

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
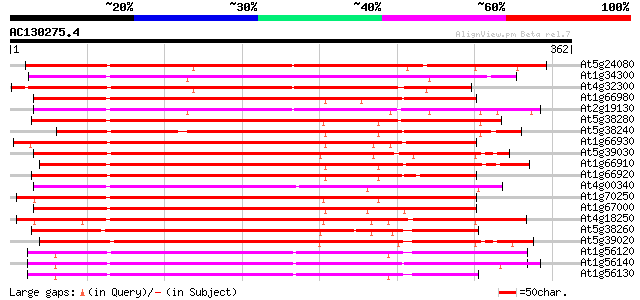
Query= AC130275.4 - phase: 0
(362 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At5g24080 receptor-like protein kinase 263 1e-70
At1g34300 putative protein 254 7e-68
At4g32300 S-receptor kinase -like protein 247 6e-66
At1g66980 putative kinase 244 4e-65
At2g19130 putative receptor-like protein kinase 243 9e-65
At5g38280 receptor serine/threonine kinase PR5K 241 6e-64
At5g38240 receptor serine/threonine protein kinase -like 239 1e-63
At1g66930 receptor serine/threonine kinase PR5K, putative 239 1e-63
At5g39030 receptor protein kinase - like protein 239 2e-63
At1g66910 receptor serine/threonine kinase PR5K, putative 238 4e-63
At1g66920 receptor serine/threonine kinase PR5K, putative 236 1e-62
At4g00340 receptor-like protein kinase 236 1e-62
At1g70250 hypothetical protein 235 3e-62
At1g67000 hypothetical protein 234 4e-62
At4g18250 receptor serine/threonine kinase-like protein 233 9e-62
At5g38260 receptor serine/threonine protein kinase - like 232 2e-61
At5g39020 receptor protein kinase - like protein 227 9e-60
At1g56120 receptor-like protein kinase, putative 225 3e-59
At1g56140 receptor-like protein kinase, putative 222 3e-58
At1g56130 receptor-like protein kinase, putative 219 2e-57
>At5g24080 receptor-like protein kinase
Length = 872
Score = 263 bits (672), Expect = 1e-70
Identities = 158/355 (44%), Positives = 217/355 (60%), Gaps = 23/355 (6%)
Query: 11 EKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMA 70
+ P+ FT + L+ T+N+S LLGSGGFGTVYKG + T+VAVK L + + + +F+
Sbjct: 515 DSPVSFTYRDLQNCTNNFSQLLGSGGFGTVYKGTVAGETLVAVKRLDRALSHG-EREFIT 573
Query: 71 EVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETK---VLGYEKLHEI 127
EV TIG +HH NLVRL G+C E + LVYEYM NGSLD+++F + +L + EI
Sbjct: 574 EVNTIGSMHHMNLVRLCGYCSEDSHRLLVYEYMINGSLDKWIFSSEQTANLLDWRTRFEI 633
Query: 128 AIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGG 187
A+ TA+GIAY HE+C++RIIH DIKP NILLD NF PKV+DFGLAK RE++H+ +T
Sbjct: 634 AVATAQGIAYFHEQCRNRIIHCDIKPENILLDDNFCPKVSDFGLAKMMGREHSHV-VTMI 692
Query: 188 RGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKD 247
RGT GY APE PIT K DVYS+GMLL EI+G RRNL + ++P W +K+
Sbjct: 693 RGTRGYLAPEWVSNRPITVKADVYSYGMLLLEIVGGRRNLDMSYDAEDFFYPGWAYKELT 752
Query: 248 AGLLGEAM--IVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGS---LEI 302
G +A+ + G+ E+ + + + +KVA WC+Q +RP M VVK+LEG+ + +
Sbjct: 753 NGTSLKAVDKRLQGVAEEEEVV--KALKVAFWCIQDEVSMRPSMGEVVKLLEGTSDEINL 810
