

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130806.4 - phase: 0
(359 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
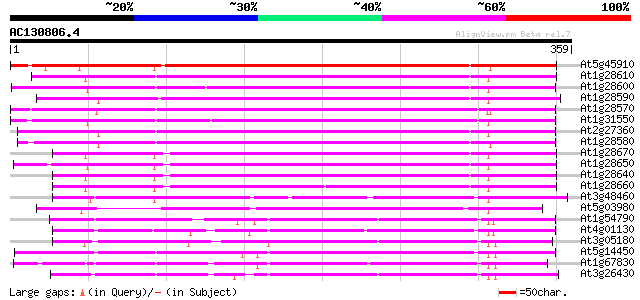
At5g45910 GDSL-motif lipase/hydrolase-like protein 306 9e-84
At1g28610 lipase, putative 280 9e-76
At1g28600 lipase like protein 274 5e-74
At1g28590 putative lipase (At1g28590) 268 3e-72
At1g28570 lipase like protein 265 2e-71
At1g31550 unknown protein 261 3e-70
At2g27360 putative lipase 261 5e-70
At1g28580 lipase like protein 259 2e-69
At1g28670 lipase 257 6e-69
At1g28650 lipase, putative 257 8e-69
At1g28640 lipase, putative 255 3e-68
At1g28660 unknown protein 251 3e-67
At3g48460 lipase - like protein 200 1e-51
At5g03980 lipase -like protein 177 8e-45
At1g54790 172 3e-43
At4g01130 putative acetyltransferase 165 4e-41
At3g05180 putative nodulin 158 5e-39
At5g14450 early nodule-specific protein - like 156 2e-38
At1g67830 ENOD8-like protein 155 4e-38
At3g26430 nodulin, putative 150 1e-36
>At5g45910 GDSL-motif lipase/hydrolase-like protein
Length = 372
Score = 306 bits (785), Expect = 9e-84
Identities = 169/375 (45%), Positives = 228/375 (60%), Gaps = 30/375 (8%)
Query: 1 MNVSTLFLITFTCGFLQNVVS--NANPLSYEAFFNFGDSISDTGN---AASIFLPMPNPI 55
M ++ LF++ F+ FL +V S L YE+ FNFGDS+SDTGN + + P +
Sbjct: 1 MRINMLFIVAFS--FLVSVRSLPMRPTLKYESIFNFGDSLSDTGNFLLSGDVDSPNIGRL 58
Query: 56 PYGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAY-----ENKSIDQDIKKGVNFAFAG 110
PYG ++F +GR S+GRLIIDFIAEA GLP++P Y N S+D K+G NFA AG
Sbjct: 59 PYGQTFFNRSTGRCSDGRLIIDFIAEASGLPYIPPYLQSLRTNDSVD--FKRGANFAVAG 116
Query: 111 ATVLNVEYYVKNGLPLPD-TNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIG 169
AT ++ GL + TN +L IQL WFK +KP LCK+K +C YF+KSLF+VGEIG
Sbjct: 117 ATANEFSFFKNRGLSVTLLTNKTLDIQLDWFKKLKPSLCKTKPECEQYFRKSLFLVGEIG 176
Query: 170 GNDI-MKHMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTL 228
GND + ++ ++VPF++ I + T+ LIEEGA+ L+VPGN P+GCSAA+
Sbjct: 177 GNDYNYPLLAFRSFKHAMDLVPFVINKIMDVTSALIEEGAMTLIVPGNLPIGCSAALLER 236
Query: 229 VNSNKKEDYDEFG-CLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQ 287
N N YD C + NNL + N +LK + LR+K+P KIIY DYY+ A + +
Sbjct: 237 FNDNSGWLYDSRNQCYMPLNNLAKLHNDKLKKGLAALRKKYPYAKIIYADYYSSAMQFFN 296
Query: 288 TPQQYGFDKDAIFKACCGG------------CGSLIATVCSDPSKRINWDGPHFTEAAYK 335
+P +YGF ++ KACCGG CG +T C DPS NWDG H TEAAY+
Sbjct: 297 SPSKYGF-TGSVLKACCGGGDGRYNVQPNVRCGEKGSTTCEDPSTYANWDGIHLTEAAYR 355
Query: 336 LIAKGLVEGPFSNPS 350
IA GL+ G F+ P+
Sbjct: 356 HIATGLISGRFTMPT 370
>At1g28610 lipase, putative
Length = 383
Score = 280 bits (716), Expect = 9e-76
Identities = 154/352 (43%), Positives = 205/352 (57%), Gaps = 18/352 (5%)
Query: 15 FLQNVVSNANPLSYEAFFNFGDSISDTGNAASI----FLPMPNPIPYGSSYFKHPSGRMS 70
F+ V S + E+ +FGDSI+DTGN + LP+ +PYG ++F HP+GR
Sbjct: 16 FVTIVSSQTQCRNLESIISFGDSITDTGNLVGLSDRNHLPVTAFLPYGETFFHHPTGRSC 75
Query: 71 NGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTN 130
NGR+IIDFIAE GLP +P + S + + +KGVNFA AGAT L K G+ P +N
Sbjct: 76 NGRIIIDFIAEFLGLPHVPPFYG-SKNGNFEKGVNFAVAGATALETSILEKRGIYYPHSN 134
Query: 131 NSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDIMKHMKHKTVIELREIVP 190
SL IQL FK P LC S DC + I+GEIGGND E++E+VP
Sbjct: 135 ISLGIQLKTFKESLPNLCGSPTDCRDMIGNAFIIMGEIGGNDFNFAFFVNKTSEVKELVP 194
Query: 191 FMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF-GCLIAYNNL 249
++ I++ L++ G +VPGNFP+GCSA TL ++ KE+YD GCL N+
Sbjct: 195 LVITKISSAIVELVDMGGRTFLVPGNFPLGCSATYLTLYQTSNKEEYDPLTGCLTWLNDF 254
Query: 250 IEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG---- 305
EY+N +L+ + L + +P V IIY DY+N RLYQ P ++GF D ACCG
Sbjct: 255 SEYYNEKLQAELNRLSKLYPHVNIIYGDYFNALLRLYQEPSKFGF-MDRPLPACCGLGGP 313
