

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
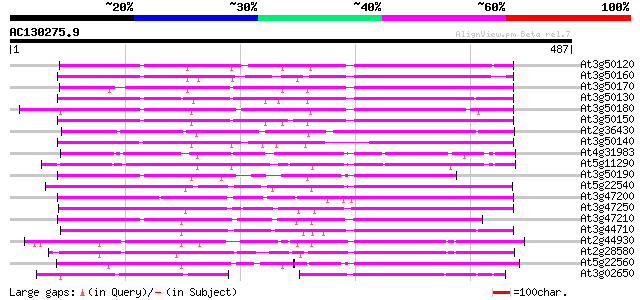
Query= AC130275.9 + phase: 0
(487 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g50120 putative protein 216 3e-56
At3g50160 putative protein 207 1e-53
At3g50170 putative protein 206 2e-53
At3g50130 putative protein 205 4e-53
At3g50180 putative protein 201 8e-52
At3g50150 putative protein 201 8e-52
At2g36430 unknown protein (At2g36430) 187 9e-48
At3g50140 putative protein 179 4e-45
At4g31983 putative protein 178 7e-45
At5g11290 putative protein 177 9e-45
At3g50190 putative protein 176 2e-44
At5g22540 unknown protein 165 6e-41
At3g47200 unknown protein 162 5e-40
At3g47250 unknown protein 155 5e-38
At3g47210 unknown protein 150 1e-36
At3g44710 unknown protein 137 1e-32
At2g44930 unknown protein 112 4e-25
At2g28580 hypothetical protein 111 1e-24
At5g22560 putative protein 94 2e-19
At3g02650 hypothetical protein 82 7e-16
>At3g50120 putative protein
Length = 531
Score = 216 bits (549), Expect = 3e-56
Identities = 143/432 (33%), Positives = 228/432 (52%), Gaps = 53/432 (12%)
Query: 44 VCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGSKL 103
+CIYRVP + + K+Y P +S+GPYH+G + L+ M+ K + +R+
Sbjct: 103 LCIYRVPYYLQENDNKSYFPQTVSLGPYHHGKKRLRSMDRHKWRAVNRVLKRTNQGIKMY 162
Query: 104 EEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLSDN-----EFKHV 158
+A + LE++ AR CY G + LSS+EF++M+++DG F+++L R + ++
Sbjct: 163 IDAMRELEEK---ARACYEGPLSLSSNEFIEMLVLDGCFVLELFRGAVEGFTELGYARND 219
Query: 159 PSLS-RWMLPTIRREMIMLENQLPMFVLTKLFEL---TDKNNSLHPQMSFYNLSFKFFYR 214
P + R + +I+R+M+MLENQLP+FVL +L EL T L Q L+ +FF
Sbjct: 220 PVFAMRGSMHSIQRDMVMLENQLPLFVLNRLLELQLGTRNQTGLVAQ-----LAIRFFDP 274
Query: 215 LLQSESRKTPECQTSYKFKIE--------------HVLDLLRYNIRPKLIGEEPRGCQS- 259
L+ ++ T Q+ + + H LD+ R ++ EPR +
Sbjct: 275 LMPTDEPLTKSGQSKLENSLARDKSFDPFADMGELHCLDVFRRSLLRSSPKPEPRLTRKR 334
Query: 260 -------------QMIHSITELKEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYI 306
Q+IH +TELKE G+K + ++ D+ F + L IP L I
Sbjct: 335 WSRNTRVADKRRQQLIHCVTELKEAGIKFRRRKTDRFWDMQFKNGY------LEIPRLLI 388
Query: 307 GDHRGTVFRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEH 366
D ++F N++AFE+CH + D+T+Y+ F++ LI+S DVS LHY G+I H LGSD
Sbjct: 389 HDGTKSLFLNLIAFEQCHIDSSNDITSYIIFMDNLIDSHEDVSYLHYCGIIEHWLGSDSE 448
Query: 367 VAELINNIAKEIVPDMNESYLYNVVNEANEYLGCWRARFRASLVHNYLTS-WAVGLSTLG 425
VA+L N + +E+V D +SYL + E N Y +RA+L H Y + WA+ +S
Sbjct: 449 VADLFNRLCQEVVFDTEDSYLSRLSIEVNRYYDHKWNAWRATLKHKYFNNPWAI-VSFCA 507
Query: 426 ALLALYFTFIQA 437
A++ L TF Q+
Sbjct: 508 AVILLVLTFSQS 519
>At3g50160 putative protein
Length = 511
Score = 207 bits (526), Expect = 1e-53
Identities = 141/417 (33%), Positives = 219/417 (51%), Gaps = 61/417 (14%)
Query: 42 ESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGS 101
+++CIYRVPP + + K+Y+P +SIGPYH+G +HL ME K + + + +
Sbjct: 102 DNLCIYRVPPYLQENDTKSYMPQIVSIGPYHHGHKHLMPMERHKWRAVNMVMARAKHDIE 161
Query: 102 KLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLSDN--EFKHVP 159
+A K LE++ AR CY G I ++ +EF++M+++DG FII++ + S+ E + P
Sbjct: 162 MYIDAMKELEEK---ARACYQGPINMNRNEFIEMLVLDGVFIIEIFKGTSEGFQEIGYAP 218
Query: 160 SLS----RWMLPTIRREMIMLENQLPMFVLTKLFELT-----DKNNSLHPQMSFYNLSFK 210
+ R ++ +IRR+M+MLENQLP VL L +L DK N L
Sbjct: 219 NDPVFGMRGLMQSIRRDMVMLENQLPWSVLKGLLQLQRPDVLDKVN--------VQLFQP 270
Query: 211 FFYRLLQSESRKTPECQTSYKFKIEHVLDLLRYNIRPK---------LIGEEPRGCQSQM 261
FF LL + T E H LD+LR + ++ ++P+ Q+
Sbjct: 271 FFQPLLPTREVLTEEGGL-------HCLDVLRRGLLQSSGTSDEDMSMVNKQPQ----QL 319
Query: 262 IHSITELKEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRNIVAFE 321
IH +TEL+ GV+ E+ DI F + L IP L I D ++F N++AFE
Sbjct: 320 IHCVTELRNAGVEFMRKETGHFWDIEFKNGY------LKIPKLLIHDGTKSLFLNLIAFE 373
Query: 322 KCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPD 381
+CH + + +T+Y+ F++ LINS+ DVS LH+ G+I + LGSD V++L N + KE++ D
Sbjct: 374 QCHIKSSKKITSYIIFMDNLINSSEDVSYLHHYGIIENWLGSDSEVSDLFNGLGKEVIFD 433
Query: 382 MNESYLYNVVNEANEYLGCWRARFRASLVHNYLTS-WAVGLSTLGALLALYFTFIQA 437
N+ YL + E N Y +A+L H Y + WA YF+FI A
Sbjct: 434 PNDGYLSALTGEVNIYYRRKWNYLKATLRHKYFNNPWA------------YFSFIAA 478
>At3g50170 putative protein
Length = 541
