

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.5 + phase: 0
(569 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
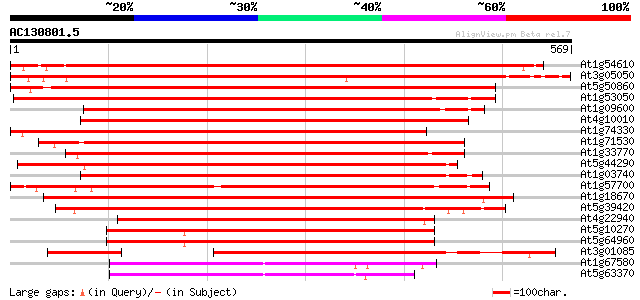
Score E
Sequences producing significant alignments: (bits) Value
At1g54610 Unknown protein (T22H22.5) 806 0.0
At3g05050 putative cyclin-dependent protein kinase 723 0.0
At5g50860 cyclin-dependent protein kinase-like 629 e-180
At1g53050 cell division-related protein, putative 597 e-171
At1g09600 putative protein kinase 556 e-158
At4g10010 protein kinase like protein 534 e-152
At1g74330 putative protein kinase 527 e-150
At1g71530 protein kinase-like 526 e-149
At1g33770 protein kinase, putative 523 e-148
At5g44290 cyclin-dependent protein kinase-like protein 522 e-148
At1g03740 putative protein kinase 518 e-147
At1g57700 516 e-146
At1g18670 hypothetical protein 516 e-146
At5g39420 cyclin-dependent kinase CDC2C (CDC2CAt) 502 e-142
At4g22940 putative cdc2 kinase homolog 400 e-111
At5g10270 cdc2-like protein kinase 358 4e-99
At5g64960 cdc2-like protein kinase 355 3e-98
At3g01085 putative protein 304 1e-82
At1g67580 putative protein kinase 245 5e-65
At5g63370 protein kinase 230 2e-60
>At1g54610 Unknown protein (T22H22.5)
Length = 572
Score = 806 bits (2082), Expect = 0.0
Identities = 399/562 (70%), Positives = 468/562 (82%), Gaps = 25/562 (4%)
Query: 1 MGCVIGRQASSN--------KGSGAQTNRIKVDEASAATTASNG------EEKNVVEIEN 46
MGCV GR+A++ K S A + + V E+S T SNG E+K E
Sbjct: 1 MGCVFGREAATTTTAEAKQAKSSKASSGVVVVGESSV--TKSNGVIADDVEKKKNEEANG 58
Query: 47 DQKKKSDDSVQRSRRSKPNPRLSNPPKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADT 106
D+++KS R R +KPNPRLSNP KH RGEQVAAGWPSWL+ CGEAL GW+PRKADT
Sbjct: 59 DKERKSSKG-DRRRSTKPNPRLSNPSKHWRGEQVAAGWPSWLSDACGEALNGWVPRKADT 117
Query: 107 FEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNV 166
FEKIDKIGQGTYSNVYKA D +TGK+VALKKVRFDNLEPES+KFMAREI++LRRLDHPNV
Sbjct: 118 FEKIDKIGQGTYSNVYKAKDMLTGKIVALKKVRFDNLEPESVKFMAREILVLRRLDHPNV 177
Query: 167 IKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRR 226
+KL+GLVTSRMSCSLYLVF YM+HDLAGLA+SPV++F+ES++KC M QL+SGLEHCH+R
Sbjct: 178 VKLEGLVTSRMSCSLYLVFQYMDHDLAGLASSPVVKFSESEVKCLMRQLISGLEHCHSRG 237
Query: 227 VLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYG 286
VLHRDIKGSNLLID+ G+LKIADFGLA+ FDPN+ PMTSRVVTLWYR PELLLGATDYG
Sbjct: 238 VLHRDIKGSNLLIDDGGVLKIADFGLATIFDPNHKRPMTSRVVTLWYRAPELLLGATDYG 297
Query: 287 VGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPR 346
VGIDLWSAGCIL ELL G+PIMPGRTEVEQLHKIYKLCGSPS++YWKK K + ++KPR
Sbjct: 298 VGIDLWSAGCILAELLAGRPIMPGRTEVEQLHKIYKLCGSPSEDYWKKGKFTHGAIYKPR 357
Query: 347 EPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKY 406
EPYKR IRETFK FPPS+LPLID LL+I+P +R+TAS AL+SEFFT+EPYAC+P+ LPKY
Sbjct: 358 EPYKRSIRETFKDFPPSSLPLIDALLSIEPEDRQTASAALKSEFFTSEPYACEPADLPKY 417
Query: 407 PPSKEMDAKRRDDEVRRQRAASKAQVDGSKKHRTRERSMKAMPAPEANAELQSNIDRRRL 466
PPSKE+DAKRRD+E RRQRAASKAQ DG++K+R R+RS +A+PAPEANAELQSN+DRRRL
Sbjct: 418 PPSKEIDAKRRDEETRRQRAASKAQGDGARKNRHRDRSNRALPAPEANAELQSNVDRRRL 477
Query: 467 ITHANAKSKSEKFPPPHQD-GQLGFPLGSSHHIDPDTIPADI--SFTSTTYTYSKE---- 519
ITHANAKSKSEKFPPPHQD G +G PLG+S HIDP IP D+ SFTS+++ +SK+
Sbjct: 478 ITHANAKSKSEKFPPPHQDGGAMGVPLGASQHIDPTFIPRDMVPSFTSSSFNFSKDEPPT 537
Query: 520 PFQAWSGPIGNAADIGVSKRKK 541
Q WSGP+G+ GVS++KK
Sbjct: 538 QVQTWSGPLGHPI-TGVSRKKK 558
>At3g05050 putative cyclin-dependent protein kinase
Length = 593
Score = 723 bits (1866), Expect = 0.0
Identities = 383/602 (63%), Positives = 439/602 (72%), Gaps = 44/602 (7%)
Query: 1 MGCVIGRQASSNKGSGA--------QTNRIKVDEASAATT-------------ASNGEEK 39
MGCV+GR SS SG+ ++NR +V+ S T ASNGEE
Sbjct: 1 MGCVLGRPGSSGSVSGSRDEVSTRIESNRHQVNNVSVTKTETTESTSAVVVASASNGEEV 60
Query: 40 NVVEIENDQKKKSDDSV----------QRSRRSKPNPRLSNPPKHLRGEQVAAGWPSWLT 89
E DQKK++ V +R R P+PR SNPPK+L GEQVAAGWPSWL+
Sbjct: 61 RNHEDVVDQKKENGFVVTEAKERKSKGERKRSKPPDPRRSNPPKNLLGEQVAAGWPSWLS 120
Query: 90 AVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIK 149
VCGEAL+GW+PRKAD+FEKIDKIG GTYSNVYKA DS+TG +VALKKVR D E ES+K
Sbjct: 121 EVCGEALSGWLPRKADSFEKIDKIGSGTYSNVYKAKDSLTGNIVALKKVRCDVNERESLK 180
Query: 150 FMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIK 209
FMAREI+ILRRLDHPNVIKL+GLVTSRMS SLYLVF YM+HDLAGLAASP I+FTE Q+K
Sbjct: 181 FMAREILILRRLDHPNVIKLEGLVTSRMSSSLYLVFRYMDHDLAGLAASPEIKFTEQQVK 240
Query: 210 CYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVV 269
CYM QLLSGLEHCHNR VLHRDIKGSNLLID+ G+L+I DFGLA+FFD + MT+RVV
