

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.7 - phase: 0 /pseudo
(833 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
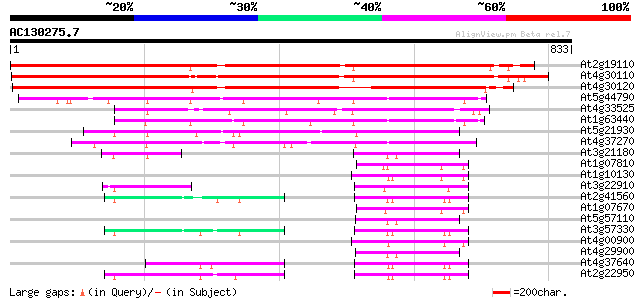
Score E
Sequences producing significant alignments: (bits) Value
At2g19110 putative heavy metal transporter (hma4) 779 0.0
At4g30110 cadmium-transporting ATPase-like protein 766 0.0
At4g30120 cadmium-transporting ATPase-like protein 702 0.0
At5g44790 COPPER-TRANSPORTING ATPASE RAN1 (RESPONSIVE-TO-ANTAGON... 230 3e-60
At4g33525 metal-transporting P-type ATPase 212 6e-55
At1g63440 ATP dependent copper transporter, putative 208 1e-53
At5g21930 metal-transporting ATPase - like protein 190 3e-48
At4g37270 Cu2+-transporting ATPase-like protein 173 3e-43
At3g21180 putative Ca2+-transporting ATPase 77 5e-14
At1g07810 ER-type Ca2+-pump protein 76 7e-14
At1g10130 putative calcium ATPase 75 2e-13
At3g22910 calmodulin-stimulated calcium-ATPase, putative 75 2e-13
At2g41560 putative Ca2+-ATPase 75 2e-13
At1g07670 endoplasmic reticulum-type calcium-transporting ATPase... 75 2e-13
At5g57110 Ca2+-transporting ATPase-like protein (emb|CAB79748.1) 73 6e-13
At3g57330 Ca2+-transporting ATPase-like protein 73 6e-13
At4g00900 Ca2+-transporting ATPase - like protein 73 7e-13
At4g29900 Ca2+-transporting ATPase - like protein 70 4e-12
At4g37640 plasma membrane-type calcium ATPase (ACA2) 67 5e-11
At2g22950 pseudogene 65 2e-10
>At2g19110 putative heavy metal transporter (hma4)
Length = 1172
Score = 779 bits (2012), Expect = 0.0
Identities = 414/796 (52%), Positives = 546/796 (68%), Gaps = 49/796 (6%)
Query: 2 VENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIA 61
V+ +++S F+VLG+CC +E ++E ILK L GVK SVIVP+RTV VVHD LLIS +IA
Sbjct: 13 VKKLQKSYFDVLGICCTSEVPIIENILKSLDGVKEYSVIVPSRTVIVVHDSLLISPFQIA 72
Query: 62 DALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGF 121
ALN ARLEA+ R GET+ + K P + GLLL LSFLK++Y PL WLA+ +VA G
Sbjct: 73 KALNEARLEANVRVNGETSFKNKWPSPFAVVSGLLLLLSFLKFVYSPLRWLAVAAVAAGI 132
Query: 122 PKVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKAL 181
+L +A ASI+ ++INILV++ V T A+QDF + ++FLF+I+ WLETRA++KA
Sbjct: 133 YPILAKAFASIKRPRIDINILVIITVIATLAMQDFMEAAAVVFLFTISDWLETRASYKAT 192
Query: 182 VAMSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKML 241
M SL ++APQ AI+AETGE V+V++VK++T++AVKAG+ IP+DGIVV+G CEVDEK L
Sbjct: 193 SVMQSLMSLAPQKAIIAETGEEVEVDEVKVDTVVAVKAGETIPIDGIVVDGNCEVDEKTL 252
Query: 242 TGESFPVTKESDSLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNYSVGKRHSSGKNV 296
TGE+FPV K+ DS VWAGTIN+N Y + D ++ K + + S + +
Sbjct: 253 TGEAFPVPKQRDSTVWAGTINLNGYICVKTTSLAGDCVVAKMAKLVEEAQSSKTKSQRLI 312
Query: 297 KTC*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALI 356
C + + +I ++ +C+A+VP + V +++ WFHLALVVL+SGCPC LI
Sbjct: 313 DKCSQYYTPAII----------LVSACVAIVPVIMKVHNLKHWFHLALVVLVSGCPCGLI 362
Query: 357 LSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDD 415
LSTPVA FCALTKAA SGLL+K DYL+TLS+IK VAFDKTGTITRGEF V DF S D
Sbjct: 363 LSTPVATFCALTKAATSGLLIKSADYLDTLSKIKIVAFDKTGTITRGEFIVIDFKSLSRD 422
Query: 416 INNETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRD 475
IN +LLYW+SS+ESKSSHP+A +VDYA+ S++P PE VE++QNFPGEGI+G IDG D
Sbjct: 423 INLRSLLYWVSSVESKSSHPMAATIVDYAKSVSVEPRPEEVEDYQNFPGEGIYGKIDGND 482
Query: 476 IYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNN------CEETLVGVFSLVDACRSGAL 529
I+IGNK++ RA C E+ TK E L G F+L DACRSG
Sbjct: 483 IFIGNKKIASRAGCS------TVPEIEVDTKGGKTVGYVYVGERLAGFFNLSDACRSGVS 536
Query: 530 EAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPI 589
+AM ELK LG+++ MLTGD+ AM+ Q QL N +D+VH +LLP +K+++I+ FKKEGP
Sbjct: 537 QAMAELKSLGIKTAMLTGDNQAAAMHAQEQLGNVLDVVHGDLLPEDKSRIIQEFKKEGPT 596
Query: 590 AMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKL 649
AM+GDG+NDAPALATADIGISMGISGSALA +T N ILMSNDIR++P+A++LAR+ RK+
Sbjct: 597 AMVGDGVNDAPALATADIGISMGISGSALATQTGNIILMSNDIRRIPQAVKLARRARRKV 656
Query: 650 VENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTK 709
VENV +S+ K ILALA AG+PL+W AVL DVGTCLLVI NSML+L+E K ++ +
Sbjct: 657 VENVCLSIILKAGILALAFAGHPLIWAAVLVDVGTCLLVIFNSMLLLREKKKIGNKKCYR 716
Query: 710 GSK---YGKFLEDKTKPLLNKQSNIDEEKGLLS---GEECGKDCCKNATHYVETSKERND 763
S G+ LE + +D E GLL+ +C CC ++N
Sbjct: 717 ASTSKLNGRKLEG------DDDYVVDLEAGLLTKSGNGQCKSSCC---------GDKKNQ 761
Query: 764 ESCGVSKLSLLNGNDH 779
E+ + K S +DH
Sbjct: 762 ENVVMMKPSSKTSSDH 777
>At4g30110 cadmium-transporting ATPase-like protein
Length = 951
Score = 766 bits (1979), Expect = 0.0
Identities = 420/829 (50%), Positives = 555/829 (66%), Gaps = 47/829 (5%)
Query: 3 ENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIAD 62
+ + +S F+VLG+CC +E L+E IL + GVK SVIVP+RTV VVHD L++S+ +I
Sbjct: 4 KKMTKSYFDVLGICCTSEVPLIENILNSMDGVKEFSVIVPSRTVIVVHDTLILSQFQIVK 63
Query: 63 ALNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFP 122
ALN A+LEA+ R GETN + K P + G+LL LSF KY+Y P WLA+ +V G
Sbjct: 64 ALNQAQLEANVRVTGETNFKNKWPSPFAVVSGILLLLSFFKYLYSPFRWLAVAAVVAGIY 123
Query: 123 KVLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALV 182
+L +A+AS+ ++INILV++ V T +QD+T+ V++FLF+IA+WL++RA++KA
Sbjct: 124 PILAKAVASLARFRIDINILVVVTVGATIGMQDYTEAAVVVFLFTIAEWLQSRASYKASA 183
Query: 183 AMSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLT 242
M SL ++APQ A++AETGE V+V+++K NT++AVKAG+ IP+DG+VV+G CEVDEK LT
Sbjct: 184 VMQSLMSLAPQKAVIAETGEEVEVDELKTNTVIAVKAGETIPIDGVVVDGNCEVDEKTLT 243
Query: 243 GESFPVTKESDSLVWAGTINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RS 302
GE+FPV K DS VWAGTIN+N Y I +T C + K +N KT +
Sbjct: 244 GEAFPVPKLKDSTVWAGTINLNGY--ITVNTTAL-AEDCVVAKMAKLVEEAQNSKTETQR 300
Query: 303 F*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTPVA 362
F K +K + ++ C +P AL V +++ W HLALVVL+S CPC LILSTPVA
