

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
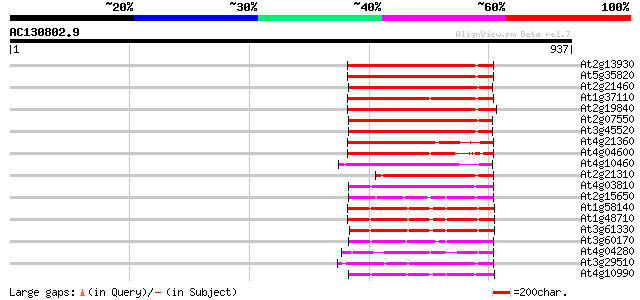
Query= AC130802.9 + phase: 0 /pseudo
(937 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At2g13930 putative retroelement pol polyprotein 258 1e-68
At5g35820 copia-like retrotransposable element 247 2e-65
At2g21460 putative retroelement pol polyprotein 239 4e-63
At1g37110 239 7e-63
At2g19840 copia-like retroelement pol polyprotein 232 6e-61
At2g07550 putative retroelement pol polyprotein 228 1e-59
At3g45520 copia-like polyprotein 226 4e-59
At4g21360 putative transposable element 198 1e-50
At4g04600 putative polyprotein 191 1e-48
At4g10460 putative retrotransposon 188 1e-47
At2g21310 putative retroelement pol polyprotein 177 2e-44
At4g03810 putative retrotransposon protein 168 1e-41
At2g15650 putative retroelement pol polyprotein 168 1e-41
At1g58140 hypothetical protein 166 6e-41
At1g48710 hypothetical protein 166 6e-41
At3g61330 copia-type polyprotein 163 5e-40
At3g60170 putative protein 159 7e-39
At4g04280 putative transposon protein 155 1e-37
At3g29510 hypothetical protein 149 9e-36
At4g10990 putative retrotransposon polyprotein 147 3e-35
>At2g13930 putative retroelement pol polyprotein
Length = 1335
Score = 258 bits (658), Expect = 1e-68
Identities = 128/243 (52%), Positives = 174/243 (70%), Gaps = 2/243 (0%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+ SLYGLKQSPR+W RFDEF+ + RS YDSCVY K N +YLLLYVDD+L+AS
Sbjct: 963 KRSLYGLKQSPRQWNLRFDEFMRGIKYTRSAYDSCVYFKKCNGDTYIYLLLYVDDMLIAS 1022
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
++K E+ +LK+ L+ EFEMKDLG AK++LG++I R+RD G L LSQ GY+KK F+M
Sbjct: 1023 ANKSEVNELKQLLSREFEMKDLGDAKKILGMEISRDRDAGLLTLSQEGYVKKVLRSFQMD 1082
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
N+K VSTPLG H KL + E++ + M+ PYA+ +GSIMY M+ +RPDLAY++ ++
Sbjct: 1083 NAKPVSTPLGIHFKLKAATDKEYEEQFERMKIVPYANTIGSIMYSMIGTRPDLAYSLGVI 1142
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SRFM+ P HWQA+KWVLRY+ G+ K L + + Q++ L GY D+DY N DTR+S+
Sbjct: 1143 SRFMSKPLKDHWQAVKWVLRYMRGTEKKKLCFRK--QEDFLLRGYCDSDYGSNFDTRRSI 1200
Query: 805 VGF 807
G+
Sbjct: 1201 TGY 1203
>At5g35820 copia-like retrotransposable element
Length = 1342
Score = 247 bits (631), Expect = 2e-65
Identities = 122/243 (50%), Positives = 168/243 (68%), Gaps = 2/243 (0%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQSPR+W +RFD F++ +G+ RS Y+ CVY + N+ +YLLLYVDD+L+AS
Sbjct: 970 KKSLYGLKQSPRQWNQRFDSFMINSGYQRSKYNPCVYTQQLNDGSYIYLLLYVDDMLIAS 1029
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
+KD+I KLKE LN EFEMKDLGPA+++LG++I RNR++G L LSQ Y+ F M
Sbjct: 1030 QNKDQIQKLKESLNREFEMKDLGPARKILGMEITRNREQGILDLSQSEYVAGVLRAFGMD 1089
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