Score = 206 bits (524), Expect = 2e-53
Identities = 138/430 (32%), Positives = 218/430 (50%), Gaps = 51/430 (11%)
Query: 44 VCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLK----NKFFHRLFDPNGAN 99
+CIYRVP + + K+Y P +S+GPYH+G + L+ ME K NK RL
Sbjct: 113 LCIYRVPHYLQENDKKSYFPQTVSLGPYHHGKKRLRPMERHKWRALNKVLKRL------- 165
Query: 100 GSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLSDN-----E 154
++E + + E AR CY G I LS +EF +M+++DG F+++L R +
Sbjct: 166 KQRIEMYTNAMRELEEKARACYEGPISLSRNEFTEMLVLDGCFVLELFRGTVEGFTEIGY 225
Query: 155 FKHVPSLS-RWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFFY 213
++ P + R ++ +I+R+MIMLENQLP+FVL +L EL + + + +++ KFF
Sbjct: 226 ARNDPVFAMRGLMHSIQRDMIMLENQLPLFVLDRLLEL--QLGTQNQTGIVAHVAVKFFD 283
Query: 214 RLLQSESRKTPECQTSYKFKIE------------HVLDLLRYNIRPKLIGEEPRGC---- 257
L+ + T Q+ +E H LD+ R ++ R
Sbjct: 284 PLMPTGEALTKPDQSKLMNWLEKSLDTLGDKGELHCLDVFRRSLLQSSPTPNTRSLLKRL 343
Query: 258 ----------QSQMIHSITELKEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYIG 307
Q Q++H +TEL+E GVK + ++ DI F + L IP L I
Sbjct: 344 TRNTRVVDKRQQQLVHCVTELREAGVKFRKRKTDRFWDIEFKNGY------LEIPKLLIH 397
Query: 308 DHRGTVFRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHV 367
D ++F N++AFE+CH + +T+Y+ F++ LINS+ DVS LHY G+I H LGSD V
Sbjct: 398 DGTKSLFSNLIAFEQCHIESSNHITSYIIFMDNLINSSEDVSYLHYCGIIEHWLGSDSEV 457
Query: 368 AELINNIAKEIVPDMNESYLYNVVNEANEYLGCWRARFRASLVHNYLTSWAVGLSTLGAL 427
A+L N + +E+V D +S+L + + N Y +A+L H Y + S A+
Sbjct: 458 ADLFNRLCQEVVFDPKDSHLSRLSGDVNRYYNRKWNVLKATLTHKYFNNPWAYFSFSAAV 517
Query: 428 LALYFTFIQA 437
+ L T Q+
Sbjct: 518 ILLLLTLCQS 527
>At3g50130 putative protein
Length = 564
Score = 205 bits (522), Expect = 4e-53
Identities = 142/429 (33%), Positives = 221/429 (51%), Gaps = 46/429 (10%)
Query: 42 ESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGS 101
+ +CIYRVP + K+Y P +S+GP+H+G++HL M+ K + + + +
Sbjct: 137 DKLCIYRVPQYLQENNKKSYFPQTVSLGPFHHGNKHLLPMDRHKWRAVNMVMARTKHDIE 196
Query: 102 KLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLSDNEFKHV--- 158
+A K LE AR CY G I LSS++F +M+++DG F+++L R +D F +
Sbjct: 197 MYIDAMKELEDR---ARACYEGPIDLSSNKFSEMLVLDGCFVLELFRG-ADEGFSELGYD 252
Query: 159 ---PSLS-RWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFFYR 214
P + R + +I+R+M+MLENQLP+FVL +L E+ + H L+ +FF
Sbjct: 253 RNDPVFAMRGSMHSIQRDMVMLENQLPLFVLNRLLEI--QLGKRHQTGLVSRLAVRFFDP 310
Query: 215 LLQSES--RKTPECQTSYKF--------KIE-HVLDLLRYNIRPKLIGEEPRGC------ 257
L+ ++ KT + KF K E H LD+ R N+ EPR
Sbjct: 311 LMPTDEPLTKTDDSLEQDKFFNPIADKDKGELHCLDVFRRNLLRPCSNPEPRLSRMRWSW 370
Query: 258 --------QSQMIHSITELKEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYIGDH 309
Q Q+IH +TEL+E G+K + ++ DI F + L IP L I D
Sbjct: 371 RTRVADKRQQQLIHCVTELREAGIKFRTRKTDRFWDIRFKNGY------LEIPKLLIHDG 424
Query: 310 RGTVFRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAE 369
++F N++AFE+CH + D+T+Y+ F++ LI+S+ DV LHY G+I H LG+D VA+
Sbjct: 425 TKSLFSNLIAFEQCHIDSSNDITSYIIFMDNLIDSSEDVRYLHYCGIIEHWLGNDYEVAD 484
Query: 370 LINNIAKEIVPDMNESYLYNVVNEAN-EYLGCWRARFRASLVHNYLTSWAVGLSTLGALL 428
L N + +E+ D SYL + N+ + Y W +A L H Y + S AL+
Sbjct: 485 LFNRLCQEVAFDPQNSYLSQLSNKVDRNYSRKWNV-LKAILKHKYFNNPWAYFSFFAALV 543
Query: 429 ALYFTFIQA 437
L T Q+
Sbjct: 544 LLVLTLFQS 552
>At3g50180 putative protein
Length = 588
Score = 201 bits (511), Expect = 8e-52
Identities = 139/451 (30%), Positives = 235/451 (51%), Gaps = 37/451 (8%)
Query: 9 IKRKFHGTAVKSSKFWRGLVEKELQSIKPEEKNE--SVCIYRVPPNMLNVEPKAYIPSNI 66
I G+ K ++ + +K Q+ + +++ +CIY+VP + + K+Y P +
Sbjct: 141 INEVVEGSGTKRLEWVISIKDKLEQAYRNDDRTSWGKLCIYKVPHYLHGNDKKSYFPQTV 200
Query: 67 SIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGSKLEEACKFLEKEEINARNCYMGEIK 126
S+GPYH+G Q Q ME K + + + +A LE++ AR CY G I
Sbjct: 201 SLGPYHHGRQQTQSMECHKWRAVNMVLKRTNQGIEVFLDAMIELEEK---ARACYEGSIV 257
Query: 127 LSSDEFLKMMLVDGSFIIQLLRDLSDNEFK-----HVPSLS-RWMLPTIRREMIMLENQL 180
LSS+EF +M+L+DG FI++LL+ +++ K + P + R + +I+R+MIMLENQL
Sbjct: 258 LSSNEFTEMLLLDGCFILELLQGVNEGFLKLGYDHNDPVFAVRGSMHSIQRDMIMLENQL 317
Query: 181 PMFVLTKLFELTDKNNSLHPQMSFYNLSFKFFYRLLQSESRKTPECQTSYKFKIE-HVLD 239
P+FVL +L EL + Q L +FF L+ + T E H LD
Sbjct: 318 PLFVLNRLLELQPGTQN---QTGLVELVVRFFIPLMPTAETLTENSPPRGVSNGELHCLD 374
