

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133341.5 - phase: 1 /pseudo
(808 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
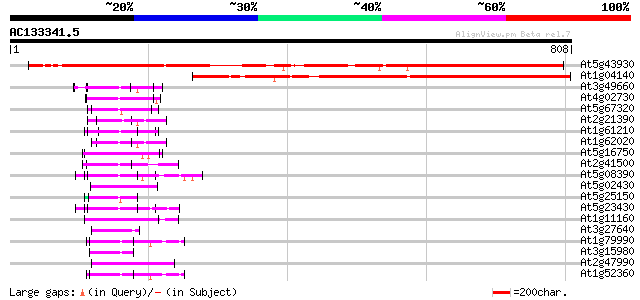
Score E
Sequences producing significant alignments: (bits) Value
At5g43930 unknown protein 742 0.0
At1g04140 unknown protein 520 e-147
At3g49660 putative WD-40 repeat - protein 69 8e-12
At4g02730 putative WD-repeat protein 59 1e-08
At5g67320 unknown protein 56 9e-08
At2g21390 coatomer alpha subunit 55 2e-07
At1g61210 54 3e-07
At1g62020 hypothetical protein 52 1e-06
At5g16750 WD40-repeat protein 51 2e-06
At2g41500 putative U4/U6 small nuclear ribonucleoprotein 51 3e-06
At5g08390 katanin p80 subunit - like protein 50 4e-06
At5g02430 putative protein 50 4e-06
At5g25150 transcription initiation factor IID-associated factor-... 50 5e-06
At5g23430 unknown protein 50 7e-06
At1g11160 hypothetical protein 49 9e-06
At3g27640 unknown protein 49 1e-05
At1g79990 coatomer protein like complex, subunit beta 2 (beta pr... 48 2e-05
At3g15980 putative coatomer complex subunit 47 3e-05
At2g47990 unknown protein 47 3e-05
At1g52360 coatomer complex subunit, putative 47 3e-05
>At5g43930 unknown protein
Length = 726
Score = 742 bits (1916), Expect = 0.0
Identities = 421/782 (53%), Positives = 506/782 (63%), Gaps = 104/782 (13%)
Query: 28 NVFNLLARREISPRTKHVARDAKRGLLSWYLNYFSL*TISTKM*AS*ITTSLKMILRVEA 87
+VFN+L +RE+SP+ K V R + G WY + S T + M T + VEA
Sbjct: 36 SVFNMLVQREMSPKAKFVPRK-RWGKSRWYTDS-SCGTNNESM----KETGQSLTSWVEA 89
Query: 88 DSLRHLSAKYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMR 147
+SL+HLSAKYCPL PRSTIAAAFS DG+ LASTHGDHTVKIIDCETG CLKVL GH R
Sbjct: 90 ESLQHLSAKYCPLGAPPRSTIAAAFSTDGRTLASTHGDHTVKIIDCETGNCLKVLTGHRR 149
Query: 148 TPWVVRFHPLHPKILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVAS 207
TPWVVRFHP H +I+ASGSLD EVRLW+ TSECI SH FYRPIASIAFHA+GE++AVAS
Sbjct: 150 TPWVVRFHPHHSEIVASGSLDLEVRLWNTTTSECIRSHLFYRPIASIAFHAEGELLAVAS 209
Query: 208 GHKLYIWHYDKKGEASYSPIFVLKTRRSLRAVHFHPHAAPYLLTAEVNDLDSSDSSMTEA 267
GHKL++WHY+++GE S SP VLKTRRSLRAVHFHPH AP LLTAEVN++DS DSSM+ A
Sbjct: 210 GHKLHMWHYNRRGEGS-SPTVVLKTRRSLRAVHFHPHGAPLLLTAEVNEIDSLDSSMSRA 268
Query: 268 TSIGYLQYPPPAVFVTNVHPTEHVTLSSEPTNVSLPFFLVPPYTVDESRAELQHASHDAG 327
TS+GYL+YPPPA+ T+ +
Sbjct: 269 TSMGYLRYPPPAILFTSTESNQ-------------------------------------- 290
Query: 328 SGRIQIESSAVAQFQADTNSTEQHDTTVSPMDTVSEIPTNSQAGTEYPAHTAFSNGMGIG 387
+S A+ + T+S T S + +P NS SN
Sbjct: 291 -------TSLAAENENRTSSPPLPLATSSGPSGPNSVPGNSP-----------SNIFLTR 332
Query: 388 IGNLT---MDGMETDETRPAEGSQHRNPTDASSLNGMLHGLSRQTANHGVHPEDGH--PF 442
G+ T +DGM+ DE +P RN G+ Q +N PE G
Sbjct: 333 AGDRTSPAVDGMDVDEAQPVG----RN------------GIPSQVSNRSDFPELGQIRQL 376
Query: 443 VSSRDPSGWELPFLQGWMMGQSQAGLPSMLPHTGVSRDTLAPQISSSVMANTLPTSNADV 502
RD WELPFLQGW+M Q ++ TG S ++ S T++ +
Sbjct: 377 FHFRDRVSWELPFLQGWLMAQGHGVANPVVTPTGSSNHGISAPSS---------TASLEA 427
Query: 503 AMPSSAMSGSINIPGSSVRSGLRSHFSH---SRTPVSESGNLAASINTPHDGSDIQTIMS 559
A+ + +N+ S R G + S SRT + E +S NT H GSD Q +++
Sbjct: 428 AVALLEIPSGVNLHAVSRRGGAQEQTSQPQFSRTGLPEG---VSSRNTQH-GSDAQPVVN 483
Query: 560 RIQSELATSVAA----AAATELPCTVKLRVWSHDIKNPCSPLNADRCRLIIPHAVLCSEM 615
R+QSELATS+AA AAA ELPCTVKLR+WSHDIK+P + L +DRC IPHAVLCSEM
Sbjct: 484 RVQSELATSIAASAAAAAAAELPCTVKLRMWSHDIKDPYAQLKSDRCLFTIPHAVLCSEM 543
Query: 616 GAHFSPCGRFLAACVACMLPHIEADPGLQTPVHQESGIATSPTRHPISAHQVMYELRIYS 675
GAH+SPCGR+LAACVAC+ PH E DPGLQT Q+SG+ATSPTRHP++AHQV+YELR+YS
Sbjct: 544 GAHYSPCGRYLAACVACVFPHGEIDPGLQTQAQQDSGLATSPTRHPVTAHQVIYELRVYS 603
Query: 676 LEEATFGLVLASRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYT 735
L++ +FG VL SRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLL+SIV DGETT +T
Sbjct: 604 LQKESFGSVLVSRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLRSIVSDGETTSHFFT 663
