGNSLD*VQMMSTQ----LGKREGL-VKIPL*LTLIQVFGRNRR 157
L E E +LA + V +++ +T +G +EG+ K + + G
Sbjct: 86 LTESERKRLAGMERVVSVFPSRKLKLQTTSSWNFMGLKEGIKTKRTRSIESDTIIGVIDS 145
Query: 158 VLTTEE*VQFH*DGVGILLRKFHVTVYLNG---LPRKLIGARFFNKAYEAFHGKLPSSQQ 214
+ E F G G +K+ T K+IGAR + +A Q
Sbjct: 146 GIYPES-DSFSDQGFGPPPKKWKGTCAGGKNFTCNNKVIGARDYTAKSKA--------NQ 196
Query: 215 TARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFG 274
TARD+ G GTHT S A GN V N+ +G+GNGT +GG P +R+A YK C D C G
Sbjct: 197 TARDYSGHGTHTASIAAGNAVANSNFYGLGNGTARGGVPAARIAVYKVC----DNEGCDG 252
Query: 275 ADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEG 334
+++A D AI DG D+IS+S N D I+IGAFHA+A +L V +AGN G
Sbjct: 253 EAMMSAFDDAIADGVDVISISI---VLDNIPPFEEDPIAIGAFHAMAVGVLTVNAAGNNG 309
Query: 335 PTPGSVTNVAPWVFTVAASTLDRDF-SSVMTINNKTLTGASLFVNLPPNQDFLIIISTDA 393
P +VT+ APWVF+VAAS +R F + V+ + K L G S+ ++ ++ A
Sbjct: 310 PKISTVTSTAPWVFSVAASVTNRAFMAKVVLGDGKILIGRSVNTYDMNGTNYPLVYGKSA 369
Query: 394 KFANVTDVDAQFCRPGTLDPSKVNGKVVACD-REGKINSIAEGQEALSAGAVGVIMRNQP 452
+ + A+ C P LD V GK+V CD +G I EA GAVG I++N P
Sbjct: 370 ALSTCSVDKARLCEPKCLDGKLVKGKIVLCDSTKGLI-------EAQKLGAVGSIVKN-P 421
Query: 453 EVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPV 512
E D + + P VS ++ D +S+ + S P+ AT+ S + + AP+
Sbjct: 422 EPDRAFIRSFP--VSFLSNDDYKSLVSYMNSTKNPK-----ATVLKSEEIS---NQRAPL 471
Query: 513 MASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMS 572
+ASFSSRGP+ + ILKPD+TAPGV ILAAYS +S + D RR +++ GTSM+
Sbjct: 472 VASFSSRGPSSIVSDILKPDITAPGVEILAAYSPDSSPTESEFDTRR-VKYSVLSGTSMA 530
Query: 573 CPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
CPHV G A +KT HP WSP+ I+SAIMTT
Sbjct: 531 CPHVAGVAAYVKTFHPQWSPSMIQSAIMTT 560
>At1g20150 unknown protein
Length = 780
Score = 278 bits (710), Expect = 8e-75
Identities = 172/427 (40%), Positives = 244/427 (56%), Gaps = 30/427 (7%)
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIK 249
RKLIGAR++N ++ L +T RDF+G GTH S A G + NA+ +G+ +G ++
Sbjct: 188 RKLIGARYYNSSFF-----LDPDYETPRDFLGHGTHVASIAAGQIIANASYYGLASGIMR 242
Query: 250 GGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFT 309
GGSP SR+A Y+AC ++ C G+ +LAA D AI DG D+IS+S G P+ +
Sbjct: 243 GGSPSSRIAMYRAC----SLLGCRGSSILAAFDDAIADGVDVISISMGLWPDN----LLE 294
Query: 310 DEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTI---N 366
D +SIG+FHA+ R I +V S GN GP+ SV N APW+ TVAAST+DR F S + +
Sbjct: 295 DPLSIGSFHAVERGITVVCSVGNSGPSSQSVFNAAPWMITVAASTIDRGFESNILLGGDE 354
Query: 367 NKTLTGASL-FVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDR 425
N+ + G + N+ Q + +I + AK + + A+ C P TLD + V GK+V CD
Sbjct: 355 NRLIEGFGINIANIDKTQAYPLIHARSAKKIDANEEAARNCAPDTLDQTIVKGKIVVCDS 414
Query: 426 EGKINSIA-EGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSE 484