Query: 240 LLRYNIR-PKLIGEEPRGCQS-----QMIHSITELKEGGVKIKACESRELMDISFGKKWG 293
+ ++ P+ G+ + ++I ++TEL++ G K K ++ DI F +
Sbjct: 375 VFHRSLLFPRSSGKANYSRVADKHLQRVIPTVTELRDAGFKFKLNKTDRFWDIKFSNGY- 433
Query: 294 IMVKELTIPPLYIGDHRGTVFRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHY 353
L IP L I D ++F N++AFE+CH + D+T+Y+ F++ LI+S D+S LH+
Sbjct: 434 -----LEIPGLLIHDGTKSLFLNLIAFEQCHIESSNDITSYIIFMDNLIDSPEDISYLHH 488
Query: 354 KGVIHHSLGSDEHVAELINNIAKEIVPDMNESYLYNVVNEANEYLGCWRARF-------R 406
G+I HSLGS+ VA++ N + +E+V D + YL ++ E + C++ + +
Sbjct: 489 CGIIEHSLGSNSEVADMFNQLCQEVVFDTKDIYLSQLLIEVHR---CYKQNYSRKLNSLK 545
Query: 407 ASLVHNYLTSWAVGLSTLGALLALYFTFIQA 437
+L+ YL + LS A++ L TF Q+
Sbjct: 546 TTLILKYLDNPWAYLSFFAAVILLILTFSQS 576
>At3g50150 putative protein
Length = 509
Score = 201 bits (511), Expect = 8e-52
Identities = 138/424 (32%), Positives = 219/424 (51%), Gaps = 42/424 (9%)
Query: 42 ESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGS 101
+ +CIYRVP + + K+Y+P +SIGPYH+G HL+ ME K + + + N
Sbjct: 88 DKLCIYRVPFYLQENDKKSYLPQTVSIGPYHHGKVHLRPMERHKWRAVNMIMARTKHNIE 147
Query: 102 KLEEACKFLEKEEINARNCYMGEIKL-SSDEFLKMMLVDGSFIIQLLRDLSDNEFK---- 156
+A K LE+E AR CY G I + +S+EF +M+++DG F+++L + K
Sbjct: 148 MYIDAMKELEEE---ARACYQGPIDMKNSNEFTEMLVLDGCFVLELFKGTIQGFQKIGYA 204
Query: 157 -HVPSLS-RWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFFYR 214
+ P + R ++ +I+R+MIMLENQLP+FVL +L L + + + ++ +FF
Sbjct: 205 RNDPVFAKRGLMHSIQRDMIMLENQLPLFVLDRLLGL--QTGTPNQTGIVAEVAVRFFKT 262
Query: 215 LLQSES--RKTPECQTSYKFKIE-------HVLDLLRYNIRPKLIGEEPRGC-------- 257
L+ + K+ S + E H LD+ R + E
Sbjct: 263 LMPTSEVLTKSERSLDSQEKSDELGDNGGLHCLDVFH---RSLIQSSETTNQGTPYEDMS 319
Query: 258 ----QSQMIHSITELKEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYIGDHRGTV 313
Q Q+IH +TEL+ GV E+ +L DI F + L IP L I D ++
Sbjct: 320 MVEKQQQLIHCVTELRGAGVNFMRKETGQLWDIEFKNGY------LKIPKLLIHDGTKSL 373
Query: 314 FRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINN 373
F N++AFE+CH + + ++T+Y+ F++ LINS+ DVS LH+ G+I H LGSD VA+L N
Sbjct: 374 FSNLIAFEQCHTQSSNNITSYIIFMDNLINSSQDVSYLHHDGIIEHWLGSDSEVADLFNR 433
Query: 374 IAKEIVPDMNESYLYNVVNEANEYLGCWRARFRASLVHNYLTSWAVGLSTLGALLALYFT 433
+ KE++ D + YL + E N Y +A+L Y + S A++ L+ T
Sbjct: 434 LCKEVIFDPKDGYLSQLSREVNRYYSRKWNSLKATLRQKYFNNPWAYFSFSAAVILLFLT 493
Query: 434 FIQA 437
F Q+
Sbjct: 494 FFQS 497
>At2g36430 unknown protein (At2g36430)
Length = 448
Score = 187 bits (476), Expect = 9e-48
Identities = 126/406 (31%), Positives = 204/406 (50%), Gaps = 26/406 (6%)
Query: 46 IYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGSKLEE 105
I+RVP +M++ + Y P +SIGPYH G L+ +E K ++ + L LE+
Sbjct: 47 IFRVPQSMIDCNGRCYEPRVVSIGPYHRGQTQLKMIEEHKWRYLNVLL--TRTQNLTLED 104
Query: 106 ACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLSDNEFKHVPS----L 161
K ++ E AR CY I + S+EF +MM++DG F+++L R ++ N P+
Sbjct: 105 YMKSVKNVEEVARECYSETIHMDSEEFNEMMVLDGCFLLELFRKVN-NLVPFEPNDPLVA 163
Query: 162 SRWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFFYRLLQSESR 221
W+LP R+ + LENQ+P FVL LF LT +N S +L+F FF ++
Sbjct: 164 MAWVLPFFYRDFLCLENQIPFFVLETLFNLTRGDNENETNASLQSLAFAFFNNMMH---- 219
Query: 222 KTPECQTSYK-FKIEHVLDLLRYNIRPKLIGEEPRGCQ-------SQMIHSITELKEGGV 273
+T E +K + +H+LDLLR + P+ P S +IHSI++L+ G+
Sbjct: 220 RTEEDLARFKELRAKHLLDLLRSSFIPESELHTPPATNPGKEKMPSHIIHSISKLRRAGI 279
Query: 274 KIKACESRELMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRNIVAFEKCHKRCNPDMTT 333
K+ REL D + +P + + D + N VA+E+CH C+ TT
Sbjct: 280 KL-----RELKDAESFLVVRFRHGTIEMPAITVDDFMSSFLENCVAYEQCHVACSMHFTT 334
Query: 334 YMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMNESYLYNVVNE 393
Y L+ L N+ DV L + +I + G+D +A+ +N++ +++ D+ + YL ++ E
Sbjct: 335 YATLLDCLTNTYKDVEYLCDQNIIENYFGTDTELAKFVNSLGRDVAFDITQCYLKDLFEE 394
Query: 394 ANEYL-GCWRARFRASLVHNYLTSWAVGLSTLGALLALYFTFIQAI 438
NEY W + A+ Y S +S L AL+ L + IQ I
Sbjct: 395 VNEYYKSSWHVEW-ATFKFTYFNSPWSFVSALAALVLLVLSVIQTI 439
>At3g50140 putative protein
Length = 508
Score = 179 bits (453), Expect = 4e-45
Identities = 127/431 (29%), Positives = 207/431 (47%), Gaps = 80/431 (18%)
Query: 42 ESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGS 101
+ +CIYRVP ++ + +Y P +S+GPYH+G +HL+ M+ K + + +