SK TPLG H KL + + M+ PY + +GSIMY M+ SRPDLAY V +V
Sbjct: 1090 QSKVSQTPLGAHFKLRAANEKTLARDAEYMKLVPYPNAIGSIMYSMIGSRPDLAYPVGVV 1149
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SRFM+ P HWQA+KWV+RY+ G+ L++ + D+ + GY D+DYA ++D R+S+
Sbjct: 1150 SRFMSKPSKEHWQAVKWVMRYMKGTQDTCLRFKK--DDKFEIRGYCDSDYATDLDRRRSI 1207
Query: 805 VGF 807
GF
Sbjct: 1208 TGF 1210
>At2g21460 putative retroelement pol polyprotein
Length = 1333
Score = 239 bits (611), Expect = 4e-63
Identities = 117/241 (48%), Positives = 170/241 (69%), Gaps = 2/241 (0%)
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASS 625
+SLYGLKQ+P++W +F+ ++ + GF+RS YDSC Y+ + ++ +YLLLYVDD+L+A+
Sbjct: 962 KSLYGLKQAPKQWNEKFNAYMSEIGFIRSLYDSCAYIKELSDGSRVYLLLYVDDMLVAAK 1021
Query: 626 SKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSN 685
+K++I +LKE L+ F+MKDLG AKR+LG++I RNR++ L+LSQ GYL K E + M+
Sbjct: 1022 NKEDISQLKEELSQRFDMKDLGAAKRILGMEIIRNREENTLWLSQNGYLNKILETYNMAE 1081
Query: 686 SKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVS 745
SK V TPLG H K+ + E ++ M+ PY+S VGSIMY M+ +RPDLAY V I+S
Sbjct: 1082 SKHVVTPLGAHLKMRAATVEKQEQDEDYMKSIPYSSAVGSIMYAMIGTRPDLAYPVGIIS 1141
Query: 746 RFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLV 805
R+M+ P HW +KWVLRY+ GSL L+Y R++ + + GY DAD+A D R+S+
Sbjct: 1142 RYMSQPAREHWLGVKWVLRYIKGSLGTKLQYKRSS--DFKVVGYCDADHAACKDRRRSIT 1199
Query: 806 G 806
G
Sbjct: 1200 G 1200
>At1g37110
Length = 1356
Score = 239 bits (609), Expect = 7e-63
Identities = 119/243 (48%), Positives = 174/243 (70%), Gaps = 5/243 (2%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQSPR+W +RFD F+ F+RS +D+CVY+ +E +YLLLYVDD+L+A
Sbjct: 986 KKSLYGLKQSPRQWNKRFDRFMSSQQFIRSEHDACVYVKHVSEHDFIYLLLYVDDMLIAG 1045
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
+SK EI ++KE+L+ EFEMKD+G A R+LGIDI R+R G L LSQ Y++K +RF MS
Sbjct: 1046 ASKAEINRVKEQLSTEFEMKDMGGASRILGIDIYRDRKGGVLKLSQEIYIRKVLDRFNMS 1105
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
+K + P+G H KL+ + EDE + PY+S VGSIMY M+ +RPDLAYA+ ++
Sbjct: 1106 GAKMTNAPVGAHFKLA---AVREEDECVDTDVVPYSSAVGSIMYAMLGTRPDLAYAICLI 1162
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+M+ PG +HW+A+KWV+RYL G+ L +T+ + + + GY D++YA ++D R+S+
Sbjct: 1163 SRYMSKPGSMHWEAVKWVMRYLKGAQDLNLVFTK--EKDFTVTGYCDSNYAADLDRRRSI 1220
Query: 805 VGF 807
G+
Sbjct: 1221 SGY 1223
>At2g19840 copia-like retroelement pol polyprotein
Length = 1137
Score = 232 bits (592), Expect = 6e-61
Identities = 121/248 (48%), Positives = 161/248 (64%), Gaps = 2/248 (0%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+ SLYGL+QSPR+W RF+EF+ K G+ RS YDSCVY + +YLLLYVDDIL+AS
Sbjct: 781 KRSLYGLRQSPRQWNNRFNEFMQKIGYERSKYDSCVYFKELQSGEYIYLLLYVDDILIAS 840
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
K + LK LN EFEMKDLG AK++LG++I R+R G + +SQ GYL K F M