Sbjct: 241 CYMKQLLSGLEHCHNRGVLHRDIKGSNLLIDDGGVLRIGDFGLATFFDASKRQEMTNRVV 300
Query: 270 TLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSD 329
TLWYR PELL G +Y VG+DLWSAGCIL ELL G+ IMPGR EVEQLH+IYKLCGSPS+
Sbjct: 301 TLWYRSPELLHGVVEYSVGVDLWSAGCILAELLAGRAIMPGRNEVEQLHRIYKLCGSPSE 360
Query: 330 EYWKKSKLPNA---TLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDAL 386
EYWKK +LP+ KP YKR IRE +K F P AL L+D LLA+DP ER+TA+D L
Sbjct: 361 EYWKKIRLPSTHKHAHHKPLPQYKRRIREVYKDFSPEALSLLDTLLALDPAERQTATDVL 420
Query: 387 RSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQVDGSKKHRTRERSMK 446
S+FFTTEP AC PS LPKYPPSKE+DAKRRD+E RRQR A KAQ + ++ R RER+ +
Sbjct: 421 MSDFFTTEPLACQPSDLPKYPPSKEIDAKRRDEEYRRQREARKAQGESGRRMRPRERAPR 480
Query: 447 AMPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQLGFPLGSSHHIDPDTIPAD 506
AMPAPEANAE QSNIDR R+ITHANAKSKSEKFPPPHQDG LGF +GSS +DP IP
Sbjct: 481 AMPAPEANAENQSNIDRMRMITHANAKSKSEKFPPPHQDGSLGFQVGSSRRLDPSEIP-- 538
Query: 507 ISFTSTTYTYSKEPFQAWSGPIGNAADIGVSKRKKYTAGEAWDLLKPHTGAPKDKFKGKA 566
S S T +YSKEPFQ WSGP+ A IG T D+ K A K KGK
Sbjct: 539 YSTNSFTSSYSKEPFQTWSGPL---APIGAPDS---TTRRRNDINKERRMA--SKVKGKR 590
Query: 567 II 568
I+
Sbjct: 591 IV 592
>At5g50860 cyclin-dependent protein kinase-like
Length = 580
Score = 629 bits (1621), Expect = e-180
Identities = 314/509 (61%), Positives = 389/509 (75%), Gaps = 24/509 (4%)
Query: 1 MGCVIGRQASSNKGSGAQT---------NRIKVDEASAATTASNGEEKNVVEIENDQKKK 51
MGCV+ ++++ +K N + +S+ TTA+ + +E +KKK
Sbjct: 1 MGCVLCKESTGDKRKHNNPDEPPPADLRNTEDLPSSSSTTTAA-------ISVEIGEKKK 53
Query: 52 SD-DSVQ--RSRRSKPNPRLSNPPKHLRGEQVAA--GWPSWLTAVCGEALTGWIPRKADT 106
D DS+Q RR+ S G + GWP WL A CG+++ PR+A T
Sbjct: 54 KDLDSIQIQPERRTWHTGDFSAGSSRRPGMSLRTPEGWPPWLIAACGDSIKDLTPRRATT 113
Query: 107 FEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNV 166
+EK++KIGQGTYSNVYKA D ++GK+VALKKVRFDNLE ES+KFMAREI++LRRL+HPNV
Sbjct: 114 YEKLEKIGQGTYSNVYKAKDLLSGKIVALKKVRFDNLEAESVKFMAREILVLRRLNHPNV 173
Query: 167 IKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRR 226
IKLQGLVTSR+SCSLYLVF+YMEHDL+GLAA+ ++F Q+KC+M QLLSGLEHCH+R
Sbjct: 174 IKLQGLVTSRVSCSLYLVFEYMEHDLSGLAATQGLKFDLPQVKCFMKQLLSGLEHCHSRG 233
Query: 227 VLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYG 286
VLHRDIKGSNLLIDN+GILKIADFGLA+F+DP MTSRVVTLWYRPPELLLGAT YG
Sbjct: 234 VLHRDIKGSNLLIDNDGILKIADFGLATFYDPKQKQTMTSRVVTLWYRPPELLLGATSYG 293
Query: 287 VGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPR 346
G+DLWSAGCI+ ELL GKP+MPGRTEVEQLHKI+KLCGSPSD YWKK +LPNATLFKP+
Sbjct: 294 TGVDLWSAGCIMAELLAGKPVMPGRTEVEQLHKIFKLCGSPSDSYWKKYRLPNATLFKPQ 353
Query: 347 EPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKY 406
PYKRC+ E F GF PS++ L++ LL IDP +R T++ AL SEFFTTEP CDPSSLPKY
Sbjct: 354 HPYKRCVAEAFNGFTPSSVHLVETLLTIDPADRGTSTSALNSEFFTTEPLPCDPSSLPKY 413
Query: 407 PPSKEMDAKRRDDEVRRQR--AASKAQVDGSKKHRTR-ERSMKAMPAPEANAELQSNIDR 463
PPSKE++ K RD+E+RRQ+ A + +DG+++ R R +R+ +A+PAPEANAE Q+N+DR
Sbjct: 414 PPSKELNVKLRDEELRRQKGLAGKGSGIDGARRIRYRGDRTGRAIPAPEANAESQANLDR 473
Query: 464 RRLITHANAKSKSEKFPPPHQDGQLGFPL 492
R I+ N KSKSEKFPPPHQDG +G+PL
Sbjct: 474 WRSISQTNGKSKSEKFPPPHQDGAVGYPL 502
>At1g53050 cell division-related protein, putative
Length = 694
Score = 597 bits (1538), Expect = e-171
Identities = 289/488 (59%), Positives = 367/488 (74%), Gaps = 5/488 (1%)
Query: 5 IGRQASSNKGSGAQTNRIKVDEASAATTASNGEEKNVVEIENDQKKKSDDSVQRSRRSKP 64
+ R +S++ + + D S SN + + + + + + ++ + P
Sbjct: 32 VSRPVASSRREEPLRIKERSDVVSVRPVLSNKQSNVSLHLRGENLSRREKRIENVAATSP 91
Query: 65 NPRLSNPPKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKA 124
K GE VAAGWP WL +V GEA+ GW+PR+AD+FEK+DKIGQGTYSNVY+A
Sbjct: 92 LAMSITIAKATEGEYVAAGWPPWLASVAGEAIRGWVPRRADSFEKLDKIGQGTYSNVYRA 151
Query: 125 IDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLV 184
D K+VALKKVRFDNLEPES++FMAREI ILRRLDHPN+IKL+GLVTSRMSCSLYLV
Sbjct: 152 RDLDQKKIVALKKVRFDNLEPESVRFMAREIQILRRLDHPNIIKLEGLVTSRMSCSLYLV 211
Query: 185 FDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGI 244
F+YMEHDLAGLA+ P I+F+ESQ+KCY+ QLL GL+HCH+R VLHRDIKGSNLLIDN G+
Sbjct: 212 FEYMEHDLAGLASHPAIKFSESQVKCYLQQLLHGLDHCHSRGVLHRDIKGSNLLIDNSGV 271
Query: 245 LKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVG 304
LKIADFGLASFFDP P+TSRVVTLWYRPPELLLGAT YG +DLWSAGCIL EL G
Sbjct: 272 LKIADFGLASFFDPRQTQPLTSRVVTLWYRPPELLLGATRYGAAVDLWSAGCILAELYAG 331
Query: 305 KPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSA 364
KPIMPGRTEVEQLHKI+KLCGSP+++YW KS+LP+AT+FKP +PYKR + ETFK FP A
Sbjct: 332 KPIMPGRTEVEQLHKIFKLCGSPTEDYWVKSRLPHATIFKPTQPYKRLVGETFKEFPQPA 391
