

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130804.6 + phase: 0 /pseudo
(99 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
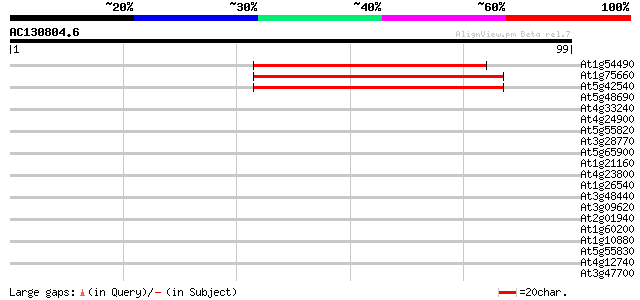
Score E
Sequences producing significant alignments: (bits) Value
At1g54490 unknown protein 66 3e-12
At1g75660 Dhp1-like protein 62 4e-11
At5g42540 5'-3' exoribonuclease 2 61 8e-11
At5g48690 unknown protein 30 0.27
At4g33240 putative protein 29 0.45
At4g24900 unknown protein 28 0.59
At5g55820 unknown protein 28 0.77
At3g28770 hypothetical protein 28 0.77
At5g65900 ATP-dependent RNA helicase-like 28 1.0
At1g21160 transcription factor, putative 28 1.0
At4g23800 98b like protein 27 1.3
At1g26540 unknown protein 27 1.3
At3g48440 unknown protein 27 1.7
At3g09620 putative RNA helicase 27 1.7
At2g01940 putative C2H2-type zinc finger protein 27 1.7
At1g60200 27 1.7
At1g10880 hypothetical protein 27 1.7
At5g55830 serine/threonine-specific kinase like protein 26 2.9
At4g12740 adenine DNA glycosylase like protein 26 2.9
At3g47700 unknown protein 26 2.9
>At1g54490 unknown protein
Length = 947
Score = 66.2 bits (160), Expect = 3e-12
Identities = 32/41 (78%), Positives = 35/41 (85%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEME 84
IDGV PRAKMN+QR+RRFR AKD EAEEERLRK+FEME
Sbjct: 103 IDGVAPRAKMNQQRSRRFRAAKDAAEAEAEEERLRKDFEME 143
>At1g75660 Dhp1-like protein
Length = 1020
Score = 62.4 bits (150), Expect = 4e-11
Identities = 30/44 (68%), Positives = 35/44 (79%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
IDGV PRAKMN+QR+RRFR+AKD AEEERLR+EFE E +
Sbjct: 102 IDGVAPRAKMNQQRSRRFRSAKDASDAAAEEERLREEFEREGRR 145
>At5g42540 5'-3' exoribonuclease 2
Length = 746
Score = 61.2 bits (147), Expect = 8e-11
Identities = 30/44 (68%), Positives = 33/44 (74%)
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
IDGV PRAKMN+QR RRFR AKD AEEE+LR+EFE E K
Sbjct: 104 IDGVAPRAKMNQQRARRFRAAKDAAEAAAEEEQLREEFEREGKK 147
>At5g48690 unknown protein
Length = 301
Score = 29.6 bits (65), Expect = 0.27
Identities = 19/72 (26%), Positives = 37/72 (51%), Gaps = 1/72 (1%)
Query: 29 KFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEM-REAEEERLRKEFEMEANK 87
+FN + +P P+ + RAK ++ R+ R ++ + RE E+ER+R E+ K
Sbjct: 43 EFNIEIESPNPSDDTAETSHARAKELSEQARKLREEQETKREREREKERIRAGKELMETK 102
Query: 88 FFLNKNVKCQNL 99
+N + +N+
Sbjct: 103 RIAEENERKRNI 114
>At4g33240 putative protein
Length = 1757
Score = 28.9 bits (63), Expect = 0.45
Identities = 13/40 (32%), Positives = 22/40 (54%)
Query: 40 TLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRK 79
T+ G +PR KMN + +RR + D+E E + + R+
Sbjct: 139 TIDSTAGPSPRPKMNPRASRRVSSNMDSEKSEQQNAKSRR 178