Sbjct: 841 RDKRTVCDLKALLNSEFEMKDLGDAKKILGMEIVRDRKAGTMSISQEGYLLKVLGNFGMD 900
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
+K V TP+G H KL + + ++M PY S VGS+MY M+ +RPDLA++V +V
Sbjct: 901 QAKPVFTPMGAHFKLKPATDEEVMRQSEVMRAVPYQSAVGSLMYSMIGTRPDLAHSVGLV 960
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
RFM+ P HWQA+KW+LRY+ GS+ L Y + E LEGY D+DYA + + R+S
Sbjct: 961 CRFMSKPLKEHWQAVKWILRYIRGSIDRKLCYKN--EGELILEGYCDSDYAADKEGRRST 1018
Query: 805 VGFCVYSL 812
G V +L
Sbjct: 1019 SGVKVVAL 1026
>At2g07550 putative retroelement pol polyprotein
Length = 1356
Score = 228 bits (581), Expect = 1e-59
Identities = 115/241 (47%), Positives = 161/241 (66%), Gaps = 2/241 (0%)
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASS 625
++LYGLKQ+P++W +FD F+ + FV+S YDSC Y + ++YLL+YVDDIL+AS
Sbjct: 985 KALYGLKQAPKQWNEKFDNFMKEICFVKSAYDSCAYTKVLPDGSVMYLLIYVDDILVASK 1044
Query: 626 SKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSN 685
+K+ I LK L FEMKDLG AK++LG++I R+R G L+LSQ GYL K E + M+
Sbjct: 1045 NKEAITALKANLGMRFEMKDLGAAKKILGMEIIRDRTLGVLWLSQEGYLNKILETYNMAE 1104
Query: 686 SKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVS 745
+K TPLG H K + ++ M+ PY+S VGSIMY M+ +RPDLAY V I+S
Sbjct: 1105 AKPAMTPLGAHFKFQAATEQKLIRDEDFMKSVPYSSAVGSIMYAMLGTRPDLAYPVGIIS 1164
Query: 746 RFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLV 805
RFM+ P HW +KWVLRY+ G+LK L Y +++ ++ GY DADYA ++D R+S+
Sbjct: 1165 RFMSQPIKEHWLGVKWVLRYIKGTLKTRLCYKKSS--SFSIVGYCDADYAADLDKRRSIT 1222
Query: 806 G 806
G
Sbjct: 1223 G 1223
>At3g45520 copia-like polyprotein
Length = 1363
Score = 226 bits (577), Expect = 4e-59
Identities = 114/241 (47%), Positives = 160/241 (66%), Gaps = 2/241 (0%)
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASS 625
+SLYGLKQ+PR+W +F+ ++ + GF RS YDSC Y K ++ +YLL YVDD+L+A++
Sbjct: 992 KSLYGLKQAPRQWNEKFNHYMTEIGFKRSDYDSCAYTKKLSDDSTMYLLFYVDDMLVAAN 1051
Query: 626 SKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSN 685
+ I LK+ L+ +FEMKDLG AK++LGI+I +R+ G L+LSQ YL K + F M
Sbjct: 1052 NMQAIDALKKELSIKFEMKDLGAAKKILGIEIIIDREAGVLWLSQESYLNKVLKTFNMLE 1111
Query: 686 SKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVS 745
SK TPLG H K+ + E++ M PY+S VGSIMY M+ +RPDLAY V +VS
Sbjct: 1112 SKPALTPLGAHLKMKSATEEKLSTEEEYMNSVPYSSAVGSIMYAMIGTRPDLAYPVGVVS 1171
Query: 746 RFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLV 805
RFM+ P HW +KWVLRY+ G++ L Y R + ++ GY DADYA ++D R+S+
Sbjct: 1172 RFMSQPAKEHWLGVKWVLRYIKGTVDTRLCYKR--NSDFSICGYCDADYAADLDKRRSIT 1229
Query: 806 G 806
G
Sbjct: 1230 G 1230
>At4g21360 putative transposable element
Length = 1308
Score = 198 bits (504), Expect = 1e-50
Identities = 110/245 (44%), Positives = 157/245 (63%), Gaps = 36/245 (14%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQ+PR+W ++F F+L F RS +DSCVY+ + N +YLLLYVDD+L+A+