Query: 365 LPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQ 424
L L++ LL+++P +R TA+ AL+SEFF+T P CDPSSLPKYPPSKE+DA+ RD+E RRQ
Sbjct: 392 LALLETLLSVNPDDRGTATAALKSEFFSTRPLPCDPSSLPKYPPSKELDARMRDEESRRQ 451
Query: 425 RAASKAQVDGSKKHRTRERSMKAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQ 484
++ D + R + +A+PAP+ANAEL +++ +R+ + + +S+SEKF P +
Sbjct: 452 VGGNR---DQRHQERRGTKESRAIPAPDANAELVASMQKRQ--SQSTNRSRSEKFNPHPE 506
Query: 485 DGQLGFPL 492
+ GFP+
Sbjct: 507 EVASGFPI 514
>At1g09600 putative protein kinase
Length = 714
Score = 556 bits (1433), Expect = e-158
Identities = 267/406 (65%), Positives = 328/406 (80%), Gaps = 6/406 (1%)
Query: 76 RGEQVAAGWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVAL 135
RG QV AGWPSWL +V GEA+ GWIPRKAD+FEK++KIGQGTYS+VYKA D T ++VAL
Sbjct: 132 RGAQVMAGWPSWLASVAGEAINGWIPRKADSFEKLEKIGQGTYSSVYKARDLETNQLVAL 191
Query: 136 KKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGL 195
KKVRF N++P+S++FMAREIIILRRLDHPNV+KL+GL+TSR+S S+YL+F+YMEHDLAGL
Sbjct: 192 KKVRFANMDPDSVRFMAREIIILRRLDHPNVMKLEGLITSRVSGSMYLIFEYMEHDLAGL 251
Query: 196 AASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASF 255
A++P I F+E+QIKCYM QLL GLEHCH+R VLHRDIKGSNLL+D+ LKI DFGLA+F
Sbjct: 252 ASTPGINFSEAQIKCYMKQLLHGLEHCHSRGVLHRDIKGSNLLLDHNNNLKIGDFGLANF 311
Query: 256 FDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVE 315
+ + P+TSRVVTLWYRPPELLLG+TDYGV +DLWS GCIL EL GKPIMPGRTEVE
Sbjct: 312 YQGHQKQPLTSRVVTLWYRPPELLLGSTDYGVTVDLWSTGCILAELFTGKPIMPGRTEVE 371
Query: 316 QLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAID 375
QLHKI+KLCGSPS+EYWK SKLP+AT+FKP++PYKRC+ ETFK P SAL L++ LLA++
Sbjct: 372 QLHKIFKLCGSPSEEYWKISKLPHATIFKPQQPYKRCVAETFKSLPSSALALVEVLLAVE 431
Query: 376 PVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQVDGS 435
P R T + AL SEFFTT P A DPSSLPKY P KE+D K +++E +R++ S Q D
Sbjct: 432 PDARGTTASALESEFFTTSPLASDPSSLPKYQPRKEIDVKAQEEEAKRKKDTSSKQNDSK 491
Query: 436 KKHRTRERSMKAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPP 481
+ R KA+PAP++NAE ++I +R+ N S S+KF P
Sbjct: 492 QV----SRESKAVPAPDSNAESLTSIQKRQ--GQHNQVSNSDKFNP 531
>At4g10010 protein kinase like protein
Length = 649
Score = 534 bits (1375), Expect = e-152
Identities = 255/396 (64%), Positives = 321/396 (80%), Gaps = 2/396 (0%)
Query: 72 PKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGK 131
P E +AAGWPSWLT+V GEA+ GW+PR+A++FEK+DKIGQGTYS+VY+A D TGK
Sbjct: 121 PHSPEAELIAAGWPSWLTSVAGEAIKGWVPRRAESFEKLDKIGQGTYSSVYRARDLETGK 180
Query: 132 VVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHD 191
+VA+KKVRF N++PES++FMAREI ILR+LDHPNV+KL+ LVTS++S SLYLVF+YMEHD
Sbjct: 181 MVAMKKVRFVNMDPESVRFMAREINILRKLDHPNVMKLECLVTSKLSGSLYLVFEYMEHD 240
Query: 192 LAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFG 251
L+GLA P ++FTESQIKCYM QLLSGLEHCH+R +LHRDIKG NLL++N+G+LKI DFG
Sbjct: 241 LSGLALRPGVKFTESQIKCYMKQLLSGLEHCHSRGILHRDIKGPNLLVNNDGVLKIGDFG 300
Query: 252 LASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGR 311
LA+ + P P+TSRVVTLWYR PELLLGAT+YG GIDLWS GCIL EL +GKPIMPGR
Sbjct: 301 LANIYHPEQDQPLTSRVVTLWYRAPELLLGATEYGPGIDLWSVGCILTELFLGKPIMPGR 360
Query: 312 TEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKL 371
TEVEQ+HKI+K CGSPSD+YW+K+KLP AT FKP++PYKR + ETFK PPSAL L+DKL
Sbjct: 361 TEVEQMHKIFKFCGSPSDDYWQKTKLPLATSFKPQQPYKRVLLETFKNLPPSALALVDKL 420
Query: 372 LAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAAS-KA 430
L+++P +R TAS L S+FFT EP C+ SSLPKYPPSKE+DAK RD+E RR+++ + K
Sbjct: 421 LSLEPAKRGTASSTLSSKFFTMEPLPCNVSSLPKYPPSKELDAKVRDEEARRKKSETVKG 480
Query: 431 QVDGSKKHRTRE-RSMKAMPAPEANAELQSNIDRRR 465
+ S + +R+ +S P A+ + + I +R
Sbjct: 481 RGPESVRRGSRDFKSTATTPEFVASGQSKDTITTKR 516
>At1g74330 putative protein kinase
Length = 445
Score = 527 bits (1358), Expect = e-150
Identities = 257/438 (58%), Positives = 329/438 (74%), Gaps = 16/438 (3%)
Query: 1 MGCVIGRQASS-------------NKGSGAQTNRIKVDEASAATTASNGEEKNVVEIEND 47
MGCV +Q S N+ + + RI V++ T ++
Sbjct: 1 MGCVSSKQTVSVTPAIDHSGVFKDNENECSGSGRIVVEDPPRPTLKKLVSWRSRSGKRRS 60
Query: 48 QKKKSDDSVQRSRRSKP-NPRLSNPPKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADT 106
QK S+ + R S + RL N ++L EQVAAGWP+WL+ V GEA+ GW+P ++D
Sbjct: 61 QKSGSELGSESGRASDSLSFRLGNVSRYLEAEQVAAGWPAWLSNVAGEAIHGWVPLRSDA 120
Query: 107 FEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNV 166
FEK++KIGQGTYSNV++A+++ TG++VALKKVRFDN EPES+KFMAREI+ILRRL+HPN+
Sbjct: 121 FEKLEKIGQGTYSNVFRAVETETGRIVALKKVRFDNFEPESVKFMAREILILRRLNHPNI 180