>At4g24900 unknown protein
Length = 421
Score = 28.5 bits (62), Expect = 0.59
Identities = 20/67 (29%), Positives = 27/67 (39%), Gaps = 8/67 (11%)
Query: 36 APPPTLPDIDGVTPRAKMNKQRTRRF--------RTAKDNEMREAEEERLRKEFEMEANK 87
APPP L DG ++N+ RF R N + A ER + E EME +
Sbjct: 295 APPPWLDANDGDFSSVQLNQSDVARFQAKVPGKNRKLNPNRVGAAWAERRKIEIEMEKSG 354
Query: 88 FFLNKNV 94
N+
Sbjct: 355 HVTKSNI 361
>At5g55820 unknown protein
Length = 1826
Score = 28.1 bits (61), Expect = 0.77
Identities = 16/41 (39%), Positives = 23/41 (56%), Gaps = 5/41 (12%)
Query: 52 KMNKQRTRRFRTAK-----DNEMREAEEERLRKEFEMEANK 87
K K+ R+ + A+ + E ++ EEER RKEFEM K
Sbjct: 1565 KKKKEEDRKKKEAEMAWKQEMEKKKKEEERKRKEFEMADRK 1605
>At3g28770 hypothetical protein
Length = 2081
Score = 28.1 bits (61), Expect = 0.77
Identities = 13/44 (29%), Positives = 26/44 (58%)
Query: 52 KMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANKFFLNKNVK 95
K K+ ++ + K+ EM+E+EE++L+K E + + +N K
Sbjct: 1180 KKEKKSSKDQQKKKEKEMKESEEKKLKKNEEDRKKQTSVEENKK 1223
>At5g65900 ATP-dependent RNA helicase-like
Length = 633
Score = 27.7 bits (60), Expect = 1.0
Identities = 13/41 (31%), Positives = 22/41 (52%)
Query: 49 PRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANKFF 89
P+ K KQR +++ E+ + EEE+ +E + NK F
Sbjct: 116 PKKKKKKQRKDTEAKSEEEEVEDKEEEKKLEETSIMTNKTF 156
>At1g21160 transcription factor, putative
Length = 1088
Score = 27.7 bits (60), Expect = 1.0
Identities = 17/35 (48%), Positives = 23/35 (65%), Gaps = 2/35 (5%)
Query: 49 PRAKMNKQRT-RRFRTAKDNEMREAEEERLRKEFE 82
P+ KQ T R++ A+D + +E EEERLRKE E
Sbjct: 219 PKHVREKQETLARWKEAEDGKKKE-EEERLRKEEE 252
>At4g23800 98b like protein
Length = 456
Score = 27.3 bits (59), Expect = 1.3
Identities = 15/52 (28%), Positives = 25/52 (47%)
Query: 41 LPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANKFFLNK 92
L ++ + + K+ K +T KD +R+ EEE ++ E E K L K
Sbjct: 48 LMEMQTMLEKMKIEKDKTEELLKEKDEILRKKEEELETRDAEQEKLKVELKK 99
>At1g26540 unknown protein
Length = 695
Score = 27.3 bits (59), Expect = 1.3
Identities = 16/44 (36%), Positives = 26/44 (58%), Gaps = 1/44 (2%)
Query: 45 DGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANKF 88
D TP++++NK + + AK E R+ E+R+ E E+E KF
Sbjct: 592 DIATPQSRINKVLSLQVGRAKKVEERKCLEKRIEAE-EIEMQKF 634
>At3g48440 unknown protein
Length = 448
Score = 26.9 bits (58), Expect = 1.7
Identities = 10/24 (41%), Positives = 17/24 (70%)
Query: 52 KMNKQRTRRFRTAKDNEMREAEEE 75
K N R+F+ A+DN++RE E++
Sbjct: 132 KFNHPLARKFQIARDNKVREKEDD 155
>At3g09620 putative RNA helicase
Length = 989
Score = 26.9 bits (58), Expect = 1.7
Identities = 16/50 (32%), Positives = 27/50 (54%), Gaps = 3/50 (6%)
