

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.12 + phase: 0
(124 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
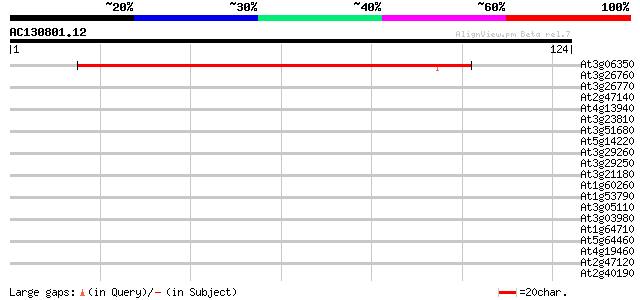
Score E
Sequences producing significant alignments: (bits) Value
At3g06350 putative dehydroquinase shikimate dehydrogenase 82 7e-17
At3g26760 putative short chain alcohol dehydrogenase 30 0.40
At3g26770 short chain alcohol dehydrogenase like protein 28 0.88
At2g47140 alcohol dehydrogenase like protein 28 0.88
At4g13940 adenosylhomocysteinase 27 2.0
At3g23810 S-adenosyl-L-homocysteinase like protein 27 2.0
At3g51680 short-chain alcohol dehydrogenase-like protein 27 2.6
At5g14220 protoporphyrinogen IX oxidase 27 3.4
At3g29260 short-chain alcohol dehydrogenase, putative 27 3.4
At3g29250 short-chain alcohol dehydrogenase, putative 27 3.4
At3g21180 putative Ca2+-transporting ATPase 27 3.4
At1g60260 beta-glucosidase like protein 26 4.4
At1g53790 26 5.7
At3g05110 hypothetical protein 25 7.5
At3g03980 putative short-chain type dehydrogenase/reductase 25 7.5
At1g64710 alcohol dehydrogenase (EC 1.1.1.1) like protein 25 7.5
At5g64460 putative phosphoglycerate mutase 25 9.8
At4g19460 unknown protein 25 9.8
At2g47120 putative alcohol dehydrogenase 25 9.8
At2g40190 unknown protein 25 9.8
>At3g06350 putative dehydroquinase shikimate dehydrogenase
Length = 603
Score = 82.0 bits (201), Expect = 7e-17
Identities = 41/92 (44%), Positives = 62/92 (66%), Gaps = 5/92 (5%)
Query: 16 VRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAILA 75
+ AGGAGKALA+GAK +GA++VI + ++++ LA A+ G+ S +L N+ P G +LA
Sbjct: 459 IGAGGAGKALAYGAKEKGAKVVIANRTYERALELAEAIGGKALSLTDLDNYHPEDGMVLA 518
Query: 76 NATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
N T +GM P+ + P+++ Y LVFDAVY
Sbjct: 519 NTTSMGMQPNVEETPISKDALKHYALVFDAVY 550
>At3g26760 putative short chain alcohol dehydrogenase
Length = 269
Score = 29.6 bits (65), Expect = 0.40
Identities = 17/44 (38%), Positives = 27/44 (60%), Gaps = 3/44 (6%)
Query: 10 LDGRLFVRAGGA---GKALAFGAKTRGARIVIFDIDFDKSRSLA 50
L+G++ V GGA GKA A ++GA+++I DID + +A
Sbjct: 5 LEGKVAVITGGASGIGKATAEEFVSQGAQVIIVDIDEEAGHMVA 48
>At3g26770 short chain alcohol dehydrogenase like protein
Length = 306
Score = 28.5 bits (62), Expect = 0.88
Identities = 18/52 (34%), Positives = 27/52 (51%), Gaps = 3/52 (5%)
Query: 10 LDGRLFVRAGGA---GKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQ 58
L+G++ + GGA GKA A GAR+VI D+D + A + E +
Sbjct: 41 LEGKVALITGGASGLGKATASEFLRHGARVVIADLDAETGTKTAKELGSEAE 92
>At2g47140 alcohol dehydrogenase like protein
Length = 257
Score = 28.5 bits (62), Expect = 0.88
Identities = 16/54 (29%), Positives = 29/54 (53%), Gaps = 3/54 (5%)
Query: 10 LDGRLFVRAGGAGKALAFGAKT---RGARIVIFDIDFDKSRSLACAVFGEVQSY 60
LDG++ + GGA A + GAR+VI D+ + +++A ++ + SY