Query: 167 IKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRR 226
IKL+GL+TS++SC++ LVF+YMEHDL GL +SP I+FT QIKCYM QLLSGL+HCH+R
Sbjct: 181 IKLEGLITSKLSCNIQLVFEYMEHDLTGLLSSPDIKFTTPQIKCYMKQLLSGLDHCHSRG 240
Query: 227 VLHRDIKGSNLLIDNEGILKIADFGLASFFDP--NYMNPMTSRVVTLWYRPPELLLGATD 284
V+HRDIKGSNLL+ NEGILK+ADFGLA+F + + P+TSRVVTLWYRPPELLLGATD
Sbjct: 241 VMHRDIKGSNLLLSNEGILKVADFGLANFSNSSGHKKKPLTSRVVTLWYRPPELLLGATD 300
Query: 285 YGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFK 344
YG +DLWS GC+ ELL+GKPI+ GRTEVEQLHKI+KLCGSP ++YWKKSKLP+A LFK
Sbjct: 301 YGASVDLWSVGCVFAELLLGKPILRGRTEVEQLHKIFKLCGSPPEDYWKKSKLPHAMLFK 360
Query: 345 PREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLP 404
P++ Y C+RET K + + LI+ LL+IDP +R TAS AL S++FTT+P+ACDPSSLP
Sbjct: 361 PQQTYDSCLRETLKDLSETEINLIETLLSIDPHKRGTASSALVSQYFTTKPFACDPSSLP 420
Query: 405 KYPPSKEMDAKRRDDEVR 422
YPPSKE+D K RD+ R
Sbjct: 421 IYPPSKEIDTKHRDEAAR 438
>At1g71530 protein kinase-like
Length = 655
Score = 526 bits (1354), Expect = e-149
Identities = 263/436 (60%), Positives = 329/436 (75%), Gaps = 9/436 (2%)
Query: 30 ATTASNGEEKNVVEI----ENDQKKKSDDSVQRSRRSKPNPRLSNPPKHLRGEQVAAGWP 85
+T++ K VV + N ++ + V + + +P +SN + E AA WP
Sbjct: 71 STSSDRNSTKPVVVVGAPTRNPTRRVTAIPVAQPAQQQPARVISN-----KTELPAAEWP 125
Query: 86 SWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEP 145
SWL +V GEA+ GW+PR A++FEK+DKIGQGTYS+VYKA D TGK+VA+KKVRF N++P
Sbjct: 126 SWLASVAGEAIKGWVPRCAESFEKLDKIGQGTYSSVYKARDLETGKIVAMKKVRFVNMDP 185
Query: 146 ESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTE 205
ES++FMAREI+ILR+LDHPNV+KL+GLVTSR+S SLYLVF+YMEHDLAGLAA+P I+F+E
Sbjct: 186 ESVRFMAREILILRKLDHPNVMKLEGLVTSRLSGSLYLVFEYMEHDLAGLAATPGIKFSE 245
Query: 206 SQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMT 265
QIKCYM QL GLEHCH R +LHRDIKGSNLLI+NEG+LKI DFGLA+F+ + +T
Sbjct: 246 PQIKCYMQQLFRGLEHCHRRGILHRDIKGSNLLINNEGVLKIGDFGLANFYRGDGDLQLT 305
Query: 266 SRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCG 325
SRVVTLWYR PELLLGAT+YG IDLWSAGCIL EL GKPIMPGRTEVEQ+HKI+KLCG
Sbjct: 306 SRVVTLWYRAPELLLGATEYGPAIDLWSAGCILTELFAGKPIMPGRTEVEQMHKIFKLCG 365
Query: 326 SPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDA 385
SPS++YW+++ LP AT FKP PYK + ETF FP SAL LI+KLLAI+P +R +A+
Sbjct: 366 SPSEDYWRRATLPLATSFKPSHPYKPVLAETFNHFPSSALMLINKLLAIEPEKRGSAAST 425
Query: 386 LRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQVDGSKKHRTRERSM 445
LRSEFFTTEP +PS+LP+YPPSKE+DAK R++E R+ RA + G R R + +
Sbjct: 426 LRSEFFTTEPLPANPSNLPRYPPSKELDAKLRNEEARKLRAEGNKRRGGETVTRGRPKDL 485
Query: 446 KAMPAPEANAELQSNI 461
K PE A QS +
Sbjct: 486 KTAQTPEFMAAGQSKV 501
>At1g33770 protein kinase, putative
Length = 614
Score = 523 bits (1347), Expect = e-148
Identities = 256/411 (62%), Positives = 319/411 (77%), Gaps = 10/411 (2%)
Query: 57 QRSRRSKPNPR-----LSNPPKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADTFEKID 111
QR RR N + +SN P+ E +AAGWP WLT+V GEA+ GW+PR+AD+FEK+D
Sbjct: 86 QRGRRVSDNGKGGGLIISNVPRSAEAELIAAGWPYWLTSVAGEAIKGWVPRRADSFEKLD 145
Query: 112 KIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQG 171
KIGQGTYS VYKA D TGK+VA+KKVRF N++PES++FMAREI ILR+LDHPNV+KLQ
Sbjct: 146 KIGQGTYSIVYKARDLETGKIVAMKKVRFANMDPESVRFMAREINILRKLDHPNVMKLQC 205
Query: 172 LVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRD 231
LVTS++S SL+LVF+YMEHDL+GLA P ++FTE QIKC+M QLL GLEHCH+R +LHRD
Sbjct: 206 LVTSKLSGSLHLVFEYMEHDLSGLALRPGVKFTEPQIKCFMKQLLCGLEHCHSRGILHRD 265
Query: 232 IKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDL 291
IKGSNLL++N+G+LKI DFGLASF+ P+ P+TSRVVTLWYR PELLLG+T+YG IDL
Sbjct: 266 IKGSNLLVNNDGVLKIGDFGLASFYKPDQDQPLTSRVVTLWYRAPELLLGSTEYGPAIDL 325
Query: 292 WSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKR 351
WS GCIL EL V KPIMPGRTEVEQ+HKI+KLCGSPS+E+W +K P AT +KP+ PYKR
Sbjct: 326 WSVGCILAELFVCKPIMPGRTEVEQMHKIFKLCGSPSEEFWNTTKFPQATSYKPQHPYKR 385
Query: 352 CIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKE 411
+ ETFK S+L L+DKLL+++P +R +AS L SEFFTTEP C SSLPKYPPSKE
Sbjct: 386 VLLETFKNLSSSSLDLLDKLLSVEPEKRCSASSTLLSEFFTTEPLPCHISSLPKYPPSKE 445
Query: 412 MDAKRRDDEVRRQRAASKAQVDGSKKHRTRERSMK-AMPAPEANAELQSNI 461
+DAK RD+E +R+ KA+ + H + R ++ + PE A SN+
Sbjct: 446 LDAKVRDEEAKRK----KAEAVKWRGHESVRRGLRDSKVTPEFIASGNSNV 492
>At5g44290 cyclin-dependent protein kinase-like protein
Length = 644
Score = 522 bits (1345), Expect = e-148
Identities = 256/449 (57%), Positives = 329/449 (73%), Gaps = 5/449 (1%)
Query: 9 ASSNKGSGAQTNRIKVDEASAATTASNGEEKNVVEIENDQKKKSDDSVQRSRRSKPNPRL 68