Query: 306 -------GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPS 350
CGS+ CSDPSK +NWDG H TEAAYK IA GL++GP++ PS
Sbjct: 314 YNFTLSKKCGSVGVKYCSDPSKYVNWDGVHMTEAAYKWIADGLLKGPYTIPS 365
>At1g28600 lipase like protein
Length = 393
Score = 274 bits (701), Expect = 5e-74
Identities = 152/365 (41%), Positives = 213/365 (57%), Gaps = 20/365 (5%)
Query: 2 NVSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIF----LPMPNPIPY 57
++ +L + F+ F+ V S ++++ +FGDSI+DTGN + LP+ PY
Sbjct: 3 SLDSLVIFLFSTLFVTIVSSETPCPNFKSIISFGDSIADTGNLVGLSDRNQLPVTAFPPY 62
Query: 58 GSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVE 117
G ++F HP+GR +GR+I+DFIAE GLP++P Y S +++ KGVNFA AGAT L
Sbjct: 63 GETFFHHPTGRSCDGRIIMDFIAEFVGLPYVPPYFG-SKNRNFDKGVNFAVAGATALKSS 121
Query: 118 YYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MKH 176
+ K G+ P TN SL +QL FK P LC S DC +L ++GEIGGND
Sbjct: 122 FLKKRGIQ-PHTNVSLGVQLKSFKKSLPNLCGSPSDCRDMIGNALILMGEIGGNDYNFPF 180
Query: 177 MKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKED 236
K V E+ E+VPF++ +I++T LI G +VPG FP+GCS TL ++ K++
Sbjct: 181 FNRKPVKEVEELVPFVIASISSTITELIGMGGKTFLVPGEFPIGCSVVYLTLYKTSNKDE 240
Query: 237 YD-EFGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFD 295
YD GCL N EY + +LK + LR+ +P V IIY DYYN R+++ P ++GF
Sbjct: 241 YDPSTGCLKWLNKFGEYHSEKLKVELNRLRKLYPHVNIIYADYYNSLLRIFKEPAKFGF- 299
Query: 296 KDAIFKACCG-----------GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEG 344
+ F ACCG CGS+ C DPSK + WDG H TEAAYK IA G++ G
Sbjct: 300 MERPFPACCGIGGPYNFNFTRKCGSVGVKSCKDPSKYVGWDGVHMTEAAYKWIADGILNG 359
Query: 345 PFSNP 349
P++NP
Sbjct: 360 PYANP 364
>At1g28590 putative lipase (At1g28590)
Length = 403
Score = 268 bits (685), Expect = 3e-72
Identities = 149/352 (42%), Positives = 208/352 (58%), Gaps = 19/352 (5%)
Query: 18 NVVSNANPLSYEAFFNFGDSISDTGNAASIFLPMPNPI----PYGSSYFKHPSGRMSNGR 73
+V S ++++ +FGDSI+DTGN + P P PYG ++F HP+GR S+GR
Sbjct: 24 SVNSQTQCRNFKSIISFGDSIADTGNLLGLSDPNDLPASAFPPYGETFFHHPTGRYSDGR 83
Query: 74 LIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSL 133
LIIDFIAE G P +P + + + KKGVNFA AGAT L + + G+ TN SL
Sbjct: 84 LIIDFIAEFLGFPLVPPFYGCQ-NANFKKGVNFAVAGATALEPSFLEERGIHSTITNVSL 142
Query: 134 SIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MKHMKHKTVIELREIVPFM 192
S+QL F P LC S DC + +L ++GEIGGND + K V E+ E+VPF+
Sbjct: 143 SVQLRSFTESLPNLCGSPSDCRDMIENALILMGEIGGNDYNFALFQRKPVKEVEELVPFV 202
Query: 193 VEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF-GCLIAYNNLIE 251
+ I++ L+ G +VPGNFP+G SA+ TL ++ KE+YD GCL N+ E
Sbjct: 203 IATISSAITELVCMGGRTFLVPGNFPIGYSASYLTLYKTSNKEEYDPLTGCLKWLNDFSE 262
Query: 252 YFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG------ 305
Y+N QL+ + LR+ +P V IIY DYYN RL+Q P ++GF + ACCG
Sbjct: 263 YYNKQLQEELNGLRKLYPHVNIIYADYYNALLRLFQEPAKFGF-MNRPLPACCGVGGSYN 321
Query: 306 -----GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPSLK 352
CGS+ C DPS+ +N+DG H TEAAY+LI++GL++GP++ P K
Sbjct: 322 FNFSRRCGSVGVEYCDDPSQYVNYDGIHMTEAAYRLISEGLLKGPYAIPPFK 373
>At1g28570 lipase like protein
Length = 389
Score = 265 bits (678), Expect = 2e-71
Identities = 151/366 (41%), Positives = 213/366 (57%), Gaps = 20/366 (5%)
Query: 1 MNVSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIFLPMPNP----IP 56
M + + FLI T L V S ++++ +FGDSI+DTGN ++ P P +P
Sbjct: 6 MKLVSFFLILSTF-CLTTVNSEPQCHNFKSIISFGDSIADTGNLLALSDPTNLPKVAFLP 64
Query: 57 YGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNV 116
YG ++F HP+GR SNGRLIIDFIAE G P +P + S + + +KGVNFA GAT L
Sbjct: 65 YGETFFHHPTGRFSNGRLIIDFIAEFLGFPLVPPFYG-SQNANFEKGVNFAVGGATALER 123
Query: 117 EYYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MK 175
+ + G+ P TN SL++QL FK P LC S DC + SL ++GEIGGND