Query: 736 VLEVYRVSDMELVRVLPSAEDEVNVACFHPFPGGGLVYGTKEGKLRILHYDGARPVNGTG 795
VLE+YRVSDMELVRVLPS+EDEVNVACFHP PGGGLVYGTKEGKLRI Y+ A N TG
Sbjct: 664 VLEIYRVSDMELVRVLPSSEDEVNVACFHPSPGGGLVYGTKEGKLRIFQYNTAATSNFTG 723
Query: 796 PS 797
P+
Sbjct: 724 PN 725
>At1g04140 unknown protein
Length = 529
Score = 520 bits (1338), Expect = e-147
Identities = 284/547 (51%), Positives = 352/547 (63%), Gaps = 28/547 (5%)
Query: 264 MTEATSIGYLQYPPPAVFVTNVHPTEHVTLSSEPTNVSLPFFLVPPYTVDESRAELQHAS 323
MT +TS GYL+YPPPA+F TN +L++E V LP+ L+P Y+ D+ R + ++S
Sbjct: 1 MTRSTSPGYLRYPPPAIFFTNTQSGSRTSLAAELPLVPLPYLLLPSYSADDPR--ILYSS 58
Query: 324 HDAGSGRIQIESSAVAQFQADTNSTEQHDTTVSPMDTVSEIPTNSQAGTEYPAHTA---F 380
G Q +FQ++ +S E T+SP + P ++A F
Sbjct: 59 GTTGPRNAQ------TRFQSNQSSVEHGSRTISPSPLPMATSADLSGSYHVPDNSASNTF 112
Query: 381 SNGMGIGIGNLTMDGMETDETRPAEGSQHRNPTDASSLNGMLHGLSRQTANHGVHPEDGH 440
+ G +D M+ DE +P ++R P+ SS +L Q H
Sbjct: 113 ATQAGARNSTTAVDAMDVDEAQPV--GRNRVPSQVSSQPDLLEFGQLQQLFH-------- 162
Query: 441 PFVSSRDPSGWELPFLQGWMMGQSQAGLPSMLPHTGVSRDTLAPQISSSVMANTLPTSNA 500
RD WELPFLQGW+M QSQAG S+ TG S + S A+ T++
Sbjct: 163 ----FRDRGSWELPFLQGWLMAQSQAGANSVALPTGSSGHVNSTPYMGSSSASHSSTASL 218
Query: 501 DVAMPSSAMSGSINIPGSSVRSGLRSHFSHSRTPVSESGNLAASINTPHDGSDIQTIMSR 560
+ + S + G +N+ G S R R SR S +S NT H+G+D Q +++R
Sbjct: 219 EAGVASLEIPGGVNLYGVSARGDSRDRILQSRFAGSGLAEGRSSRNTQHEGADAQPVVNR 278
Query: 561 IQSELATSVAAAAATELPCTVKLRVWSHDIKNPCSPLNADRCRLIIPHAVLCSEMGAHFS 620
I SELA+S+AAA ELPCTVKLRVWSHDIK+PCS L +D+CRL I HAVLCSEMGAHFS
Sbjct: 279 IPSELASSIAAA---ELPCTVKLRVWSHDIKDPCSILKSDKCRLTIHHAVLCSEMGAHFS 335
Query: 621 PCGRFLAACVACMLPHIEADPGLQTPVHQESGIATSPTRHPISAHQVMYELRIYSLEEAT 680
PCGR+LAACVAC++PH E DP LQT V Q+SG+ATSPTRHP++AHQVMYELR+YSLE+ +
Sbjct: 336 PCGRYLAACVACVIPHAETDPSLQTLVQQDSGLATSPTRHPVTAHQVMYELRVYSLEKES 395
Query: 681 FGLVLASRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYTVLEVY 740
FG VL SRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIV DGETT +TVLE+Y
Sbjct: 396 FGSVLVSRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVSDGETTSHFFTVLEIY 455
Query: 741 RVSDMELVRVLPSAEDEVNVACFHPFPGGGLVYGTKEGKLRILHYDGARPVNGTGPSYFP 800
RVSDMELVRVLPS+EDEVNVACFHP PGGGLVYGTKEGKLRI Y+ A N T P+ P
Sbjct: 456 RVSDMELVRVLPSSEDEVNVACFHPSPGGGLVYGTKEGKLRIFRYNTAAASNLTAPNSSP 515
Query: 801 EETIVGV 807
+E + V
Sbjct: 516 DENLAEV 522
>At3g49660 putative WD-40 repeat - protein
Length = 317
Score = 69.3 bits (168), Expect = 8e-12
Identities = 35/98 (35%), Positives = 55/98 (55%), Gaps = 2/98 (2%)
Query: 111 AFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQE 170
AFS D + + S D T+K+ D ETG +K LIGH + V F+P ++ SGS D+
Sbjct: 78 AFSSDARFIVSASDDKTLKLWDVETGSLIKTLIGHTNYAFCVNFNP-QSNMIVSGSFDET 136
Query: 171 VRLWDANTSECITSHHFYR-PIASIAFHAKGEIIAVAS 207
VR+WD T +C+ + P+ ++ F+ G +I +S
Sbjct: 137 VRIWDVTTGKCLKVLPAHSDPVTAVDFNRDGSLIVSSS 174
Score = 50.1 bits (118), Expect = 5e-06
Identities = 38/133 (28%), Positives = 66/133 (49%), Gaps = 8/133 (6%)
Query: 94 SAKYCPLLPAPRSTIAAA-FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTP-WV 151
+ K +LPA + A F+ DG ++ S+ D +I D TG C+K LI P
Sbjct: 144 TGKCLKVLPAHSDPVTAVDFNRDGSLIVSSSYDGLCRIWDSGTGHCVKTLIDDENPPVSF 203
Query: 152 VRFHPLHPKILASGSLDQEVRLWDANTSECI---TSHHFYRPIASIAFHAKG--EIIAVA 206
VRF P + K + G+LD +RLW+ ++++ + T H + S AF I++ +
Sbjct: 204 VRFSP-NGKFILVGTLDNTLRLWNISSAKFLKTYTGHVNAQYCISSAFSVTNGKRIVSGS 262
Query: 207 SGHKLYIWHYDKK 219
+ +++W + K
Sbjct: 263 EDNCVHMWELNSK 275
Score = 46.6 bits (109), Expect = 6e-05
Identities = 30/84 (35%), Positives = 47/84 (55%), Gaps = 11/84 (13%)
Query: 92 HLSAKYCPLLPAPRSTIAAAFS-PDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPW 150
H++A+YC I++AFS +GK + S D+ V + + + + L+ L GH T