Sbjct: 301 FIDK--CSKYYTPAIILISICFVAIPFALKVHNLKHWVHLALVVLVSACPCGLILSTPVA 358
Query: 363 IFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDDINNETL 421
FCALTKAA SGLL+KG DYLETL++IK VAFDKTGTITRGEF V DF S +DI+ ++L
Sbjct: 359 TFCALTKAATSGLLIKGADYLETLAKIKIVAFDKTGTITRGEFIVMDFQSLSEDISLQSL 418
Query: 422 LYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNK 481
LYW+SS ESKSSHP+A A+VDYAR S++P PE VE++QNFPGEGI+G IDG+++YIGNK
Sbjct: 419 LYWVSSTESKSSHPMAAAVVDYARSVSVEPKPEAVEDYQNFPGEGIYGKIDGKEVYIGNK 478
Query: 482 RVGVRAICKRDNCEVQFQRPEISTKKNN------CEETLVGVFSLVDACRSGALEAMEEL 535
R+ RA C + ++ TK ETL GVF+L DACRSG +AM+EL
Sbjct: 479 RIASRAGC------LSVPDIDVDTKGGKTIGYVYVGETLAGVFNLSDACRSGVAQAMKEL 532
Query: 536 KLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKK-EGPIAMIGD 594
K LG++ MLTGD+ AM+ Q QL NA+DIV AELLP +K+++I+ K+ EGP AM+GD
Sbjct: 533 KSLGIKIAMLTGDNHAAAMHAQEQLGNAMDIVRAELLPEDKSEIIKQLKREEGPTAMVGD 592
Query: 595 GINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVI 654
G+NDAPALATADIGISMG+SGSALA ET N ILMSNDIR++P+AI+LA++ RK+VENV+
Sbjct: 593 GLNDAPALATADIGISMGVSGSALATETGNIILMSNDIRRIPQAIKLAKRAKRKVVENVV 652
Query: 655 ISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTKGSKYG 714
IS+ K AILALA AG+PL+W AVL DVGTCLLVILNSML+L + HK + + S
Sbjct: 653 ISITMKGAILALAFAGHPLIWAAVLADVGTCLLVILNSMLLLSDKHKTGNKCYRESSSSS 712
Query: 715 KFLEDKTKPLLNKQSNIDEEKGLL---SGEECGKDCCKNATH-----------------Y 754
+ +K L + D E GLL S + C CC T
Sbjct: 713 VLIAEK----LEGDAAGDMEAGLLPKISDKHCKPGCCGTKTQEKAMKPAKASSDHSHSGC 768
Query: 755 VETSKERN----DESCGVSKLSLLNGNDHGMFMFVEVHVVKHCVMIQDS 799
ET ++ N +SC + L +G+D G +H V +Q S
Sbjct: 769 CETKQKDNVTVVKKSCCAEPVDLGHGHDSGCCGDKSQQPHQHEVQVQQS 817
>At4g30120 cadmium-transporting ATPase-like protein
Length = 711
Score = 702 bits (1811), Expect = 0.0
Identities = 380/756 (50%), Positives = 499/756 (65%), Gaps = 79/756 (10%)
Query: 4 NIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADA 63
N++ S F+V+G+CC++E ++V +L+ + GVK SVIVP+RTV VVHD LIS +I A
Sbjct: 11 NLQTSYFDVVGICCSSEVSIVGNVLRQVDGVKEFSVIVPSRTVIVVHDTFLISPLQIVKA 70
Query: 64 LNTARLEASFRPQGETNNEKKCPDILTMACGLLLALSFLKYIYPPLGWLALGSVAIGFPK 123
LN ARLEAS RP GET+ + + P + G+LL LSF KY Y PL WLA+ +V G
Sbjct: 71 LNQARLEASVRPYGETSLKSQWPSPFAIVSGVLLVLSFFKYFYSPLEWLAIVAVVAGVFP 130
Query: 124 VLFRAIASIRALTLNINILVLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVA 183
+L +A+AS+ L+IN L L+AV T +QDFT+ I+FLFS+A WLE+ A HKA +
Sbjct: 131 ILAKAVASVTRFRLDINALTLIAVIATLCMQDFTEAATIVFLFSVADWLESSAAHKASIV 190
Query: 184 MSSLTNMAPQMAIVAETGERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTG 243
MSSL ++AP+ A++A+TG VDV++V +NT+++VKAG++IP+DG+VV+G C+VDEK LTG
Sbjct: 191 MSSLMSLAPRKAVIADTGLEVDVDEVGINTVVSVKAGESIPIDGVVVDGSCDVDEKTLTG 250
Query: 244 ESFPVTKESDSLVWAGTINMNAYPKI-----IFDTYDFRIYKCKNYSVGKRHSSGKNVKT 298
ESFPV+K+ +S V A TIN+N Y K+ D ++ K + + + + +
Sbjct: 251 ESFPVSKQRESTVMAATINLNGYIKVKTTALARDCVVAKMTKLVEEAQKSQTKTQRFIDK 310
Query: 299 C*RSF*QKVISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILS 358
C R + V+ V +C AV+P L V D+ WFHLALVVL+SGCPC LILS
Sbjct: 311 CSRYYTPAVV----------VSAACFAVIPVLLKVQDLSHWFHLALVVLVSGCPCGLILS 360
Query: 359 TPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDF-SAGDDIN 417
TPVA FCALTKAA SG L+K GD LETL++IK VAFDKTGTIT+ EF V+DF S IN
Sbjct: 361 TPVATFCALTKAATSGFLIKTGDCLETLAKIKIVAFDKTGTITKAEFMVSDFRSLSPSIN 420
Query: 418 NETLLYWISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIY 477
LLYW+SSIE KSSHP+A AL+DYAR S++P P+ VENFQNFPGEG++G IDG+DIY
Sbjct: 421 LHKLLYWVSSIECKSSHPMAAALIDYARSVSVEPKPDIVENFQNFPGEGVYGRIDGQDIY 480
Query: 478 IGNKRVGVRAICKRDNCEVQFQRPEISTKKNNCEETLVGVFSLVDACRSGALEAMEELKL 537
IGNKR+ RA C L
Sbjct: 481 IGNKRIAQRAGC-----------------------------------------------L 493
Query: 538 LGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPIAMIGDGIN 597
G+++ MLTGD+ AM Q QL+NA+DIVH+ELLP +KA++I++FK +GP M+GDG+N
Sbjct: 494 TGIQTAMLTGDNQDAAMSTQEQLENALDIVHSELLPQDKARIIDDFKIQGPTMMVGDGLN 553
Query: 598 DAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISV 657
DAPALA ADIGISMGISGSALA ET + ILMSNDIRK+P+ +RLA+++ +K++ENV++SV
Sbjct: 554 DAPALAKADIGISMGISGSALATETGDIILMSNDIRKIPKGMRLAKRSHKKVIENVVLSV 613
Query: 658 GFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEK---RESTKGSKYG 714
K AI+ L GYPLVW AVL D GTCLLVILNSM++L++ + R ST S
Sbjct: 614 SIKGAIMVLGFVGYPLVWAAVLADAGTCLLVILNSMMLLRDEREAVSTCYRAST--SSPV 671
Query: 715 KFLEDKTKPLLNKQSNIDEEKGLL--SGEECGKDCC 748
K ED+ + D E GLL S E K CC
Sbjct: 672 KLEEDEVE---------DLEVGLLQKSEETSKKSCC 698
>At5g44790 COPPER-TRANSPORTING ATPASE RAN1 (RESPONSIVE-TO-ANTAGONIST
1)
Length = 1001
Score = 230 bits (586), Expect = 3e-60
Identities = 224/796 (28%), Positives = 366/796 (45%), Gaps = 118/796 (14%)
Query: 14 GMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALNTARL---- 69
G+ +A ++E IL L+GV+ + + + VV D ++S + D +
Sbjct: 215 GILNELDAQVLEGILTRLNGVRQFRLDRISGELEVVFDPEVVSSRSLVDGIEEDGFGKFK 274
Query: 70 --------EASFRPQGETNNEKK------------------CPDI------LTMACGLLL 97
S + GE +N + CP I L CG +
Sbjct: 275 LRVMSPYERLSSKDTGEASNMFRRFISSLVLSIPLFFIQVICPHIALFDALLVWRCGPFM 334
Query: 98 ALSFLKYIYPPLGWLALGSVAIGFPKVLFRAIASIRALTLNINILVLL------------ 145
+LK+ + +G + A ++R + N+++LV L
Sbjct: 335 MGDWLKWALVSVIQFVIGK------RFYVAAWRALRNGSTNMDVLVALGTSASYFYSVGA 388
Query: 146 ----AVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAETG 201
AV G + F ++I + ++LE+ A K AM L + P AI+ G
Sbjct: 389 LLYGAVTGFWSPTYFDASAMLITFVLLGKYLESLAKGKTSDAMKKLVQLTPATAILLTEG 448
Query: 202 E--------RVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESD 253
+ +D ++ L V G IP DG+VV G V+E M+TGES PV+KE D
Sbjct: 449 KGGKLVGEREIDALLIQPGDTLKVHPGAKIPADGVVVWGSSYVNESMVTGESVPVSKEVD 508