Query: 303 PKTFNPFQHLI-DGTEFTTHSVQES----------NTYTTSVSSVMVSDSSIVCA 346
P LI +G E +++ NT TTS S S S C+
Sbjct: 811 PPMPQTILELIEEGLEDVYRAMRREFNNQLSSLTVNTITTSQSYRSSSRSHATCS 865
>At1g34300 putative protein
Length = 829
Score = 254 bits (648), Expect = 7e-68
Identities = 139/319 (43%), Positives = 193/319 (59%), Gaps = 8/319 (2%)
Query: 13 PIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEV 72
P++FT ++L+ T ++ LG+GGFGTVY+G+ +N T+VAVK L G ++QF EV
Sbjct: 471 PVQFTYKELQRCTKSFKEKLGAGGFGTVYRGVLTNRTVVAVKQLEGIEQG--EKQFRMEV 528
Query: 73 GTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF--HETKVLGYEKLHEIAIG 130
TI HH NLVRL GFC + LVYE+M NGSLD +LF K L +E IA+G
Sbjct: 529 ATISSTHHLNLVRLIGFCSQGRHRLLVYEFMRNGSLDNFLFTTDSAKFLTWEYRFNIALG 588
Query: 131 TARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGT 190
TA+GI YLHEEC+ I+H DIKP NIL+D NF KV+DFGLAK N ++ M+ RGT
Sbjct: 589 TAKGITYLHEECRDCIVHCDIKPENILVDDNFAAKVSDFGLAKLLNPKDNRYNMSSVRGT 648
Query: 191 PGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDAGL 250
GY APE PIT K DVYS+GM+L E++ +RN + + + F IW +++ + G
Sbjct: 649 RGYLAPEWLANLPITSKSDVYSYGMVLLELVSGKRNFDVSEKTNHKKFSIWAYEEFEKGN 708
Query: 251 LGEAMIVCGIEEKNKEIAE--RMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIPKTFNP 308
+ E++ ++ + RM+K + WC+Q +P RP M VV+MLEG EI P
Sbjct: 709 TKAILDTRLSEDQTVDMEQVMRMVKTSFWCIQEQPLQRPTMGKVVQMLEGITEIKNPLCP 768
Query: 309 FQHLIDGTEFTTHSVQESN 327
I F+ +S+ S+
Sbjct: 769 --KTISEVSFSGNSMSTSH 785
>At4g32300 S-receptor kinase -like protein
Length = 821
Score = 247 bits (631), Expect = 6e-66
Identities = 141/304 (46%), Positives = 191/304 (62%), Gaps = 14/304 (4%)
Query: 2 DKFLNDMEREKPIRFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSN 61
D FL ++ PIRF + L+ AT+N+S LG GGFG+VY+G +G+ +AVK L G
Sbjct: 470 DNFLENLSG-MPIRFAYKDLQSATNNFSVKLGQGGFGSVYEGTLPDGSRLAVKKLEGIGQ 528
Query: 62 KKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETK---V 118
K ++F AEV IG IHH +LVRL GFC E L YE++ GSL+R++F + +
Sbjct: 529 GK--KEFRAEVSIIGSIHHLHLVRLRGFCAEGAHRLLAYEFLSKGSLERWIFRKKDGDVL 586
Query: 119 LGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRE 178
L ++ IA+GTA+G+AYLHE+C RI+H DIKP NILLD NF KV+DFGLAK RE
Sbjct: 587 LDWDTRFNIALGTAKGLAYLHEDCDARIVHCDIKPENILLDDNFNAKVSDFGLAKLMTRE 646
Query: 179 NTHITMTGGRGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWF 238
+H+ T RGT GY APE + I+ K DVYS+GM+L E+IG R+N T + F
Sbjct: 647 QSHV-FTTMRGTRGYLAPEWITNYAISEKSDVYSYGMVLLELIGGRKNYDPSETSEKCHF 705