Sbjct: 239 HVNAQYC---------ISSAFSVTNGKRIVSGSEDNCVHMWELNSKKLLQKLEGHTETVM 289
Query: 151 VVRFHPLHPKILASGSLDQEVRLW 174
V HP ++ASGSLD+ VR+W
Sbjct: 290 NVACHPTE-NLIASGSLDKTVRIW 312
Score = 43.1 bits (100), Expect = 6e-04
Identities = 29/112 (25%), Positives = 55/112 (48%), Gaps = 4/112 (3%)
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVR--FHPLHPKILASGSLDQ 169
FSP+GK + D+T+++ + + + LK GH+ + + F + K + SGS D
Sbjct: 206 FSPNGKFILVGTLDNTLRLWNISSAKFLKTYTGHVNAQYCISSAFSVTNGKRIVSGSEDN 265
Query: 170 EVRLWDANTSECITSHHFY-RPIASIAFHAKGEIIAVASGHK-LYIWHYDKK 219
V +W+ N+ + + + + ++A H +IA S K + IW K+
Sbjct: 266 CVHMWELNSKKLLQKLEGHTETVMNVACHPTENLIASGSLDKTVRIWTQKKE 317
>At4g02730 putative WD-repeat protein
Length = 333
Score = 58.5 bits (140), Expect = 1e-08
Identities = 34/99 (34%), Positives = 53/99 (53%), Gaps = 3/99 (3%)
Query: 111 AFSPDGKVLASTHGDHTVKIIDCETG-RCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQ 169
A+S D S D T++I D + CLKVL GH + V F+P ++ SGS D+
Sbjct: 92 AWSSDSHYTCSASDDCTLRIWDARSPYECLKVLRGHTNFVFCVNFNP-PSNLIVSGSFDE 150
Query: 170 EVRLWDANTSECITSHHFY-RPIASIAFHAKGEIIAVAS 207
+R+W+ T +C+ + PI+S+ F+ G +I AS
Sbjct: 151 TIRIWEVKTGKCVRMIKAHSMPISSVHFNRDGSLIVSAS 189
Score = 45.1 bits (105), Expect = 2e-04
Identities = 33/114 (28%), Positives = 51/114 (43%), Gaps = 6/114 (5%)
Query: 110 AAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVV--RFHPLHPKILASGSL 167
A FSP+GK + D T+K+ + TG+ LKV GH + + F + K + SGS
Sbjct: 219 AKFSPNGKFILVATLDSTLKLSNYATGKFLKVYTGHTNKVFCITSAFSVTNGKYIVSGSE 278
Query: 168 DQEVRLWDANTSECITSHHFYR-PIASIAFHAKGEIIAVASGH---KLYIWHYD 217
D V LWD + + + S++ H I+ + H + IW D
Sbjct: 279 DNCVYLWDLQARNILQRLEGHTDAVISVSCHPVQNEISSSGNHLDKTIRIWKQD 332
Score = 33.1 bits (74), Expect = 0.64
Identities = 22/64 (34%), Positives = 35/64 (54%), Gaps = 4/64 (6%)
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWV--VRFHPLHPKILASGSLDQ 169
F+ DG ++ S D + KI D + G CLK LI ++P V +F P + K + +LD
Sbjct: 178 FNRDGSLIVSASHDGSCKIWDAKEGTCLKTLIDD-KSPAVSFAKFSP-NGKFILVATLDS 235
Query: 170 EVRL 173
++L
Sbjct: 236 TLKL 239
>At5g67320 unknown protein
Length = 613
Score = 55.8 bits (133), Expect = 9e-08
Identities = 38/113 (33%), Positives = 56/113 (48%), Gaps = 10/113 (8%)
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHP--------KILA 163
+ P G +LAS D T KI + + + L H + + +R+ P P LA
Sbjct: 456 WDPTGSLLASCSDDSTAKIWNIKQSTFVHDLREHTKEIYTIRWSPTGPGTNNPNKQLTLA 515
Query: 164 SGSLDQEVRLWDANTSECITSHHFYR-PIASIAFHAKGEIIAVASGHK-LYIW 214
S S D V+LWDA + + S + +R P+ S+AF GE IA S K ++IW
Sbjct: 516 SASFDSTVKLWDAELGKMLCSFNGHREPVYSLAFSPNGEYIASGSLDKSIHIW 568
Score = 45.1 bits (105), Expect = 2e-04
Identities = 26/86 (30%), Positives = 44/86 (50%), Gaps = 1/86 (1%)
Query: 119 LASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANT 178
LAS D TVK+ D E G+ L GH + + F P + + +ASGSLD+ + +W
Sbjct: 514 LASASFDSTVKLWDAELGKMLCSFNGHREPVYSLAFSP-NGEYIASGSLDKSIHIWSIKE 572
Query: 179 SECITSHHFYRPIASIAFHAKGEIIA 204
+ + ++ I + ++ +G IA
Sbjct: 573 GKIVKTYTGNGGIFEVCWNKEGNKIA 598
Score = 36.6 bits (83), Expect = 0.058
Identities = 31/119 (26%), Positives = 46/119 (38%), Gaps = 24/119 (20%)
Query: 109 AAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPK-------- 160
A A+SP +LAS GD T +I G V G +++ H K
Sbjct: 270 ACAWSPSASLLASGSGDATARIWSIPEGSFKAVHTGRNINALILK----HAKGKSNEKSK 325
Query: 161 ------------ILASGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVAS 207
+LA+GS D + R+W N T PI S+ ++ KG+ + S
Sbjct: 326 DVTTLDWNGEGTLLATGSCDGQARIWTLNGELISTLSKHKGPIFSLKWNKKGDYLLTGS 384
>At2g21390 coatomer alpha subunit
Length = 1218
Score = 54.7 bits (130), Expect = 2e-07
Identities = 36/106 (33%), Positives = 57/106 (52%), Gaps = 7/106 (6%)
Query: 125 DHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANTSECI-- 182