Query: 254 SLVWAGTINMNAY-----PKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVI 308
S V GTINM+ K+ D +I + + K F VI
Sbjct: 509 SPVIGGTINMHGALHMKATKVGSDAVLSQIISLVETAQMSKAPIQKFADYVASIFVPVVI 568
Query: 309 STKIHR*FRKVLYSCIAVVPAALS--VPDMEPWFHLALV----VLLSGCPCALILSTPVA 362
+ + F V +S V A +P+ F +L+ V++ CPCAL L+TP A
Sbjct: 569 TLAL---FTLVGWSIGGAVGAYPDEWLPENGTHFVFSLMFSISVVVIACPCALGLATPTA 625
Query: 363 IFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSAGDDINNETLL 422
+ A A +G+L+KGGD LE ++K V FDKTGT+T+G+ +VT +++ L
Sbjct: 626 VMVATGVGATNGVLIKGGDALEKAHKVKYVIFDKTGTLTQGKATVTTTKVFSEMDRGEFL 685
Query: 423 YWISSIESKSSHPVAGALVDYAR-LHSIKPVPENVE----------------NFQNFPGE 465
++S E+ S HP+A A+V YAR H E+ E +F PG+
Sbjct: 686 TLVASAEASSEHPLAKAIVAYARHFHFFDESTEDGETNNKDLQNSGWLLDTSDFSALPGK 745
Query: 466 GIFGTIDGRDIYIGNKR-VGVRAICKRDNCEVQFQRPEISTKKN---NCEETLVGVFSLV 521
GI ++ + I +GN++ + AI D+ E + E S K LVGV +
Sbjct: 746 GIQCLVNEKMILVGNRKLMSENAINIPDHVEKFVEDLEESGKTGVIVAYNGKLVGVMGIA 805
Query: 522 DACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIE 581
D + A +E L +GVR +M+TGD+ + A V ++ I+ V AE++P KA +I
Sbjct: 806 DPLKREAALVVEGLLRMGVRPIMVTGDNWRTARAVAKEV--GIEDVRAEVMPAGKADVIR 863
Query: 582 NFKKEG-PIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIR 640
+ +K+G +AM+GDGIND+PALA AD+G+++G +G+ +A E ++ +LM N++ V AI
Sbjct: 864 SLQKDGSTVAMVGDGINDSPALAAADVGMAIG-AGTDVAIEAADYVLMRNNLEDVITAID 922
Query: 641 LARKTTRKLVENVIISVGFKCAILALAIAG--YPLV------WLAVLTDVGTCLLVILNS 692
L+RKT ++ N + ++ + + +A AG +P++ W A + + V+ +S
Sbjct: 923 LSRKTLTRIRLNYVFAMAYNVVSIPIA-AGVFFPVLRVQLPPWAAGACMALSSVSVVCSS 981
Query: 693 MLILQENHKYEKREST 708
+L+ +Y+K T
Sbjct: 982 LLL----RRYKKPRLT 993
Score = 38.5 bits (88), Expect = 0.016
Identities = 24/78 (30%), Positives = 36/78 (45%)
Query: 2 VENIKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIA 61
V +++ V GM CA + VE L ++GV SV + VV D L+ E I
Sbjct: 52 VSGLRKIQVGVTGMTCAACSNSVEAALMNVNGVFKASVALLQNRADVVFDPNLVKEEDIK 111
Query: 62 DALNTARLEASFRPQGET 79
+A+ A EA + +T
Sbjct: 112 EAIEDAGFEAEILAEEQT 129
Score = 35.4 bits (80), Expect = 0.13
Identities = 22/63 (34%), Positives = 31/63 (48%)
Query: 10 FEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALNTARL 69
F + GM CA VE IL+ L GVK V + V +D +I++ I +A+ A
Sbjct: 137 FTIGGMTCAACVNSVEGILRDLPGVKRAVVALSTSLGEVEYDPNVINKDDIVNAIEDAGF 196
Query: 70 EAS 72
E S
Sbjct: 197 EGS 199
>At4g33525 metal-transporting P-type ATPase
Length = 632
Score = 212 bits (540), Expect = 6e-55
Identities = 177/615 (28%), Positives = 285/615 (45%), Gaps = 83/615 (13%)
Query: 156 FTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAETGE------RVDVNDV 209
F + ++I + + LE RA KA M+ L ++ P A + G+ V N +
Sbjct: 31 FEEPVMLIAFVLLGRNLEQRAKIKATSDMTGLLSVLPSKARLLLDGDLQNSTVEVPCNSL 90
Query: 210 KMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAYPKI 269
+ ++ + GD +P DG+V G+ +DE TGE PVTKES S V AG+IN+N
Sbjct: 91 SVGDLVVILPGDRVPADGVVKSGRSTIDESSFTGEPLPVTKESGSQVAAGSINLNG---- 146
Query: 270 IFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKV-ISTKIHR*FRKVLYSCIA--- 325
T +++ G + G ++ + ++ + + + + Y +A
Sbjct: 147 ---TLTVEVHRS-----GGETAVGDIIRLVEEAQSREAPVQQLVDKVAGRFTYGVMALSA 198
Query: 326 ------------VVPAAL-SVPDMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAI 372
V+P+AL + M L+ VL+ CPCAL L+TP A+ + A
Sbjct: 199 ATFTFWNLFGAHVLPSALHNGSPMSLALQLSCSVLVVACPCALGLATPTAMLVGTSLGAR 258
Query: 373 SGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTD--------FSAGDDINNETLLYW 424
GLLL+GGD LE S + TV FDKTGT+T+G VT+ + D + +L
Sbjct: 259 RGLLLRGGDILEKFSLVDTVVFDKTGTLTKGHPVVTEVIIPENPRHNLNDTWSEVEVLML 318
Query: 425 ISSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGN---- 480
+++ES ++HPV A+V AR + + + F PG G ++ + + +G
Sbjct: 319 AAAVESNTTHPVGKAIVKAARARNCQTMKAEDGTFTEEPGSGAVAIVNNKRVTVGTLEWV 378
Query: 481 KRVGVRAICKRDNCEVQFQRPEISTKK---NNCEETLVGVFSLVDACRSGALEAMEELKL 537
KR G N + + EI+ + + TL V D R A + +E L
Sbjct: 379 KRHGATG-----NSLLALEEHEINNQSVVYIGVDNTLAAVIRFEDKVREDAAQVVENLTR 433
Query: 538 LGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPI-AMIGDGI 596
G+ ML+GD A YV S + + V A + P EK I +K I AM+GDGI
Sbjct: 434 QGIDVYMLSGDKRNAANYVASVVGINHERVIAGVKPAEKKNFINELQKNKKIVAMVGDGI 493
Query: 597 NDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIIS 656
NDA ALA++++G++MG G+ A+E S +LM N + ++ +A+ L+R+T + + +N+ +
Sbjct: 494 NDAAALASSNVGVAMG-GGAGAASEVSPVVLMGNRLTQLLDAMELSRQTMKTVKQNLWWA 552
Query: 657 VGFKCAILALAIAGYPLVWLAVLTDVGTCLL--------------VILNSMLI-----LQ 697
G+ I G P+ +L GT L V+ NS+L+
Sbjct: 553 FGYN-------IVGIPIAAGVLLPLTGTMLTPSMAGALMGVSSLGVMTNSLLLRYRFFSN 605
Query: 698 ENHKYEKRESTKGSK 712
N K K E +G+K
Sbjct: 606 RNDKNVKPEPKEGTK 620
>At1g63440 ATP dependent copper transporter, putative
Length = 995
Score = 208 bits (529), Expect = 1e-53
Identities = 165/584 (28%), Positives = 283/584 (48%), Gaps = 42/584 (7%)
Query: 156 FTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAE-------TGER-VDVN 207
F ++I + ++LE A K A++ L N+AP AI+ TGE +D
Sbjct: 406 FETSAMLISFIILGKYLEVMAKGKTSQAIAKLMNLAPDTAILLSLDKEGNVTGEEEIDGR 465
Query: 208 DVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVWAGTINMNAY- 266
++ N ++ + G + DG V+ G+ V+E M+TGE+ PV K V GT+N N
Sbjct: 466 LIQKNDVIKIVPGAKVASDGYVIWGQSHVNESMITGEARPVAKRKGDTVIGGTLNENGVL 525
Query: 267 ----PKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*FRKVLYS 322
++ ++ +I + + + K + F VI L
Sbjct: 526 HVKVTRVGSESALAQIVRLVESAQLAKAPVQKLADRISKFFVPLVIFLSFSTWLAWFLAG 585
Query: 323 CIAVVPAALSVPDMEPWFHLALV----VLLSGCPCALILSTPVAIFCALTKAAISGLLLK 378
+ P + +P F LAL V++ CPCAL L+TP A+ A G+L+K
Sbjct: 586 KLHWYPESW-IPSSMDSFELALQFGISVMVIACPCALGLATPTAVMVGTGVGASQGVLIK 644
Query: 379 GGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSAGDDINNETLLYWISSIESKSSHPVAG 438