Query: 239 PIWVWKKKDAGLLGEAMIVCGIEEKNKEI----AERMIKVALWCVQYRPELRPIMSVVVK 294
P + +KK + G L M + + KN ++ +R +K ALWC+Q + RP MS VV+
Sbjct: 706 PSFAFKKMEEGKL---MDIVDGKMKNVDVTDERVQRAMKTALWCIQEDMQTRPSMSKVVQ 762
Query: 295 MLEG 298
MLEG
Sbjct: 763 MLEG 766
>At1g66980 putative kinase
Length = 1109
Score = 244 bits (624), Expect = 4e-65
Identities = 130/293 (44%), Positives = 189/293 (64%), Gaps = 10/293 (3%)
Query: 16 FTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTI 75
+T Q++ T +++ ++G GGFG VYKG S+G +VAVKVL+ + E F+ EV T+
Sbjct: 786 YTYAQVKRITKSFAEVVGRGGFGIVYKGTLSDGRVVAVKVLKDTKGN--GEDFINEVATM 843
Query: 76 GRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV-LGYEKLHEIAIGTARG 134
R H N+V L GFC E + A++YE++ NGSLD+++ +T V + + L+ IA+G A G
Sbjct: 844 SRTSHLNIVSLLGFCSEGSKRAIIYEFLENGSLDKFILGKTSVNMDWTALYRIALGVAHG 903
Query: 135 IAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYA 194
+ YLH C+ RI+H+DIKP N+LLD +F PKV+DFGLAK C ++ + ++M RGT GY
Sbjct: 904 LEYLHHSCKTRIVHFDIKPQNVLLDDSFCPKVSDFGLAKLCEKKESILSMLDTRGTIGYI 963
Query: 195 APELWMPF--PITHKCDVYSFGMLLFEIIGRRR----NLAIKNTESQEWFPIWVWKKKDA 248
APE+ ++HK DVYS+GML+ EIIG R N A + S +FP WV++ ++
Sbjct: 964 APEMISRVYGNVSHKSDVYSYGMLVLEIIGARNKEKANQACASNTSSMYFPEWVYRDLES 1023
Query: 249 GLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLE 301
G I GI + E+A++M V LWC+Q P RP M+ VV+M+EGSLE
Sbjct: 1024 CKSGR-HIEDGINSEEDELAKKMTLVGLWCIQPSPVDRPAMNRVVEMMEGSLE 1075
>At2g19130 putative receptor-like protein kinase
Length = 828
Score = 243 bits (621), Expect = 9e-65
Identities = 153/349 (43%), Positives = 209/349 (59%), Gaps = 27/349 (7%)
Query: 16 FTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTI 75
F+ ++L+ AT N+S+ LG GGFG+V+KG + + +AVK L G S ++QF EV TI
Sbjct: 483 FSYRELQNATKNFSDKLGGGGFGSVFKGALPDSSDIAVKRLEGISQG--EKQFRTEVVTI 540
Query: 76 GRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLF----HETKVLGYEKLHEIAIGT 131
G I H NLVRL GFC E + LVY+YM NGSLD +LF E VLG++ +IA+GT
Sbjct: 541 GTIQHVNLVRLRGFCSEGSKKLLVYDYMPNGSLDSHLFLNQVEEKIVLGWKLRFQIALGT 600
Query: 132 ARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTP 191
ARG+AYLH+EC+ IIH DIKP NILLD F PKVADFGLAK R+ + + +T RGT
Sbjct: 601 ARGLAYLHDECRDCIIHCDIKPENILLDSQFCPKVADFGLAKLVGRDFSRV-LTTMRGTR 659
Query: 192 GYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWK--KKDAG 249
GY APE IT K DVYS+GM+LFE++ RRN E +FP W KD
Sbjct: 660 GYLAPEWISGVAITAKADVYSYGMMLFELVSGRRNTEQSENEKVRFFPSWAATILTKDGD 719
Query: 250 LLGEAMIVCGIEEKNKEIAE--RMIKVALWCVQYRPELRPIMSVVVKMLEGSLEI--PKT 305