+ I + E G +G+++ + +D + + +V+ I D I + S
Sbjct: 415 DLDNQVIQWKSDEVKRLGGIGMVLVDDESMD-LSFIDPSFLVTIIKPEDGIQIMSYINS- 472
Query: 485 ITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAY 544
T E I T + P + G AP + SFSSRGP + ILKPD+ APGVNILA++
Sbjct: 473 -TREPIAT-----IMPTRSRTGHMLAPSIPSFSSRGPYLLTRSILKPDIAAPGVNILASW 526
Query: 545 SLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTTDC 604
L N + + FNI+ GTSMSCPHV G A +K+ +P+WSPAAI+SAIMTT
Sbjct: 527 --LVGDRNAAPEGKPPPLFNIESGTSMSCPHVSGIAARLKSRYPSWSPAAIRSAIMTTAV 584
Query: 605 NGQMENT 611
QM NT
Sbjct: 585 --QMTNT 589
Score = 31.2 bits (69), Expect = 1.8
Identities = 23/68 (33%), Positives = 36/68 (52%), Gaps = 11/68 (16%)
Query: 45 YIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAILE 104
YI+Y+GA +S D T H +LL S+L + + + ++ Y +GFAA L
Sbjct: 33 YIIYMGA-------ASSDGSTDN-DHVELLSSLL---QRSGKTPMHRYKHGFSGFAAHLS 81
Query: 105 EEEAAQLA 112
E+EA +A
Sbjct: 82 EDEAHLIA 89
>At5g45650 subtilisin-like protease
Length = 791
Score = 276 bits (705), Expect = 3e-74
Identities = 171/425 (40%), Positives = 234/425 (54%), Gaps = 25/425 (5%)
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQ----TARDFVGPGTHTLSTAGGNFVQNATIFG-IG 244
RK+IGAR++ K YE ++G ++ + RD G G+HT STA G V A+ G
Sbjct: 199 RKIIGARYYVKGYERYYGAFNATANKDFLSPRDPDGHGSHTASTAVGRRVLGASALGGFA 258
Query: 245 NGTIKGGSPRSRVATYKACWSLTDVVD-----CFGADVLAAIDQAIYDGADLISVSAGGK 299
G+ GG+P +R+A YKACW+ + C D+LAAID AI DG +IS+S G
Sbjct: 259 KGSASGGAPLARLAIYKACWAKPNAEKVEGNICLEEDMLAAIDDAIADGVHVISISIG-- 316
Query: 300 PNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDF 359
T P D I++GA HA+ RNI++ ASAGN GP PG+++N+APW+ TV ASTLDR F
Sbjct: 317 -TTEPFPFTQDGIAMGALHAVKRNIVVAASAGNSGPKPGTLSNLAPWIITVGASTLDRAF 375
Query: 360 SSVMTINNKTLTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGK 419
+ + N ++ ++ +++ + + C P +L P V+GK
Sbjct: 376 VGGLVLGNGYTIKTDSITAFKMDKFAPLVYASNVVVPGIALNETSQCLPNSLKPELVSGK 435
Query: 420 VVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITT 479
VV C R G + I +G E AG G+I+ N +G + ++ H V T T
Sbjct: 436 VVLCLR-GAGSRIGKGMEVKRAGGAGMILGN-IAANGNEVPSDSHFVPTAG-------VT 486
Query: 480 PKGSEITPEDIKT--NATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPG 537
P + E IKT N + P + + AP M FSSRGPN V P ILKPD+TAPG
Sbjct: 487 PTVVDKILEYIKTDKNPKAFIKPGKTVYKYQAAPSMTGFSSRGPNVVDPNILKPDITAPG 546
Query: 538 VNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKS 597
+ ILAA+S S S + D R +NI GTSMSCPHV G L+K +HP WS AAI+S
Sbjct: 547 LYILAAWSGADSPSKMSVDQRVA-GYNIYSGTSMSCPHVAGAIALLKAIHPKWSSAAIRS 605
Query: 598 AIMTT 602
A+MTT
Sbjct: 606 ALMTT 610
Score = 46.2 bits (108), Expect = 5e-05
Identities = 26/76 (34%), Positives = 39/76 (51%), Gaps = 5/76 (6%)
