

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133139.3 - phase: 0
(185 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
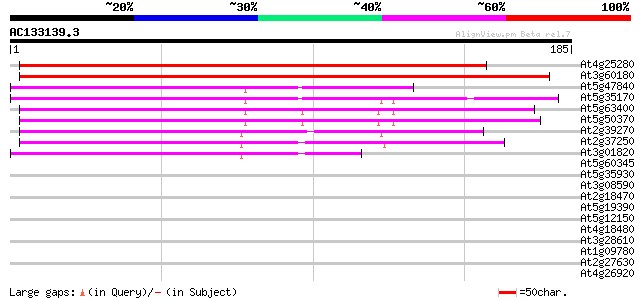
Score E
Sequences producing significant alignments: (bits) Value
At4g25280 UMP/CMP kinase like protein 229 8e-61
At3g60180 uridylate kinase like protein 219 7e-58
At5g47840 putative protein 94 4e-20
At5g35170 adenylate kinase -like protein 89 1e-18
At5g63400 adenylate kinase 86 1e-17
At5g50370 adenylate kinase 80 8e-16
At2g39270 putative adenylate kinase 75 3e-14
At2g37250 putative adenylate kinase 74 4e-14
At3g01820 putative adenylate kinase 48 3e-06
At5g60345 unknown protein 30 0.57
At5g35930 unknown protein 29 1.3
At3g08590 putative 2,3-bisphosphoglycerate-independent phosphogl... 29 1.6
At2g18470 putative protein kinase 29 1.6
At5g19390 unknown protein 28 2.2
At5g12150 unknown protein 28 2.2
At4g18480 protein ch-42 precursor, chloroplast 28 2.2
At3g28610 unknown protein 28 2.2
At1g09780 putative 2,3-bisphosphoglycerate-independent phosphogl... 28 2.2
At2g27630 hypothetical protein 28 2.8
At4g26920 putative homeodomain protein 28 3.7
>At4g25280 UMP/CMP kinase like protein
Length = 227
Score = 229 bits (583), Expect = 8e-61
Identities = 108/154 (70%), Positives = 131/154 (84%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC +IVETFG +HLSAGDLLR+ + +E GAMIL I++G+IVPS VTV+
Sbjct: 50 GGPGSGKGTQCEKIVETFGLQHLSAGDLLRREIAMHTENGAMILNLIKDGKIVPSEVTVK 109
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 123
LI +E++ DNRKFLIDGFPR+EENR+AFE I +PD VL+FDCPEEEMVKRVL+RNQG
Sbjct: 110 LIQKELESSDNRKFLIDGFPRTEENRVAFERIIRADPDVVLFFDCPEEEMVKRVLNRNQG 169
Query: 124 RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLH 157
RIDDNI T+KKRLK+F ALN PVID+Y +G+L+
Sbjct: 170 RIDDNITTMKKRLKIFNALNRPVIDYYKNKGKLY 203
>At3g60180 uridylate kinase like protein
Length = 204
Score = 219 bits (558), Expect = 7e-58
Identities = 104/175 (59%), Positives = 133/175 (75%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQCA +V+ F + H SAGDLLR + S SE+GAMI I EGRIVPS +TV+
Sbjct: 28 GGPGSGKGTQCANVVKHFSYTHFSAGDLLRAEIKSGSEFGAMIQSMIAEGRIVPSEITVK 87
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 123
L+ + M+ N KFLIDGFPR+EENR FE++ EP FVL+FDCPEEE+ +R++SRNQG
Sbjct: 88 LLCKAMEESGNDKFLIDGFPRNEENRNVFENVARIEPAFVLFFDCPEEELERRIMSRNQG 147
Query: 124 RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVFAA 178
R DDNI+TIKKR KVF LP+I +Y +G+L +INA + +E+FE VR +FA+
Sbjct: 148 REDDNIETIKKRFKVFVESTLPIISYYESKGKLRKINAAKSSEEVFEAVRVLFAS 202
>At5g47840 putative protein
Length = 283
Score = 94.0 bits (232), Expect = 4e-20
Identities = 49/135 (36%), Positives = 76/135 (56%), Gaps = 3/135 (2%)
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
M+SG P SGKGTQC I +G H+SAGDLLR + S SE G E + +G++VP +
Sbjct: 68 MISGAPASGKGTQCELITHKYGLVHISAGDLLRAEIASGSENGRRAKEHMEKGQLVPDEI 127
Query: 61 TVRLILREMQYGDNRK--FLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVL 118
V ++ + D+ + +L+DG+PRS A + G +PD + + PEE +++RV+
Sbjct: 128 VVMMVKDRLSQTDSEQKGWLLDGYPRSASQATALKGF-GFQPDLFIVLEVPEEILIERVV 186
Query: 119 SRNQGRIDDNIDTIK 133
R + I +K
Sbjct: 187 GRRLDPVTGKIYHLK 201
>At5g35170 adenylate kinase -like protein