+ +++ +E +I E R KVA WC+Q RP MS VV++LEG LE+ P
Sbjct: 720 I--RSLVDPRLEGDAVDIEEVTRACKVACWCIQDEESHRPAMSQVVQILEGVLEVNPPPF 777
Query: 306 FNPFQHLI----------DGTEFTTHSVQESNTYTTSVSS--VMVSDSS 342
Q L+ + + ++H+ +++ +++S SS M +D+S
Sbjct: 778 PRSIQALVVSDEDVVFFTESSSSSSHNSSQNHKHSSSSSSSKKMTNDNS 826
>At5g38280 receptor serine/threonine kinase PR5K
Length = 665
Score = 241 bits (614), Expect = 6e-64
Identities = 128/316 (40%), Positives = 205/316 (64%), Gaps = 16/316 (5%)
Query: 15 RFTGQQLRIATDNYSNLLGSGGFGTVYKG-IFSNGTMVAVKVLRGSSNKKIDEQFMAEVG 73
R++ +++ T++++++LG GGFGTVYKG + +G VAVK+L+ S E+F+ EV
Sbjct: 320 RYSYTRVKKMTNSFAHVLGKGGFGTVYKGKLADSGRDVAVKILKVSEGN--GEEFINEVA 377
Query: 74 TIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV-LGYEKLHEIAIGTA 132
++ R H N+V L GFC+E+N A++YE+M NGSLD+Y+ + +E+L+++A+G +
Sbjct: 378 SMSRTSHVNIVSLLGFCYEKNKRAIIYEFMPNGSLDKYISANMSTKMEWERLYDVAVGIS 437
Query: 133 RGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPG 192
RG+ YLH C RI+H+DIKP NIL+D+N PK++DFGLAK C + + I+M RGT G
Sbjct: 438 RGLEYLHNRCVTRIVHFDIKPQNILMDENLCPKISDFGLAKLCKNKESIISMLHMRGTFG 497
Query: 193 YAAPELWMP--FPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQE---WFPIWVWKKKD 247
Y APE++ ++HK DVYS+GM++ E+IG + ++ + S +FP WV+K +
Sbjct: 498 YIAPEMFSKNFGAVSHKSDVYSYGMVVLEMIGAKNIEKVEYSGSNNGSMYFPEWVYKDFE 557
Query: 248 AGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEI----- 302
G + + I ++ ++IA++++ VALWC+Q P RP M V++MLEG+LE
Sbjct: 558 KGEI-TRIFGDSITDEEEKIAKKLVLVALWCIQMNPSDRPPMIKVIEMLEGNLEALQVPP 616
Query: 303 -PKTFNPFQHLIDGTE 317
P F+P + + D E
Sbjct: 617 NPLLFSPEETVPDTLE 632
>At5g38240 receptor serine/threonine protein kinase -like
Length = 588
Score = 239 bits (611), Expect = 1e-63
Identities = 132/311 (42%), Positives = 197/311 (62%), Gaps = 21/311 (6%)
Query: 31 LLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGFC 90
++G GGFGTVYKG +G VAVK+L+ S+ E F+ EV +I + H N+V L GFC
Sbjct: 286 VVGRGGFGTVYKGNLRDGRKVAVKILKDSNGNC--EDFINEVASISQTSHVNIVSLLGFC 343
Query: 91 FERNLIALVYEYMGNGSLDRYLFHETKVLGYEKLHEIAIGTARGIAYLHEECQHRIIHYD 150
FE++ A+VYE++ NGSLD ++ L L+ IA+G ARGI YLH C+ RI+H+D
Sbjct: 344 FEKSKRAIVYEFLENGSLD-----QSSNLDVSTLYGIALGVARGIEYLHFGCKKRIVHFD 398
Query: 151 IKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPELWMPF--PITHKC 208
IKP N+LLD+N PKVADFGLAK C ++ + +++ RGT GY APEL+ ++HK
Sbjct: 399 IKPQNVLLDENLKPKVADFGLAKLCEKQESILSLLDTRGTIGYIAPELFSRVYGNVSHKS 458
Query: 209 DVYSFGMLLFEIIGRRRNLAIKNTESQE---WFPIWVWKKKDAGLLGEAMIVCGIEEKNK 265