D+ +K+ + +T RCL L+GH+ V+FH +P I+ S S DQ +R+W+ + CI
Sbjct: 72 DYKIKVWNYKTHRCLFTLLGHLDYIRTVQFHHENPWIV-SASDDQTIRIWNWQSRTCISV 130
Query: 183 -TSHHFYRPIASIAFHAKGEIIAVAS-GHKLYIWHYDKKGEASYSP 226
T H+ Y AS FH K +++ AS + +W + S SP
Sbjct: 131 LTGHNHYVMCAS--FHPKEDLVVSASLDQTVRVWDIGALKKKSASP 174
Score = 43.9 bits (102), Expect = 4e-04
Identities = 22/64 (34%), Positives = 33/64 (51%), Gaps = 1/64 (1%)
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
F + + S D T++I + ++ C+ VL GH FHP ++ S SLDQ V
Sbjct: 101 FHHENPWIVSASDDQTIRIWNWQSRTCISVLTGHNHYVMCASFHP-KEDLVVSASLDQTV 159
Query: 172 RLWD 175
R+WD
Sbjct: 160 RVWD 163
Score = 42.0 bits (97), Expect = 0.001
Identities = 29/83 (34%), Positives = 43/83 (50%), Gaps = 5/83 (6%)
Query: 141 VLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANTS---ECITSHHFYRPIASIAFH 197
VL GH R FHP P ++ SG+ D++V+LW N + E T ++S+ FH
Sbjct: 199 VLEGHDRGVNWASFHPTLP-LIVSGADDRQVKLWRMNETKAWEVDTLRGHMNNVSSVMFH 257
Query: 198 AKGEIIAVASGHK-LYIWHYDKK 219
AK +II S K + +W K+
Sbjct: 258 AKQDIIVSNSEDKSIRVWDATKR 280
Score = 33.5 bits (75), Expect = 0.49
Identities = 25/106 (23%), Positives = 47/106 (43%), Gaps = 8/106 (7%)
Query: 118 VLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDAN 177
+LAS H +++ D G + H V FH P + SG D ++++W+
Sbjct: 24 ILASLHSG-VIQLWDYRMGTLIDRFDEHEGPVRGVHFHNSQP-LFVSGGDDYKIKVWNYK 81
Query: 178 TSEC---ITSHHFYRPIASIAFHAKGE-IIAVASGHKLYIWHYDKK 219
T C + H Y I ++ FH + I++ + + IW++ +
Sbjct: 82 THRCLFTLLGHLDY--IRTVQFHHENPWIVSASDDQTIRIWNWQSR 125
>At1g61210
Length = 282
Score = 53.9 bits (128), Expect = 3e-07
Identities = 33/103 (32%), Positives = 49/103 (47%), Gaps = 2/103 (1%)
Query: 109 AAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLD 168
A F P G+ LAS D +KI D C++ GH R +RF P + + SG LD
Sbjct: 135 AVEFHPFGEFLASGSSDANLKIWDIRKKGCIQTYKGHSRGISTIRFTP-DGRWVVSGGLD 193
Query: 169 QEVRLWDANTSECITSHHFYR-PIASIAFHAKGEIIAVASGHK 210
V++WD + + F+ PI S+ FH ++A S +
Sbjct: 194 NVVKVWDLTAGKLLHEFKFHEGPIRSLDFHPLEFLLATGSADR 236
Score = 46.6 bits (109), Expect = 6e-05
Identities = 28/73 (38%), Positives = 39/73 (53%), Gaps = 1/73 (1%)
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
F+PDG+ + S D+ VK+ D G+ L H + FHPL +LA+GS D+ V
Sbjct: 180 FTPDGRWVVSGGLDNVVKVWDLTAGKLLHEFKFHEGPIRSLDFHPLE-FLLATGSADRTV 238
Query: 172 RLWDANTSECITS 184
+ WD T E I S
Sbjct: 239 KFWDLETFELIGS 251
Score = 43.9 bits (102), Expect = 4e-04
Identities = 25/89 (28%), Positives = 45/89 (50%), Gaps = 3/89 (3%)
Query: 128 VKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANTSECITSHHF 187
+K+ D E + ++ GH V FHP + LASGS D +++WD CI ++
Sbjct: 112 IKLWDVEEAKMVRAFTGHRSNCSAVEFHPF-GEFLASGSSDANLKIWDIRKKGCIQTYKG 170
Query: 188 Y-RPIASIAFHAKGE-IIAVASGHKLYIW 214
+ R I++I F G +++ + + +W
Sbjct: 171 HSRGISTIRFTPDGRWVVSGGLDNVVKVW 199
>At1g62020 hypothetical protein
Length = 1216
Score = 52.0 bits (123), Expect = 1e-06
Identities = 33/106 (31%), Positives = 56/106 (52%), Gaps = 7/106 (6%)
Query: 125 DHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANTSECI-- 182
D+ +K+ + + RCL L+GH+ V+FH +P I+ S S DQ +R+W+ + C+
Sbjct: 72 DYKIKVWNYKNHRCLFTLLGHLDYIRTVQFHHEYPWIV-SASDDQTIRIWNWQSRTCVSV 130
Query: 183 -TSHHFYRPIASIAFHAKGEIIAVAS-GHKLYIWHYDKKGEASYSP 226
T H+ Y AS FH K +++ AS + +W + + SP
Sbjct: 131 LTGHNHYVMCAS--FHPKEDLVVSASLDQTVRVWDIGALRKKTVSP 174
Score = 43.1 bits (100), Expect = 6e-04
Identities = 21/57 (36%), Positives = 31/57 (53%), Gaps = 1/57 (1%)
Query: 119 LASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWD 175
+ S D T++I + ++ C+ VL GH FHP ++ S SLDQ VR+WD
Sbjct: 108 IVSASDDQTIRIWNWQSRTCVSVLTGHNHYVMCASFHP-KEDLVVSASLDQTVRVWD 163
Score = 42.0 bits (97), Expect = 0.001
Identities = 29/83 (34%), Positives = 43/83 (50%), Gaps = 5/83 (6%)