Length = 588
Score = 89.4 bits (220), Expect = 1e-18
Identities = 56/208 (26%), Positives = 103/208 (48%), Gaps = 30/208 (14%)
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
M+SG P SGKGTQC IV FG H+S GDLLR + S ++ G E + G +VP +
Sbjct: 83 MISGAPASGKGTQCELIVHKFGLVHISTGDLLRAEVSSGTDIGKRAKEFMNSGSLVPDEI 142
Query: 61 TVRLILREMQYGDNRK--FLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVL 118
+ ++ + D ++ +L+DGFPRS + + + +PD + D P+E ++ R +
Sbjct: 143 VIAMVAGRLSREDAKEHGWLLDGFPRSFAQAQSLDKL-NVKPDIFILLDVPDEILIDRCV 201
Query: 119 SRN----QGRI---------------------DDNIDTIKKRLKVFEALNLPVIDHYARR 153
R G+I DD + +K RL++++ + +I Y+
Sbjct: 202 GRRLDPVTGKIYHIKNYPPESDEIKARLVTRPDDTEEKVKARLQIYKQNSEAIISAYS-- 259
Query: 154 GRLHRINAVGTEDEIFEQVRPVFAACEV 181
+ +I+A ++ +FE+ + + + ++
Sbjct: 260 DVMVKIDANRPKEVVFEETQTLLSQIQL 287
>At5g63400 adenylate kinase
Length = 246
Score = 85.5 bits (210), Expect = 1e-17
Identities = 62/203 (30%), Positives = 92/203 (44%), Gaps = 33/203 (16%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
G PGSGKGTQ + + + HLS GD+LR A+ S + G E + +G +V + V
Sbjct: 40 GPPGSGKGTQSPVVKDEYCLCHLSTGDMLRAAVASKTPLGVKAKEAMEKGELVSDDLVVG 99
Query: 64 LILREMQYGDNRK-FLIDGFPRSEENRIAFEHI---TGTEPDFVLYFDCPEEEMVKRVLS 119
+I M +K F++DGFPR+ + + GTE D VL F + + +R+
Sbjct: 100 IIDEAMNKPKCQKGFILDGFPRTVTQAEKLDEMLKRRGTEIDKVLNFAIDDAILEERITG 159
Query: 120 R----NQGRI-------------------------DDNIDTIKKRLKVFEALNLPVIDHY 150
R + GR DDN D +K RL F + PVID+Y
Sbjct: 160 RWIHPSSGRSYHTKFAPPKTPGVDDITGEPLIQRKDDNADVLKSRLAAFHSQTQPVIDYY 219
Query: 151 ARRGRLHRINAVGTEDEIFEQVR 173
A++ L I A E+ +V+
Sbjct: 220 AKKAVLTNIQAEKAPQEVTSEVK 242
>At5g50370 adenylate kinase
Length = 248
Score = 79.7 bits (195), Expect = 8e-16
Identities = 60/205 (29%), Positives = 92/205 (44%), Gaps = 33/205 (16%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
G PGSGKGTQ I + F HLS GD+LR A+ + + G E + +G +V + V
Sbjct: 41 GPPGSGKGTQSPVIKDEFCLCHLSTGDMLRAAVAAKTPLGVKAKEAMDKGELVSDDLVVG 100
Query: 64 LILREMQYGDNRK-FLIDGFPRSEENRIAFEHI---TGTEPDFVLYFDCPEEEMVKRVLS 119
++ M +K F++DGFPR+ + + G + D VL F + + +R+
Sbjct: 101 IMDEAMNRPKCQKGFILDGFPRTVTQAEKLDEMLNRRGAQIDKVLNFAIDDSVLEERITG 160
Query: 120 R----NQGRI-------------------------DDNIDTIKKRLKVFEALNLPVIDHY 150
R + GR DDN D ++ RL F PVID+Y
Sbjct: 161 RWIHPSSGRSYHTKFAPPKVPGVDDLTGEPLIQRKDDNADVLRSRLDAFHKQTQPVIDYY 220
Query: 151 ARRGRLHRINAVGTEDEIFEQVRPV 175
A++ L I A +E+ + V+ V
Sbjct: 221 AKKENLVNIPAEKAPEEVTKVVKKV 245
>At2g39270 putative adenylate kinase
Length = 295
Score = 74.7 bits (182), Expect = 3e-14
Identities = 47/196 (23%), Positives = 84/196 (41%), Gaps = 45/196 (22%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
G PG GKGT +R+ G H++ GDL+R+ + S + + E + G++VP +
Sbjct: 71 GCPGVGKGTYASRLSSLLGVPHIATGDLVREELSSSGLLSSQLKELVNHGKLVPDEFIIS 130
Query: 64 LILREMQYGDNR---KFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSR 120
L+ + +Q G ++ +++DGFPR+ E +T D V+ EE ++ + L R
Sbjct: 131 LLSKRLQAGKDKGESGYILDGFPRTVTQAEILEGVTNI--DLVINLKLREEALLAKCLGR 188
Query: 121 N----------------------------------------QGRIDDNIDTIKKRLKVFE 140
R DD + +K+RL+++
Sbjct: 189 RICSECGGNYNVACIDIKGDDDTPRMYMPPLLPPPNCESKLISRADDTEEVVKERLRIYN 248