Sbjct: 967 KKSLYGLKQAPRQWNKKFHAFMLSLQFARSEHDSCVYVKEVNPGEFVYLLLYVDDMLLAA 1026
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
SK EI KLKE L+ +FEMKD+G A R+LGIDI RNR +G L LSQ Y+ K +RFRM+
Sbjct: 1027 KSKSEISKLKEALSLKFEMKDMGAASRILGIDIIRNRKEGTLRLSQTRYVDKVIQRFRMA 1086
Query: 685 NSKTVSTPLGHHTKLS--IQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVS 742
++K VSTP+G H KL+ I + + E PY+S VGS+MY M+ + PD+AYA+
Sbjct: 1087 DAKVVSTPMGAHFKLTSLIDEIGSVDPEV-----VPYSSAVGSVMYAMIGTIPDVAYAMG 1141
Query: 743 IVSRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRK 802
+VSRFM+ PG +L+ ++GY D+D+A ++D R+
Sbjct: 1142 LVSRFMSRPG---------------ANLE--------------VQGYCDSDHAADLDKRR 1172
Query: 803 SLVGF 807
S+ G+
Sbjct: 1173 SISGY 1177
>At4g04600 putative polyprotein
Length = 922
Score = 191 bits (486), Expect = 1e-48
Identities = 107/243 (44%), Positives = 157/243 (64%), Gaps = 30/243 (12%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++SLYGLKQ+PR+W ++F+ F++ GF RS +DSCVY+ + + +YLL+YVDD+L+A
Sbjct: 633 KKSLYGLKQAPRQWNKKFNAFMMDQGFTRSLHDSCVYVKEVSPDQFVYLLVYVDDMLIAG 692
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
S E+ K+KE L+ FEMKD+G A R+LGIDI+RNR++G L LSQ YL+K +RFRM+
Sbjct: 693 KSMAEVNKVKEGLSLNFEMKDMGAASRILGIDIERNREEGTLCLSQSKYLEKVIQRFRMA 752
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
++K VSTP+G H KLS ++ DE E PY+S VGS+MY M+ +RPD+ YA+ +V
Sbjct: 753 DAKGVSTPIGAHFKLS---AVRNNDESVDTEVCPYSSAVGSVMYAMIGTRPDVVYALGLV 809
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
NGS +GGL Y R ++ + D+DY ++D R+S+
Sbjct: 810 ----------------------NGS-EGGL-YKRKRHEDTWI---CDSDYVADLDKRRSI 842
Query: 805 VGF 807
G+
Sbjct: 843 SGY 845
>At4g10460 putative retrotransposon
Length = 1230
Score = 188 bits (478), Expect = 1e-47
Identities = 105/258 (40%), Positives = 151/258 (57%), Gaps = 30/258 (11%)
Query: 549 GTTGRFCGGQV*GVSFEESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEK 608
G +F G+V +++YGLKQSPR W ++FD ++L+ GF RS + C Y+ +
Sbjct: 870 GYESQFKQGEV--CLLNKTMYGLKQSPRRWNQKFDSYMLEIGFERSPRNKCAYIKSLEDG 927
Query: 609 VILYLLLYVDDILMASSSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFL 668
+YLL+YVDD+L+A+ I +LK++L+ +FEMKDLG AKR+LG++I R+R KG L L
Sbjct: 928 SKVYLLIYVDDMLVAARDMQVISELKQKLSEKFEMKDLGAAKRILGMEISRDRVKGTLTL 987
Query: 669 SQLGYLKKGGERFRMSNSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMY 728
SQ YL K E + + K V TPLG H K+ Q +++ M+ PY++ VGSIMY
Sbjct: 988 SQEDYLSKVLETYNVDQCKFVVTPLGAHLKMHAATEQQLLSDEEYMKSVPYSNAVGSIMY 1047
Query: 729 GMVCSRPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEG 788
M+ +RPDLAY V I+SRFM+ P KG + L G
Sbjct: 1048 SMIDTRPDLAYCVGIISRFMSKP-------------------KGA---------DLTLRG 1079