Query: 141 VLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANTS---ECITSHHFYRPIASIAFH 197
VL GH R FHP P ++ SG+ D++V+LW N + E T ++S+ FH
Sbjct: 199 VLEGHDRGVNWAAFHPTLP-LIVSGADDRQVKLWRMNETKAWEVDTLRGHMNNVSSVMFH 257
Query: 198 AKGEIIAVASGHK-LYIWHYDKK 219
AK +II S K + +W K+
Sbjct: 258 AKQDIIVSNSEDKSIRVWDATKR 280
>At5g16750 WD40-repeat protein
Length = 876
Score = 51.2 bits (121), Expect = 2e-06
Identities = 34/113 (30%), Positives = 55/113 (48%), Gaps = 3/113 (2%)
Query: 105 RSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILAS 164
R + FS + + + GD TVKI G CLK GH + F + ++
Sbjct: 542 RRIFSVEFSTVDQCVMTASGDKTVKIWAISDGSCLKTFEGHTSSVLRASFITDGTQFVSC 601
Query: 165 GSLDQEVRLWDANTSECITSHHFYR-PIASIAFHAKGEIIAVASGHK-LYIWH 215
G+ D ++LW+ NTSECI ++ + + ++A K E+IA G + +WH
Sbjct: 602 GA-DGLLKLWNVNTSECIATYDQHEDKVWALAVGKKTEMIATGGGDAVINLWH 653
Score = 44.7 bits (104), Expect = 2e-04
Identities = 32/118 (27%), Positives = 56/118 (47%), Gaps = 11/118 (9%)
Query: 109 AAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLD 168
A A SPD K+L S +++ D ET +C++ GH + H +LA+ D
Sbjct: 65 ALALSPDDKLLFSAGHSRQIRVWDLETLKCIRSWKGHEGPVMGMACH-ASGGLLATAGAD 123
Query: 169 QEVRLWDANTSECITSHHFYR----PIASIAFHA---KGEIIAVASGHKLYIWHYDKK 219
++V +WD + C H++R ++SI FH K +I+ + + +W + K
Sbjct: 124 RKVLVWDVDGGFCT---HYFRGHKGVVSSILFHPDSNKNILISGSDDATVRVWDLNAK 178
Score = 40.8 bits (94), Expect = 0.003
Identities = 40/120 (33%), Positives = 55/120 (45%), Gaps = 11/120 (9%)
Query: 128 VKIIDCETGRCLKVLIGHMRTPWVVR--FHPLHPKILASGSLDQEVRLWDANTSECI--- 182
V++ D T C VL GH + ++ +GS D+ VRLW+A + CI
Sbjct: 383 VRVYDVATMSCSYVLAGHKEVVLSLDTCVSSSGNVLIVTGSKDKTVRLWNATSKSCIGVG 442
Query: 183 TSHHFYRPIASIAFHAKG-EIIAVASGHK-LYIWHYDKKGEASYSPIFVLKTRRSLRAVH 240
T H+ I ++AF K SG + L +W D E S PI LKT RS+ A H
Sbjct: 443 TGHN--GDILAVAFAKKSFSFFVSGSGDRTLKVWSLDGISEDSEEPI-NLKT-RSVVAAH 498
>At2g41500 putative U4/U6 small nuclear ribonucleoprotein
Length = 554
Score = 50.8 bits (120), Expect = 3e-06
Identities = 39/133 (29%), Positives = 58/133 (43%), Gaps = 15/133 (11%)
Query: 111 AFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQE 170
AF P GK L +T D T ++ D TG L + GH R+ + + F + AS LD
Sbjct: 346 AFHPSGKYLGTTSYDKTWRLWDINTGAELLLQEGHSRSVYGIAFQQ-DGALAASCGLDSL 404
Query: 171 VRLWDANTSECITSHHFY-RPIASIAFHAKGEIIAVASGHKLYIWHYDKKGEASYSPIFV 229
R+WD T I + +P+ S+ F G +H GE + I+
Sbjct: 405 ARVWDLRTGRSILVFQGHIKPVFSVNFSPNG-------------YHLASGGEDNQCRIWD 451
Query: 230 LKTRRSLRAVHFH 242
L+ R+SL + H
Sbjct: 452 LRMRKSLYIIPAH 464
Score = 47.8 bits (112), Expect = 3e-05
Identities = 27/71 (38%), Positives = 37/71 (52%), Gaps = 1/71 (1%)
Query: 105 RSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILAS 164
RS AF DG + AS D ++ D TGR + V GH++ + V F P + LAS
Sbjct: 382 RSVYGIAFQQDGALAASCGLDSLARVWDLRTGRSILVFQGHIKPVFSVNFSP-NGYHLAS 440
Query: 165 GSLDQEVRLWD 175
G D + R+WD
Sbjct: 441 GGEDNQCRIWD 451
Score = 39.7 bits (91), Expect = 0.007
Identities = 28/99 (28%), Positives = 44/99 (44%), Gaps = 5/99 (5%)
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
FSP+G LAS D+ +I D + L ++ H V++ P LA+ S D +V
Sbjct: 431 FSPNGYHLASGGEDNQCRIWDLRMRKSLYIIPAHANLVSQVKYEPQEGYFLATASYDMKV 490
Query: 172 RLW---DANTSECITSHHFYRPIASIAFHAKGEIIAVAS 207
+W D + + + H +AS+ A IA S
Sbjct: 491 NIWSGRDFSLVKSLAGHE--SKVASLDITADSSCIATVS 527
Score = 39.7 bits (91), Expect = 0.007
Identities = 31/119 (26%), Positives = 51/119 (42%), Gaps = 3/119 (2%)
Query: 105 RSTIAAAFSPDGKVLASTHGDHTVKIIDC-ETGRCLKVLIGHMRTPWVVRFHPLHPKILA 163
R +FS DGK+LA+ K+ + + + VL H V F P+ LA
Sbjct: 256 RPLTGCSFSRDGKILATCSLSGVTKLWEMPQVTNTIAVLKDHKERATDVVFSPVDD-CLA 314
Query: 164 SGSLDQEVRLWDANTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLY-IWHYDKKGE 221
+ S D+ +LW + + T +A +AFH G+ + S K + +W + E