Sbjct: 124 SFLEERGIHFPYTNVSLAVQLSSFKESLPNLCVSPSDCRDMIENSLILMGEIGGNDYNYA 183
Query: 176 HMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKE 235
K + E++E+VP ++E I++ LI G +VPG FP+GCS A +L ++ E
Sbjct: 184 FFVGKNIEEIKELVPLVIETISSAITELIGMGGKTFLVPGEFPLGCSVAYLSLYQTSNIE 243
Query: 236 DYDEF-GCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGF 294
+YD GCL N EY + QL+ + L++ +P V IIY DYYN RL Q P ++GF
Sbjct: 244 EYDPLTGCLKWLNKFSEYHDEQLQAELNRLQKLYPHVNIIYADYYNTLLRLAQEPAKFGF 303
Query: 295 DKDAIFKACC--GG---------CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVE 343
+ ACC GG G+ + C DPSK ++WDG H TEAAY+L+A+G+++
Sbjct: 304 ISRPL-PACCALGGPFNFTLGRKRGTQVPECCDDPSKYVSWDGVHMTEAAYRLMAEGILK 362
Query: 344 GPFSNP 349
GP++ P
Sbjct: 363 GPYAIP 368
>At1g31550 unknown protein
Length = 394
Score = 261 bits (668), Expect = 3e-70
Identities = 153/366 (41%), Positives = 209/366 (56%), Gaps = 23/366 (6%)
Query: 1 MNVSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIF----LPMPNPIP 56
M + +LFL T F+ V S + ++E+ +FGDSI+DTGN + LPM P
Sbjct: 10 MKLGSLFLSTL---FVSIVSSESQCRNFESIISFGDSIADTGNLLGLSDHNNLPMSAFPP 66
Query: 57 YGSSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNV 116
YG ++F HP+GR S+GRLIIDFIAE GLP++P Y S + + +KGVNFA A AT L
Sbjct: 67 YGETFFHHPTGRFSDGRLIIDFIAEFLGLPYVPPYFG-STNGNFEKGVNFAVASATALES 125
Query: 117 EYYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MK 175
+ + G P N SL +QL FK P LC DC +L ++GEIG ND
Sbjct: 126 SFLEEKGYHCPH-NFSLGVQLKIFKQSLPNLCGLPSDCRDMIGNALILMGEIGANDYNFP 184
Query: 176 HMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKE 235
+ + + E++E+VP ++ I++ LI G +VPG FP+GCS A TL ++ E
Sbjct: 185 FFQLRPLDEVKELVPLVISTISSAITELIGMGGRTFLVPGGFPLGCSVAFLTLHQTSNME 244
Query: 236 DYDEF-GCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGF 294
+YD GCL N EY + QL+ + LR+ +P V IIY DYYN + RL + P +YGF
Sbjct: 245 EYDPLTGCLKWLNKFGEYHSEQLQEELNRLRKLNPHVNIIYADYYNASLRLGREPSKYGF 304
Query: 295 DKDAIFKACCG-----------GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVE 343
+ ACCG CGS+ CSDPSK + WDG H TEAA+K +A GLV+
Sbjct: 305 INRHL-SACCGVGGPYNFNLSRSCGSVGVEACSDPSKYVAWDGLHMTEAAHKSMADGLVK 363
Query: 344 GPFSNP 349
GP++ P
Sbjct: 364 GPYAIP 369
>At2g27360 putative lipase
Length = 394
Score = 261 bits (666), Expect = 5e-70
Identities = 151/363 (41%), Positives = 204/363 (55%), Gaps = 21/363 (5%)
Query: 6 LFLITFTCGFLQNVV-SNANPLSYEAFFNFGDSISDTGNAASIFLPMPNPI----PYGSS 60
+ L F FL VV S ++++ +FGDSI+DTGN + P P PYG +
Sbjct: 8 MLLSFFISTFLITVVTSQTRCRNFKSIISFGDSITDTGNLLGLSSPNDLPESAFPPYGET 67
Query: 61 YFKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYV 120
+F HPSGR S+GRLIIDFIAE G+P +P + S + + +KGVNFA GAT L
Sbjct: 68 FFHHPSGRFSDGRLIIDFIAEFLGIPHVPPFYG-SKNGNFEKGVNFAVGGATALECSVLE 126
Query: 121 KNGLPLPDTNNSLSIQLGWFKNIKPLLC-KSKEDCNIYFKKSLFIVGEIGGNDI-MKHMK 178
+ G +N SL QL FK P LC S DC + + ++GEIGGND
Sbjct: 127 EKGTHCSQSNISLGNQLKSFKESLPYLCGSSSPDCRDMIENAFILIGEIGGNDYNFPLFD 186
Query: 179 HKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYD 238
K + E++E+VP ++ I++ + L++ GA +VPGNFP+GCS A TL + KE+Y+
Sbjct: 187 RKNIEEVKELVPLVITTISSAISELVDMGARTFLVPGNFPLGCSVAYLTLYETPNKEEYN 246
Query: 239 EF-GCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKD 297
GCL N+ Y N QL+ ++ LR +P V IIY DYYN RL Q P ++G D
Sbjct: 247 PLTGCLTWLNDFSVYHNEQLQAELKRLRNLYPHVNIIYGDYYNTLLRLMQEPSKFGL-MD 305
Query: 298 AIFKACCG-----------GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPF 346
ACCG CGS CSDPSK +NWDG H TEAAYK I++G++ GP+
Sbjct: 306 RPLPACCGLGGPYNFTFSIKCGSKGVEYCSDPSKYVNWDGIHMTEAAYKWISEGVLTGPY 365
Query: 347 SNP 349
+ P
Sbjct: 366 AIP 368
>At1g28580 lipase like protein
Length = 390
Score = 259 bits (662), Expect = 2e-69