+S K ++N++++ E+ +++ E+ + D + DD K P +
Sbjct: 36 SSQTKEEQDRSNKVRLIESEKFSSSRFSEKHQEIAEIGDTDEDEDDDHHPPEELKREPSV 95
Query: 69 SNPPKH---LRGEQVAAGWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKAI 125
PP + ++AAGWP+WL +V GEAL W PR+A TFEK++KIGQGTYS+VYKA
Sbjct: 96 VIPPSPETVSKEAELAAGWPAWLVSVAGEALVNWTPRRASTFEKLEKIGQGTYSSVYKAR 155
Query: 126 DSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVF 185
D K+VALK+VRFD + ES+KFMAREII++RRLDHPNV+KL+GL+T+ +S SLYLVF
Sbjct: 156 DLTNNKIVALKRVRFDLSDLESVKFMAREIIVMRRLDHPNVLKLEGLITASVSSSLYLVF 215
Query: 186 DYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGIL 245
+YM+HDL GLA+ P I+F+E Q+KCYM QLLSGL HCH+R VLHRDIKGSNLLID+ G+L
Sbjct: 216 EYMDHDLVGLASIPGIKFSEPQVKCYMQQLLSGLHHCHSRGVLHRDIKGSNLLIDSNGVL 275
Query: 246 KIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGK 305
KIADFGLA+FFDP P+TSRVVTLWYRPPELLLGA YGVG+DLWS GCILGEL GK
Sbjct: 276 KIADFGLATFFDPQNCVPLTSRVVTLWYRPPELLLGACHYGVGVDLWSTGCILGELYSGK 335
Query: 306 PIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSAL 365
PI+ G+TEVEQLHKI+KLCGSP+++YW+K KLP + F+P PY R + E FK P + L
Sbjct: 336 PILAGKTEVEQLHKIFKLCGSPTEDYWRKLKLPPSAAFRPALPYGRRVAEMFKDLPTNVL 395
Query: 366 PLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQR 425
L++ LL+IDP R +A+ AL SE+F TEP+ACDPSSLPKYPPSKE+DAK RDD R++
Sbjct: 396 SLLEALLSIDPDRRGSAARALESEYFRTEPFACDPSSLPKYPPSKEIDAKIRDDAKRQRP 455
Query: 426 AASKAQVDGSKKHRTRERSMKAMPAPEAN 454
K + S+ R+ ER K +P +AN
Sbjct: 456 TQEKHERQDSQTRRSHER--KLIPPVKAN 482
>At1g03740 putative protein kinase
Length = 740
Score = 518 bits (1333), Expect = e-147
Identities = 250/409 (61%), Positives = 320/409 (78%), Gaps = 7/409 (1%)
Query: 72 PKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGK 131
PK +QVAAGWPSWL +V GE+L W PR+A+TFEK++KIGQGTYS+VY+A D + K
Sbjct: 178 PKDAERKQVAAGWPSWLVSVAGESLVDWAPRRANTFEKLEKIGQGTYSSVYRARDLLHNK 237
Query: 132 VVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHD 191
+VALKKVRFD + ES+KFMAREII++RRLDHPNV+KL+GL+T+ +S SLYLVF+YM+HD
Sbjct: 238 IVALKKVRFDLNDMESVKFMAREIIVMRRLDHPNVLKLEGLITAPVSSSLYLVFEYMDHD 297
Query: 192 LAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFG 251
L GL++ P ++FTE Q+KCYM QLLSGLEHCH+R VLHRDIKGSNLLID++G+LKIADFG
Sbjct: 298 LLGLSSLPGVKFTEPQVKCYMRQLLSGLEHCHSRGVLHRDIKGSNLLIDSKGVLKIADFG 357
Query: 252 LASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGR 311
LA+FFDP +TS VVTLWYRPPELLLGA+ YGVG+DLWS GCILGEL GKPI+PG+
Sbjct: 358 LATFFDPAKSVSLTSHVVTLWYRPPELLLGASHYGVGVDLWSTGCILGELYAGKPILPGK 417
Query: 312 TEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKL 371
TEVEQLHKI+KLCGSP++ YW+K KLP++ FK PY+R + E FK FP S L L++ L
Sbjct: 418 TEVEQLHKIFKLCGSPTENYWRKQKLPSSAGFKTAIPYRRKVSEMFKDFPASVLSLLETL 477
Query: 372 LAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQ-RAASKA 430
L+IDP R +A AL SE+F T+P+ACDPS+LPKYPPSKE+DAK RD+ R+Q A K
Sbjct: 478 LSIDPDHRSSADRALESEYFKTKPFACDPSNLPKYPPSKEIDAKMRDEAKRQQPMRAEKQ 537
Query: 431 QVDGSKKHRTRERSMKAMPAPEANAELQSNIDRRRLITHANAKSKSEKF 479
+ S + ER K +P +AN L ++++ + + +S+++ F
Sbjct: 538 ERQDSMTRISHER--KFVPPVKANNSLSMTMEKQ----YKDLRSRNDSF 580
>At1g57700
Length = 686
Score = 516 bits (1328), Expect = e-146
Identities = 268/526 (50%), Positives = 356/526 (66%), Gaps = 53/526 (10%)
Query: 1 MGCVIGRQASSNKGSGAQTNRIKVD---------EASAATTASNGEEKNVVEIENDQKKK 51
MGC+ + +N +TN + + ++ + T+ NG E + I +D K
Sbjct: 1 MGCICSKGVRTNDDY-IETNHVSIGKENPKASKKQSDSEETSVNGNEATLRLIPDDVKDT 59
Query: 52 -SDDSVQRSRRSKPN---------------------------PRLSNPPKHLRGEQVA-- 81
SD+ V+ K + PR+S G++ A
Sbjct: 60 FSDEEVEELEEKKESSFEMKSCESVLQKGNVLEIVDNVGPLQPRMSRIGSVSNGDRAAKV 119
Query: 82 -AGWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRF 140
AGWPSWL +V GEA+ GWIPR AD+FEK++ IGQGTYS+VY+A D T ++VALKKVRF
Sbjct: 120 IAGWPSWLVSVAGEAINGWIPRSADSFEKLEMIGQGTYSSVYRARDLETNQIVALKKVRF 179
Query: 141 DNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPV 200
N++PES++FMAREIIILRRL+HPNV+KL+GL+ S+ S S+YL+F+YM+HDLAGLA++P
Sbjct: 180 ANMDPESVRFMAREIIILRRLNHPNVMKLEGLIISKASGSMYLIFEYMDHDLAGLASTPG 239
Query: 201 IRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNY 260
I+F+++Q QLL GLEHCH+ VLHRDIK SNLL+D LKI DFGL++F+
Sbjct: 240 IKFSQAQ------QLLLGLEHCHSCGVLHRDIKCSNLLLDRNNNLKIGDFGLSNFYRGQR 293
Query: 261 MNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKI 320
P+TSRVVTLWYRPPELLLG+TDYGV +DLWS GCIL EL GKP++PGRTEVEQ+HKI
Sbjct: 294 KQPLTSRVVTLWYRPPELLLGSTDYGVTVDLWSTGCILAELFTGKPLLPGRTEVEQMHKI 353