Sbjct: 315 TASADRTAKLWKTDGTLLQTFEGHLDRLARVAFHPSGKYLGTTSYDKTWRLWDINTGAE 373
>At5g08390 katanin p80 subunit - like protein
Length = 823
Score = 50.4 bits (119), Expect = 4e-06
Identities = 28/73 (38%), Positives = 39/73 (53%), Gaps = 1/73 (1%)
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
F+PDG+ + S D+ VK+ D G+ L H + FHP H +LA+GS D+ V
Sbjct: 244 FTPDGRWIVSGGEDNVVKVWDLTAGKLLHEFKSHEGKIQSLDFHP-HEFLLATGSADKTV 302
Query: 172 RLWDANTSECITS 184
+ WD T E I S
Sbjct: 303 KFWDLETFELIGS 315
Score = 48.9 bits (115), Expect = 1e-05
Identities = 44/182 (24%), Positives = 81/182 (44%), Gaps = 21/182 (11%)
Query: 109 AAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLD 168
+ F ++A+ T+K+ D E + ++ L GH V FHP + ASGSLD
Sbjct: 157 SVTFDASEGLVAAGAASGTIKLWDLEEAKVVRTLTGHRSNCVSVNFHPF-GEFFASGSLD 215
Query: 169 QEVRLWDANTSECITSHHFY-RPIASIAFHAKGE-IIAVASGHKLYIWHYDKKGEASYSP 226
+++WD CI ++ + R + + F G I++ + + +W +
Sbjct: 216 TNLKIWDIRKKGCIHTYKGHTRGVNVLRFTPDGRWIVSGGEDNVVKVWDL-----TAGKL 270
Query: 227 IFVLKTRR-SLRAVHFHPHAAPYLL-------TAEVNDLDSSD---SSMTEATSIGYLQY 275
+ K+ ++++ FHPH +LL T + DL++ + S TE T + L +
Sbjct: 271 LHEFKSHEGKIQSLDFHPH--EFLLATGSADKTVKFWDLETFELIGSGGTETTGVRCLTF 328
Query: 276 PP 277
P
Sbjct: 329 NP 330
Score = 44.3 bits (103), Expect = 3e-04
Identities = 37/120 (30%), Positives = 52/120 (42%), Gaps = 7/120 (5%)
Query: 95 AKYCPLLPAPRST-IAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVR 153
AK L RS ++ F P G+ AS D +KI D C+ GH R V+R
Sbjct: 184 AKVVRTLTGHRSNCVSVNFHPFGEFFASGSLDTNLKIWDIRKKGCIHTYKGHTRGVNVLR 243
Query: 154 FHPLHPKILASGSLDQEVRLWDANTSECITSHHFYR---PIASIAFHAKGEIIAVASGHK 210
F P + + SG D V++WD + + H F I S+ FH ++A S K
Sbjct: 244 FTP-DGRWIVSGGEDNVVKVWDLTAGKLL--HEFKSHEGKIQSLDFHPHEFLLATGSADK 300
>At5g02430 putative protein
Length = 905
Score = 50.4 bits (119), Expect = 4e-06
Identities = 31/99 (31%), Positives = 52/99 (52%), Gaps = 4/99 (4%)
Query: 117 KVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDA 176
++L S+ D TV++ D ET CLK L H V+F+PL SGSLD ++R+W+
Sbjct: 531 QLLLSSSMDKTVRLWDIETQSCLK-LFAHNDYVTCVQFNPLDEDYFISGSLDAKIRIWNI 589
Query: 177 NTSECITSHHFYRPIASIAFHAKGEIIAVAS--GH-KLY 212
+ + + + + ++ + G+ V S GH +LY
Sbjct: 590 SNRQVVEWNDLKEMVTAVCYTPDGQAAFVGSINGHCRLY 628
>At5g25150 transcription initiation factor IID-associated
factor-like protein
Length = 700
Score = 50.1 bits (118), Expect = 5e-06
Identities = 23/70 (32%), Positives = 39/70 (54%), Gaps = 1/70 (1%)
Query: 114 PDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRL 173
P+ +A+ D TV++ D +TG C+++ IGH + P + +ASG D + +
Sbjct: 509 PNCNYIATGSSDKTVRLWDVQTGECVRIFIGHRSMVLSLAMSP-DGRYMASGDEDGTIMM 567
Query: 174 WDANTSECIT 183
WD +T+ CIT
Sbjct: 568 WDLSTARCIT 577
Score = 44.3 bits (103), Expect = 3e-04
Identities = 29/111 (26%), Positives = 44/111 (39%), Gaps = 35/111 (31%)
Query: 108 IAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPL---------- 157
++ A SPDG+ +AS D T+ + D T RC+ L+GH W + + +
Sbjct: 545 LSLAMSPDGRYMASGDEDGTIMMWDLSTARCITPLMGHNSCVWSLSYRVVQGYDIEWYGF 604
Query: 158 -------------------------HPKILASGSLDQEVRLWDANTSECIT 183
+LASGS D V+LWD +S +T
Sbjct: 605 LSWVVECCIGLMCWVVNGMTGFGCGEGSLLASGSADCTVKLWDVTSSTKLT 655
Score = 40.8 bits (94), Expect = 0.003
Identities = 36/147 (24%), Positives = 52/147 (34%), Gaps = 39/147 (26%)
Query: 97 YCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHP 156
Y LL +A FSP G + S+ D T+++ + L GH W +F P
Sbjct: 411 YTLLLGHSGPVYSATFSPPGDFVLSSSADTTIRLWSTKLNANLVCYKGHNYPVWDAQFSP 470
Query: 157 L------------------------------------HP--KILASGSLDQEVRLWDANT 178
HP +A+GS D+ VRLWD T
Sbjct: 471 FGHYFASCSHDRTARIWSMDRIQPLRIMAGHLSDVDWHPNCNYIATGSSDKTVRLWDVQT 530
Query: 179 SECITSHHFYRP-IASIAFHAKGEIIA 204
EC+ +R + S+A G +A
Sbjct: 531 GECVRIFIGHRSMVLSLAMSPDGRYMA 557
>At5g23430 unknown protein