Sbjct: 111 DKICIYRVPLSLKKSDKNSYFPQAVSLGPYHHGDEHLRPMDYHKWRAVNMVMKRTKQGIE 170
Query: 102 KLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLSDNEFK----- 156
+A K LE+ AR CY G I LSS++F +M+++DG F++ L R + K
Sbjct: 171 MYIDAMKELEE---RARACYEGPIGLSSNKFTQMLVLDGCFVLDLFRGAYEGFSKLGYDR 227
Query: 157 HVPSLS-RWMLPTIRREMIMLENQLPMFVLTKLFEL---TDKNNSLHPQMSFYNLSFKFF 212
+ P + R + +IRR+M+MLENQLP+FVL +L EL T L Q L+ +FF
Sbjct: 228 NDPVFAMRGSMHSIRRDMLMLENQLPLFVLNRLLELQLGTQYQTGLVAQ-----LAVRFF 282
Query: 213 YRLLQS--ESRKTPECQTSY---------KFKIE-HVLDLLRYNIRPKLIGEEPR----- 255
L+ + S K Q + K K E H LD+ R ++ + +PR
Sbjct: 283 NPLMPTYMSSTKIENSQENNNKFFNPIADKEKEELHCLDVFRRSLLQPSLKPDPRLSRSR 342
Query: 256 ---------GCQSQMIHSITELKEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYI 306
Q Q++H +TEL+E G K
Sbjct: 343 WSRKPLVADKRQQQLLHCVTELREAGTK-------------------------------- 370
Query: 307 GDHRGTVFRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEH 366
++F N++A+E+CH D+T+Y+ F++ LI+SA D+ LHY +I H LG+D
Sbjct: 371 -----SLFSNLIAYEQCHIDSTNDITSYIIFMDNLIDSAEDIRYLHYYDIIEHWLGNDSE 425
Query: 367 VAELINNIAKEIVPDMNESYLYNVVNEANEYLGCWRARFRASLVHNYLTSWAVGLSTLGA 426
VA++ N + +E+ D+ +YL + N+ + Y +A+L H Y ++ S A
Sbjct: 426 VADVFNRLCQEVAFDLENTYLSELSNKVDRYYNRKWNVLKATLKHKYFSNPWAYFSFFAA 485
Query: 427 LLALYFTFIQA 437
++ L T Q+
Sbjct: 486 VILLLLTLFQS 496
>At4g31983 putative protein
Length = 414
Score = 178 bits (451), Expect = 7e-45
Identities = 138/413 (33%), Positives = 207/413 (49%), Gaps = 49/413 (11%)
Query: 45 CIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGSKLE 104
CIY+VP + + P AY P +S GP H G + LQ ME K ++ F P S LE
Sbjct: 28 CIYKVPNKLRRLNPDAYTPRLVSFGPLHRGKEELQAMEDQKYRYLLS-FIPR--TNSSLE 84
Query: 105 EACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLSDNEFKHVPSL--- 161
+ + E NAR+CY ++KL SDEF++M++VDGSF+++LL H P L
Sbjct: 85 DLVRLARTWEQNARSCYAEDVKLHSDEFVEMLVVDGSFLVELLLR------SHYPRLRGE 138
Query: 162 ------SRWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFF-YR 214
+ M+ + R+MI++ENQLP FV+ ++F L N S L+ + F Y
Sbjct: 139 NDRIFGNSMMITDVCRDMILIENQLPFFVVKEIFLLL-LNYYQQGTPSIIQLAQRHFSYF 197
Query: 215 LLQSESRKTPECQTSYKFKIEHVLDLLRYNIRPKL-IGEEPRGCQSQMIHSITELKEGGV 273
L + + K + + EH +DLLR P+ I E + TEL GV
Sbjct: 198 LSRIDDEK-------FITEPEHFVDLLRSCYLPQFPIKLEYTTVKVDNAPEATELHTAGV 250
Query: 274 KIKACESRE-LMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRNIVAFEKCHKRC-NPDM 331
+ K E+ L+DISF G+ L IP + + D ++++NI+ FE+C RC N +
Sbjct: 251 RFKPAETSSCLLDISFAD--GV----LKIPTIVVDDLTESLYKNIIGFEQC--RCSNKNF 302
Query: 332 TTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMNESYLYNVV 391
Y+ L I S D L + G+I + LG+ V+ L N+I+KE++ D + ++++
Sbjct: 303 LDYIMLLGCFIKSPTDADLLIHSGIIVNYLGNSVDVSNLFNSISKEVIYD--RRFYFSML 360
Query: 392 NE-----ANEYLGCWRARFRASLVHNYLTSWAVGLSTLGALLALYFTFIQAIC 439
+E N W+A R HN WAV S ALL L TFIQ++C
Sbjct: 361 SENLQAYCNTPWNRWKAILRRDYFHN---PWAVA-SVFAALLLLLLTFIQSVC 409
>At5g11290 putative protein
Length = 422
Score = 177 bits (450), Expect = 9e-45
Identities = 136/422 (32%), Positives = 212/422 (50%), Gaps = 33/422 (7%)
Query: 28 VEKELQSIKPEEKNESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNK 87
+++ L+S+ N CIY+ P + + P AY P +SIGP HYG + LQ ME K +
Sbjct: 19 IKENLESLSSLSTN--CCIYKAPERLRRLNPDAYSPRVVSIGPLHYGKEELQAMEDHKLR 76
Query: 88 FFHRLFDPNGANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLL 147
+ + F P A LE+ + E AR CY +++LSSDE++KM++VD SF+++LL
Sbjct: 77 YL-QSFIPRTA--LSLEDLVRVARTWEERARFCYTEDVRLSSDEYVKMLIVDASFLVELL 133
Query: 148 RDLSDNEFKHVPSL---SRWMLPTIRREMIMLENQLPMFVLTKLFEL--TDKNNSLHPQM 202
+ ++ + + M+ + ++++LENQLP FV+ +F L D + L P
Sbjct: 134 LRSQFDVYRGMLDRIYGKQKMIVDVNHDVMLLENQLPYFVVEGMFGLLHVDYHRELPPLT 193
Query: 203 SFYNLSFKFFYRLLQSESRKTPECQTSYKFKIEHVLDLLRYNIRPKLIGEEPRGCQSQM- 261
+ FK F+ + S SR + KI H +DLLR +I L+ P G M
Sbjct: 194 RIIHNHFKKFWMSIPSFSRSISDS------KICHFVDLLR-SIHLPLVLSFPGGSMRMMD 246
Query: 262 -IHSITELKEGGVKIKACESRE-LMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRNIVA 319
+ S E++ GVK++ ++ +DISF G+ LTIP + I D +++RNI+
Sbjct: 247 SVLSAKEIQNAGVKLQPADNNTCALDISFAN--GV----LTIPKIKINDITESLYRNIIL 300
Query: 320 FEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIV 379