GG LE ++ + FDKTGT+T G+ V ++ +++ E S HP+A
Sbjct: 645 GGQALERAHKVNCIVFDKTGTLTMGKPVVVKTKLLKNMVLREFYELVAATEVNSEHPLAK 704
Query: 439 ALVDYA---RLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNKR-VGVRAICKRDNC 494
A+V+YA R P +F + G+G+ T+ GR+I +GNK + + D+
Sbjct: 705 AIVEYAKKFRDDEENPAWPEACDFVSITGKGVKATVKGREIMVGNKNLMNDHKVIIPDDA 764
Query: 495 EVQFQRPEISTKKN---NCEETLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQ 551
E E + + L+GV S+ D + A EA+ LK + ++S+M+TGD+
Sbjct: 765 EELLADSEDMAQTGILVSINSELIGVLSVSDPLKPSAREAISILKSMNIKSIMVTGDNWG 824
Query: 552 VAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEG-PIAMIGDGINDAPALATADIGIS 610
A + ++ ID V AE P +KA+ ++ + G +AM+GDGIND+PAL AD+G++
Sbjct: 825 TANSIAREV--GIDSVIAEAKPEQKAEKVKELQAAGHVVAMVGDGINDSPALVAADVGMA 882
Query: 611 MGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAIAG 670
+G +G+ +A E ++ +LM +++ V AI L+RKT ++ N + ++G+ ++ + IA
Sbjct: 883 IG-AGTDIAIEAADIVLMKSNLEDVITAIDLSRKTFSRIRLNYVWALGYN--LMGIPIAA 939
Query: 671 YPLV---------WLAVLTDVGTCLLVILNSMLILQENHKYEKR 705
L W+A + + V+ S+L+ +N+K K+
Sbjct: 940 GVLFPGTRFRLPPWIAGAAMAASSVSVVCCSLLL--KNYKRPKK 981
Score = 34.3 bits (77), Expect = 0.30
Identities = 17/65 (26%), Positives = 33/65 (50%)
Query: 14 GMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADALNTARLEASF 73
GM C + ++ +ER+L+ ++GV+ V + + +D L S ++ + + A EA
Sbjct: 137 GMTCTSCSSTIERVLQSVNGVQRAHVALAIEEAEIHYDPRLSSYDRLLEEIENAGFEAVL 196
Query: 74 RPQGE 78
GE
Sbjct: 197 ISTGE 201
Score = 30.4 bits (67), Expect = 4.3
Identities = 19/68 (27%), Positives = 31/68 (44%)
Query: 5 IKRSNFEVLGMCCATEATLVERILKPLHGVKAVSVIVPARTVTVVHDVLLISESKIADAL 64
I R+ F+VLGM C+ A VE+ +K L G+ + ++ + I + +
Sbjct: 50 ISRAVFQVLGMTCSACAGSVEKAIKRLPGIHDAVIDALNNRAQILFYPNSVDVETIRETI 109
Query: 65 NTARLEAS 72
A EAS
Sbjct: 110 EDAGFEAS 117
>At5g21930 metal-transporting ATPase - like protein
Length = 856
Score = 190 bits (482), Expect = 3e-48
Identities = 173/620 (27%), Positives = 284/620 (44%), Gaps = 66/620 (10%)
Query: 110 GWLALGSVAIGFPKVLFRAIASIRALTLNINILVLLAVCGTAALQ-------------DF 156
G LA+G++ ++LF I + + N+N LV L ++ F
Sbjct: 194 GGLAVGALLGPGRELLFDGIKAFGKRSPNMNSLVGLGSMAAFSISLISLVNPELEWDASF 253
Query: 157 TDGGVIIFLFSI-AQWLETRATHKALVAMSSLTNM-APQMAIVAETGER----------- 203
D V++ F + + LE RA +A M+ L ++ + Q +V + +
Sbjct: 254 FDEPVMLLGFVLLGRSLEERAKLQASTDMNELLSLISTQSRLVITSSDNNTPVDSVLSSD 313
Query: 204 -----VDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESDSLVWA 258
V V+D+++ L V G+ P+DG V+ G+ VDE MLTGES PV KE V A
Sbjct: 314 SICINVSVDDIRVGDSLLVLPGETFPVDGSVLAGRSVVDESMLTGESLPVFKEEGCSVSA 373
Query: 259 GTINMNAYPKIIFDTYDF-----RIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKIH 313
GTIN + +I + +I + + G + F ++S
Sbjct: 374 GTINWDGPLRIKASSTGSNSTISKIVRMVEDAQGNAAPVQRLADAIAGPFVYTIMSLSAM 433
Query: 314 R*FRKVLYSCIAVVPAAL----SVPDMEPW---FHLALVVLLSGCPCALILSTPVAIFCA 366
F Y + P L + PD + LA+ VL+ CPCAL L+TP AI
Sbjct: 434 T-FAFWYYVGSHIFPDVLLNDIAGPDGDALALSLKLAVDVLVVSCPCALGLATPTAILIG 492
Query: 367 LTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSAGDDINNETLLYWIS 426
+ A G L++GGD LE L+ I VA DKTGT+T G V A + +L +
Sbjct: 493 TSLGAKRGYLIRGGDVLERLASIDCVALDKTGTLTEGR-PVVSGVASLGYEEQEVLKMAA 551
Query: 427 SIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGN-KRVGV 485
++E ++HP+A A+V+ A ++K PE PG G IDGR + +G+ + V
Sbjct: 552 AVEKTATHPIAKAIVNEAESLNLK-TPETRGQLTE-PGFGTLAEIDGRFVAVGSLEWVSD 609
Query: 486 RAICKRDNCEVQFQRPEISTKKNNCEET----------------LVGVFSLVDACRSGAL 529
R + K D+ ++ + K +N T ++G ++ D R A
Sbjct: 610 RFLKKNDSSDMVKLESLLDHKLSNTSSTSRYSKTVVYVGREGEGIIGAIAISDCLRQDAE 669
Query: 530 EAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEG-P 588
+ L+ G+++V+L+GD V + + + L P +K + I N + G
Sbjct: 670 FTVARLQEKGIKTVLLSGDREGAVATVAKNVGIKSESTNYSLSPEKKFEFISNLQSSGHR 729
Query: 589 IAMIGDGINDAPALATADIGISMGISGSA-LANETSNAILMSNDIRKVPEAIRLARKTTR 647
+AM+GDGINDAP+LA AD+GI++ I A+ ++ IL+ N + V +A+ LA+ T
Sbjct: 730 VAMVGDGINDAPSLAQADVGIALKIEAQENAASNAASVILVRNKLSHVVDALSLAQATMS 789
Query: 648 KLVENVIISVGFKCAILALA 667
K+ +N+ ++ + + +A
Sbjct: 790 KVYQNLAWAIAYNVISIPIA 809
>At4g37270 Cu2+-transporting ATPase-like protein
Length = 819
Score = 173 bits (439), Expect = 3e-43
Identities = 170/664 (25%), Positives = 301/664 (44%), Gaps = 76/664 (11%)
Query: 92 ACGLLLALSFLKYIYPP--LGWLALGSVAIGFPKV----LFRAIASIRALTLNINILVLL 145
A + LA + Y+ P + L + +GFP V A+ I +NI++L+ L
Sbjct: 132 AAAMFLAAAVCPYLAPEPYIKSLQNAFMIVGFPLVGVSASLDALMDIAGGKVNIHVLMAL 191
Query: 146 AVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAETG---- 201
A + + + +GG+++ +F++A E T +++V + L P A++ E
Sbjct: 192 AAFASVFMGNALEGGLLLAMFNLAHIAEEFFTSRSMVDVKELKESNPDSALLIEVHNGNV 251
Query: 202 --------ERVDVNDVKMNTILAVKAGDAIPLDGIVVEGKCEVDEKMLTGESFPVTKESD 253
+ V V+ V++ + + V G+ +P+D V +G + + LTGE P+ ++
Sbjct: 252 PNISDLSYKSVPVHSVEVGSYVLVGTGEIVPVDCEVYQGSATITIEHLTGEVKPLEAKAG 311
Query: 254 SLVWAGTINMNA--YPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTK 311
V G N++ K D + K + + HS+ ++ F +
Sbjct: 312 DRVPGGARNLDGRMIVKATKAWNDSTLNKIVQLTE-EAHSNKPKLQRWLDEFGENYSKVV 370
Query: 312 IHR*FRKVLYSCIAVVPAAL------SVPDMEPWFHLALVVLLSGCPCALILSTPVAIFC 365
+ VL IA + L S + AL ++++ PCAL ++ P+A
Sbjct: 371 V------VLSLAIAFLGPFLFKWPFLSTAACRGSVYRALGLMVAASPCALAVA-PLAYAT 423
Query: 366 ALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFS------VTDFSAGDDIN-- 417
A++ A G+LLKG L+ L+ T+AFDKTGT+T G + + G + +
Sbjct: 424 AISSCARKGILLKGAQVLDALASCHTIAFDKTGTLTTGGLTCKAIEPIYGHQGGTNSSVI 483
Query: 418 -------NETLLYWISSIESKSSHPVAGALVDYARLHSI-KPVPEN-VENFQNFPGEGIF 468
+ L +++E ++HP+ A+VD HS+ K +P VE+F+ FPG G+
Sbjct: 484 TCCIPNCEKEALAVAAAMEKGTTHPIGRAVVD----HSVGKDLPSIFVESFEYFPGRGLT 539
Query: 469 GTIDGRDIYIGNKRVGVRAICKRDNCEVQFQRPEISTKKNNCE----------------E 512