DVYS+GML+ E+ G R ++N +S +FP W++K + G + ++ G+ + +
Sbjct: 459 DVYSYGMLVLEMTGARNKERVQNADSNNSSAYFPDWIFKDLENGDYVK-LLADGLTREEE 517
Query: 266 EIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEI------PKTFNPFQHLIDGTEFT 319
+IA++MI V LWC+Q+RP RP M+ VV M+EG+L+ P P Q+ + E +
Sbjct: 518 DIAKKMILVGLWCIQFRPSDRPSMNKVVGMMEGNLDSLDPPPKPLLHMPMQN--NNAESS 575
Query: 320 THSVQESNTYT 330
S ++S+ Y+
Sbjct: 576 QPSEEDSSIYS 586
>At1g66930 receptor serine/threonine kinase PR5K, putative
Length = 876
Score = 239 bits (611), Expect = 1e-63
Identities = 135/314 (42%), Positives = 197/314 (61%), Gaps = 18/314 (5%)
Query: 3 KFLNDMEREK-----PIR-FTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVL 56
K +D +EK P++ +T Q++ T +++ ++G GGFG VY+G +G MVAVKVL
Sbjct: 519 KTSDDRRQEKLKALIPLKHYTYAQVKRMTKSFAEVVGRGGFGIVYRGTLCDGRMVAVKVL 578
Query: 57 RGSSNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET 116
+ S E F+ EV ++ + H N+V L GFC E + A++YE++ NGSLD+++ +T
Sbjct: 579 KESKGNN-SEDFINEVSSMSQTSHVNIVSLLGFCSEGSRRAIIYEFLENGSLDKFISEKT 637
Query: 117 KV-LGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNC 175
V L L+ IA+G ARG+ YLH C+ RI+H+DIKP N+LLD N PKV+DFGLAK C
Sbjct: 638 SVILDLTALYGIALGVARGLEYLHYGCKTRIVHFDIKPQNVLLDDNLSPKVSDFGLAKLC 697
Query: 176 NRENTHITMTGGRGTPGYAAPELWMPF--PITHKCDVYSFGMLLFEIIGRRRNLAIKNTE 233
++ + +++ RGT GY APE+ ++HK DVYS+GML+FE+IG R+
Sbjct: 698 EKKESVMSLMDTRGTIGYIAPEMISRVYGSVSHKSDVYSYGMLVFEMIGARKKERFGQNS 757
Query: 234 ---SQEWFPIWVWK---KKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRP 287
S +FP W++K K D G L I GI + +EIA++M V LWC+Q P RP
Sbjct: 758 ANGSSMYFPEWIYKDLEKADNGDLEH--IEIGISSEEEEIAKKMTLVGLWCIQSSPSDRP 815
Query: 288 IMSVVVKMLEGSLE 301
M+ VV+M+EGSL+
Sbjct: 816 PMNKVVEMMEGSLD 829
>At5g39030 receptor protein kinase - like protein
Length = 806
Score = 239 bits (610), Expect = 2e-63
Identities = 140/318 (44%), Positives = 197/318 (61%), Gaps = 20/318 (6%)
Query: 16 FTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTI 75
+T +L+ T ++S ++G GGFGTVY G SNG VAVKVL+ E F+ EV ++
Sbjct: 488 YTYAELKKITKSFSYIIGKGGFGTVYGGNLSNGRKVAVKVLKDLKGSA--EDFINEVASM 545
Query: 76 GRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVL-GYEKLHEIAIGTARG 134
+ H N+V L GFCFE + A+VYE++ NGSLD+++ + L+ IA+G ARG
Sbjct: 546 SQTSHVNIVSLLGFCFEGSKRAIVYEFLENGSLDQFMSRNKSLTQDVTTLYGIALGIARG 605
Query: 135 IAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYA 194
+ YLH C+ RI+H+DIKP NILLD N PKV+DFGLAK C + + +++ RGT GY