Query: 50 RAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANKFFLNK---NVKC 96
R + + R R +D E RE E+E+ RK+ + E ++ N+ +VKC
Sbjct: 75 RGRRERDRDRGKYLKRDRERREREKEKGRKKQKKERSREDCNEESDDVKC 124
>At2g01940 putative C2H2-type zinc finger protein
Length = 439
Score = 26.9 bits (58), Expect = 1.7
Identities = 13/45 (28%), Positives = 25/45 (54%)
Query: 51 AKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANKFFLNKNVK 95
A+ K+ +R R +NE A++ R + + E+E KF +++K
Sbjct: 344 AEEAKREAKRQREIAENEFANAKKIRQKAQAELERAKFLKEQSMK 388
>At1g60200
Length = 781
Score = 26.9 bits (58), Expect = 1.7
Identities = 18/36 (50%), Positives = 22/36 (61%), Gaps = 5/36 (13%)
Query: 51 AKMNKQRTRRFRTAKDNEM----REAEEERLRKEFE 82
+K N +R+R R K+ EM REAE ER RKE E
Sbjct: 288 SKRNDRRSRE-RGEKEQEMDRYEREAERERSRKERE 322
>At1g10880 hypothetical protein
Length = 651
Score = 26.9 bits (58), Expect = 1.7
Identities = 14/49 (28%), Positives = 25/49 (50%)
Query: 43 DIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANKFFLN 91
DID + A++ +Q+ T + E R ++EFE E ++ F+N
Sbjct: 509 DIDSGSSLAQVEEQKWPYLNTTSTITIDEETYSRKQREFECERDQVFVN 557
>At5g55830 serine/threonine-specific kinase like protein
Length = 681
Score = 26.2 bits (56), Expect = 2.9
Identities = 13/36 (36%), Positives = 20/36 (55%)
Query: 61 FRTAKDNEMREAEEERLRKEFEMEANKFFLNKNVKC 96
+R + + EA +ERL+ EF+ E K L +KC
Sbjct: 586 WRLHSEGRVLEAVDERLKGEFDEEMMKKLLLVGLKC 621
>At4g12740 adenine DNA glycosylase like protein
Length = 608
Score = 26.2 bits (56), Expect = 2.9
Identities = 16/42 (38%), Positives = 23/42 (54%), Gaps = 3/42 (7%)
Query: 49 PRAKMNK---QRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
PR K+ + ++ R A+ REAEEE +E E EA+K
Sbjct: 49 PREKLMRKCREKKEAEREAEREAEREAEEEEKAEEAEAEADK 90
>At3g47700 unknown protein
Length = 795
Score = 26.2 bits (56), Expect = 2.9
Identities = 17/64 (26%), Positives = 27/64 (41%), Gaps = 3/64 (4%)
Query: 24 ETLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEM 83
E L+ F W + P P + KM +++ + + D +R E R+R E
Sbjct: 727 EVLYGVFRTWCVRPEGFFPKLSEGLTLLKMEEKQVKDGLSRGDKWLR---ENRIRYLSEA 783
Query: 84 EANK 87
EA K
Sbjct: 784 EAKK 787
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,303,888
Number of Sequences: 26719
Number of extensions: 84462
Number of successful extensions: 419
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 379
Number of HSP's gapped (non-prelim): 60
length of query: 99
length of database: 11,318,596
effective HSP length: 75
effective length of query: 24
effective length of database: 9,314,671
effective search space: 223552104
effective search space used: 223552104
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC130804.6