Identities = 146/361 (40%), Positives = 201/361 (55%), Gaps = 22/361 (6%)
Query: 6 LFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIFLPMPNPI----PYGSSY 61
+FL TF + NV S +++ +FGDSI+DTGN + P P PYG ++
Sbjct: 16 IFLSTFV---VTNVSSETKCREFKSIISFGDSIADTGNLLGLSDPKDLPHMAFPPYGENF 72
Query: 62 FKHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVK 121
F HP+GR SNGRLIIDFIAE GLP +P + S + + +KGVNFA GAT L +
Sbjct: 73 FHHPTGRFSNGRLIIDFIAEFLGLPLVPPFYG-SHNANFEKGVNFAVGGATALERSFLED 131
Query: 122 NGLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDI-MKHMKHK 180
G+ P TN SL +QL FK P +C S DC + +L ++GEIGGND K
Sbjct: 132 RGIHFPYTNVSLGVQLNSFKESLPSICGSPSDCRDMIENALILMGEIGGNDYNYAFFVDK 191
Query: 181 TVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF 240
+ E++E++P ++ I++ LI G +VPG FP+GCS T ++ E+YD
Sbjct: 192 GIEEIKELMPLVITTISSAITELIGMGGRTFLVPGEFPVGCSVLYLTSHQTSNMEEYDPL 251
Query: 241 -GCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAI 299
GCL N E QL+ + L++ +P V IIY DYYN LYQ P ++GF +
Sbjct: 252 TGCLKWLNKFGENHGEQLRAELNRLQKLYPHVNIIYADYYNALFHLYQEPAKFGF-MNRP 310
Query: 300 FKACCGG-----------CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSN 348
ACCG CG+ I C DPSK + WDG H TEAAY+L+A+G++ GP++
Sbjct: 311 LSACCGAGGPYNYTVGRKCGTDIVESCDDPSKYVAWDGVHMTEAAYRLMAEGILNGPYAI 370
Query: 349 P 349
P
Sbjct: 371 P 371
>At1g28670 lipase
Length = 384
Score = 257 bits (657), Expect = 6e-69
Identities = 147/344 (42%), Positives = 202/344 (57%), Gaps = 26/344 (7%)
Query: 28 YEAFFNFGDSISDTGNAASI----FLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAY 83
+++ +FGDSI+DTGN + LP +PYG S+F PSGR SNGRLIIDFIAE
Sbjct: 33 FKSIISFGDSIADTGNYLHLSDVNHLPQSAFLPYGESFFHPPSGRASNGRLIIDFIAEFL 92
Query: 84 GLPFLPAY---ENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWF 140
GLP++P Y +N S +Q G+NFA GAT L+ + + G+ TN SLS+QL F
Sbjct: 93 GLPYVPPYFGSQNVSFEQ----GINFAVYGATALDRAFLLGKGIESDFTNVSLSVQLDTF 148
Query: 141 KNIKPLLC-KSKEDCNIYFKKSLFIVGEIGGNDI-MKHMKHKTVIELREIVPFMVEAITN 198
K I P LC S DC SL ++GEIGGND + K++ E++E+VP +V+AI++
Sbjct: 149 KQILPNLCASSTRDCKEMLGDSLILMGEIGGNDYNYPFFEGKSINEIKELVPLIVKAISS 208
Query: 199 TTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF-GCLIAYNNLIEYFNGQL 257
LI+ G +VPG FP GCSAA TL + ++D D GC N E+ N QL
Sbjct: 209 AIVDLIDLGGKTFLVPGGFPTGCSAAYLTLFQTVAEKDQDPLTGCYPLLNEFGEHHNEQL 268
Query: 258 KNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG-----------G 306
K ++ L++ +P V IIY DY+N R YQ P +YGF K+ ACCG
Sbjct: 269 KTELKRLQKFYPHVNIIYADYHNSLYRFYQEPAKYGF-KNKPLAACCGVGGKYNFTIGKE 327
Query: 307 CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPS 350
CG C +PS+ +NWDG H TEAAY+ + +G++ GP++ P+
Sbjct: 328 CGYEGVNYCQNPSEYVNWDGYHLTEAAYQKMTEGILNGPYATPA 371
>At1g28650 lipase, putative
Length = 385
Score = 257 bits (656), Expect = 8e-69
Identities = 151/368 (41%), Positives = 212/368 (57%), Gaps = 27/368 (7%)
Query: 3 VSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIF----LPMPNPIPYG 58
VS L ++ +T + + S Y++ +FGDSI+DTGN + LP +PYG
Sbjct: 12 VSFLLILYYTTIVVASSESRCR--RYKSIISFGDSIADTGNYVHLSNVNNLPQAAFLPYG 69
Query: 59 SSYFKHPSGRMSNGRLIIDFIAEAYGLPFLPAY---ENKSIDQDIKKGVNFAFAGATVLN 115
S+F PSGR S+GRL+IDFIAE GLP++P Y +N S +Q G+NFA GAT L+
Sbjct: 70 ESFFHPPSGRYSDGRLVIDFIAEFLGLPYVPPYFGSQNVSFNQ----GINFAVYGATALD 125
Query: 116 VEYYVKNGLPLPDTNNSLSIQLGWFKNIKPLLCKSK-EDCNIYFKKSLFIVGEIGGNDI- 173
+ VK G+ TN SLS+QL FK I P LC S DC SL ++GEIGGND
Sbjct: 126 RAFLVKQGIKSDFTNISLSVQLNTFKQILPNLCASSTRDCREMLGDSLILMGEIGGNDYN 185
Query: 174 MKHMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNK 233
+ K++ E++E+VP +++AI++ LI+ G +VPGNFP+GCS A TL +
Sbjct: 186 YPFFEGKSINEIKELVPLIIKAISSAIVDLIDLGGKTFLVPGNFPIGCSTAYLTLFQTAT 245
Query: 234 KEDYDEFGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYG 293