Sbjct: 6 LDGKIVIITGGASGIGAESVRLFTEHGARVVIVDVQDELGQNVAVSIGEDKASY 59
>At4g13940 adenosylhomocysteinase
Length = 485
Score = 27.3 bits (59), Expect = 2.0
Identities = 11/24 (45%), Positives = 15/24 (61%)
Query: 19 GGAGKALAFGAKTRGARIVIFDID 42
G GK A KT GAR+++ +ID
Sbjct: 271 GDVGKGCAAAMKTAGARVIVTEID 294
>At3g23810 S-adenosyl-L-homocysteinase like protein
Length = 485
Score = 27.3 bits (59), Expect = 2.0
Identities = 11/24 (45%), Positives = 15/24 (61%)
Query: 19 GGAGKALAFGAKTRGARIVIFDID 42
G GK A KT GAR+++ +ID
Sbjct: 271 GDVGKGCAAAMKTAGARVIVTEID 294
>At3g51680 short-chain alcohol dehydrogenase-like protein
Length = 303
Score = 26.9 bits (58), Expect = 2.6
Identities = 17/44 (38%), Positives = 23/44 (51%), Gaps = 3/44 (6%)
Query: 10 LDGRLFVRAGGA---GKALAFGAKTRGARIVIFDIDFDKSRSLA 50
L+G++ + GGA GKA GA +VI D+D SLA
Sbjct: 32 LEGKVAIITGGAHGIGKATVMLFARHGATVVIADVDNVAGSSLA 75
>At5g14220 protoporphyrinogen IX oxidase
Length = 508
Score = 26.6 bits (57), Expect = 3.4
Identities = 13/39 (33%), Positives = 22/39 (56%), Gaps = 2/39 (5%)
Query: 6 QVHPLDGR--LFVRAGGAGKALAFGAKTRGARIVIFDID 42
Q+ + G+ V AG +G A A+ K+RG + +F+ D
Sbjct: 10 QIEAVSGKRVAVVGAGVSGLAAAYKLKSRGLNVTVFEAD 48
>At3g29260 short-chain alcohol dehydrogenase, putative
Length = 259
Score = 26.6 bits (57), Expect = 3.4
Identities = 14/47 (29%), Positives = 27/47 (56%), Gaps = 3/47 (6%)
Query: 10 LDGRLFVRAGGAGKALAFGAKT---RGARIVIFDIDFDKSRSLACAV 53
LDG++ + GGA A A+ GA++VI D+ + +++A ++
Sbjct: 6 LDGKIVIITGGASGIGAEAARLFTDHGAKVVIVDLQEELGQNVAVSI 52
>At3g29250 short-chain alcohol dehydrogenase, putative
Length = 260
Score = 26.6 bits (57), Expect = 3.4
Identities = 15/47 (31%), Positives = 26/47 (54%), Gaps = 3/47 (6%)
Query: 10 LDGRLFVRAGGAGKALAFGAKT---RGARIVIFDIDFDKSRSLACAV 53
LDG++ + GGA A + GA++VI DI + ++LA ++
Sbjct: 6 LDGKIAIITGGASGIGAEAVRLFTDHGAKVVIVDIQEELGQNLAVSI 52
>At3g21180 putative Ca2+-transporting ATPase
Length = 1090
Score = 26.6 bits (57), Expect = 3.4
Identities = 21/58 (36%), Positives = 28/58 (48%), Gaps = 7/58 (12%)
Query: 14 LFVRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPVKG 71
LF AG + +AFG+ T A FDID +K S+ Q+ NL + VKG
Sbjct: 105 LFKLAGE--QQIAFGSSTPAASTGNFDIDLEKLVSMT-----RNQNMSNLQQYGGVKG 155
>At1g60260 beta-glucosidase like protein
Length = 449
Score = 26.2 bits (56), Expect = 4.4
Identities = 19/61 (31%), Positives = 29/61 (47%), Gaps = 3/61 (4%)
Query: 30 KTRGARIVIFDIDFDKSRSLACAVFGEVQSY-KNLVNFQPVKGAILANATPIGMHPDTDR 88
+T G+R+ +F + + +A V Y K + PV IL N TP+ H DT R
Sbjct: 299 RTIGSRLPVFSEEESEQYPVAPWTMEAVLEYIKQSYDNPPVY--ILENGTPMTQHKDTHR 356
Query: 89 I 89
+
Sbjct: 357 V 357
>At1g53790
Length = 381
Score = 25.8 bits (55), Expect = 5.7
Identities = 17/64 (26%), Positives = 30/64 (46%), Gaps = 7/64 (10%)
Query: 10 LDGRLFVRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLVNFQPV 69
+DG L+ AG F + R + +V FD+ +K + VF + L+N++