Query: 141 ALNLPVIDHYARRGRL 156
+ PV + Y +RG+L
Sbjct: 249 KMTQPVEEFYKKRGKL 264
>At2g37250 putative adenylate kinase
Length = 284
Score = 73.9 bits (180), Expect = 4e-14
Identities = 48/203 (23%), Positives = 86/203 (41%), Gaps = 45/203 (22%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
G PG GKGT +R+ G H++ GDL+R+ + S + E + +G++V + V
Sbjct: 58 GCPGVGKGTYASRLSTLLGVPHIATGDLVREELASSGPLSQKLSEIVNQGKLVSDEIIVD 117
Query: 64 LILREMQYGDNR---KFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSR 120
L+ + ++ G+ R F++DGFPR+ + T+ D V+ PEE +V + L R
Sbjct: 118 LLSKRLEAGEARGESGFILDGFPRTMRQAEILGDV--TDIDLVVNLKLPEEVLVDKCLGR 175
Query: 121 NQ----------------------------------------GRIDDNIDTIKKRLKVFE 140
R DD + +K RL+++
Sbjct: 176 RTCSQCGKGFNVAHINLKGENGRPGISMDPLLPPHQCMSKLVTRADDTEEVVKARLRIYN 235
Query: 141 ALNLPVIDHYARRGRLHRINAVG 163
+ P+ ++Y +G+L + G
Sbjct: 236 ETSQPLEEYYRTKGKLMEFDLPG 258
>At3g01820 putative adenylate kinase
Length = 219
Score = 48.1 bits (113), Expect = 3e-06
Identities = 30/119 (25%), Positives = 55/119 (46%), Gaps = 5/119 (4%)
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
+L G PG+ + R+ + H+S G L+R+ + S I + E ++VP +V
Sbjct: 66 VLMGAPGAWRHVFAERLSKLLEVPHISMGSLVRQELNPRSSLYKEIASAVNERKLVPKSV 125
Query: 61 TVRLILREMQYGDNR---KFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKR 116
L+ + ++ G R F++ G PR+ + I + D V+ C E+ +V R
Sbjct: 126 VFALLSKRLEEGYARGETGFILHGIPRTRFQAETLDQI--AQIDLVVNLKCSEDHLVNR 182
>At5g60345 unknown protein
Length = 178
Score = 30.4 bits (67), Expect = 0.57
Identities = 10/34 (29%), Positives = 22/34 (64%)
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRK 34
+++G PG+GK T + + E +++ GDL+++
Sbjct: 17 LITGTPGTGKSTTASALAEATNLRYICIGDLVKE 50
>At5g35930 unknown protein
Length = 1175
Score = 29.3 bits (64), Expect = 1.3
Identities = 17/38 (44%), Positives = 22/38 (57%)
Query: 25 HLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTV 62
+L G RK D++ G+M ETI EGRI SA+ V
Sbjct: 797 YLFIGSHSRKFSCIDAKSGSMYWETILEGRIEGSAMVV 834
>At3g08590 putative 2,3-bisphosphoglycerate-independent
phosphoglycerate mutase
Length = 560
Score = 28.9 bits (63), Expect = 1.6
Identities = 13/49 (26%), Positives = 26/49 (52%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIRE 52
GGPG G + + +ET G +++A + V+ S+Y ++E + +
Sbjct: 512 GGPGLSAGVRFRQDIETPGLANVAATVMNLHGFVAPSDYETSLIEVVEK 560
>At2g18470 putative protein kinase
Length = 633
Score = 28.9 bits (63), Expect = 1.6
Identities = 18/50 (36%), Positives = 30/50 (60%), Gaps = 5/50 (10%)
Query: 16 RIVETFGF---KHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTV 62
R++ TFG+ ++ S+G L K+ V YG M+LE I R V +++T+
Sbjct: 443 RVMGTFGYLAPEYASSGKLTEKSDVFS--YGVMLLELITGKRPVDNSITM 490
>At5g19390 unknown protein
Length = 870
Score = 28.5 bits (62), Expect = 2.2
Identities = 24/80 (30%), Positives = 35/80 (43%), Gaps = 7/80 (8%)
Query: 51 REGRIVPSAVTVRLILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPE 110
RE R + S V R IL ++ IDG P E + F GT+ + +L
Sbjct: 151 REKRPLKSLVVGRPILLALED-------IDGSPSFLEKALQFIEKYGTKIEGILRQSADV 203
Query: 111 EEMVKRVLSRNQGRIDDNID 130
EE+ +RV QG+ + D
Sbjct: 204 EEVERRVQEYEQGKTEFTFD 223
>At5g12150 unknown protein
Length = 827