Query: 789 YVDADYAGNVDTRKSLVG 806
Y D+DYA N++ R+S+ G
Sbjct: 1080 YCDSDYAANLENRRSISG 1097
>At2g21310 putative retroelement pol polyprotein
Length = 838
Score = 177 bits (450), Expect = 2e-44
Identities = 87/196 (44%), Positives = 130/196 (65%), Gaps = 4/196 (2%)
Query: 612 YLLLYVDDILMASSSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQL 671
Y + ++DD+ +K+ + +LKERL+ EFEMKDLGPA+++LG++I RNR++ L LSQ
Sbjct: 515 YFITFIDDL--TRKNKETVKELKERLSSEFEMKDLGPARKILGMEITRNREENILELSQR 572
Query: 672 GYLKKGGERFRMSNSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMV 731
YL+K + FRM K V TPL H K ++E++ M+ PYA+ VGSIMY M+
Sbjct: 573 SYLQKVLKTFRMDECKPVKTPLAPHMKFVAATETEAEEQADQMKSIPYANAVGSIMYSMI 632
Query: 732 CSRPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVD 791
SRPDLA+ V ++SRFM+ P + HW +KWVLRY+ GS+ L+Y R + + + GY D
Sbjct: 633 GSRPDLAHPVGVISRFMSKPLMNHWLGVKWVLRYIKGSIDSRLQYKR--EGDFVVTGYCD 690
Query: 792 ADYAGNVDTRKSLVGF 807
+D++G+ D +S G+
Sbjct: 691 SDHSGDRDRSRSTTGY 706
>At4g03810 putative retrotransposon protein
Length = 964
Score = 168 bits (426), Expect = 1e-41
Identities = 94/241 (39%), Positives = 139/241 (57%), Gaps = 3/241 (1%)
Query: 567 SLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASSS 626
S+YGLKQ+ R W RF+E + + F+R+ + CVY K + + +L+LYVDDIL+ +
Sbjct: 590 SIYGLKQASRSWNLRFNEAIKEFDFIRNEEEPCVYK-KTSGSAVAFLVLYVDDILLLGND 648
Query: 627 KDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSNS 686
+ +K L F MKD+G A +LGI I R+R + LSQ Y+ K RF M +S
Sbjct: 649 IPLLQSVKTWLGSCFSMKDMGEAAYILGIRIYRDRLNKIIGLSQDTYIDKVLHRFNMHDS 708
Query: 687 KTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVSR 746
K P+ H LS QCP + DE++ M PYAS +GSIMY M+ +RPD+A A+S+ SR
Sbjct: 709 KKGFIPMSHGITLSKTQCPSTHDERERMSKIPYASAIGSIMYAMLYTRPDVACALSMTSR 768
Query: 747 FMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLVG 806
+ ++PG HW ++ + +YL + L Y +E + GY DA + + D +S G
Sbjct: 769 YQSDPGESHWIVVRNIFKYLRRTKDKFLVY--GGSEELVVSGYTDASFQTDKDDFRSQSG 826
Query: 807 F 807
F
Sbjct: 827 F 827
>At2g15650 putative retroelement pol polyprotein
Length = 1347
Score = 168 bits (425), Expect = 1e-41
Identities = 94/242 (38%), Positives = 147/242 (59%), Gaps = 10/242 (4%)
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASS 625
++LYGLKQ+PR WY R D + ++ GF RS D+ +Y KK E V++ + LYVDD+++ +
Sbjct: 981 KALYGLKQAPRAWYERIDSYFIQNGFARSMNDAALYSKKKGEDVLI-VSLYVDDLIITGN 1039
Query: 626 SKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSN 685
+ I K+ + EFEM DLG LG+++ N+D +FLSQ Y K ++F M
Sbjct: 1040 NTHLINTFKKNMKDEFEMTDLGLLNYFLGMEV--NQDDSGIFLSQEKYANKLIDKFGMKE 1097
Query: 686 SKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVS 745
SK+VSTPL K + D+K+ + T Y VG ++Y + SRPD+ YA S +S
Sbjct: 1098 SKSVSTPLTPQGK----RKGVEGDDKEFADPTKYRRIVGGLLY-LCASRPDVMYASSYLS 1152