Query: 43 QCYIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAI 102
Q YIVY G H ++ H+ L S+ S E+A +++YSY INGFAA
Sbjct: 25 QVYIVYFGEHKGDKAFHEIEEH-----HHSYLQSVKESEEDARASLLYSYKHSINGFAAE 79
Query: 103 LEEEEAAQLANCTQHV 118
L ++A++L + V
Sbjct: 80 LTPDQASKLEKLAEVV 95
>At4g26330 subtilisin protease - like
Length = 746
Score = 275 bits (703), Expect = 5e-74
Identities = 200/573 (34%), Positives = 289/573 (49%), Gaps = 83/573 (14%)
Query: 80 SHEEAEEAIIYSYNKQINGFAAILEEEEAAQLANCTQHV------------------LGN 121
S ++AE++++YSYN GF+A L +AA LA Q + LG
Sbjct: 13 SKDDAEQSMLYSYNNGFLGFSAKLNSTQAASLAKLNQVITVFKSKSLKLHTTRSWDFLGL 72
Query: 122 SLD*VQMMST-QLGKREGLVK--------IPL*LTLIQVFG-----RNRRVLTTEE*VQF 167
++D + QL +V I L L L+ + G + R + +
Sbjct: 73 AVDNARRTPPPQLAYGSDIVVGIFDTGLFISLKLLLLSILGIWPESESFRETPEAKPIPS 132
Query: 168 H*DGVGILLRKFHVTVYLNGLPRKLIGARFFNKAYEAFHGKLPSSQ----QTARDFVGPG 223
+G + F +V+ N RKLIGARF+ + +E +G + ++ ++ RD++G G
Sbjct: 133 SWNGKCVGGEDFDPSVHCN---RKLIGARFYLRGFEETYGTIDFTRDPEYRSPRDYLGHG 189
Query: 224 THTLSTAGGNFVQNAT-IFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAID 282
THT STA G+ V+N + FG+G GT +GG+P +R+A +K CW C AD+LAA D
Sbjct: 190 THTASTAVGSVVRNVSGFFGLGRGTARGGAPLARLAVFKTCWGKDLEGVCTEADILAAFD 249
Query: 283 QAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTN 342
AI+DG +IS S G P +P F IGAFHA R I +V S GN+GP PG V N
Sbjct: 250 DAIHDGVHVISASFGYSPPLSP--FFESSADIGAFHAAERGISVVFSTGNDGPDPGVVQN 307
Query: 343 VAPWVFTVAASTLDRDFSSVMTINNK-TLTGASLFVNLPPNQDFLIIISTDAKFANVTDV 401
VAPW +VAAST+DR F + + I+ TLTG SL +Q+ ++ + N
Sbjct: 308 VAPWAVSVAASTVDRSFPTRIVIDGSFTLTGQSLI-----SQEITGTLALATTYFN---- 358
Query: 402 DAQFCRPGTLDPSKVNGKVVAC-DREGKINSIAEGQ-EALSAGAVGVIMRNQPEVDGKTL 459
C+ N ++ C G + I E Q A+ A A+ +I P + L
Sbjct: 359 -GGVCKWENWMKKLANETIILCFSTLGPVQFIEEAQAAAIRANALALIFAASPT---RQL 414
Query: 460 LAEPHVVST----------INYYDARSITTPKGSEITPEDIKTNATIRMSPANALNGRKP 509
E ++ T I Y ARS T P +++ P+ + G
Sbjct: 415 AEEVDMIPTVRVDILHGTRIRNYLARSPTVP--------------MVKIGPSKTVIGETT 460
Query: 510 APVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGT 569
AP +A FSSRGP+ + P ILKPD+TAPG+ ILAA+ + L+ + R +N Q GT
Sbjct: 461 APSVAYFSSRGPSSLSPDILKPDITAPGIGILAAWP-PRTPPTLLPGDHRSIEWNFQSGT 519
Query: 570 SMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
SMSCPHV G L+++ HP+WSP+AI+SAIMTT
Sbjct: 520 SMSCPHVAGVMALLQSAHPDWSPSAIRSAIMTT 552
>At1g04110 putative subtilisin protease
Length = 775
Score = 275 bits (703), Expect = 5e-74
Identities = 199/554 (35%), Positives = 284/554 (50%), Gaps = 52/554 (9%)
Query: 76 SILGSHEEAEEA---IIYSYNKQINGFAAILEEEEAAQLANCTQHVLGNSLD*VQMMSTQ 132
++LG EE EE ++YSY I GFAA L E EA L + V +Q+ +T
Sbjct: 56 AVLGVEEEEEEPSSRLLYSYGSAIEGFAAQLTESEAEILRYSPEVVAVRPDHVLQVQTTY 115