Length = 837
Score = 49.7 bits (117), Expect = 7e-06
Identities = 28/73 (38%), Positives = 39/73 (53%), Gaps = 1/73 (1%)
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
F+PDG+ + S D+ VK+ D G+ L H + FHP H +LA+GS D+ V
Sbjct: 151 FTPDGRWVVSGGEDNIVKVWDLTAGKLLTEFKSHEGQIQSLDFHP-HEFLLATGSADRTV 209
Query: 172 RLWDANTSECITS 184
+ WD T E I S
Sbjct: 210 KFWDLETFELIGS 222
Score = 45.4 bits (106), Expect = 1e-04
Identities = 36/118 (30%), Positives = 52/118 (43%), Gaps = 3/118 (2%)
Query: 95 AKYCPLLPAPRST-IAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVR 153
AK L RS I+ F P G+ AS D +KI D C+ GH R V+R
Sbjct: 91 AKIVRTLTGHRSNCISVDFHPFGEFFASGSLDTNLKIWDIRKKGCIHTYKGHTRGVNVLR 150
Query: 154 FHPLHPKILASGSLDQEVRLWDANTSECITSHHFYR-PIASIAFHAKGEIIAVASGHK 210
F P + + SG D V++WD + +T + I S+ FH ++A S +
Sbjct: 151 FTP-DGRWVVSGGEDNIVKVWDLTAGKLLTEFKSHEGQIQSLDFHPHEFLLATGSADR 207
Score = 45.1 bits (105), Expect = 2e-04
Identities = 33/137 (24%), Positives = 60/137 (43%), Gaps = 5/137 (3%)
Query: 109 AAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLD 168
+ F ++A+ T+K+ D E + ++ L GH V FHP + ASGSLD
Sbjct: 64 SVTFDASEVLVAAGAASGTIKLWDLEEAKIVRTLTGHRSNCISVDFHPF-GEFFASGSLD 122
Query: 169 QEVRLWDANTSECITSHHFY-RPIASIAFHAKGEIIAVASGHKLYIWHYDKKGEASYSPI 227
+++WD CI ++ + R + + F G + V+ G + +D +
Sbjct: 123 TNLKIWDIRKKGCIHTYKGHTRGVNVLRFTPDGRWV-VSGGEDNIVKVWDLTAGKLLTEF 181
Query: 228 FVLKTRRSLRAVHFHPH 244
++++ FHPH
Sbjct: 182 --KSHEGQIQSLDFHPH 196
>At1g11160 hypothetical protein
Length = 974
Score = 49.3 bits (116), Expect = 9e-06
Identities = 36/138 (26%), Positives = 62/138 (44%), Gaps = 9/138 (6%)
Query: 109 AAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLD 168
A F P G+ LAS D +++ D C++ GH R + F P + + SG LD
Sbjct: 54 AVEFHPFGEFLASGSSDTNLRVWDTRKKGCIQTYKGHTRGISTIEFSP-DGRWVVSGGLD 112
Query: 169 QEVRLWDANTSECITSHHFYR-PIASIAFHAKGEIIAVASGHK-LYIWHYDKKGEASYSP 226
V++WD + + + PI S+ FH ++A S + + W + ++
Sbjct: 113 NVVKVWDLTAGKLLHEFKCHEGPIRSLDFHPLEFLLATGSADRTVKFWDLE-----TFEL 167
Query: 227 IFVLKTRRS-LRAVHFHP 243
I + + +RA+ FHP
Sbjct: 168 IGTTRPEATGVRAIAFHP 185
Score = 47.4 bits (111), Expect = 3e-05
Identities = 28/108 (25%), Positives = 53/108 (48%), Gaps = 3/108 (2%)
Query: 109 AAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLD 168
+ AF+ + ++ + +K+ D E + ++ GH V FHP + LASGS D
Sbjct: 12 SVAFNSEEVLVLAGASSGVIKLWDLEESKMVRAFTGHRSNCSAVEFHPF-GEFLASGSSD 70
Query: 169 QEVRLWDANTSECITSHHFY-RPIASIAFHAKGE-IIAVASGHKLYIW 214
+R+WD CI ++ + R I++I F G +++ + + +W
Sbjct: 71 TNLRVWDTRKKGCIQTYKGHTRGISTIEFSPDGRWVVSGGLDNVVKVW 118
>At3g27640 unknown protein
Length = 535
Score = 48.9 bits (115), Expect = 1e-05
Identities = 27/68 (39%), Positives = 35/68 (50%), Gaps = 3/68 (4%)
Query: 119 LASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANT 178
L + GD T+K+ D E +C VLIGH T + HP + +L SGS D LWD
Sbjct: 143 LLTASGDQTIKVWDVEENKCTGVLIGHTGTVKSMCSHPTNSDLLVSGSRDGCFALWDL-- 200
Query: 179 SECITSHH 186
C +S H
Sbjct: 201 -RCKSSSH 207
>At1g79990 coatomer protein like complex, subunit beta 2 (beta
prime)
Length = 920
Score = 48.1 bits (113), Expect = 2e-05
Identities = 26/64 (40%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Query: 115 DGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLW 174
D L + DHT K+ D +T C++ L GH V FHP P I+ +GS D VR+W
Sbjct: 198 DKPYLITGSDDHTAKVWDYQTKSCVQTLEGHTHNVSAVSFHPELP-IIITGSEDGTVRIW 256
Query: 175 DANT 178
A T
Sbjct: 257 HATT 260
Score = 43.1 bits (100), Expect = 6e-04
Identities = 36/147 (24%), Positives = 64/147 (43%), Gaps = 12/147 (8%)
Query: 111 AFSPDGKVLASTHGDHTVKIIDCETG-RCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQ 169
A P + S+ D +K+ D E G C ++ GH V F+P AS SLD+
Sbjct: 106 AVHPTLPYVLSSSDDMLIKLWDWEKGWLCTQIFEGHSHYVMQVTFNPKDTNTFASASLDR 165