Query: 321 YKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERE 380
+KLCGSPS+EYW++S+L +AT+FKP+ PYKRC+ +TFK P SAL L++ LLA++P R
Sbjct: 354 FKLCGSPSEEYWRRSRLRHATIFKPQHPYKRCVADTFKDLPSSALALLEVLLAVEPDARG 413
Query: 381 TASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQVDGSKKHRT 440
TAS AL+SEFFTT+P+ +PSSLP+Y P KE DAK R++E RR++ +S Q + +
Sbjct: 414 TASSALQSEFFTTKPFPSEPSSLPRYQPRKEFDAKLREEEARRRKGSSSKQ----NEQKR 469
Query: 441 RERSMKAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDG 486
R KA+PAP ANAEL ++I +R + N S SEKF P G
Sbjct: 470 LARESKAVPAPSANAELLASIQKR--LGETNRTSISEKFNPEGDSG 513
>At1g18670 hypothetical protein
Length = 662
Score = 516 bits (1328), Expect = e-146
Identities = 260/488 (53%), Positives = 347/488 (70%), Gaps = 11/488 (2%)
Query: 35 NGEEKNVVEIEND--QKKKSDDSVQRSRRSKPNPRLSNPPKHLRGEQVAAGWPSWLTAVC 92
+ +K+ E+ +D + +S + R + RL N K+L EQVAAGWP+WL+ V
Sbjct: 57 SSSKKSGSELGSDFGELSESGRASSNCRSESVSFRLGNLSKYLEAEQVAAGWPAWLSNVA 116
Query: 93 GEALTGWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA 152
GEA+ GW+P ++D FEK++KIGQGTYS+V++A ++ TG++VALKKVRFDN EPES++FMA
Sbjct: 117 GEAIHGWVPFRSDAFEKLEKIGQGTYSSVFRARETETGRIVALKKVRFDNFEPESVRFMA 176
Query: 153 REIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYM 212
REI+ILR+L+HPN+IKL+G+VTS++SCS++LVF+YMEHDL GL +SP I FT QIKCYM
Sbjct: 177 REILILRKLNHPNIIKLEGIVTSKLSCSIHLVFEYMEHDLTGLLSSPDIDFTTPQIKCYM 236
Query: 213 NQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPN-YMNPMTSRVVTL 271
QLLSGL+HCH R V+HRDIKGSNLL++NEGILK+ADFGLA+F + + P+TSRVVTL
Sbjct: 237 KQLLSGLDHCHARGVMHRDIKGSNLLVNNEGILKVADFGLANFCNASGNKQPLTSRVVTL 296
Query: 272 WYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEY 331
WYRPPELLLGAT+YG +DLWS GC+ ELL+GKP++ GRTEVEQLHKI+KLCGSP ++Y
Sbjct: 297 WYRPPELLLGATEYGASVDLWSVGCVFAELLIGKPVLQGRTEVEQLHKIFKLCGSPPEDY 356
Query: 332 WKKSKLPNATLFKPREPYKRCIRET--FKGFPPSALPLIDKLLAIDPVERETASDALRSE 389
WKKSKLP+A LFKP++ Y C+RET KG + + LI+ LL+I P +R TAS AL S+
Sbjct: 357 WKKSKLPHAMLFKPQQHYDGCLRETLKLKGLSDADINLIETLLSIQPHKRGTASTALVSQ 416
Query: 390 FFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAAS-KAQVDGSKKHRTRERSMKAM 448
+FT++P+ACDPSSLP Y PSKE+DAK R+D R++ + + + + K R K
Sbjct: 417 YFTSKPFACDPSSLPVYSPSKEIDAKHREDTTRKKISGNGRRGTESRKPTRKPPAFAKLA 476
Query: 449 PAPEANAELQSNIDRRRLITHANAKSKSEKF-----PPPHQDGQLGFPLGSSHHIDPDTI 503
PA + Q R H + S S F P H+ + +S P +
Sbjct: 477 PAEDVRHHSQKFQKRNGHSVHNSIDSDSTLFEKMQKPSNHEKDEASHVKNASQGDVPFSG 536
Query: 504 PADISFTS 511
P +S +S
Sbjct: 537 PLQVSVSS 544
>At5g39420 cyclin-dependent kinase CDC2C (CDC2CAt)
Length = 644
Score = 502 bits (1293), Expect = e-142
Identities = 251/475 (52%), Positives = 336/475 (69%), Gaps = 21/475 (4%)
Query: 47 DQKKKSDDSVQRSRRSKP--------NPRLSNPPKHLRGEQVAAGWPSWLTAVCGEALTG 98
D ++ + +RSR+S+ L + +++ EQ AAGWP+WL + EA+ G
Sbjct: 37 DHSLEASHNSKRSRKSRRLGGSDLRIGVSLGSSHRNIEAEQAAAGWPAWLCSAAAEAVHG 96
Query: 99 WIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIIL 158
W+P KA+ F+K++KIGQGTYS+V++A + TGK+VALKKV+FDNL+PESI+FMAREI+IL
Sbjct: 97 WVPLKAEAFQKLEKIGQGTYSSVFRAREVETGKMVALKKVKFDNLQPESIRFMAREILIL 156
Query: 159 RRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSG 218
R+L+HPN++KL+G+VTSR S S+YLVF+YMEHDLAGL+++P IRFTE QIKCYM QLL G
Sbjct: 157 RKLNHPNIMKLEGIVTSRASSSIYLVFEYMEHDLAGLSSNPDIRFTEPQIKCYMKQLLWG 216
Query: 219 LEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPEL 278
LEHCH R V+HRDIK SN+L++N+G+LK+ DFGLA+ P+ N +TSRVVTLWYR PEL
Sbjct: 217 LEHCHMRGVIHRDIKASNILVNNKGVLKLGDFGLANVVTPSNKNQLTSRVVTLWYRAPEL 276
Query: 279 LLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLP 338
L+G+T YGV +DLWS GC+ E+L+GKPI+ GRTE+EQLHKIYKLCGSP D +WK++KLP
Sbjct: 277 LMGSTSYGVSVDLWSVGCVFAEILMGKPILKGRTEIEQLHKIYKLCGSPQDSFWKRTKLP 336
Query: 339 NATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYAC 398
+AT FKP+ Y+ +RE K + + L++ LL+++P +R TAS AL SE+F T PYAC
Sbjct: 337 HATSFKPQHTYEATLRERCKDLSATGVYLLETLLSMEPDKRGTASSALNSEYFLTRPYAC 396
Query: 399 DPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQVDG-SKKHRTRER------SMKAMPAP 451
DPSSLPKYPP+KEMDAK RDD +RR+RA K + G +KH+ R + +P
Sbjct: 397 DPSSLPKYPPNKEMDAKYRDD-MRRKRANLKLRDSGVGRKHKRPHRAEYDPKNYAKLPIR 455
Query: 452 EANAELQ---SNIDRRRLITHANAKSKSEKFPPPHQDGQLGFPLGSSHHIDPDTI 503
+ E++ + R TH N S+ P GF DPD I
Sbjct: 456 KDTLEVKNIPNEASRATTTTHGNYYKVSDL--PMTTGPASGFAWAVKRRKDPDNI 508
>At4g22940 putative cdc2 kinase homolog
Length = 353
Score = 400 bits (1027), Expect = e-111
Identities = 194/329 (58%), Positives = 247/329 (74%), Gaps = 7/329 (2%)