T++G R+ ++ + F+ + S + + +
Sbjct: 540 ATVNGVKTVAEESRLRKASLGSIEFITSLFKSEDESKQIKDAVNASSYGKDFVHAALSVD 599
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLG-VRSVMLTGDSSQVAMYVQSQLKNAIDIVHAEL 571
V + L D R G + ELK +R +MLTGD A V + + I V+ L
Sbjct: 600 QKVTLIHLEDQPRPGVSGVIAELKSWARLRVMMLTGDHDSSAWRVANAV--GITEVYCNL 657
Query: 572 LPHEKAKLIENFKKE--GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMS 629
P +K ++N +E G + M+G+GINDAPALA A +GI + SA A ++ +L+
Sbjct: 658 KPEDKLNHVKNIAREAGGGLIMVGEGINDAPALAAATVGIVLAQRASATAIAVADILLLR 717
Query: 630 NDIRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVI 689
++I VP + +R+TT + +NV +++ ++ G+ +WL VL G LLV
Sbjct: 718 DNITGVPFCVAKSRQTTSLVKQNVALALTSIFLAALPSVLGFVPLWLTVLLHEGGTLLVC 777
Query: 690 LNSM 693
LNS+
Sbjct: 778 LNSV 781
>At3g21180 putative Ca2+-transporting ATPase
Length = 1090
Score = 76.6 bits (187), Expect = 5e-14
Identities = 52/189 (27%), Positives = 89/189 (46%), Gaps = 31/189 (16%)
Query: 511 EETLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQ---LKNAIDIV 567
E L+ + + D CR G EA+ GV+ M+TGD+ Q A + + L + + V
Sbjct: 689 ELILLAIVGIKDPCRPGVREAVRICTSAGVKVRMVTGDNLQTAKAIALECGILSSDTEAV 748
Query: 568 HAELL---------------------------PHEKAKLIENFKKEGPI-AMIGDGINDA 599
++ P++K L++ +K G + A+ GDG NDA
Sbjct: 749 EPTIIEGKVFRELSEKEREQVAKKITVMGRSSPNDKLLLVQALRKNGDVVAVTGDGTNDA 808
Query: 600 PALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGF 659
PAL ADIG+SMGISG+ +A E+S+ I++ ++ V + +R R + + + +
Sbjct: 809 PALHEADIGLSMGISGTEVAKESSDIIILDDNFASVVKVVRWGRSVYANIQKFIQFQLTV 868
Query: 660 KCAILALAI 668
A L + +
Sbjct: 869 NVAALIINV 877
Score = 44.3 bits (103), Expect = 3e-04
Identities = 31/129 (24%), Positives = 60/129 (46%), Gaps = 11/129 (8%)
Query: 137 LNINILVLLAVCGTAA-------LQDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTN 189
L + IL++ AV A + + DGG I F + + + ++ + +L +
Sbjct: 206 LTLIILIIAAVTSLALGIKTEGLKEGWLDGGSIAFAVLLVIVVTAVSDYRQSLQFQNLND 265
Query: 190 MAPQMAIVAETGER---VDVNDVKMNTILAVKAGDAIPLDGIVVEG-KCEVDEKMLTGES 245
+ + G R + + DV + ++ ++ GD +P DG+++ G +DE +TGES
Sbjct: 266 EKRNIQLEVMRGGRTVKISIYDVVVGDVIPLRIGDQVPADGVLISGHSLAIDESSMTGES 325
Query: 246 FPVTKESDS 254
V K+ S
Sbjct: 326 KIVHKDQKS 334
Score = 36.6 bits (83), Expect = 0.060
Identities = 21/81 (25%), Positives = 39/81 (47%)
Query: 333 VPDMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTV 392
V D F +A+ +++ P L L+ + + ++ K L++ ET+ T+
Sbjct: 433 VDDCVKIFTIAVTIVVVAVPEGLPLAVTLTLAYSMRKMMADKALVRRLSACETMGSATTI 492
Query: 393 AFDKTGTITRGEFSVTDFSAG 413
DKTGT+T + +V + AG
Sbjct: 493 CSDKTGTLTLNQMTVVETYAG 513
>At1g07810 ER-type Ca2+-pump protein
Length = 1061
Score = 76.3 bits (186), Expect = 7e-14
Identities = 58/208 (27%), Positives = 97/208 (45%), Gaps = 41/208 (19%)
Query: 515 VGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVA--------------------- 553
VG L D R +A+ + + G+R +++TGD+ A
Sbjct: 622 VGFVGLRDPPRKEVRQAIADCRTAGIRVMVITGDNKSTAEAICREIGVFEADEDISSRSL 681
Query: 554 -----MYVQSQ---LKNAIDIVHAELLPHEKAKLIENFKKEGPI-AMIGDGINDAPALAT 604
M VQ Q L+ ++ + P K +++ K++G + AM GDG+NDAPAL
Sbjct: 682 TGIEFMDVQDQKNHLRQTGGLLFSRAEPKHKQEIVRLLKEDGEVVAMTGDGVNDAPALKL 741
Query: 605 ADIGISMGISGSALANETSNAILMSNDIRKVPEAI---RLARKTTRKLVENVIIS-VGFK 660
ADIG++MGISG+ +A E S+ +L ++ + A+ R + + +I S +G
Sbjct: 742 ADIGVAMGISGTEVAKEASDMVLADDNFSTIVAAVGEGRSIYNNMKAFIRYMISSNIGEV 801
Query: 661 CAILALAIAGYP-------LVWLAVLTD 681
+I A G P L+W+ ++TD
Sbjct: 802 ASIFLTAALGIPEGMIPVQLLWVNLVTD 829
Score = 42.0 bits (97), Expect = 0.001
Identities = 70/299 (23%), Positives = 118/299 (39%), Gaps = 62/299 (20%)
Query: 161 VIIFLFSIAQ-----WLETRATHKALVAMSSLTNMAPQMAIVAETGERVD---VNDVKMN 212
++IFL I W ET A KAL A+ + + Q A V G +V ++
Sbjct: 117 LVIFLILIVNAIVGIWQETNA-EKALEALKEIQS---QQATVMRDGTKVSSLPAKELVPG 172
Query: 213 TILAVKAGDAIPLDGIVV---EGKCEVDEKMLTGESFPVTKESDS------------LVW 257
I+ ++ GD +P D VV V++ LTGES V+K + +V+
Sbjct: 173 DIVELRVGDKVPADMRVVALISSTLRVEQGSLTGESEAVSKTTKHVDENADIQGKKCMVF 232
Query: 258 AGTINMNAYPK-IIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*F 316
AGT +N ++ DT N +G+ HS ++ + + K++ F
Sbjct: 233 AGTTVVNGNCICLVTDTG-------MNTEIGRVHSQ---IQEAAQHEEDTPLKKKLNE-F 281
Query: 317 RKVLYSCIAVVPAA---------LSVPDMEPW--------------FHLALVVLLSGCPC 353
+VL I ++ A LS ++ W F +A+ + ++ P
Sbjct: 282 GEVLTMIIGLICALVWLINVKYFLSWEYVDGWPRNFKFSFEKCTYYFEIAVALAVAAIPE 341
Query: 354 ALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSA 412
L + K A L++ +ETL + DKTGT+T + +V+ A
Sbjct: 342 GLPAVITTCLALGTRKMAQKNALVRKLPSVETLGCTTVICSDKTGTLTTNQMAVSKLVA 400
>At1g10130 putative calcium ATPase
Length = 992
Score = 75.1 bits (183), Expect = 2e-13
Identities = 58/215 (26%), Positives = 100/215 (45%), Gaps = 42/215 (19%)
Query: 508 NNCEETLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQL---KNAI 564
N + T +G+ ++D R +AM G+R +++TGD+ A + ++ N +
Sbjct: 570 NENDLTFIGLVGMLDPPREEVRDAMLACMTAGIRVIVVTGDNKSTAESLCRKIGAFDNLV 629
Query: 565 DI--------------------------VHAELLPHEKAKLIENFKKEGPI-AMIGDGIN 597
D + + + P K L+E +K+ + AM GDG+N
Sbjct: 630 DFSGMSYTASEFERLPAVQQTLALRRMTLFSRVEPSHKRMLVEALQKQNEVVAMTGDGVN 689
Query: 598 DAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAI---RLARKTTRKLVENVI 654
DAPAL ADIGI+MG SG+A+A S+ +L ++ + A+ R T++ + +I
Sbjct: 690 DAPALKKADIGIAMG-SGTAVAKSASDMVLADDNFASIVAAVAEGRAIYNNTKQFIRYMI 748
Query: 655 IS-VGFKCAILALAIAGYP-------LVWLAVLTD 681
S +G I A+ G P L+W+ ++TD
Sbjct: 749 SSNIGEVVCIFVAAVLGIPDTLAPVQLLWVNLVTD 783
Score = 32.3 bits (72), Expect = 1.1
Identities = 60/288 (20%), Positives = 106/288 (35%), Gaps = 23/288 (7%)
Query: 143 VLLAVCGTAALQDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMAPQMAIVAETG- 201
VL G L F + VI+ + + + A A+ L +A V G