Score = 28.5 bits (62), Expect = 2.2
Identities = 22/76 (28%), Positives = 35/76 (45%), Gaps = 7/76 (9%)
Query: 51 REGRIVPSAVTVRLILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPE 110
R+ R + S+V R IL ++ IDG P E + F GT+ + +L
Sbjct: 156 RDQRPLKSSVVGRPILLALEE-------IDGSPSFLEKALQFLETYGTKVEGILRQSADV 208
Query: 111 EEMVKRVLSRNQGRID 126
EE+ +RV QG+ +
Sbjct: 209 EEVERRVQEYEQGKTE 224
>At4g18480 protein ch-42 precursor, chloroplast
Length = 424
Score = 28.5 bits (62), Expect = 2.2
Identities = 16/61 (26%), Positives = 26/61 (42%)
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
M+ G G+GK T +V+ ++ AGD + G + E + +G VP
Sbjct: 116 MIMGDRGTGKSTTVRSLVDLLPEINVVAGDPYNSDPIDPEFMGVEVRERVEKGEQVPVIA 175
Query: 61 T 61
T
Sbjct: 176 T 176
>At3g28610 unknown protein
Length = 473
Score = 28.5 bits (62), Expect = 2.2
Identities = 37/162 (22%), Positives = 70/162 (42%), Gaps = 19/162 (11%)
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
+L G PG+GK T A + + S DL A+ ++SE ++ T + IV +
Sbjct: 240 LLYGPPGTGKSTMIAAMANLLNY---SIYDLELTAIQNNSELRKILTATSNKSIIVIEDI 296
Query: 61 TVRLILREMQYGDNRKFLI---DGFPRSEENRIAFEHITGTEPDFV--LYFDCPEEEMVK 115
L L + +I DG +EEN+ +F ++G +F+ ++ C +E ++
Sbjct: 297 DCSLDLTGKRKKKESNLMIWRKDGDQDNEENK-SFVTLSGL-LNFIDGIWSACGQERIIV 354
Query: 116 RVLSR---------NQGRIDDNIDTIKKRLKVFEALNLPVID 148
+ +GR+D +I+ + F+ L +D
Sbjct: 355 FTTNHLAKLDPALIRRGRMDMHIELSYCTFEAFKTLAKNYLD 396
>At1g09780 putative 2,3-bisphosphoglycerate-independent
phosphoglycerate mutase
Length = 557
Score = 28.5 bits (62), Expect = 2.2
Identities = 13/47 (27%), Positives = 26/47 (54%)
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETI 50
GGPG +G + + +ET G +++A + V+ S+Y ++E +
Sbjct: 510 GGPGLAQGVRFRKDLETPGLANVAATVMNLHGFVAPSDYEPTLIEVV 556
>At2g27630 hypothetical protein
Length = 653
Score = 28.1 bits (61), Expect = 2.8
Identities = 23/105 (21%), Positives = 43/105 (40%), Gaps = 15/105 (14%)
Query: 36 MVSDSEYGAMILET------IREGRIVPSAVTVRLILREMQYGDNRKFLIDGFPRSEENR 89
+ SD E G ++ E R+ +I+P +V ++ ++Y G +
Sbjct: 336 LASDDERGKLLKEIKNLLVMFRDLKILPCSVRDCVMQYPLRYF--------GELEVSKQS 387
Query: 90 IAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQGRIDDNIDTIKK 134
+ H+ T P + + DC E + L R + DD ID + +
Sbjct: 388 LVDSHLVET-PQSICFLDCHELNKILNFLKRIKSERDDGIDVVSR 431
>At4g26920 putative homeodomain protein
Length = 461
Score = 27.7 bits (60), Expect = 3.7
Identities = 13/42 (30%), Positives = 22/42 (51%)
Query: 6 PGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMIL 47
PG G + A V + GF ++ +++ +VSD E G +L
Sbjct: 412 PGQNSGDEDAGCVVSVGFLTIATEEIVANTLVSDVERGIYLL 453
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.142 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,168,451
Number of Sequences: 26719
Number of extensions: 174827
Number of successful extensions: 445
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 415
Number of HSP's gapped (non-prelim): 27
length of query: 185
length of database: 11,318,596
effective HSP length: 93
effective length of query: 92
effective length of database: 8,833,729
effective search space: 812703068
effective search space used: 812703068
T: 11
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC133139.3