Query: 133 LGKREGLVKIPL*LTLIQVFGRNR-------RVLTT---EE*VQFH*DGVGILLRKFH-- 180
K GL V+ ++R VL T E F G+ + RK+
Sbjct: 116 SYKFLGLDGFGN----SGVWSKSRFGQGTIIGVLDTGVWPESPSFDDTGMPSIPRKWKGI 171
Query: 181 ----VTVYLNGLPRKLIGARFFNKAYEAFHG-----KLPSSQQTARDFVGPGTHTLSTAG 231
+ + RKLIGARFF + + + +P +ARD G GTHT ST G
Sbjct: 172 CQEGESFSSSSCNRKLIGARFFIRGHRVANSPEESPNMPREYISARDSTGHGTHTASTVG 231
Query: 232 GNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADL 291
G+ V A + G G G +G +P + +A YK CW C+ +D+LAAID AI D D+
Sbjct: 232 GSSVSMANVLGNGAGVARGMAPGAHIAVYKVCW----FNGCYSSDILAAIDVAIQDKVDV 287
Query: 292 ISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVA 351
+S+S GG P ++ D I+IG F A+ R I ++ +AGN GP SV N APWV T+
Sbjct: 288 LSLSLGGFPIP----LYDDTIAIGTFRAMERGISVICAAGNNGPIESSVANTAPWVSTIG 343
Query: 352 ASTLDRDFSSVMTI-NNKTLTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGT 410
A TLDR F +V+ + N K L G SL+ P + + D ++FC G+
Sbjct: 344 AGTLDRRFPAVVRLANGKLLYGESLY---PGKGIKNAGREVEVIYVTGGDKGSEFCLRGS 400
Query: 411 LDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVV--ST 468
L ++ GK+V CDR G +G+ AG V +I+ N E++ + + H++ +
Sbjct: 401 LPREEIRGKMVICDR-GVNGRSEKGEAVKEAGGVAMILAN-TEINQEEDSIDVHLLPATL 458
Query: 469 INYYDARSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYI 528
I Y ++ + + + P+ R+ + GR AP +A FS+RGP+ P I
Sbjct: 459 IGYTESVLLKAYVNATVKPK-------ARIIFGGTVIGRSRAPEVAQFSARGPSLANPSI 511
Query: 529 LKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHP 588
LKPD+ APGVNI+AA+ + L D+RR F + GTSMSCPHV G LI++ +P
Sbjct: 512 LKPDMIAPGVNIIAAWPQNLGPTGLPYDSRR-VNFTVMSGTSMSCPHVSGITALIRSAYP 570
Query: 589 NWSPAAIKSAIMTT 602
NWSPAAIKSA+MTT
Sbjct: 571 NWSPAAIKSALMTT 584
>At1g01900 putative subtilisin-like serine protease
Length = 774
Score = 272 bits (696), Expect = 3e-73
Identities = 173/418 (41%), Positives = 237/418 (56%), Gaps = 29/418 (6%)
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQ--QTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGT 247
+K+IGA F K YE+ GK+ + ++ RD G GTHT STA G+ V A FG G
Sbjct: 191 KKIIGASAFYKGYESIVGKINETTDFRSTRDAQGHGTHTASTAAGDIVPKANYFGQAKGL 250
Query: 248 IKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVI 307
G SR+A YKACW+L C DV+AAID+AI DG D+IS+S GG
Sbjct: 251 ASGMRFTSRIAAYKACWAL----GCASTDVIAAIDRAILDGVDVISLSLGGSSRP----F 302
Query: 308 FTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTINN 367
+ D I+I F A+ +NI + SAGN GPT +V+N APW+ TVAAS DR F +++ I N
Sbjct: 303 YVDPIAIAGFGAMQKNIFVSCSAGNSGPTASTVSNGAPWLMTVAASYTDRTFPAIVRIGN 362
Query: 368 -KTLTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDRE 426
K+L G+SL+ L T + + A FC +L V GK+V C R
Sbjct: 363 RKSLVGSSLYKGKSLKNLPLAFNRTAGE-----ESGAVFCIRDSLKRELVEGKIVICLR- 416