Sbjct: 68 VLALANGETGLTAFLEPFVILLILAANAAVGVITETNAEKALEELRAYQANIATVLRNGC 127
Query: 202 -ERVDVNDVKMNTILAVKAGDAIPLDGIVVE---GKCEVDEKMLTGESFPVTKESDSLVW 257
+ ++ I+ V G IP D ++E VD+ +LTGES V K+ D +
Sbjct: 128 FSILPATELVPGDIVEVTVGCKIPADLRMIEMSSNTFRVDQAILTGESCSVEKDVDCTLT 187
Query: 258 AGTINMNAYPKIIFDTYDFRIYKCKN--YSVGKRHSSGKNVKTC*RSF*QKVISTKIHR* 315
+ + I+F D + + VG + G + ++ + K
Sbjct: 188 TNAVYQDK-KNILFSGTDVVAGRGRAVVIGVGSNTAMGSIHDSMLQTDDEATPLKKKLDE 246
Query: 316 FRKVLYSCIAVVPAALSVPDM----EP-----------WFHLALVVLLSGCPCALILSTP 360
F L IA + + V ++ +P +F +A+ + ++ P L
Sbjct: 247 FGSFLAKVIAGICVLVWVVNIGHFSDPSHGGFFKGAIHYFKIAVALAVAAIPEGLPAVVT 306
Query: 361 VAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVT 408
+ K A +++ +ETL + DKTGT+T SV+
Sbjct: 307 TCLALGTKKMARLNAIVRSLPSVETLGCTTVICSDKTGTLTTNMMSVS 354
>At3g22910 calmodulin-stimulated calcium-ATPase, putative
Length = 1017
Score = 74.7 bits (182), Expect = 2e-13
Identities = 54/211 (25%), Positives = 96/211 (44%), Gaps = 42/211 (19%)
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAM------------------ 554
+L+G+ + D CR G +A+E+ + GV M+TGD+ A
Sbjct: 635 SLLGIIGIKDPCRPGVKKAVEDCQFAGVNIKMITGDNIFTARAIAVECGILTPEDEMNSE 694
Query: 555 ----------YVQSQLKNAID--IVHAELLPHEKAKLIENFKKEG-PIAMIGDGINDAPA 601
Y Q + ++ V A P +K +++ K+ G +A+ GDG NDAPA
Sbjct: 695 AVLEGEKFRNYTQEERLEKVERIKVMARSSPFDKLLMVKCLKELGHVVAVTGDGTNDAPA 754
Query: 602 LATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKL---------VEN 652
L ADIG+SMGI G+ +A E+S+ +++ ++ V ++ R + V
Sbjct: 755 LKEADIGLSMGIQGTEVAKESSDIVILDDNFASVATVLKWGRCVYNNIQKFIQFQLTVNV 814
Query: 653 VIISVGFKCAILA--LAIAGYPLVWLAVLTD 681
+ + F A+ A + + L+W+ ++ D
Sbjct: 815 AALVINFVAAVSAGDVPLTAVQLLWVNLIMD 845
Score = 43.9 bits (102), Expect = 4e-04
Identities = 37/148 (25%), Positives = 70/148 (47%), Gaps = 17/148 (11%)
Query: 138 NINILVLLAVCGTAAL----------QDFTDGGVIIFLFSIAQWLETRATHKALVAMSSL 187
++ IL+LL C T +L + + DGG I + + + + L
Sbjct: 158 DLTILILLG-CATLSLGFGIKEHGLKEGWYDGGSIFVAVFLVVAVSAVSNFRQNRQFDKL 216
Query: 188 TNMAPQMAI-VAETGERVDVN--DVKMNTILAVKAGDAIPLDGIVVEGK-CEVDEKMLTG 243
+ ++ + I V G R +++ D+ + I+ + GD +P DG+ VEG VDE +TG
Sbjct: 217 SKVSSNIKIDVVRNGRRQEISIFDIVVGDIVCLNIGDQVPADGVFVEGHLLHVDESSMTG 276
Query: 244 ES--FPVTKESDSLVWAGTINMNAYPKI 269
ES V+ ++ +++GT + + K+
Sbjct: 277 ESDHVEVSLTGNTFLFSGTKIADGFGKM 304
Score = 33.5 bits (75), Expect = 0.50
Identities = 17/71 (23%), Positives = 34/71 (46%)
Query: 343 ALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITR 402
A+ +++ P L L+ + + ++ + +++ ET+ + DKTGT+T
Sbjct: 397 AVTIIVVAIPEGLPLAVTLTLAYSMKRMMKDNAMVRKLSACETMGSATVICTDKTGTLTL 456
Query: 403 GEFSVTDFSAG 413
+ VTDF G
Sbjct: 457 NQMKVTDFWFG 467
>At2g41560 putative Ca2+-ATPase
Length = 1030
Score = 74.7 bits (182), Expect = 2e-13
Identities = 56/206 (27%), Positives = 95/206 (45%), Gaps = 37/206 (17%)
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQ----LKNAIDI-- 566
T+V V + D R G EA++ + G+ M+TGD+ A + + + + I
Sbjct: 642 TMVAVVGIKDPVRPGVREAVQTCQAAGITVRMVTGDNISTAKAIAKECGIYTEGGLAIEG 701
Query: 567 -------------------VHAELLPHEKAKLIENFKKEGP-IAMIGDGINDAPALATAD 606
V A LP +K L+ N +K G +A+ GDG NDAPAL AD
Sbjct: 702 SEFRDLSPHEMRAIIPKIQVMARSLPLDKHTLVSNLRKIGEVVAVTGDGTNDAPALHEAD 761
Query: 607 IGISMGISGSALANETSNAILMSNDIRKVPEAIRLARK---TTRKLVE--------NVII 655
IG++MGI+G+ +A E ++ I+M ++ + + R R +K V+ +II
Sbjct: 762 IGLAMGIAGTEVAKENADVIIMDDNFKTIVNVARWGRAVYINIQKFVQFQLTVNVVALII 821
Query: 656 SVGFKCAILALAIAGYPLVWLAVLTD 681
+ C + + L+W+ ++ D
Sbjct: 822 NFVSACITGSAPLTAVQLLWVNMIMD 847
Score = 48.1 bits (113), Expect = 2e-05
Identities = 67/306 (21%), Positives = 119/306 (37%), Gaps = 53/306 (17%)
Query: 142 LVLLAVCGTAAL----------QDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMA 191
L++L VC ++ + DG I+ + + + +K + L
Sbjct: 171 LIILMVCAVVSIGVGVATEGFPRGMYDGTGILLSILLVVMVTAISDYKQSLQFRDLDREK 230
Query: 192 PQMAI-VAETGER--VDVNDVKMNTILAVKAGDAIPLDGIVVEG-KCEVDEKMLTGESFP 247
++ + V G R + ++D+ + ++ + GD +P DGI + G E+DE L+GES P
Sbjct: 231 KKIIVQVTRDGSRQEISIHDLVVGDVVHLSIGDQVPADGIFISGYNLEIDESSLSGESEP 290
Query: 248 --VTKESDSLVWAGTINMNAYPKIIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*Q 305
V KE L+ +GT N K++ T VG R GK ++T
Sbjct: 291 SHVNKEKPFLL-SGTKVQNGSAKMLVTT------------VGMRTEWGKLMETLVDGGED 337
Query: 306 K-----------VISTKIHR*FRKVLY--SCIAVVPAALSVPDMEPW-----------FH 341
+ I KI F + + CI V + W F
Sbjct: 338 ETPLQVKLNGVATIIGKIGLSFAVLTFVVLCIRFVLDKATSGSFTNWSSEDALTLLDYFA 397
Query: 342 LALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTIT 401
+++ +++ P L L+ +++ A+ K L++ ET+ + DKTGT+T
Sbjct: 398 ISVTIIVVAVPEGLPLAVTLSLAFAMKKLMSDRALVRHLAACETMGSSTCICTDKTGTLT 457
Query: 402 RGEFSV 407
V
Sbjct: 458 TNHMVV 463
>At1g07670 endoplasmic reticulum-type calcium-transporting ATPase 4
(ECA4)
Length = 1061
Score = 74.7 bits (182), Expect = 2e-13
Identities = 54/208 (25%), Positives = 96/208 (45%), Gaps = 41/208 (19%)
Query: 515 VGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYV------------------ 556
VG L D R +A+ + + G+R +++TGD+ A +
Sbjct: 622 VGFVGLRDPPRKEVRQAIADCRTAGIRVMVITGDNKSTAEAICREIGVFEADEDISSRSL 681
Query: 557 -----------QSQLKNAIDIVHAELLPHEKAKLIENFKKEGPI-AMIGDGINDAPALAT 604
++ L+ ++ + P K +++ K++G + AM GDG+NDAPAL
Sbjct: 682 TGKEFMDVKDQKNHLRQTGGLLFSRAEPKHKQEIVRLLKEDGEVVAMTGDGVNDAPALKL 741
Query: 605 ADIGISMGISGSALANETSNAILMSNDIRKVPEAI---RLARKTTRKLVENVIIS-VGFK 660
ADIG++MGISG+ +A E S+ +L ++ + A+ R + + +I S +G
Sbjct: 742 ADIGVAMGISGTEVAKEASDLVLADDNFSTIVAAVGEGRSIYNNMKAFIRYMISSNIGEV 801
Query: 661 CAILALAIAGYP-------LVWLAVLTD 681
+I A G P L+W+ ++TD
Sbjct: 802 ASIFLTAALGIPEGMIPVQLLWVNLVTD 829
Score = 42.0 bits (97), Expect = 0.001
Identities = 70/299 (23%), Positives = 118/299 (39%), Gaps = 62/299 (20%)