Query: 746 RFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLV 805
R+M++P I H+Q K VLRY+ G+ G+ +T +++ L GY D+D+ G+++ +KS
Sbjct: 1153 RYMSSPSIQHYQEAKRVLRYVKGTSNFGVLFT--SKETPRLVGYSDSDWGGSLEDKKSTT 1210
Query: 806 GF 807
G+
Sbjct: 1211 GY 1212
>At1g58140 hypothetical protein
Length = 1320
Score = 166 bits (420), Expect = 6e-41
Identities = 92/246 (37%), Positives = 151/246 (60%), Gaps = 12/246 (4%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+++LYGLKQ+PR W R D++ + F++ Y+ +Y +K ++ IL LYVDD++
Sbjct: 949 KKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALY-IKIQKEDILIACLYVDDLIFTG 1007
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
++ + K+ + EFEM D+G LGI++K+ D G +F++Q GY K+ ++F+M
Sbjct: 1008 NNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQE-DNG-IFITQEGYAKEVLKKFKMD 1065
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
+S V TP+ KLS ++E + ++ T + S VGS+ Y + C+RPD+ YAV +V
Sbjct: 1066 DSNPVCTPMECGIKLS------KKEEGEGVDPTTFKSLVGSLRY-LTCTRPDILYAVGVV 1118
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+M +P H++A K +LRY+ G++ GL Y+ + + L GY D+D+ G+VD RKS
Sbjct: 1119 SRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTS--DYKLVGYSDSDWGGDVDDRKST 1176
Query: 805 VGFCVY 810
GF Y
Sbjct: 1177 SGFVFY 1182
>At1g48710 hypothetical protein
Length = 1352
Score = 166 bits (420), Expect = 6e-41
Identities = 92/246 (37%), Positives = 151/246 (60%), Gaps = 12/246 (4%)
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+++LYGLKQ+PR W R D++ + F++ Y+ +Y +K ++ IL LYVDD++
Sbjct: 981 KKALYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALY-IKIQKEDILIACLYVDDLIFTG 1039
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
++ + K+ + EFEM D+G LGI++K+ D G +F++Q GY K+ ++F+M
Sbjct: 1040 NNPSMFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQE-DNG-IFITQEGYAKEVLKKFKMD 1097
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
+S V TP+ KLS ++E + ++ T + S VGS+ Y + C+RPD+ YAV +V
Sbjct: 1098 DSNPVCTPMECGIKLS------KKEEGEGVDPTTFKSLVGSLRY-LTCTRPDILYAVGVV 1150
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
SR+M +P H++A K +LRY+ G++ GL Y+ + + L GY D+D+ G+VD RKS
Sbjct: 1151 SRYMEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTS--DYKLVGYSDSDWGGDVDDRKST 1208
Query: 805 VGFCVY 810
GF Y
Sbjct: 1209 SGFVFY 1214
>At3g61330 copia-type polyprotein
Length = 1352
Score = 163 bits (412), Expect = 5e-40
Identities = 91/243 (37%), Positives = 148/243 (60%), Gaps = 12/243 (4%)
Query: 568 LYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASSSK 627
LYGLKQ+PR W R D++ + F++ Y+ +Y +K ++ IL LYVDD++ ++
Sbjct: 984 LYGLKQAPRAWNTRIDKYFKEKDFIKCPYEHALY-IKIQKEDILIACLYVDDLIFTGNNP 1042
Query: 628 DEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSNSK 687