Query: 427 GKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTIN--YYDARSITTPKGSE 484
G A+G+E +G +++ + E +G+ LLA+PHV+ ++ + D +++
Sbjct: 417 GASGRTAKGEEVKRSGGAAMLLVST-EAEGEELLADPHVLPAVSLGFSDGKTLLNYLAGA 475
Query: 485 ITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAY 544
NAT + G AP++A+FSSRGP+ P I KPD+ APG+NILA +
Sbjct: 476 -------ANATASVRFRGTAYGAT-APMVAAFSSRGPSVAGPEIAKPDIAAPGLNILAGW 527
Query: 545 SLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
S +S S L +D RR FNI GTSM+CPH+ G A LIK++H +WSPA IKSAIMTT
Sbjct: 528 SPFSSPSLLRSDPRR-VQFNIISGTSMACPHISGIAALIKSVHGDWSPAMIKSAIMTT 584
>At1g20160 unknown protein
Length = 769
Score = 270 bits (690), Expect = 2e-72
Identities = 163/417 (39%), Positives = 238/417 (56%), Gaps = 26/417 (6%)
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIK 249
RK+IGAR++ + S T RD +G G+H ST G+ V+NA+ +G+ +GT K
Sbjct: 184 RKIIGARYYKNPDD------DSEYYTTRDVIGHGSHVSSTIAGSAVENASYYGVASGTAK 237
Query: 250 GGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFT 309
GGS +R+A YK C + C G+ +LAA D AI DG D++S+S G + + T
Sbjct: 238 GGSQNARIAMYKVC----NPGGCTGSSILAAFDDAIADGVDVLSLSLGAPAYARID-LNT 292
Query: 310 DEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDF-SSVMTINNK 368
D I+IGAFHA+ + IL++ SAGN+GP G+VTN APW+ TVAA+T+DRDF S V+ NK
Sbjct: 293 DPIAIGAFHAVEQGILVICSAGNDGPDGGTVTNTAPWIMTVAANTIDRDFESDVVLGGNK 352
Query: 369 TLTGASL-FVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDR-E 426
+ G + F N+ + + +I AK A+ ++ A+ C +LD KV GK+V C+
Sbjct: 353 VIKGEGIHFSNVSKSPVYPLIHGKSAKSADASEGSARACDSDSLDQEKVKGKIVLCENVG 412
Query: 427 GKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEIT 486
G + + E S G G + VD +T V S + I + + +EI
Sbjct: 413 GSYYASSARDEVKSKGGTGCVF-----VDDRTRA----VASAYGSFPTTVIDSKEAAEIF 463
Query: 487 PEDIKTNATI-RMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYS 545
T + + P + PAP +A FSSRGP+ + ILKPD+TAPGV+ILAA++
Sbjct: 464 SYLNSTKDPVATILPTATVEKFTPAPAVAYFSSRGPSSLTRSILKPDITAPGVSILAAWT 523
Query: 546 LLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
+ S++ + + +N+ GTSM+ PHV A LIK+ HP W P+AI+SAIMTT
Sbjct: 524 --GNDSSISLEGKPASQYNVISGTSMAAPHVSAVASLIKSQHPTWGPSAIRSAIMTT 578
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,104,204
Number of Sequences: 26719
Number of extensions: 554930
Number of successful extensions: 1546
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 57
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1165
Number of HSP's gapped (non-prelim): 133
length of query: 611
length of database: 11,318,596
effective HSP length: 105
effective length of query: 506
effective length of database: 8,513,101
effective search space: 4307629106
effective search space used: 4307629106
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC133779.1