Query: 161 VIIFLFSIAQ-----WLETRATHKALVAMSSLTNMAPQMAIVAETGERVD---VNDVKMN 212
++IFL I W ET A KAL A+ + + Q A V G +V ++
Sbjct: 117 LVIFLILIVNAIVGIWQETNA-EKALEALKEIQS---QQATVMRDGTKVSSLPAKELVPG 172
Query: 213 TILAVKAGDAIPLDGIVV---EGKCEVDEKMLTGESFPVTKESDS------------LVW 257
I+ ++ GD +P D VV V++ LTGES V+K + +V+
Sbjct: 173 DIVELRVGDKVPADMRVVALISSTLRVEQGSLTGESEAVSKTTKHVDENADIQGKKCMVF 232
Query: 258 AGTINMNAYPK-IIFDTYDFRIYKCKNYSVGKRHSSGKNVKTC*RSF*QKVISTKIHR*F 316
AGT +N ++ DT N +G+ HS ++ + + K++ F
Sbjct: 233 AGTTVVNGNCICLVTDTG-------MNTEIGRVHSQ---IQEAAQHEEDTPLKKKLNE-F 281
Query: 317 RKVLYSCIAVVPAA---------LSVPDMEPW--------------FHLALVVLLSGCPC 353
+VL I ++ A LS ++ W F +A+ + ++ P
Sbjct: 282 GEVLTMIIGLICALVWLINVKYFLSWEYVDGWPRNFKFSFEKCTYYFEIAVALAVAAIPE 341
Query: 354 ALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSVTDFSA 412
L + K A L++ +ETL + DKTGT+T + +V+ A
Sbjct: 342 GLPAVITTCLALGTRKMAQKNALVRKLPSVETLGCTTVICSDKTGTLTTNQMAVSKLVA 400
>At5g57110 Ca2+-transporting ATPase-like protein (emb|CAB79748.1)
Length = 1074
Score = 73.2 bits (178), Expect = 6e-13
Identities = 47/186 (25%), Positives = 91/186 (48%), Gaps = 31/186 (16%)
Query: 514 LVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQ---LKNAIDIVHAE 570
L+ + + D CR G +++ + GV+ M+TGD+ Q A + + L + D+
Sbjct: 674 LLAIVGIKDPCRPGVKDSVVLCQNAGVKVRMVTGDNVQTARAIALECGILSSDADLSEPT 733
Query: 571 LL---------------------------PHEKAKLIENFKKEGPI-AMIGDGINDAPAL 602
L+ P++K L+++ +++G + A+ GDG NDAPAL
Sbjct: 734 LIEGKSFREMTDAERDKISDKISVMGRSSPNDKLLLVQSLRRQGHVVAVTGDGTNDAPAL 793
Query: 603 ATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCA 662
ADIG++MGI+G+ +A E+S+ I++ ++ V + +R R + + + + A
Sbjct: 794 HEADIGLAMGIAGTEVAKESSDIIILDDNFASVVKVVRWGRSVYANIQKFIQFQLTVNVA 853
Query: 663 ILALAI 668
L + +
Sbjct: 854 ALVINV 859
Score = 35.0 bits (79), Expect = 0.17
Identities = 22/97 (22%), Positives = 47/97 (47%), Gaps = 1/97 (1%)
Query: 333 VPDMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTV 392
+ D+ +A+ +++ P L L+ + + ++ K L++ ET+ T+
Sbjct: 420 IDDVVKVLTVAVTIVVVAVPEGLPLAVTLTLAYSMRKMMADKALVRRLSACETMGSATTI 479
Query: 393 AFDKTGTITRGEFSVTD-FSAGDDINNETLLYWISSI 428
DKTGT+T + +V + ++ G + E L I+S+
Sbjct: 480 CSDKTGTLTLNQMTVVESYAGGKKTDTEQLPATITSL 516
>At3g57330 Ca2+-transporting ATPase-like protein
Length = 1025
Score = 73.2 bits (178), Expect = 6e-13
Identities = 56/206 (27%), Positives = 93/206 (44%), Gaps = 37/206 (17%)
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQ----LKNAIDI-- 566
TLV V + D R G EA++ + G+ M+TGD+ A + + + I
Sbjct: 639 TLVAVVGIKDPVRPGVREAVQTCQAAGITVRMVTGDNISTAKAIAKECGILTAGGVAIEG 698
Query: 567 -------------------VHAELLPHEKAKLIENFKKEGP-IAMIGDGINDAPALATAD 606
V A LP +K L+ N +K G +A+ GDG NDAPAL AD
Sbjct: 699 SDFRNLPPHEMRAILPKIQVMARSLPLDKHTLVNNLRKMGEVVAVTGDGTNDAPALHEAD 758
Query: 607 IGISMGISGSALANETSNAILMSNDIRKVPEAIRLARK---TTRKLVE--------NVII 655
IG++MGI+G+ +A E ++ I+M ++ + + R +K V+ +II
Sbjct: 759 IGLAMGIAGTEVAKENADVIIMDDNFATIVNVAKWGRAVYINIQKFVQFQLTVNVVALII 818
Query: 656 SVGFKCAILALAIAGYPLVWLAVLTD 681
+ C + + L+W+ ++ D
Sbjct: 819 NFVSACITGSAPLTAVQLLWVNMIMD 844
Score = 52.4 bits (124), Expect = 1e-06
Identities = 69/298 (23%), Positives = 119/298 (39%), Gaps = 37/298 (12%)
Query: 142 LVLLAVCGTAAL----------QDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMA 191
L++L VC ++ + DG I+ + + + +K + L
Sbjct: 171 LIILMVCAVVSIGVGVATEGFPKGMYDGTGILLSIILVVMVTAISDYKQSLQFRDLDREK 230
Query: 192 PQMAI-VAETGER--VDVNDVKMNTILAVKAGDAIPLDGIVVEG-KCEVDEKMLTGESFP 247
++ I V G R V ++D+ + ++ + GD +P DGI + G E+DE L+GES P
Sbjct: 231 KKIIIQVTRDGSRQEVSIHDLVVGDVVHLSIGDQVPADGIFISGYNLEIDESSLSGESEP 290
Query: 248 --VTKESDSLVWAGTINMNAYPKIIFDTYDFRIYKCK---NYSVGKRHSSGKNVKTC*RS 302
V KE L+ +GT N K++ T R K S G + VK +
Sbjct: 291 SHVNKEKPFLL-SGTKVQNGSAKMLVTTVGMRTEWGKLMDTLSEGGEDETPLQVKLNGVA 349
Query: 303 F*QKVISTKIHR*FRKVLY--SCIAVVPAALSVPDMEPW-----------FHLALVVLLS 349
I KI F + + CI V + + W F +A+ +++
Sbjct: 350 ----TIIGKIGLGFAVLTFVVLCIRFVVEKATAGSITEWSSEDALTLLDYFAIAVTIIVV 405
Query: 350 GCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSV 407
P L L+ +++ A+ + L++ ET+ + DKTGT+T V
Sbjct: 406 AVPEGLPLAVTLSLAFAMKQLMSDRALVRHLAACETMGSSTCICTDKTGTLTTNHMVV 463
>At4g00900 Ca2+-transporting ATPase - like protein
Length = 1054
Score = 72.8 bits (177), Expect = 7e-13
Identities = 61/219 (27%), Positives = 99/219 (44%), Gaps = 45/219 (20%)
Query: 508 NNCEETL--VGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLK---- 561
+N E L VGV L D R A+E+ + G+R +++TGD+ A + +++
Sbjct: 607 SNIETNLIFVGVVGLRDPPREEVGRAIEDCRDAGIRVMVITGDNKSTAEAICCEIRLFSE 666
Query: 562 ---------------------------NAIDIVHAELLPHEKAKLIENFKKEGPI-AMIG 593
+ V + P K +++ K+ G I AM G
Sbjct: 667 NEDLSQSSFTGKEFMSLPASRRSEILSKSGGKVFSRAEPRHKQEIVRMLKEMGEIVAMTG 726
Query: 594 DGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLAR---KTTRKLV 650
DG+NDAPAL ADIGI+MGI+G+ +A E S+ +L ++ + A+ R + +
Sbjct: 727 DGVNDAPALKLADIGIAMGITGTEVAKEASDMVLADDNFSTIVSAVAEGRSIYNNMKAFI 786
Query: 651 ENVIIS-VGFKCAILALAIAGYP-------LVWLAVLTD 681
+I S VG +I A G P L+W+ ++TD
Sbjct: 787 RYMISSNVGEVISIFLTAALGIPECMIPVQLLWVNLVTD 825
Score = 38.1 bits (87), Expect = 0.020
Identities = 24/105 (22%), Positives = 46/105 (42%), Gaps = 3/105 (2%)
Query: 339 WFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTG 398
+F +A+ + ++ P L + K A +++ +ETL + DKTG
Sbjct: 312 YFKIAVALAVAAIPEGLPAVITTCLALGTRKMAQKNAIVRKLPSVETLGCTTVICSDKTG 371
Query: 399 TITRGEFSVTDFSAGDDINNETLLYWISSIESKSSHPVAGALVDY 443
T+T + S T+F + +T + S+ + P G +VD+
Sbjct: 372 TLTTNQMSATEFFT---LGGKTTTTRVFSVSGTTYDPKDGGIVDW 413
>At4g29900 Ca2+-transporting ATPase - like protein
Length = 1069
Score = 70.5 bits (171), Expect = 4e-12
Identities = 48/186 (25%), Positives = 86/186 (45%), Gaps = 31/186 (16%)