Query: 170 EVRLWDANTSE-CITSHHFYRPIASIAFHAKGE---IIAVASGHKLYIWHYDKKGEASYS 225
+++W+ + + T + + + + G+ +I + H +W Y K S
Sbjct: 166 TIKIWNLGSPDPNFTLDAHLKGVNCVDYFTGGDKPYLITGSDDHTAKVWDYQTK-----S 220
Query: 226 PIFVLKTR-RSLRAVHFHPHAAPYLLT 251
+ L+ ++ AV FHP P ++T
Sbjct: 221 CVQTLEGHTHNVSAVSFHPE-LPIIIT 246
>At3g15980 putative coatomer complex subunit
Length = 921
Score = 47.4 bits (111), Expect = 3e-05
Identities = 26/64 (40%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Query: 115 DGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLW 174
D L + DHT K+ D +T C++ L GH V FHP P I+ +GS D VR+W
Sbjct: 201 DKPYLITGSDDHTAKVWDYQTKSCVQTLDGHTHNVSAVCFHPELP-IIITGSEDGTVRIW 259
Query: 175 DANT 178
A T
Sbjct: 260 HATT 263
Score = 41.6 bits (96), Expect = 0.002
Identities = 36/147 (24%), Positives = 63/147 (42%), Gaps = 12/147 (8%)
Query: 111 AFSPDGKVLASTHGDHTVKIIDCETG-RCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQ 169
A P + S+ D +K+ D E G C ++ GH V F+P AS SLD+
Sbjct: 109 AVHPTLPYVLSSSDDMLIKLWDWENGWACTQIFEGHSHYVMQVVFNPKDTNTFASASLDR 168
Query: 170 EVRLWDANTSE-CITSHHFYRPIASIAFHAKGE---IIAVASGHKLYIWHYDKKGEASYS 225
+++W+ + + T + + + + G+ +I + H +W Y K S
Sbjct: 169 TIKIWNLGSPDPNFTLDAHQKGVNCVDYFTGGDKPYLITGSDDHTAKVWDYQTK-----S 223
Query: 226 PIFVLKTR-RSLRAVHFHPHAAPYLLT 251
+ L ++ AV FHP P ++T
Sbjct: 224 CVQTLDGHTHNVSAVCFHPE-LPIIIT 249
>At2g47990 unknown protein
Length = 530
Score = 47.4 bits (111), Expect = 3e-05
Identities = 35/121 (28%), Positives = 53/121 (42%), Gaps = 2/121 (1%)
Query: 119 LASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDA-- 176
L S D VK D + L+GH P++ +L +GS D V++WDA
Sbjct: 151 LVSGGDDGVVKYWDVAGATVISDLLGHKDYVRCGDCSPVNDSMLVTGSYDHTVKVWDARV 210
Query: 177 NTSECITSHHFYRPIASIAFHAKGEIIAVASGHKLYIWHYDKKGEASYSPIFVLKTRRSL 236
+TS I + P+ + + G +IA A G+ + +W G+ S KT SL
Sbjct: 211 HTSNWIAEINHGLPVEDVVYLPSGGLIATAGGNSVKVWDLIGGGKMVCSMESHNKTVTSL 270
Query: 237 R 237
R
Sbjct: 271 R 271
>At1g52360 coatomer complex subunit, putative
Length = 926
Score = 47.4 bits (111), Expect = 3e-05
Identities = 26/64 (40%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Query: 115 DGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLW 174
D L + DHT K+ D +T C++ L GH V FHP P I+ +GS D VR+W
Sbjct: 198 DKPYLITGSDDHTAKVWDYQTKSCVQTLEGHTHNVSAVCFHPELP-IIITGSEDGTVRIW 256
Query: 175 DANT 178
A T
Sbjct: 257 HATT 260
Score = 42.7 bits (99), Expect = 8e-04
Identities = 36/147 (24%), Positives = 64/147 (43%), Gaps = 12/147 (8%)
Query: 111 AFSPDGKVLASTHGDHTVKIIDCETG-RCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQ 169
A P + S+ D +K+ D E G C ++ GH V F+P AS SLD+
Sbjct: 106 AVHPTLPYVLSSSDDMLIKLWDWEKGWACTQIFEGHSHYVMQVTFNPKDTNTFASASLDR 165
Query: 170 EVRLWDANTSE-CITSHHFYRPIASIAFHAKGE---IIAVASGHKLYIWHYDKKGEASYS 225
+++W+ + + T + + + + G+ +I + H +W Y K S
Sbjct: 166 TIKIWNLGSPDPNFTLDAHQKGVNCVDYFTGGDKPYLITGSDDHTAKVWDYQTK-----S 220
Query: 226 PIFVLKTR-RSLRAVHFHPHAAPYLLT 251
+ L+ ++ AV FHP P ++T
Sbjct: 221 CVQTLEGHTHNVSAVCFHPE-LPIIIT 246
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.134 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,944,388
Number of Sequences: 26719
Number of extensions: 764879
Number of successful extensions: 2493
Number of sequences better than 10.0: 144
Number of HSP's better than 10.0 without gapping: 76
Number of HSP's successfully gapped in prelim test: 68
Number of HSP's that attempted gapping in prelim test: 2052
Number of HSP's gapped (non-prelim): 392
length of query: 808
length of database: 11,318,596
effective HSP length: 107
effective length of query: 701
effective length of database: 8,459,663
effective search space: 5930223763
effective search space used: 5930223763
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC133341.5