FE+CH R + YM FL+ I S D G+I + G+ E V+ L N+I KE
Sbjct: 301 FEQCH-RLDAYFIHYMRFLSCFIRSPMDAELFIDHGIIVNRFGNAEDVSRLFNSILKETS 359
Query: 380 PD--MNESYLYNVVNEANEYLGCWRARFRASLVHNYLTSWAVGLSTLGALLALYFTFIQA 437
++ N+ N W+A R HN W+ S + A + L TF+QA
Sbjct: 360 YSGFYYKTVYGNLQAHCNAPWNKWKATLRRDYFHN---PWSAA-SVVAACVLLLLTFVQA 415
Query: 438 IC 439
IC
Sbjct: 416 IC 417
>At3g50190 putative protein
Length = 463
Score = 176 bits (447), Expect = 2e-44
Identities = 118/369 (31%), Positives = 188/369 (49%), Gaps = 55/369 (14%)
Query: 42 ESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGS 101
+ +CIYRVP + + +Y P +S+GPYH+G +HL+ M+ K + + +
Sbjct: 94 DKLCIYRVPLSFQKSDKNSYFPHTVSLGPYHHGDEHLRPMDYHKWRAVNMVMKRTKQGIE 153
Query: 102 KLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLSDNEFK----- 156
+A K LE+ AR CY G I LSS++F +M+++DG F++ L R + K
Sbjct: 154 MYIDAMKELEER---ARACYEGPIGLSSNKFTQMLVLDGCFVLDLFRGAYEGFSKLGYDR 210
Query: 157 HVPSLS-RWMLPTIRREMIMLENQLPM---FVLTKLFELTDKNNSLHPQMSFYNLSFKFF 212
+ P + R + +IRR+M+MLENQLP+ ++ + E + +NN+ KFF
Sbjct: 211 NDPVFAMRGSMHSIRRDMLMLENQLPLMPTYMPSTKIENSQENNN------------KFF 258
Query: 213 YRLLQSESRKTPECQTSYKFKIEHVLDLLRYNIRPKLIGEEPRGC-------------QS 259
+ + + + H LD+ R ++ + EPR Q
Sbjct: 259 HPIADKDKEEL------------HCLDVFRRSVLQPSLKPEPRLSRRWSWKPVVADKRQQ 306
Query: 260 QMIHSITELKEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRNIVA 319
+++H +TEL+E G+K K +S L DI F K G L IP L I D ++ N++A
Sbjct: 307 KLLHCVTELREAGIKFKRRKSDRLWDIQF--KNGC----LEIPKLLIHDGTKSLLSNLIA 360
Query: 320 FEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIV 379
FE+CH +T+Y+ F+ LINS DV L Y G+I H L SD V +L + + +++V
Sbjct: 361 FEQCHIDSTKQITSYIIFVENLINSNEDVRYLQYCGIIEHWLESDSEVVDLFHKLCQDVV 420
Query: 380 PDMNESYLY 388
D +SYLY
Sbjct: 421 FDPIDSYLY 429
>At5g22540 unknown protein
Length = 440
Score = 165 bits (417), Expect = 6e-41
Identities = 121/426 (28%), Positives = 204/426 (47%), Gaps = 40/426 (9%)
Query: 32 LQSIKPEEKNESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHR 91
L+ ++ +E CI R+P ++ + KAY P +SIGPYH+G +HL+ + K +F
Sbjct: 19 LKLLRESAGSELCCIVRIPQSLARINLKAYEPKIVSIGPYHHGKEHLKMTQQHKRRFLKF 78
Query: 92 LFDPNGANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLS 151
G +E K + E R Y ++ L S+ ++MM++DG FI+ L +S
Sbjct: 79 FVAKMEEKGFVPQELVKAVSSLEGVIRGSYSEDLGLDSENLVQMMVLDGCFILTLFFVVS 138
Query: 152 --------DNEFKHVPSLSRWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMS 203
D+ +P W+LP+IR ++++LENQ+P +L LFE + L
Sbjct: 139 GKVEYTNLDDPIFRMP----WILPSIRADLLLLENQVPYVLLQTLFE----TSKLVTCSG 190
Query: 204 FYNLSFKFFYRLLQSESRKTPEC--QTSYKFKIEHVLDLLRYNIRPKLIGEEPRGCQSQ- 260
++F+FF LQ PE + Y + +H+LDL+R P + S+
Sbjct: 191 LNEIAFEFFNYSLQK-----PETFWEKHYGLEAKHLLDLIRKTFVPVPSQRRIKDHSSKS 245
Query: 261 ---------MIHSITELKEGGVKIKACESRE-LMDISFGKKWGIMVKELTIPPLYIGDHR 310
+ S +L G+K K ++ + ++DIS+ G+ L IPP+ + D
Sbjct: 246 SFNDHEYLGFVLSAKKLHLRGIKFKPRKNTDSILDISYSN--GV----LHIPPVVMDDFT 299
Query: 311 GTVFRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAEL 370
++F N VAFE+ + + +T+Y+ F+ LIN +D S L + ++ + G+++ V+
Sbjct: 300 ASIFLNCVAFEQLYADSSNHITSYVAFMACLINEESDASFLSERRILENYFGTEDEVSRF 359
Query: 371 INNIAKEIVPDMNESYLYNVVNEANEYLGCWRARFRASLVHNYLTSWAVGLSTLGALLAL 430
I K+I D+ +SYL V NEY A +H + S S+ ALL L
Sbjct: 360 YKRIGKDIALDLEKSYLAKVFEGVNEYTSQGFHVHCAEFIHTHFDSPWTFASSFAALLLL 419
Query: 431 YFTFIQ 436
F +Q
Sbjct: 420 LFAALQ 425
>At3g47200 unknown protein
Length = 476
Score = 162 bits (409), Expect = 5e-40
Identities = 122/417 (29%), Positives = 194/417 (46%), Gaps = 30/417 (7%)
Query: 42 ESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGS 101
ES CI+RVP + + + PKAY P +SIGPYHYG +HLQ ++ K + D
Sbjct: 44 ESCCIFRVPESFVALNPKAYKPKVVSIGPYHYGEKHLQMIQQHKPRLLQLFLDEAKKKDV 103
Query: 102 KLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLSDN-EFKHVPS 160
+ K + E R Y E+K D + MM++DG FI+ + +S N E P
Sbjct: 104 EENVLVKAVVDLEDKIRKSYSEELKTGHD-LMFMMVLDGCFILMVFLIMSGNIELSEDPI 162
Query: 161 LS-RWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFFYRLLQSE 219
S W+L +I+ ++++LENQ+P FVL L+ + + ++F FF + E
Sbjct: 163 FSIPWLLSSIQSDLLLLENQVPFFVLQTLY----VGSKIGVSSDLNRIAFHFFKNPIDKE 218
Query: 220 SRKTPECQTSYKFKIEHVLDLLRYNIRPKLIGEEPRGCQSQMIHSITELKEGGVK----- 274
+ +K +H+LDL+R P E + + + E K G V