Query: 110 IDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKL 169
+ +IG GT+S V+KA D + K VALK++RFD ESIK +AREIIILR+LDHPNVIKL
Sbjct: 1 LQQIGGGTFSKVFKARDLLRNKTVALKRIRFDINNSESIKCIAREIIILRKLDHPNVIKL 60
Query: 170 QGLV-TSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVL 228
+GL+ S +LYL+F+YMEHDL GL++ + F+E Q+KCYM QLL GL+HCH VL
Sbjct: 61 EGLMLVDHDSSTLYLIFEYMEHDLLGLSSLLGVHFSEPQVKCYMRQLLRGLDHCHTNHVL 120
Query: 229 HRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYGVG 288
HRD+K SNLLI+ +G+LKIADFGLA+FFDP+ P+T+ V TLWYRPPELLLGA+ YG+G
Sbjct: 121 HRDMKSSNLLINGDGVLKIADFGLATFFDPHNSVPLTTHVATLWYRPPELLLGASHYGIG 180
Query: 289 IDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREP 348
+DLWS GC++GEL GKPI+PG+ E +QLHKI+KLCGSPSD+YW K KL +T +P P
Sbjct: 181 VDLWSTGCVIGELYAGKPILPGKNETDQLHKIFKLCGSPSDDYWTKLKLQLSTPLRPIYP 240
Query: 349 YKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPP 408
Y I ETFK FP S + L++ LL+IDP R TA+ AL+S++F TEP ACDPS LPKYP
Sbjct: 241 YGSHIAETFKQFPASVISLLETLLSIDPDFRGTAASALKSKYFKTEPLACDPSCLPKYPS 300
Query: 409 SKEMDAKRRDD------EVRRQRAASKAQ 431
SKE++ K RD+ ++RR A Q
Sbjct: 301 SKEINIKMRDNTRKQASQIRRTDEAQAVQ 329
>At5g10270 cdc2-like protein kinase
Length = 505
Score = 358 bits (919), Expect = 4e-99
Identities = 183/349 (52%), Positives = 237/349 (67%), Gaps = 17/349 (4%)
Query: 99 WIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIII 157
W R D FEK+++IG+GTY VY A + TG++VALKK+R DN E E A REI I
Sbjct: 18 WGSRSVDCFEKLEQIGEGTYGQVYMAKEIKTGEIVALKKIRMDN-EREGFPITAIREIKI 76
Query: 158 LRRLDHPNVIKLQGLVTS--------------RMSCSLYLVFDYMEHDLAGLAASPVIRF 203
L++L H NVI+L+ +VTS + +Y+VF+YM+HDL GLA P +RF
Sbjct: 77 LKKLHHENVIQLKEIVTSPGRDRDDQGKPDNNKYKGGIYMVFEYMDHDLTGLADRPGLRF 136
Query: 204 TESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNP 263
T QIKCYM QLL+GL +CH +VLHRDIKGSNLLIDNEG LK+ADFGLA + ++
Sbjct: 137 TVPQIKCYMKQLLTGLHYCHVNQVLHRDIKGSNLLIDNEGNLKLADFGLARSYSHDHTGN 196
Query: 264 MTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKL 323
+T+RV+TLWYRPPELLLGAT YG ID+WS GCI ELL KPI+PG+ E EQL+KI++L
Sbjct: 197 LTNRVITLWYRPPELLLGATKYGPAIDMWSVGCIFAELLHAKPILPGKNEQEQLNKIFEL 256
Query: 324 CGSPSDEYWK-KSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETA 382
CGSP ++ W SK+P FKP P KR +RE F+ F AL L++K+L +DP +R +A
Sbjct: 257 CGSPDEKLWPGVSKMPWFNNFKPARPLKRRVREFFRHFDRHALELLEKMLVLDPAQRISA 316
Query: 383 SDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQ 431
DAL +E+F T+P CDP SLP Y S E K++ + R+ A+K Q
Sbjct: 317 KDALDAEYFWTDPLPCDPKSLPTYESSHEFQTKKKRQQQRQNEEAAKRQ 365
>At5g64960 cdc2-like protein kinase
Length = 513
Score = 355 bits (912), Expect = 3e-98
Identities = 183/349 (52%), Positives = 234/349 (66%), Gaps = 17/349 (4%)
Query: 99 WIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIII 157
W R D FEK+++IG+GTY VY A + TG++VALKK+R DN E E A REI I
Sbjct: 18 WGSRSVDCFEKLEQIGEGTYGQVYMAKEIKTGEIVALKKIRMDN-EREGFPITAIREIKI 76
Query: 158 LRRLDHPNVIKLQGLVTS--------------RMSCSLYLVFDYMEHDLAGLAASPVIRF 203
L++L H NVI L+ +VTS + +Y+VF+YM+HDL GLA P +RF
Sbjct: 77 LKKLHHENVIHLKEIVTSPGRDRDDQGKPDNNKYKGGIYMVFEYMDHDLTGLADRPGLRF 136
Query: 204 TESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNP 263
T QIKCYM QLL+GL +CH +VLHRDIKGSNLLIDNEG LK+ADFGLA + ++
Sbjct: 137 TVPQIKCYMKQLLTGLHYCHVNQVLHRDIKGSNLLIDNEGNLKLADFGLARSYSHDHTGN 196
Query: 264 MTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKL 323
+T+RV+TLWYRPPELLLGAT YG ID+WS GCI ELL GKPI+PG+TE EQL+KIY+L
Sbjct: 197 LTNRVITLWYRPPELLLGATKYGPAIDMWSVGCIFAELLNGKPILPGKTENEQLNKIYEL 256
Query: 324 CGSPSDEYWK-KSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETA 382
CGSP + W SK+P K P KR +RE ++ F AL L++K+L +DP +R A
Sbjct: 257 CGSPDESNWPGVSKMPWYNQMKSSRPLKRRVREIYRHFDRHALELLEKMLVLDPSQRICA 316
Query: 383 SDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQ 431
DAL +E+F T+P CDP SLP Y S E K++ ++R A+K Q
Sbjct: 317 KDALDAEYFWTDPLPCDPKSLPTYESSHEFQTKKKRQQMRHNEEAAKKQ 365
>At3g01085 putative protein
Length = 536
Score = 304 bits (778), Expect = 1e-82
Identities = 156/351 (44%), Positives = 218/351 (61%), Gaps = 36/351 (10%)
Query: 207 QIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTS 266
+IKCYM QLL G+EHCH R ++HRDIK +N+L++N+G+LK+ADFGLA+ P N +TS
Sbjct: 120 KIKCYMKQLLWGVEHCHLRGIMHRDIKAANILVNNKGVLKLADFGLANIVTPRNKNQLTS 179
Query: 267 RVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGS 326
RVVTLWYR PELL+G+T Y V +DLWS GC+ E+L G+P++ GRTE+EQLHKIYKL GS
Sbjct: 180 RVVTLWYRAPELLMGSTSYSVSVDLWSVGCVFAEILTGRPLLKGRTEIEQLHKIYKLSGS 239