+ K+ + EFEM D+G LGI++K+ D G +F++Q GY K+ ++F++ +S
Sbjct: 1043 SIFEEFKKEMTKEFEMTDIGLMSYYLGIEVKQE-DNG-IFITQEGYAKEVLKKFKIDDSN 1100
Query: 688 TVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVSRF 747
V TP+ KLS ++E + ++ T + S VGS+ Y + C+RPD+ YAV +VSR+
Sbjct: 1101 PVCTPMECGIKLS------KKEEGEGVDPTTFKSLVGSLRY-LTCTRPDILYAVGVVSRY 1153
Query: 748 MANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLVGF 807
M +P H++A K +LRY+ G++ GL Y+ + + L GY D+D+ G+VD RKS GF
Sbjct: 1154 MEHPTTTHFKAAKRILRYIKGTVNFGLHYSTTS--DYKLVGYSDSDWGGDVDDRKSTSGF 1211
Query: 808 CVY 810
Y
Sbjct: 1212 VFY 1214
>At3g60170 putative protein
Length = 1339
Score = 159 bits (402), Expect = 7e-39
Identities = 93/242 (38%), Positives = 138/242 (56%), Gaps = 10/242 (4%)
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASS 625
++LYGLKQ+PR WY R + + LK F R + ++ K IL + LYVDD++ S
Sbjct: 971 KALYGLKQAPRAWYSRIEAYFLKEEFERCPSEHTLFT-KTRVGNILIVSLYVDDLIFTGS 1029
Query: 626 SKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSN 685
K + K+ + EFEM DLG K LGI++K++ G +F+ Q Y ++ RF M
Sbjct: 1030 DKAMCDEFKKSMMLEFEMSDLGKMKHFLGIEVKQS--DGGIFICQRRYAREVLARFGMDE 1087
Query: 686 SKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVS 745
S V P+ TKL+ + + DE T + VGS+MY + +RPDL Y V ++S
Sbjct: 1088 SNAVKNPIVPGTKLTKDENGEKVDE------TMFKQLVGSLMY-LTVTRPDLMYGVCLIS 1140
Query: 746 RFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLV 805
RFM+NP + HW A K +LRYL G+++ G+ Y R L + D+DYAG+++ R+S
Sbjct: 1141 RFMSNPRMSHWLAAKRILRYLKGTVELGIFYRRRKNRSLKLMAFTDSDYAGDLNDRRSTS 1200
Query: 806 GF 807
GF
Sbjct: 1201 GF 1202
>At4g04280 putative transposon protein
Length = 1104
Score = 155 bits (392), Expect = 1e-37
Identities = 97/254 (38%), Positives = 140/254 (54%), Gaps = 41/254 (16%)
Query: 554 FCGGQV*GVSFEESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYL 613
F G++ F E G ++PR+W +RF+ F++ F RS DSCVY+ + +
Sbjct: 792 FLHGELDQTLFMEQPEGF-EAPRQWNKRFNAFMMDQKFSRSVSDSCVYVKEVSN------ 844
Query: 614 LLYVDDILMASSSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGY 673
+ S EI KLK+ L+ EFEMKD+G A R LGIDI RNR +G L LSQ Y
Sbjct: 845 ----------AKSMTEIKKLKKVLSREFEMKDMGAASRKLGIDIIRNRSEGTLCLSQTSY 894
Query: 674 LKKGGERFRMSNSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCS 733
L++ ++FRM +K V+TP+G H KLS ++DE+ E PY+S VGS+M
Sbjct: 895 LERVIQKFRMDGAKVVNTPIGAHFKLS---SVHNDDERVGSEKVPYSSVVGSLM------ 945
Query: 734 RPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDAD 793
FM+ G VHW A+KW+LRYL S+ L YT+ + ++GY D+D
Sbjct: 946 -------------FMSKQGEVHWTAVKWLLRYLKWSIGLNLMYTKGF--DFKVQGYCDSD 990
Query: 794 YAGNVDTRKSLVGF 807
+A ++D S+ G+
Sbjct: 991 HAADLDKNMSISGY 1004
>At3g29510 hypothetical protein
Length = 1158
Score = 149 bits (375), Expect = 9e-36