Sbjct: 606 LEYLHYGCKTRIVHFDIKPQNILLDGNLCPKVSDFGLAKLCEKRESVLSLMDTRGTIGYI 665
Query: 195 APELW--MPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTE---SQEWFPIWVWKKKDAG 249
APE++ M ++HK DVYSFGML+ ++IG R ++ + S +FP W++K +
Sbjct: 666 APEVFSRMYGRVSHKSDVYSFGMLVIDMIGARSKEIVETVDSAASSTYFPDWIYKDLED- 724
Query: 250 LLGEAMIVCG--IEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGS---LEIPK 304
GE + G I ++ KEIA++MI V LWC+Q P RP M+ VV+M+EGS LEIP
Sbjct: 725 --GEQTWIFGDEITKEEKEIAKKMIVVGLWCIQPCPSDRPSMNRVVEMMEGSLDALEIPP 782
Query: 305 TFNPFQHLIDGTEFTTHS 322
P H+ TE T S
Sbjct: 783 --KPSMHI--STEVITES 796
>At1g66910 receptor serine/threonine kinase PR5K, putative
Length = 655
Score = 238 bits (607), Expect = 4e-63
Identities = 129/322 (40%), Positives = 205/322 (63%), Gaps = 13/322 (4%)
Query: 20 QLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIH 79
Q+ T +++ ++G GGFGTVY+G +G VAVKVL+ S E F+ EV ++ +
Sbjct: 331 QVTSITKSFAEVIGKGGFGTVYRGTLYDGRSVAVKVLKESQGN--GEDFINEVASMSQTS 388
Query: 80 HFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHE-TKVLGYEKLHEIAIGTARGIAYL 138
H N+V L GFC E A++YE+M NGSLD+++ + + + + +L+ IA+G ARG+ YL
Sbjct: 389 HVNIVTLLGFCSEGYKRAIIYEFMENGSLDKFISSKKSSTMDWRELYGIALGVARGLEYL 448
Query: 139 HEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPEL 198
H C+ RI+H+DIKP N+LLD N PKV+DFGLAK C R+ + +++ RGT GY APE+
Sbjct: 449 HHGCRTRIVHFDIKPQNVLLDDNLSPKVSDFGLAKLCERKESILSLMDTRGTIGYIAPEV 508
Query: 199 WMPF--PITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQE---WFPIWVWKKKDAGLLGE 253
+ ++HK DVYS+GML+ +IIG R + ++T S +FP W+++ + G+
Sbjct: 509 FSRVYGRVSHKSDVYSYGMLVLDIIGARNKTSTEDTTSSTSSMYFPEWIYRDLEKAHNGK 568
Query: 254 AMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIPKTFNPFQHLI 313
+ I I + EIA++M V LWC+Q P RP M+ VV+M+EG+L+ + P + ++
Sbjct: 569 S-IETAISNEEDEIAKKMTLVGLWCIQPWPLDRPAMNRVVEMMEGNLDALEV--PPRPVL 625
Query: 314 DGTEFTTHSVQESNTYTTSVSS 335
+ T ++QES+T++ +S+
Sbjct: 626 Q--QIPTATLQESSTFSEDISA 645
>At1g66920 receptor serine/threonine kinase PR5K, putative
Length = 609
Score = 236 bits (603), Expect = 1e-62
Identities = 126/293 (43%), Positives = 193/293 (65%), Gaps = 10/293 (3%)
Query: 15 RFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGT 74
+++ +Q++ T++++ ++G GGFG VY+G S+G MVAVKVL+ E F+ EV +
Sbjct: 288 QYSYEQVKRITNSFAEVVGRGGFGIVYRGTLSDGRMVAVKVLKDLKGNN-GEDFINEVAS 346
Query: 75 IGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHE-TKVLGYEKLHEIAIGTAR 133
+ + H N+V L GFC E A++YE+M NGSLD+++ + + + + +L+ IA+G AR