Sbjct: 219 G---SYWEKHRNYKAKHLLDLIRETFLPN-TSESDKASSPHVQVQLHEGKSGNVPSVDSK 274
Query: 275 ----IKACESRELMDISF----GKKWGIM-----VKELTIPPLYIGDHRGTVFRNIVAFE 321
I + + L I F K+ I+ +L IP L + F N VAFE
Sbjct: 275 AVPLILSAKRLRLQGIKFRLRRSKEDSILNVRLKKNKLQIPQLRFDGFISSFFLNCVAFE 334
Query: 322 KCHKRCNPDMTTYMFFLNRLINSANDVSAL-HYKGVIHHSLGSDEHVAELINNIAKEIVP 380
+ + + ++TTY+ F+ L+N+ DV+ L + K +I + GS+ V+E I+K++V
Sbjct: 335 QFYTDSSNEITTYIVFMGCLLNNEEDVTFLRNDKLIIENHFGSNNEVSEFFKTISKDVVF 394
Query: 381 DMNESYLYNVVNEANEYLGCWRARFRASLVHNYLTSWAVGLSTLGALLALYFTFIQA 437
+++ SYL NV NEY W A H + S LS+ L + T +Q+
Sbjct: 395 EVDTSYLNNVFKGVNEYTKKWYNGLWAGFRHTHFESPWTFLSSCAVLFVILLTMLQS 451
>At3g47250 unknown protein
Length = 480
Score = 155 bits (392), Expect = 5e-38
Identities = 118/422 (27%), Positives = 190/422 (44%), Gaps = 40/422 (9%)
Query: 43 SVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGSK 102
S CI+R+P ++ V PKAY P +SIGPYHYG HLQ ++ K +F D G
Sbjct: 60 SCCIFRIPDSLAEVNPKAYKPKVVSIGPYHYGENHLQMIQQHKFRFLELFVDRATKKGMD 119
Query: 103 LEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLLRDLS---DNEFKHVP 159
+ + R Y E+++ E + MM++DG FI+ LL +S D + P
Sbjct: 120 ENVLYAAVGALQHKIRASYSEELRIEKSELVSMMILDGCFILMLLLIVSRKIDLDMNKDP 179
Query: 160 SLS-RWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFFYRLLQS 218
+ W+L +I+ ++++LENQ+P FVL +F DK+ P ++F FF +
Sbjct: 180 IFTIPWILASIQSDLLLLENQVPFFVLRTIF---DKSGIGSPG-DLNRMAFSFFNLSIDK 235
Query: 219 ESRKTPECQTSYKFKIEHVLDLLRYNIRPKL--IGEEPRGCQSQMIH------------- 263
+ + +H+LDL R P + +GE S+ H
Sbjct: 236 PDTYWAKHRDR---GAKHLLDLNRKTFIPSMRSMGESETTPSSKFQHSKGKSGEVSSSES 292
Query: 264 ------SITELKEGGVKIK-ACESRELMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRN 316
S L+ G+K + ++ ++DI K +L IP L + ++F N
Sbjct: 293 TFPLILSAKRLRLQGIKFRLRSDAESILDIKLKK------NKLQIPLLRLDGFISSIFLN 346
Query: 317 IVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHY-KGVIHHSLGSDEHVAELINNIA 375
VAFE+ + D+T+Y+ F+ L+N D + L+ K +I + G++ V++ I
Sbjct: 347 CVAFEQFYTESTNDITSYVVFMGCLLNDQEDATFLNNDKRIIENYFGNENEVSQFFKTIC 406
Query: 376 KEIVPDMNESYLYNVVNEANEYLGCWRARFRASLVHNYLTSWAVGLSTLGALLALYFTFI 435
K++V D SYL NV NEY A H + S LS+ + + T
Sbjct: 407 KDVVFDTRRSYLRNVFEGVNEYTSKTYNGVWAGFRHTHFESPWTALSSFAVVFVILLTMT 466
Query: 436 QA 437
QA
Sbjct: 467 QA 468
>At3g47210 unknown protein
Length = 474
Score = 150 bits (380), Expect = 1e-36
Identities = 111/384 (28%), Positives = 186/384 (47%), Gaps = 31/384 (8%)
Query: 42 ESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGS 101
ES CI+RVP + + P+AY P +SIGPYH+G +HL+ ++ K +F H +
Sbjct: 91 ESCCIFRVPKSFAEMNPEAYKPKVVSIGPYHHGRKHLEMIQQHKLRFLHLFLRTASVDRD 150
Query: 102 KLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLL----RDLSDNEFKH 157
L A E E R Y ++ S E + MM++DG FI+ LL R + E +
Sbjct: 151 VLFNAVVDWEDE---IRKSYSEGLEGSPHELVYMMILDGCFILMLLLIVSRKIELYESED 207
Query: 158 VPSLSRWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFFYRLLQ 217
W+LP+I+ ++++LENQ+P FVL LF+ ++ + ++F FF +
Sbjct: 208 PILTIPWILPSIQSDLLLLENQVPFFVLQTLFDKSE----IGVPGDLNRMAFSFFNLSMD 263
Query: 218 SESRKTPECQTSYKFKIEHVLDLLRYNIRP-------KLIGEEPRGCQSQ---MIHSITE 267
R + + F +H+LDL+R + P +L + R S ++ S T
Sbjct: 264 KPERYWVKHRN---FNAKHLLDLIRMSFLPMDGYEDFQLTKGKSRKKSSSGLTLLLSATR 320
Query: 268 LKEGGVKIKACESRE-LMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRNIVAFEKCHKR 326
L G+ + ++DI K L IP L + ++ N VAFE+ + +
Sbjct: 321 LSLQGIDFSLRSGADSMLDIRLKKN------RLQIPVLRLDGFIISILLNCVAFEQFYAK 374
Query: 327 CNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMNESY 386
+T+Y+ F+ L+N D + L + +I + GS++ V++ I K++V D++ SY
Sbjct: 375 STNHITSYVVFMGCLLNGKEDATFLSRRRIIENYFGSEKEVSKFFKTICKDVVFDIHASY 434
Query: 387 LYNVVNEANEYLGCWRARFRASLV 410
L NV E NE W + R+ L+
Sbjct: 435 LRNVFVEINENTSKWYSICRSFLL 458
>At3g44710 unknown protein
Length = 504
Score = 137 bits (345), Expect = 1e-32
Identities = 123/459 (26%), Positives = 199/459 (42%), Gaps = 80/459 (17%)
Query: 45 CIYRVPPNMLNVEPKA-YIPSNISIGPYHYGSQHLQEMEVLKNKFFHRLFDPNGANGSKL 103
CI+++ N + KA Y P +S+GPYH+G ++LQ +E K +F D G
Sbjct: 48 CIFKISHKPENKKYKAAYEPRVVSLGPYHHGKKNLQMIEEHKLRFLKIFMDEAKRKGVDT 107
Query: 104 EEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQLL----RDLSDNEFKHVP 159