Identities = 90/263 (34%), Positives = 146/263 (55%), Gaps = 15/263 (5%)
Query: 548 HGTTGRFCGGQV*GVSFEESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNE 607
+G+ ++C + SLYGLKQS R WY R E+L+K G+ C+++ K
Sbjct: 766 NGSRDQYC------IRLNRSLYGLKQSGRMWYNRLSEYLVKEGYKNDPISPCIFIKKFAS 819
Query: 608 KVILYLLLYVDDILMASSSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELF 667
K + + +YVDD+ + +S EI + E L EFEMKDLG K LG+ ++ DKG +
Sbjct: 820 KGFVIIAVYVDDLNILGTS-GEIAQTVEYLKKEFEMKDLGKTKFCLGLQLEYI-DKG-IL 876
Query: 668 LSQLGYLKKGGERFRMSNSKTVSTPLGHHTKLSIQQCP---QSEDEKQLMEGTPYASGVG 724
+ Q Y + +RF M + +++P+ + L + P + +DE+ L PY S +G
Sbjct: 877 VHQKAYTETVLKRFNMDKAHPLTSPMQVRS-LGLDSDPFGPKKDDEEILGPEMPYLSAIG 935
Query: 725 SIMYGMVCSRPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDED 784
++MY +RPD+ +AV+++SRF + P HW+ +K +LRYL G++ GL YT +++
Sbjct: 936 ALMYLSSHTRPDICFAVNLLSRFSSCPTKRHWEGIKHLLRYLQGTIDFGLYYTN--HNKE 993
Query: 785 ALEGYVDADYAGNVDTRKSLVGF 807
L G+ DA Y + KS G+
Sbjct: 994 GLVGFADAGYLSDPHNGKSQTGY 1016
>At4g10990 putative retrotransposon polyprotein
Length = 1203
Score = 147 bits (371), Expect = 3e-35
Identities = 89/245 (36%), Positives = 141/245 (57%), Gaps = 12/245 (4%)
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASS 625
+SLYGLKQ+ R+WY+R L F++S D+ +++ K + I+ +L+YVDD+++AS+
Sbjct: 673 KSLYGLKQASRQWYKRLSSVFLGANFIQSPADNTMFV-KVSCTSIIVVLVYVDDLMIASN 731
Query: 626 SKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSN 685
+ LKE L EF++KDLGPA+ LG++I R+ + + + Q Y + E +S
Sbjct: 732 DSSAVENLKELLRSEFKIKDLGPARFFLGLEIARSSE--GISVCQRKYAQNLLEDVGLSG 789
Query: 686 SKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIVS 745
K S P+ + L+ E L T Y VG ++Y + +RPD+ +AV +S
Sbjct: 790 CKPSSIPMDPNLHLT------KEMGTLLPNATSYRELVGRLLY-LCITRPDITFAVHTLS 842
Query: 746 RFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLV 805
+F++ P +H QA VLRYL G+ GL Y +A E L G+ DAD+ D+R+S+
Sbjct: 843 QFLSAPTDIHMQAAHKVLRYLKGNPGQGLMY--SASSELCLNGFSDADWGTCKDSRRSVT 900
Query: 806 GFCVY 810
GFC+Y
Sbjct: 901 GFCIY 905
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.356 0.161 0.595
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,769,020
Number of Sequences: 26719
Number of extensions: 774040
Number of successful extensions: 3081
Number of sequences better than 10.0: 91
Number of HSP's better than 10.0 without gapping: 86
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 2762
Number of HSP's gapped (non-prelim): 130
length of query: 937
length of database: 11,318,596
effective HSP length: 109
effective length of query: 828
effective length of database: 8,406,225
effective search space: 6960354300
effective search space used: 6960354300
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 65 (29.6 bits)
Medicago: description of AC130802.9