Query: 327 PSDEYWKKSKL-PNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDA 385
P +E+W+K+KL P +F+P+ Y+ C+RE F FP +A+ L++ LL+IDP +R TAS A
Sbjct: 240 PDEEFWEKNKLHPQTKMFRPQHQYEGCLRERFDEFPKTAINLLENLLSIDPEKRGTASSA 299
Query: 386 LRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQVDGSKKHRTRERSM 445
L SE+F T+PYACDPS+LPKYPP+KEMDAK R++ RR+R + K + + + K + R
Sbjct: 300 LMSEYFNTQPYACDPSTLPKYPPNKEMDAKYREELQRRRRVSIKKRDNLATKKLGKSR-- 357
Query: 446 KAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQLGFPLGSSHHIDPDTIPA 505
A ++ + RL TH K ++E + +
Sbjct: 358 --------RATVKEPTNLNRLPTHQETKKEAE----------------------TEIVVQ 387
Query: 506 DISFTSTTYTYSKEPFQAWS---GPIGNAADIGVSKRKKYTAGEAWDLLKP 553
S TS T S+ P+ + S P A G KRK+ ++P
Sbjct: 388 TPSETSQATTRSEFPYNSLSQTTAPASGFAWAGTKKRKENDVASTLTYIQP 438
Score = 66.2 bits (160), Expect = 5e-11
Identities = 30/75 (40%), Positives = 46/75 (61%)
Query: 39 KNVVEIENDQKKKSDDSVQRSRRSKPNPRLSNPPKHLRGEQVAAGWPSWLTAVCGEALTG 98
+ VV + + +DD +SR + + R P ++ EQVAAGWPSWL++ EA+ G
Sbjct: 47 RRVVNSSPKKHRNNDDDAPKSRTTGVSLRSGLPHSNVEAEQVAAGWPSWLSSAAPEAVHG 106
Query: 99 WIPRKADTFEKIDKI 113
W+P +A+ FEK +KI
Sbjct: 107 WVPLRAEDFEKREKI 121
>At1g67580 putative protein kinase
Length = 752
Score = 245 bits (625), Expect = 5e-65
Identities = 136/345 (39%), Positives = 199/345 (57%), Gaps = 14/345 (4%)
Query: 102 RKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRL 161
R D FE+++KI +GTY VY+A D TG++VALKKV+ + REI IL
Sbjct: 401 RSVDEFERLNKIDEGTYGVVYRAKDKKTGEIVALKKVKMEKEREGFPLTSLREINILLSF 460
Query: 162 DHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEH 221
HP+++ ++ +V S+++V +YMEHDL L + RF++S++KC M QLL G+++
Sbjct: 461 HHPSIVDVKEVVVGSSLDSIFMVMEYMEHDLKALMETMKQRFSQSEVKCLMLQLLEGVKY 520
Query: 222 CHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLG 281
H+ VLHRD+K SNLL++N G LKI DFGLA + + + P T VVTLWYR PELLLG
Sbjct: 521 LHDNWVLHRDLKTSNLLLNNRGELKICDFGLARQYG-SPLKPYTHLVVTLWYRAPELLLG 579
Query: 282 ATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKK-SKLPNA 340
A Y ID+WS GCI+ ELL+ P+ G+TE +QL KI+++ G+P++ W SKLP
Sbjct: 580 AKQYSTAIDMWSLGCIMAELLMKAPLFNGKTEFDQLDKIFRILGTPNESIWPGFSKLPGV 639
Query: 341 TLFKPREPY----KRCIRETFKGFP---PSALPLIDKLLAIDPVERETASDALRSEFFTT 393
+ + Y K+ +F G P + L++KLL DP R T ++AL+ ++F
Sbjct: 640 KVNFVKHQYNLLRKKFPATSFTGAPVLSDAGFDLLNKLLTYDPERRITVNEALKHDWFRE 699
Query: 394 EPYACDPSSLPKYPPSKEMDAKRR-----DDEVRRQRAASKAQVD 433
P +P +P D + R D + QR Q +
Sbjct: 700 VPLPKSKDFMPTFPAQHAQDRRGRRMVKSPDPLEEQRRKELTQTE 744
>At5g63370 protein kinase
Length = 612
Score = 230 bits (586), Expect = 2e-60
Identities = 127/323 (39%), Positives = 189/323 (58%), Gaps = 16/323 (4%)
Query: 102 RKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRF--DNLEPESIKFMA--REIII 157
R + F+K++KI +GTY VYKA D T ++VALKK++ D E E + REI I
Sbjct: 292 RSVNEFQKLNKINEGTYGIVYKARDEKTKEIVALKKIKMKEDRFEEEYGFPLTSLREINI 351
Query: 158 LRRLDHPNVIKLQGLVTS-RMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLL 216
L +HP ++ ++ +V + +Y+V +++EHDL G+ F+ S++KC M QLL
Sbjct: 352 LLSCNHPAIVNVKEVVVGGKNDNDVYMVMEHLEHDLRGVMDRRKEPFSTSEVKCLMMQLL 411
Query: 217 SGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPP 276
GL++ H ++HRD+K SNLL++N G LKI DFG+A + + + P T V+T WYRPP
Sbjct: 412 DGLKYLHTNWIIHRDLKPSNLLMNNCGELKICDFGMARQYG-SPIKPYTQMVITQWYRPP 470
Query: 277 ELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKK-S 335
ELLLGA +Y +D+WS GCI+ ELL KP+ PG++E++QL KI+ + G+P++ W S
Sbjct: 471 ELLLGAKEYSTAVDMWSVGCIMAELLSQKPLFPGKSELDQLQKIFAVLGTPNEAIWPGFS 530
Query: 336 KLPNATLFKPREPYKRCIRETFKG--------FPPSALPLIDKLLAIDPVERETASDALR 387
PNA P +PY +R+ F L++ LL +DP +R T DAL
Sbjct: 531 SFPNAKAKFPTQPY-NMLRKKFPAISFVGGQILSERGFDLLNSLLTLDPEKRLTVEDALN 589
Query: 388 SEFFTTEPYACDPSSLPKYPPSK 410
+F P +P YPP +
Sbjct: 590 HGWFHEVPLPKSKDFMPTYPPKR 612
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,715,452
Number of Sequences: 26719
Number of extensions: 623705
Number of successful extensions: 4395
Number of sequences better than 10.0: 999
Number of HSP's better than 10.0 without gapping: 864
Number of HSP's successfully gapped in prelim test: 136
Number of HSP's that attempted gapping in prelim test: 2142
Number of HSP's gapped (non-prelim): 1202
length of query: 569
length of database: 11,318,596
effective HSP length: 105
effective length of query: 464
effective length of database: 8,513,101
effective search space: 3950078864
effective search space used: 3950078864
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC130801.5