Sbjct: 219 IDGILYYTAG-------FDMRARVSMVVCFDVRSEKFSFINIHVFMLMNDSCTLINYKGK 271
Query: 70 KGAI 73
GA+
Sbjct: 272 LGAL 275
>At3g05110 hypothetical protein
Length = 372
Score = 25.4 bits (54), Expect = 7.5
Identities = 14/50 (28%), Positives = 22/50 (44%)
Query: 41 IDFDKSRSLACAVFGEVQSYKNLVNFQPVKGAILANATPIGMHPDTDRIP 90
+ FD S + A G+VQ + ++ G I TP M + R+P
Sbjct: 296 VSFDDSLGVGVAYLGKVQGFASVYKQAVQHGVISFMITPEEMQRFSHRVP 345
>At3g03980 putative short-chain type dehydrogenase/reductase
Length = 270
Score = 25.4 bits (54), Expect = 7.5
Identities = 15/33 (45%), Positives = 19/33 (57%), Gaps = 3/33 (9%)
Query: 9 PLDGRLFVRAG---GAGKALAFGAKTRGARIVI 38
PL GR+ + G G G+A+A GARIVI
Sbjct: 13 PLAGRVAIVTGSSRGIGRAIAIHLAELGARIVI 45
>At1g64710 alcohol dehydrogenase (EC 1.1.1.1) like protein
Length = 380
Score = 25.4 bits (54), Expect = 7.5
Identities = 14/28 (50%), Positives = 19/28 (67%), Gaps = 1/28 (3%)
Query: 19 GGAGKALAFGAKTRGA-RIVIFDIDFDK 45
G G A+A GA+ RGA +I+ DI+ DK
Sbjct: 207 GAVGLAVAEGARARGASKIIGIDINPDK 234
>At5g64460 putative phosphoglycerate mutase
Length = 282
Score = 25.0 bits (53), Expect = 9.8
Identities = 9/22 (40%), Positives = 15/22 (67%)
Query: 80 IGMHPDTDRIPVAEYQLVFDAV 101
+G+HP R +++YQ +F AV
Sbjct: 137 LGVHPCDQRRSISDYQFLFPAV 158
>At4g19460 unknown protein
Length = 796
Score = 25.0 bits (53), Expect = 9.8
Identities = 7/27 (25%), Positives = 17/27 (62%)
Query: 78 TPIGMHPDTDRIPVAEYQLVFDAVYLH 104
+P+ P+T++IP Q+++ ++ H
Sbjct: 125 SPLDQSPETNKIPPVSDQIIYPIIHSH 151
>At2g47120 putative alcohol dehydrogenase
Length = 258
Score = 25.0 bits (53), Expect = 9.8
Identities = 14/54 (25%), Positives = 29/54 (52%), Gaps = 3/54 (5%)
Query: 10 LDGRLFVRAGGAGKALAFGAKT---RGARIVIFDIDFDKSRSLACAVFGEVQSY 60
L+G++ + GGA A A+ GA++VI D+ + +++A + + S+
Sbjct: 6 LEGKIVIITGGASGIGADAARLFTDHGAKVVIVDVQEELGQNVAVLIGKDKASF 59
>At2g40190 unknown protein
Length = 180
Score = 25.0 bits (53), Expect = 9.8
Identities = 20/53 (37%), Positives = 28/53 (52%), Gaps = 6/53 (11%)
Query: 45 KSRSLACAVFGEVQSYKNLVNFQPVKGAILANATPIGMHPDTDR---IPVAEY 94
K R++ V G+VQ YKN + + V+ +L NA G+H D I V EY
Sbjct: 42 KDRAVELKVDGDVQFYKNAMYRELVE--LLGNAV-AGLHGMIDEHFGISVVEY 91
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.143 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,701,462
Number of Sequences: 26719
Number of extensions: 96724
Number of successful extensions: 253
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 242
Number of HSP's gapped (non-prelim): 22
length of query: 124
length of database: 11,318,596
effective HSP length: 87
effective length of query: 37
effective length of database: 8,994,043
effective search space: 332779591
effective search space used: 332779591
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 53 (25.0 bits)
Medicago: description of AC130801.12