Query: 514 LVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQ---LKNAIDIVHAE 570
L+ + + D CR G ++ + GV+ M+TGD+ Q A + + L + D
Sbjct: 678 LLAIVGIKDPCRPGVKNSVLLCQQAGVKVRMVTGDNIQTAKAIALECGILASDSDASEPN 737
Query: 571 LL---------------------------PHEKAKLIENFKKEGPI-AMIGDGINDAPAL 602
L+ P++K L+++ K+ G + A+ GDG NDAPAL
Sbjct: 738 LIEGKVFRSYSEEERDRICEEISVMGRSSPNDKLLLVQSLKRRGHVVAVTGDGTNDAPAL 797
Query: 603 ATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCA 662
ADIG++MGI G+ +A E S+ I++ ++ V + +R R + + + + A
Sbjct: 798 HEADIGLAMGIQGTEVAKEKSDIIILDDNFESVVKVVRWGRSVYANIQKFIQFQLTVNVA 857
Query: 663 ILALAI 668
L + +
Sbjct: 858 ALVINV 863
Score = 41.2 bits (95), Expect = 0.002
Identities = 22/59 (37%), Positives = 34/59 (57%), Gaps = 3/59 (5%)
Query: 197 VAETGERVDVN--DVKMNTILAVKAGDAIPLDGIVVEG-KCEVDEKMLTGESFPVTKES 252
V G RV+++ D+ + ++ + GD +P DG++V G VDE +TGES V K S
Sbjct: 260 VTRDGRRVEISIYDIVVGDVIPLNIGDQVPADGVLVAGHSLAVDESSMTGESKIVQKNS 318
Score = 37.4 bits (85), Expect = 0.035
Identities = 21/79 (26%), Positives = 39/79 (48%)
Query: 335 DMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAF 394
D+ F +A+ +++ P L L+ + + ++ K L++ ET+ T+
Sbjct: 422 DLVEIFTVAVTIVVVAVPEGLPLAVTLTLAYSMRKMMADKALVRRLSACETMGSATTICS 481
Query: 395 DKTGTITRGEFSVTDFSAG 413
DKTGT+T E +V + AG
Sbjct: 482 DKTGTLTLNEMTVVECYAG 500
>At4g37640 plasma membrane-type calcium ATPase (ACA2)
Length = 1014
Score = 66.6 bits (161), Expect = 5e-11
Identities = 54/207 (26%), Positives = 97/207 (46%), Gaps = 38/207 (18%)
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQL------------ 560
T VG+ + D R G E++E + G+ M+TGD+ A + +
Sbjct: 647 TCVGIVGIKDPVRPGVKESVELCRRAGITVRMVTGDNINTAKAIARECGILTDDGIAIEG 706
Query: 561 -----KNAIDI--------VHAELLPHEKAKLIENFKK--EGPIAMIGDGINDAPALATA 605
KN ++ V A P +K L++ + + +A+ GDG NDAPAL A
Sbjct: 707 PVFREKNQEELLELIPKIQVMARSSPMDKHTLVKQLRTTFDEVVAVTGDGTNDAPALHEA 766
Query: 606 DIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARK---TTRKLVE-----NVI-IS 656
DIG++MGI+G+ +A E+++ I++ ++ + + R +K V+ NV+ +
Sbjct: 767 DIGLAMGIAGTEVAKESADVIILDDNFSTIVTVAKWGRSVYINIQKFVQFQLTVNVVALV 826
Query: 657 VGFKCAIL--ALAIAGYPLVWLAVLTD 681
V F A L + + L+W+ ++ D
Sbjct: 827 VNFSSACLTGSAPLTAVQLLWVNMIMD 853
Score = 43.9 bits (102), Expect = 4e-04
Identities = 48/220 (21%), Positives = 91/220 (40%), Gaps = 14/220 (6%)
Query: 202 ERVDVNDVKMNTILAVKAGDAIPLDGIVVEG-KCEVDEKMLTGESFPV-TKESDSLVWAG 259
+++ + D+ I+ + GD +P DG+ + G +DE LTGES PV + + +G
Sbjct: 247 QKLSIYDLLPGDIVHLAIGDQVPADGLFLSGFSVVIDESSLTGESEPVMVNAQNPFLMSG 306
Query: 260 TINMNAYPKIIFDTYDFRIYKCK---NYSVGKRHSSGKNVK---------TC*RSF*QKV 307
T + K++ T R K + G + VK F
Sbjct: 307 TKVQDGSCKMMITTVGMRTQWGKLMATLTEGGDDETPLQVKLNGVATIIGKIGLFFAVVT 366
Query: 308 ISTKIHR*FRKVLYSCIAVVPAALSVPDMEPWFHLALVVLLSGCPCALILSTPVAIFCAL 367
+ + F + L + V + ++ +F +A+ +++ P L L+ +++ A+
Sbjct: 367 FAVLVQGMFMRKLSTGTHWVWSGDEALELLEYFAIAVTIVVVAVPEGLPLAVTLSLAFAM 426
Query: 368 TKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSV 407
K L++ ET+ T+ DKTGT+T +V
Sbjct: 427 KKMMNDKALVRHLAACETMGSATTICSDKTGTLTTNHMTV 466
>At2g22950 pseudogene
Length = 1015
Score = 65.1 bits (157), Expect = 2e-10
Identities = 53/207 (25%), Positives = 96/207 (45%), Gaps = 38/207 (18%)
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQL------------ 560
T +G+ + D R G E++E + G+ M+TGD+ A + +
Sbjct: 648 TCIGIVGIKDPVRPGVRESVELCRRAGIMVRMVTGDNINTAKAIARECGILTDDGIAIEG 707
Query: 561 -----KNAIDI--------VHAELLPHEKAKLIENFKK--EGPIAMIGDGINDAPALATA 605
KN ++ V A P +K L++ + + +A+ GDG NDAPAL A
Sbjct: 708 PVFREKNQEEMLELIPKIQVMARSSPMDKHTLVKQLRTTFDEVVAVTGDGTNDAPALHEA 767
Query: 606 DIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARK---TTRKLVE-----NVI-IS 656
DIG++MGI+G+ +A E ++ I++ ++ + + R +K V+ NV+ +
Sbjct: 768 DIGLAMGIAGTEVAKEIADVIILDDNFSTIVTVAKWGRSVYINIQKFVQFQLTVNVVALI 827
Query: 657 VGFKCAIL--ALAIAGYPLVWLAVLTD 681
V F A L + + L+W+ ++ D
Sbjct: 828 VNFSSACLTGSAPLTAVQLLWVNMIMD 854
Score = 43.1 bits (100), Expect = 6e-04
Identities = 63/297 (21%), Positives = 121/297 (40%), Gaps = 35/297 (11%)
Query: 142 LVLLAVCGTAAL----------QDFTDGGVIIFLFSIAQWLETRATHKALVAMSSLTNMA 191
L++L VC +L Q DG I+ + ++ + ++ + L
Sbjct: 175 LMILGVCAFVSLIVGIATEGWPQGSHDGLGIVASILLVVFVTATSDYRQSLQFRDLDKEK 234
Query: 192 PQMAI-VAETG--ERVDVNDVKMNTILAVKAGDAIPLDGIVVEG-KCEVDEKMLTGESFP 247
++ + V G +++ + D+ ++ + GD +P DG+ + G +DE LTGES P
Sbjct: 235 KKITVQVTRNGFRQKMSIYDLLPGDVVHLAIGDQVPADGLFLSGFSVVIDESSLTGESEP 294
Query: 248 V-TKESDSLVWAGTINMNAYPKIIFDTYDFRIYKCK---NYSVGKRHSSGKNVKTC*RSF 303
V + + +GT + K++ T R K S G + VK +
Sbjct: 295 VMVTAQNPFLLSGTKVQDGSCKMLVTTVGMRTQWGKLMATLSEGGDDETPLQVKLNGVA- 353
Query: 304 *QKVISTKIHR*FRKVLYSCI--AVVPAALSVP-----------DMEPWFHLALVVLLSG 350
I KI F V ++ + + LS+ ++ +F +A+ +++
Sbjct: 354 ---TIIGKIGLSFAIVTFAVLVQGMFMRKLSLGPHWWWSGDDALELLEYFAIAVTIVVVA 410
Query: 351 CPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRGEFSV 407
P L L+ +++ A+ K L++ ET+ T+ DKTGT+T +V
Sbjct: 411 VPEGLPLAVTLSLAFAMKKMMNDKALVRHLAACETMGSATTICSDKTGTLTTNHMTV 467
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.137 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,132,191
Number of Sequences: 26719
Number of extensions: 703235
Number of successful extensions: 2031
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 39
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 1821
Number of HSP's gapped (non-prelim): 171
length of query: 833
length of database: 11,318,596
effective HSP length: 108
effective length of query: 725
effective length of database: 8,432,944
effective search space: 6113884400
effective search space used: 6113884400
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC130275.7