E GC+ N E+ N QLK ++ L++ +P V IIY DYYN L+Q P +YG
Sbjct: 246 VEHDPFTGCIPWLNKFGEHHNEQLKIELKQLQKLYPHVNIIYADYYNSLYGLFQEPAKYG 305
Query: 294 FDKDAIFKACCG-----------GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLV 342
F K+ ACCG CG + C +PS+ +NWDG H TEA Y+ +A+GL+
Sbjct: 306 F-KNRPLAACCGVGGQYNFTIGKECGENGVSYCQNPSEYVNWDGYHLTEATYQKMAQGLL 364
Query: 343 EGPFSNPS 350
G ++ P+
Sbjct: 365 NGRYTTPA 372
>At1g28640 lipase, putative
Length = 390
Score = 255 bits (651), Expect = 3e-68
Identities = 145/344 (42%), Positives = 205/344 (59%), Gaps = 26/344 (7%)
Query: 28 YEAFFNFGDSISDTGNAASI----FLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAY 83
+++ +FGDSI+DTGN + LP +PYG S+F PSGR S+GRLIIDFIAE
Sbjct: 33 FKSIISFGDSIADTGNYLHLSDVNHLPQSAFLPYGESFFHPPSGRYSDGRLIIDFIAEFL 92
Query: 84 GLPFLPAY---ENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWF 140
GLP++P+Y +N S DQ G+NFA GAT L+ + V G+ TN SLS+QL F
Sbjct: 93 GLPYVPSYFGSQNVSFDQ----GINFAVYGATALDRVFLVGKGIESDFTNVSLSVQLNIF 148
Query: 141 KNIKPLLC-KSKEDCNIYFKKSLFIVGEIGGNDI-MKHMKHKTVIELREIVPFMVEAITN 198
K I P LC S DC SL ++GEIG ND + K++ E++++VP +++AI++
Sbjct: 149 KQILPNLCTSSSRDCREMLGDSLILMGEIGVNDYNYPFFEGKSINEIKQLVPLVIKAISS 208
Query: 199 TTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF-GCLIAYNNLIEYFNGQL 257
LI+ G +VPGNFP+GC A TL + +ED+D F GC+ N EY N QL
Sbjct: 209 AIVDLIDLGGKTFLVPGNFPLGCYPAYLTLFQTAAEEDHDPFTGCIPRLNEFGEYHNEQL 268
Query: 258 KNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG-----------G 306
K ++ L++ + V IIY DYYN RLYQ P +YGF K+ ACCG
Sbjct: 269 KTELKRLQELYDHVNIIYADYYNSLFRLYQEPVKYGF-KNRPLAACCGVGGQYNFTIGKE 327
Query: 307 CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPS 350
CG + C +PS+ +NWDG H TEA ++ +A+ ++ G +++P+
Sbjct: 328 CGHRGVSCCQNPSEYVNWDGYHLTEATHQKMAQVILNGTYASPA 371
>At1g28660 unknown protein
Length = 382
Score = 251 bits (642), Expect = 3e-67
Identities = 145/343 (42%), Positives = 197/343 (57%), Gaps = 26/343 (7%)
Query: 28 YEAFFNFGDSISDTGNAASI----FLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAY 83
+ + +FGDSI+DTGN + LP PYG S+F PSGR S+GRLIIDFIAE
Sbjct: 33 FTSIISFGDSIADTGNILHLSDVNHLPQTAFFPYGESFFHPPSGRASDGRLIIDFIAEFL 92
Query: 84 GLPFLPAY---ENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWF 140
GLP++P Y +N S +Q G+NFA GAT L+ Y+V G+ TN SL +QL F
Sbjct: 93 GLPYVPPYFGSQNVSFEQ----GINFAVYGATALDRAYFVAKGIESDFTNVSLGVQLDIF 148
Query: 141 KNIKPLLC-KSKEDCNIYFKKSLFIVGEIGGNDIMKHMKHKTVIELREIVPFMVEAITNT 199
K I P LC S DC SL ++GEIGGND I ++ +++AI++
Sbjct: 149 KQILPNLCASSSRDCREMLGDSLILMGEIGGNDFFYPSSEGKSINETKLQDLIIKAISSA 208
Query: 200 TNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEF-GCLIAYNNLIEYFNGQLK 258
+ LI G +VPG FP GCSAA T + +EDYD GC+ N L E+ N QLK
Sbjct: 209 ID-LIALGGKTFLVPGGFPAGCSAACLTQYQNATEEDYDPLTGCIPRLNELGEHDNEQLK 267
Query: 259 NSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG-----------GC 307
++ L++ +P+V IIY DY+N R YQ P +YGF K+ ACCG C
Sbjct: 268 TELKRLQKLYPDVNIIYADYHNSLYRFYQEPAKYGF-KNKPLAACCGVGGKYNFTIGKEC 326
Query: 308 GSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPS 350
G + C +PS+ +NWDG H TEAAY+ +A+G++ GP++ P+
Sbjct: 327 GYEGVSYCQNPSEYVNWDGYHLTEAAYQKMAEGILNGPYATPA 369
>At3g48460 lipase - like protein
Length = 381
Score = 200 bits (508), Expect = 1e-51
Identities = 123/351 (35%), Positives = 183/351 (52%), Gaps = 31/351 (8%)
Query: 28 YEAFFNFGDSISDTGNAASIFLP-----MPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEA 82
+ + FGDS +DTGN+ S P + +P PYG ++F+ P+ R S+GRL IDF+AE+
Sbjct: 36 FNKIYAFGDSFTDTGNSRSGEGPAGFGHLSSP-PYGMTFFRRPTNRYSDGRLTIDFVAES 94
Query: 83 YGLPFLPAY-----ENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQL 137