K + E + R+ Y + + ++MM++DG FI+ + +S +E ++ P
Sbjct: 108 NGLIKAVSVLEEDIRDSYSESLYSDGKKLIEMMVLDGCFILMIFLVVAGVVSHSEIENDP 167
Query: 160 SLS-RWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFFYRLLQS 218
+ W+LP IR ++I+LENQ+P F+L +F+ + + + F FF LQ
Sbjct: 168 IFAIPWILPAIRNDLILLENQVPFFLLQTIFD----RSKIEKSSGLNEIIFHFFNYSLQK 223
Query: 219 ESRKTPECQTSYKFKIEHVLDLLRYNIRPKLIGEE-----------------PRGCQSQM 261
+ + Q K + H+LDL+R P + EE PR + ++
Sbjct: 224 SNTFWLKHQ---KVEANHLLDLIRNIYMPDVPKEEKKSNHLLDCMMIRKICMPRESKQEL 280
Query: 262 ---------IHSITELKEG-------------------------------GVKIKACESR 281
I+ L+EG G+K KA +
Sbjct: 281 DVMLKKGKQEVDISMLEEGIPKSDSLEEEDSTTGRHHLKMVLSARKLQLKGIKFKARKKA 340
Query: 282 E-LMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRNIVAFEKCHKRCNPD-MTTYMFFLN 339
E LMDI K L IPPL + D V N VAFE+ + C MT+Y+ F+
Sbjct: 341 ETLMDIRHKGKL------LQIPPLILDDFLIAVLLNCVAFEQYYSYCTKQHMTSYVAFMG 394
Query: 340 RLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMNESYLYNVVNEANEYLG 399
L+ S D L G++ + GS + V+ + K+++ D++ESYL + N+Y
Sbjct: 395 CLLKSEADAMFLSEVGILENYFGSGDEVSRFFKVVGKDVLFDIDESYLAGIFEGVNKYTS 454
Query: 400 C-WRARFRASLVHNYLTSWAVGLSTLGALLALYFTFIQA 437
W ++ A H + S LS+ AL + T IQA
Sbjct: 455 SGWHVQW-AGFKHTHFDSPWTALSSCAALTLVILTIIQA 492
>At2g44930 unknown protein
Length = 515
Score = 112 bits (281), Expect = 4e-25
Identities = 123/479 (25%), Positives = 199/479 (40%), Gaps = 67/479 (13%)
Query: 14 HGTAVKS------SKFWR--GLVEKELQSIKPEE-KNESVCIYRVPPNMLNVEPKAYIPS 64
HGTA K+ +K ++ E E+ I P E CIYRVP + V P+AY P
Sbjct: 45 HGTAEKAKIVSVTTKPYKPDSCYESEISGIWPSSFSKEYCCIYRVPNRLRRVNPEAYTPQ 104
Query: 65 NISIGPYHYGSQ--------------HLQEMEVLKNKFFHRLFDPNGANGSKLEEACKFL 110
+ IGP + + ME+ K K+ + + + G +E + +
Sbjct: 105 MLLIGPLQHSKKANALELSKTDLRYLDYMNMELHKKKYLNAIANKYG--DQIIEVFKEII 162
Query: 111 EKEEINARNCYMGEIK-LSSDEFLKMMLVDGSFIIQLL--RDLSDNEFKHVPSLSR---- 163
E E R Y + +F+ M+L+D FI+ + + + N K L
Sbjct: 163 ETSEKFVRESYAESTDWIKPQDFVDMILLDSVFILGVFIQAETTQNIKKKEEILFHEEDI 222
Query: 164 -----WMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQMSFYNLSFKFFYR-LLQ 217
++ TI ++I+LENQLP +L KL + F NL K +R ++
Sbjct: 223 IFQEPCLITTILEDLILLENQLPYVLLEKL-----------SKPFFANLKTKETFRDIIL 271
Query: 218 SESRKTPECQTSYKFKIEHVLDLLRYNIRPK---LIGEEPRGCQSQ------MIHSITEL 268
R E + F+ H DL R +R + L E+ + + + +H+ +L
Sbjct: 272 RAFRLKGEIKEGTNFR--HFTDLFR-RVRVETLCLTKEQIKSAKDKPPEIIKSLHNADKL 328
Query: 269 KEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYIGDHRGTVFRNIVAFEKCHKRCN 328
GV E + + + + GI L IP D+ + RN++A E+CH
Sbjct: 329 DSAGVDFVRLERKNDLSLVITFERGI----LEIPCFLADDNTERIMRNLMALEQCHYPLT 384
Query: 329 PDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELINNIAKEIVPDMNESYLY 388
+ Y+ FL+ LI++ DV L KGVI + LG VAE++N + +V D Y Y
Sbjct: 385 AYVCNYIAFLDFLIDTDQDVDLLVKKGVIKNWLGHQASVAEMVNKLCLGLV-DFGSHY-Y 442
Query: 389 NVVNEANEYLGCWRARFRASLVHNYLTSWAVGLSTLGALLALYFTFIQAICGFADAATK 447
+ + N++ R R A+L Y G +T+ A + L T I + K
Sbjct: 443 GIADRLNKHYESRRNRSIATLRRVYFKDLWTGTATIAAAVILVLTLIGTVASVLQVTQK 501
>At2g28580 hypothetical protein
Length = 468
Score = 111 bits (277), Expect = 1e-24
Identities = 114/437 (26%), Positives = 197/437 (44%), Gaps = 59/437 (13%)
Query: 34 SIKPEEKNESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQ--------------HLQ 79
S++PE CIYRVP + V P+AY P + IGP H+ + +
Sbjct: 35 SVRPE----FCCIYRVPDRLRKVNPEAYTPQMLLIGPLHHSKKAEALKRYKTDLRYLNYL 90
Query: 80 EMEVLKNKFFHRLFDPNGANGSKLEEACKFLEKEEINARNCYM-GEIKLSSDEFLKMMLV 138
ME+ K K + D G ++E + +E E R+ Y I +++ +F++M+L
Sbjct: 91 NMELHKKKCLDSIADIYG--DQPVKEFRRIIEINEKFIRDSYAESTIWINTKDFVEMILH 148
Query: 139 DGSFI----IQLLRDLSDNEFKHVP-SLSRWMLPT-IRREMIMLENQLPMFVLTKLFELT 192
D FI IQ L+ N+ + + + SR + T I ++I+LENQLP +L KLFE
Sbjct: 149 DSVFILLFFIQTGSTLNFNKKEDILFNQSRLINATAILEDLILLENQLPYALLEKLFEPF 208
Query: 193 DKNNSLHPQMSFYNLSFKFFYRLLQSESRKTPECQTSYKFKIEHVLDLLRYNIRPKLIG- 251
N ++ + +F +++ + F E + + + +H DL R +R +
Sbjct: 209 SSN--VNTKETFRDITLRAFRF----------EGKIKEEVRFQHFTDLFRC-VRVSTLSL 255
Query: 252 ----------EEPRGCQSQMIHSITELKEGGVKIKACESRELMDISFGKKWGIMVKELTI 301
E P+ +++++ +L GV + + + K GI L +
Sbjct: 256 TEEQINIAKNEPPKS--RKIMYNADKLDSAGVNFVNVDEENDLSLVITFKDGI----LKM 309
Query: 302 PPLYIGDHRGTVFRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSL 361
P + D+ V RN++A E+CH + Y+ FL+ LIN+ DV L KG++ + L
Sbjct: 310 PCFTVEDNTERVVRNLMALEQCHYPRTTFVCDYISFLDFLINTDQDVDLLAKKGIVKNWL 369
Query: 362 GSDEHVAELINNIAKEIVPDMNESYLYNVVNEANEYLGCWRARFRASLVHNYLTSWAVGL 421
G V E++N + +V D Y ++V N++ R +L Y G
Sbjct: 370 GHQGSVTEMVNKLCLGLV-DFGSHY-SDIVENLNKHYDNRLNRSVGTLRRVYFKDLWTGT 427
Query: 422 STLGALLALYFTFIQAI 438
+T+ A++ L T IQ +
Sbjct: 428 ATIAAVVLLVLTLIQTV 444
>At5g22560 putative protein
Length = 517
Score = 93.6 bits (231), Expect = 2e-19
Identities = 62/227 (27%), Positives = 108/227 (47%), Gaps = 29/227 (12%)
Query: 41 NESVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEMEVL--------------KN 86
++S CI+R+P ++ AY P +SIGPYH+ +++ K
Sbjct: 44 HQSCCIFRIPHTLVQSNDTAYKPKIVSIGPYHHDHGKADKVDKADKADKARFKMIQQHKQ 103
Query: 87 KFFHRLFDPNGANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGSFIIQL 146
++ G +E+ + K+E R+ Y E++L+ E + +M++DG FI+ L
Sbjct: 104 RYLDIFLSKTTKKGVGIEDLQNVVWKKEHLIRDSYSEELQLNQVELIDLMVLDGCFILML 163
Query: 147 L----RDLSDNEFKHVPSLSRWMLPTIRREMIMLENQLPMFVLTKLFELTDKNNSLHPQM 202
R + F+ + RW+LPT+R ++++LENQ+P+F+L LF T L +
Sbjct: 164 FLLVSRKVPHKAFEDPIFMLRWILPTLRSDLLLLENQVPLFLLRDLFNTT----KLATKT 219
Query: 203 SFYNLSFKFFYRLLQSESRKTPECQTSYKFKIE--HVLDLLRYNIRP 247
S + F FF S K P+ + ++ H+LDL+R P
Sbjct: 220 SLNEMIFNFF-----GYSIKRPQKFWDERMNLDASHLLDLIRKTFVP 261
Score = 78.6 bits (192), Expect = 7e-15
Identities = 61/198 (30%), Positives = 101/198 (50%), Gaps = 9/198 (4%)
Query: 247 PKLIGEEPRGCQSQMIHSITELKEGGVKIKACESREL-MDISFGKKWGIMVKELTIPPLY 305
PK + PR +++ S +L+ G+K + + E +DI+ K G+ L IPPL
Sbjct: 321 PKPLTPPPRRFL-KLVVSARKLRLRGIKFQQKKKFETPLDITL--KNGV----LKIPPLL 373
Query: 306 IGDHRGTVFRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDE 365
D ++ N VAFE+ + + +MT+Y+ F+ LIN+A+D + L KG+I + G+ E
Sbjct: 374 FDDFFSSLLINCVAFEQFNVQGTTEMTSYVTFMGCLINTADDATFLSEKGIIENYFGTGE 433
Query: 366 HVAELINNIAKEIVPDMNESYLYNVVNEANEYLGCWRARFRASLVHNYLTSWAVGLSTLG 425
++ N K+I +++SYL NV N+Y A + + Y S LS+
Sbjct: 434 QLSVFFKNTGKDISFTISKSYLANVFEGVNKYTSQGYHVHWAGVKYTYFKSPWTFLSSFA 493
Query: 426 ALLALYFTFIQA-ICGFA 442
AL+ + T QA G+A
Sbjct: 494 ALVLILLTIFQAFFAGYA 511
>At3g02650 hypothetical protein
Length = 1077
Score = 82.0 bits (201), Expect = 7e-16
Identities = 58/170 (34%), Positives = 87/170 (51%), Gaps = 8/170 (4%)
Query: 24 WRGLVEKELQSIKPEEKNE--SVCIYRVPPNMLNVEPKAYIPSNISIGPYHYGSQHLQEM 81
W V+K L + E E +V I+ VP ++ P +Y P +SIGPYH L EM
Sbjct: 21 WVINVQKSLDAELEEHDLEEVTVSIFNVPKALMCSHPDSYTPHRVSIGPYHCLKPELHEM 80
Query: 82 EVLKNKFFHRLFDPNGANGSKLEEACKFLEKEEINARNCYMGEIKLSSDEFLKMMLVDGS 141
E K ++ N N + + + L+ EI R CY I + + L +M VD S
Sbjct: 81 ERYKLMIARKI--RNQYNSFRFHDLVEKLQSMEIKIRACYHKYIGFNGETLLWIMAVDSS 138
Query: 142 FIIQLLRDLSDNEFKHVPSL-SRWMLPTIRREMIMLENQLPMFVLTKLFE 190
F+I+ L+ S F+ V +L +R I R+++M+ENQ+P+FVL K E
Sbjct: 139 FLIEFLKIYS---FRKVETLINRVGHNEILRDIMMIENQIPLFVLRKTLE 185
Score = 63.2 bits (152), Expect = 3e-10
Identities = 51/181 (28%), Positives = 87/181 (47%), Gaps = 9/181 (4%)
Query: 252 EEPRGCQSQMIHSITELKEGGVKIKACESRELMDISFGKKWGIMVKELTIPPLYIGDHRG 311
E+P + I S+++L + GV+ K + ++F G + +P + + +
Sbjct: 335 EKPPLVEELTIPSVSDLHKAGVRFKPTAHGNISTVTFDSNSG----QFYLPVINLDINTE 390
Query: 312 TVFRNIVAFEKCHKRCNPDMTTYMFFLNRLINSANDVSALHYKGVIHHSLGSDEHVAELI 371
TV RN+VA+E + T Y +N +I+S DV L +GV+ L SD+ AE+
Sbjct: 391 TVLRNLVAYEATNTSGPLVFTRYTELINGIIDSEEDVRLLREQGVLVSRLKSDQEAAEMW 450
Query: 372 NNIAKEIVPDMNESYLYNVVNEANE-YLGCWRARFRASLVHNYL-TSWAVGLSTLGALLA 429
N ++K V +L + + N Y G W+ + LV Y+ SW + L+ L A+L
Sbjct: 451 NGMSKS-VRLTKVGFLDKTIEDVNRYYTGRWKVKI-GRLVEVYVYGSWQI-LAFLAAVLL 507
Query: 430 L 430
L
Sbjct: 508 L 508
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,221,463
Number of Sequences: 26719
Number of extensions: 486511
Number of successful extensions: 1307
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 1190
Number of HSP's gapped (non-prelim): 43
length of query: 487
length of database: 11,318,596
effective HSP length: 103
effective length of query: 384
effective length of database: 8,566,539
effective search space: 3289550976
effective search space used: 3289550976
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC130275.9