LPFLP Y N + GVNFA +G+TV+ ++VKN L L T S+ +L
Sbjct: 95 MNLPFLPPYLSLKTTNANGTATDTHGVNFAVSGSTVIKHAFFVKNNLSLDMTPQSIETEL 154
Query: 138 GWFKNIKPLLCKSKEDCNIYFKKSLFIVGEIGGNDIMKHMKHKTVIELREIVPFMVEAIT 197
WF+ L +++ FK SLF +GEIG ND + + + I + T
Sbjct: 155 AWFEKYLETLGTNQKVS--LFKDSLFWIGEIGVNDYAYTLG--STVSSDTIRELSISTFT 210
Query: 198 NTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIEYFNGQL 257
L+ +G ++V G+ GC +L ++D D GC+ + NN N L
Sbjct: 211 RFLETLLNKGVKYMLVQGHPATGCLTLAMSLA---AEDDRDSLGCVQSANNQSYTHNLAL 267
Query: 258 KNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG-----------G 306
++ ++ LR K+P I+Y DY+N + + + P +YG + FKACCG
Sbjct: 268 QSKLKQLRIKYPSATIVYADYWNAYRAVIKHPSKYGITEK--FKACCGIGEPYNFQVFQT 325
Query: 307 CGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPSLKSPLFK 357
CG+ ATVC DP++ INWDG H TEA YK++A ++G F+ P L K
Sbjct: 326 CGTDAATVCKDPNQYINWDGVHLTEAMYKVMADMFLDGTFTRPRFSDLLIK 376
>At5g03980 lipase -like protein
Length = 323
Score = 177 bits (449), Expect = 8e-45
Identities = 114/340 (33%), Positives = 163/340 (47%), Gaps = 62/340 (18%)
Query: 18 NVVSNANPLSYEAFFNFGDSISDTGNA---ASIFLPMPNPIPYGSSYFKHPSGRMSNGRL 74
++V +++ + + FGDSISDTGN P P P+P
Sbjct: 17 SLVHSSDQCPINSIYQFGDSISDTGNLIRNGPASSPTPKPLP------------------ 58
Query: 75 IIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLS 134
++ VNF +G+T LN ++ + L +P TN LS
Sbjct: 59 ----------------------QREHNVFVNFGVSGSTALNSSFFSERNLHVPATNTPLS 96
Query: 135 IQLGWFKNIKPLLCK-SKEDCNIYFKKSLFIVGEIGGNDI-MKHMKHKTVIELREIVPFM 192
+QL WFK C S DC K SLF+VGEIGGND + K + E+R +P +
Sbjct: 97 MQLAWFKGHLRSTCHGSSSDC---LKHSLFMVGEIGGNDYNYGFFQGKPMEEIRSYIPHV 153
Query: 193 VEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIEY 252
V AIT +I GAV +VVPGNFP+GC T +DYD+ GCL N
Sbjct: 154 VGAITAAAREVIRAGAVNVVVPGNFPVGCFPIYLTSFPVKDTKDYDDNGCLTHLNEFAMD 213
Query: 253 FNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG------- 305
N QL+ +I +LR++ P+V I+Y DYYN + + ++ + FDK K+CCG
Sbjct: 214 HNNQLQEAIASLRKEFPDVAIVYGDYYNAFQYVLRSER---FDKSVALKSCCGTGGAYNY 270
Query: 306 ----GCGSLIATVCSDPSKRINWDGPHFTEAAYKLIAKGL 341
G++ VC +P K I+WDG H T+ AY+ ++K L
Sbjct: 271 DGKRPYGAVGVPVCQNPHKFISWDGVHLTQKAYRFMSKFL 310
>At1g54790
Length = 383
Score = 172 bits (435), Expect = 3e-43
Identities = 115/354 (32%), Positives = 172/354 (48%), Gaps = 42/354 (11%)
Query: 26 LSYEAFFNFGDSISDTGNAASIFLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAYGL 85
L Y A NFGDS SDTGN S + NP PYG +YF PSGR +GRLI+DF+ + L
Sbjct: 28 LKYPAIINFGDSNSDTGNLISAGIENVNP-PYGQTYFNLPSGRYCDGRLIVDFLLDEMDL 86
Query: 86 PFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWFKNIK- 144
PFL Y + + KKG NFA AG+T+L P + S +Q+ F K
Sbjct: 87 PFLNPYLDSLGLPNFKKGCNFAAAGSTILPAN-------PTSVSPFSFDLQISQFIRFKS 139
Query: 145 ---PLLCKSKEDCN------IYFKKSLFIVGEIGGNDIMKHMKHKTVIELREIVPFMVEA 195
LL K+ Y+ K L+++ +IG NDI KT+ ++ +P ++E
Sbjct: 140 RAIELLSKTGRKYEKYLPPIDYYSKGLYMI-DIGQNDIAGAFYSKTLDQVLASIPSILET 198
Query: 196 ITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIEYFNG 255
L EEG + + P+GC A ++ + DEFGC+ ++N + FN
Sbjct: 199 FEAGLKRLYEEGGRNIWIHNTGPLGCLAQNIAKFGTDSTK-LDEFGCVSSHNQAAKLFNL 257
Query: 256 QLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG---------- 305
QL + ++P+ + Y D ++ L ++GF+K + ACCG
Sbjct: 258 QLHAMSNKFQAQYPDANVTYVDIFSIKSNLIANYSRFGFEKPLM--ACCGVGGAPLNYDS 315
Query: 306 --GCG--------SLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNP 349
CG S+ A C+D S+ INWDG H+TEAA + ++ ++ G +S+P
Sbjct: 316 RITCGQTKVLDGISVTAKACNDSSEYINWDGIHYTEAANEFVSSQILTGKYSDP 369
>At4g01130 putative acetyltransferase
Length = 382
Score = 165 bits (417), Expect = 4e-41
Identities = 119/353 (33%), Positives = 168/353 (46%), Gaps = 43/353 (12%)
Query: 28 YEAFFNFGDSISDTGNAASIFLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAYGLPF 87
+EA FNFGDS SDTG + F P +G +YFK P+GR S+GRLIIDF+A++ G+PF
Sbjct: 32 FEAIFNFGDSNSDTGGFWAAFPAQSGP--WGMTYFKKPAGRASDGRLIIDFLAKSLGMPF 89
Query: 88 LPAYENKSIDQDIKKGVNFAFAGATVL--NVEYYVKNGLPLPDTNNSLSIQLGWFKNIKP 145
L Y +SI D + G NFA +TVL N +V P SL+IQL K K
Sbjct: 90 LSPY-LQSIGSDFRHGANFATLASTVLLPNTSLFVSGISPF-----SLAIQLNQMKQFKV 143
Query: 146 LLCKSKE---------DCNIYFKKSLFIVGEIGGNDIMKHMKHKTVIELREIVPFMVEAI 196
+ +S I F KSL+ IG ND ++ V ++ +P ++ I
Sbjct: 144 NVDESHSLDRPGLKILPSKIVFGKSLYTF-YIGQNDFTSNLASIGVERVKLYLPQVIGQI 202
Query: 197 TNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIEYFNGQ 256
T + G +V P+GC A+ T ++ D D++GCLI N ++Y+N
Sbjct: 203 AGTIKEIYGIGGRTFLVLNLAPVGCYPAILT-GYTHTDADLDKYGCLIPVNKAVKYYNTL 261
Query: 257 LKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG----------- 305
L ++ R + +IY D + L+Q P+ YG KACCG
Sbjct: 262 LNKTLSQTRTELKNATVIYLDTHKILLDLFQHPKSYGMKHG--IKACCGYGGRPYNFNQK 319
Query: 306 -GCG--------SLIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNP 349
CG S A C DP ++WDG H TEAA I+ +++G S P
Sbjct: 320 LFCGNTKVIGNFSTTAKACHDPHNYVSWDGIHATEAANHHISMAILDGSISYP 372
>At3g05180 putative nodulin
Length = 379
Score = 158 bits (399), Expect = 5e-39
Identities = 113/350 (32%), Positives = 167/350 (47%), Gaps = 42/350 (12%)
Query: 28 YEAFFNFGDSISDTGNAAS--IFLPMPNPIPYGSSYFKHP-SGRMSNGRLIIDFIAEAYG 84
+ A FNFGDS SDTG +S FLP P+ Y ++F+ P SGR NGRLI+DF+ EA
Sbjct: 34 FPAVFNFGDSNSDTGELSSGLGFLPQPS---YEITFFRSPTSGRFCNGRLIVDFLMEAID 90
Query: 85 LPFLPAYENKSIDQDIKKGVNFAFAGATV--LNVEYYVKNGLPLPDTNNSLSIQLGWFKN 142
P+L Y + Q ++G NFA A +T+ N Y G + + Q FK+
Sbjct: 91 RPYLRPYLDSISRQTYRRGCNFAAAASTIQKANAASYSPFGFGVQVS------QFITFKS 144
Query: 143 IKPLLCKSKEDCNIYFKKSLFI-----VGEIGGNDIMKHMKHKTVIELREIVPFMVEAIT 197
L + E+ Y F + +IG NDI KTV ++ +VP +++
Sbjct: 145 KVLQLIQQDEELQRYLPSEYFFSNGLYMFDIGQNDIAGAFYTKTVDQVLALVPIILDIFQ 204
Query: 198 NTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIEYFNGQL 257
+ L EGA + P+GC A + ++ +K + DEFGC+ +N + FN QL
Sbjct: 205 DGIKRLYAEGARNYWIHNTGPLGCLAQVVSIFGEDKSK-LDEFGCVSDHNQAAKLFNLQL 263
Query: 258 KNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG------------ 305
+ L Q++P + Y D ++ L +YGFD + CCG
Sbjct: 264 HGLFKKLPQQYPNSRFTYVDIFSIKSDLILNHSKYGFDHSIM--VCCGTGGPPLNYDDQV 321
Query: 306 GCGS--------LIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFS 347
GCG + A C D SK +NWDG H+TEAA + +A ++ G +S
Sbjct: 322 GCGKTARSNGTIITAKPCYDSSKYVNWDGIHYTEAANRFVALHILTGKYS 371
>At5g14450 early nodule-specific protein - like
Length = 389
Score = 156 bits (394), Expect = 2e-38
Identities = 112/371 (30%), Positives = 173/371 (46%), Gaps = 31/371 (8%)
Query: 4 STLFLITFTCGFLQNVVSNANPL-SYEAFFNFGDSISDTGNAASIFLPMPNPIPYGSSYF 62
ST+F C F + P ++ A +NFGDS SDTG ++ F P+ +P YG +F
Sbjct: 14 STVFSWLLLCLFAVTTSVSVQPTCTFPAIYNFGDSNSDTGGISAAFEPIRDP--YGQGFF 71
Query: 63 KHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKN 122
P+GR S+GRL IDFIAE GLP+L AY N S+ + + G NFA G+T+ +
Sbjct: 72 HRPTGRDSDGRLTIDFIAERLGLPYLSAYLN-SLGSNFRHGANFATGGSTIRRQNETIFQ 130
Query: 123 GLPLPDTNNSLSIQLGWFKNIKPLL---CKSKEDCNIY-----FKKSLFIVGEIGGNDIM 174
P + + Q FK LL KS+ D F K+L+ +IG ND+
Sbjct: 131 YGISPFSLDMQIAQFDQFKARSALLFTQIKSRYDREKLPRQEEFAKALYTF-DIGQNDLS 189
Query: 175 KHMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKK 234
+ +V +L+ +P +V + + + ++G V P GC + +
Sbjct: 190 VGFRTMSVDQLKATIPDIVNHLASAVRNIYQQGGRTFWVHNTGPFGCLPVNMFYMGTPAP 249
Query: 235 EDYDEFGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGF 294
D+ GC+ A N + FN +LK ++ LR++ + I Y D Y + P++ GF
Sbjct: 250 GYLDKSGCVKAQNEMAMEFNRKLKETVINLRKELTQAAITYVDVYTAKYEMMSNPKKLGF 309
Query: 295 DKDAIFKACCG--------GCGS--------LIATVCSDPSKRINWDGPHFTEAAYKLIA 338
K CCG CG+ + C +P ++WDG H+TEAA K +A
Sbjct: 310 ANP--LKVCCGYHEKYDHIWCGNKGKVNNTEIYGGSCPNPVMAVSWDGVHYTEAANKHVA 367
Query: 339 KGLVEGPFSNP 349
+ G ++P
Sbjct: 368 DRTLNGLLTDP 378
>At1g67830 ENOD8-like protein
Length = 372
Score = 155 bits (391), Expect = 4e-38
Identities = 115/372 (30%), Positives = 175/372 (46%), Gaps = 42/372 (11%)
Query: 3 VSTLFLITFTCGFLQNVVSNANPLSYEAFFNFGDSISDTGNAASIFLPMPNPIPYGSSYF 62
+S+LF ++ + ++A+ + A FNFGDS SDTG ++ F P P+GSS+F
Sbjct: 5 LSSLFALSLLSSLSPS--THAHQCHFPAIFNFGDSNSDTGGLSAAF-GQAGP-PHGSSFF 60
Query: 63 KHPSGRMSNGRLIIDFIAEAYGLPFLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKN 122
P+GR +GRL+IDFIAE+ GLP+L A+ + S+ + G NFA AG+ + + ++
Sbjct: 61 GSPAGRYCDGRLVIDFIAESLGLPYLSAFLD-SVGSNFSHGANFATAGSPIRALNSTLRQ 119
Query: 123 GLPLPDTNNSLSIQLGWFKNIKPLLCKSKEDCNIY---------FKKSLFIVGEIGGNDI 173
P SL +Q F N + +Y F K+L+ +IG ND+
Sbjct: 120 SGFSP---FSLDVQFVQFYNFHNRSQTVRSRGGVYKTMLPESDSFSKALYTF-DIGQNDL 175
Query: 174 MK-HMKHKTVIELREIVPFMVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSN 232
+ +KTV ++ VP ++ N + +G + P+GC A + N
Sbjct: 176 TAGYFANKTVEQVETEVPEIISQFMNAIKNIYGQGGRYFWIHNTGPIGCLAYVIERF-PN 234
Query: 233 KKEDYDEFGCLIAYNNLIEYFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQY 292
K D+D GC+ N+L + FN LK ++ LR E I Y D Y+ L+ Q +
Sbjct: 235 KASDFDSHGCVSPLNHLAQQFNHALKQAVIELRSSLSEAAITYVDVYSLKHELFVHAQGH 294
Query: 293 GFDKDAIFKACCG-----------GCGS---------LIATVCSDPSKRINWDGPHFTEA 332
GF + +CCG GCG I C +P K + WDG HFT+A
Sbjct: 295 GFKGSLV--SCCGHGGKYNYNKGIGCGMKKIVKGKEVYIGKPCDEPDKAVVWDGVHFTQA 352
Query: 333 AYKLIAKGLVEG 344
A K I + G
Sbjct: 353 ANKFIFDKIAPG 364
>At3g26430 nodulin, putative
Length = 380
Score = 150 bits (379), Expect = 1e-36
Identities = 113/360 (31%), Positives = 170/360 (46%), Gaps = 50/360 (13%)
Query: 27 SYEAFFNFGDSISDTGNAASIFLPMPNPIPYGSSYFKHPSGRMSNGRLIIDFIAEAYGLP 86
++ A FNFGDS SDTG ++ F P P G ++F PSGR S+GRLIIDFIAE GLP
Sbjct: 28 NFPAIFNFGDSNSDTGGLSASFGQAP--YPNGQTFFHSPSGRFSDGRLIIDFIAEELGLP 85
Query: 87 FLPAYENKSIDQDIKKGVNFAFAGATVLNVEYYVKNGLPLPDTNNSLSIQLGWFKN---- 142
+L A+ + SI + G NFA AG+TV + P SL +QL F +
Sbjct: 86 YLNAFLD-SIGSNFSHGANFATAGSTVRPPNATIAQSGVSP---ISLDVQLVQFSDFITR 141
Query: 143 ----------IKPLLCKSKEDCNIYFKKSLFIVGEIGGNDIMKHMK-HKTVIELREIVPF 191
K LL K + YF ++L+ +IG ND+ +K + T +++ +P
Sbjct: 142 SQLIRNRGGVFKKLLPKKE-----YFSQALYTF-DIGQNDLTAGLKLNMTSDQIKAYIPD 195
Query: 192 MVEAITNTTNVLIEEGAVELVVPGNFPMGCSAAMFTLVNSNKKEDYDEFGCLIAYNNLIE 251
+ + ++N + +G + P+GC + + D GC I N +
Sbjct: 196 VHDQLSNVIRKVYSKGGRRFWIHNTAPLGCLPYVLDRFPVPASQ-IDNHGCAIPRNEIAR 254
Query: 252 YFNGQLKNSIETLRQKHPEVKIIYFDYYNDAKRLYQTPQQYGFDKDAIFKACCG------ 305
Y+N +LK + LR++ E Y D Y+ L ++ GF + ACCG
Sbjct: 255 YYNSELKRRVIELRKELSEAAFTYVDIYSIKLTLITQAKKLGFRYPLV--ACCGHGGKYN 312
Query: 306 -----GCGS---------LIATVCSDPSKRINWDGPHFTEAAYKLIAKGLVEGPFSNPSL 351
CG+ ++A C+D S R++WDG HFTE I + + +G FS+P L
Sbjct: 313 FNKLIKCGAKVMIKGKEIVLAKSCNDVSFRVSWDGIHFTETTNSWIFQQINDGAFSDPPL 372
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.140 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,760,849
Number of Sequences: 26719
Number of extensions: 404581
Number of successful extensions: 1352
Number of sequences better than 10.0: 123
Number of HSP's better than 10.0 without gapping: 102
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 912
Number of HSP's gapped (non-prelim): 135
length of query: 359
length of database: 11,318,596
effective HSP length: 100
effective length of query: 259
effective length of database: 8,646,696
effective search space: 2239494264
effective search space used: 2239494264
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC130806.4