Sbjct: 347 MSQTSHVNIVTLLGFCSEGYKRAIIYEFMENGSLDKFISSKKSSTMDWRELYGIALGVAR 406
Query: 134 GIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGY 193
G+ YLH C+ RI+H+DIKP N+LLD N PKV+DFGLAK C R+ + +++ RGT GY
Sbjct: 407 GLEYLHHGCRTRIVHFDIKPQNVLLDDNLSPKVSDFGLAKLCERKESILSLMDTRGTIGY 466
Query: 194 AAPELWMPF--PITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQE---WFPIWVWKKKDA 248
APE++ ++HK DVYS+GML+ +IIG R + ++T S +FP W++K +
Sbjct: 467 IAPEVFSRVYGSVSHKSDVYSYGMLVLDIIGARNKTSTEDTTSSTSSMYFPEWIYKDLEK 526
Query: 249 GLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLE 301
G G +IV EE EIA++M V LWC+Q P RP M+ VV+M+EG+L+
Sbjct: 527 GDNGR-LIVNRSEE--DEIAKKMTLVGLWCIQPWPLDRPAMNRVVEMMEGNLD 576
>At4g00340 receptor-like protein kinase
Length = 797
Score = 236 bits (602), Expect = 1e-62
Identities = 145/315 (46%), Positives = 187/315 (59%), Gaps = 15/315 (4%)
Query: 16 FTGQQLRIATDNYSNLLGSGGFGTVYKGIF-SNGTMVAVKVLRGSSNKKIDEQFMAEVGT 74
F+ ++L+ AT+ +S+ +G GGFG V+KG + T VAVK L + + +F AEV T
Sbjct: 451 FSFKELQSATNGFSDKVGHGGFGAVFKGTLPGSSTFVAVKRLERPGSG--ESEFRAEVCT 508
Query: 75 IGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHET-KVLGYEKLHEIAIGTAR 133
IG I H NLVRL GFC E LVY+YM GSL YL + K+L +E IA+GTA+
Sbjct: 509 IGNIQHVNLVRLRGFCSENLHRLLVYDYMPQGSLSSYLSRTSPKLLSWETRFRIALGTAK 568
Query: 134 GIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGY 193
GIAYLHE C+ IIH DIKP NILLD ++ KV+DFGLAK R+ + + T RGT GY
Sbjct: 569 GIAYLHEGCRDCIIHCDIKPENILLDSDYNAKVSDFGLAKLLGRDFSRVLAT-MRGTWGY 627
Query: 194 AAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAI-------KNTESQEW-FPIWVWKK 245
APE PIT K DVYSFGM L E+IG RRN+ + K TE ++W FP W ++
Sbjct: 628 VAPEWISGLPITTKADVYSFGMTLLELIGGRRNVIVNSDTLGEKETEPEKWFFPPWAARE 687
Query: 246 KDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLE--IP 303
G + + E N E RM VA+WC+Q E+RP M VVKMLEG +E +P
Sbjct: 688 IIQGNVDSVVDSRLNGEYNTEEVTRMATVAIWCIQDNEEIRPAMGTVVKMLEGVVEVTVP 747
Query: 304 KTFNPFQHLIDGTEF 318
Q L+ G +
Sbjct: 748 PPPKLIQALVSGDSY 762
>At1g70250 hypothetical protein
Length = 676
Score = 235 bits (599), Expect = 3e-62
Identities = 128/307 (41%), Positives = 191/307 (61%), Gaps = 12/307 (3%)
Query: 5 LNDMEREKPI---RFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTM-VAVKVLRGSS 60
LN+ E + RF+ Q++ T ++ N+LG GGFGTVYKG +G+ VAVK+L+ S+
Sbjct: 312 LNEKNMEAVVMLKRFSYVQVKKMTKSFENVLGKGGFGTVYKGKLPDGSRDVAVKILKESN 371
Query: 61 NKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV-L 119
E F+ E+ ++ R H N+V L GFC+E A++YE M NGSLD+++ +
Sbjct: 372 ED--GEDFINEIASMSRTSHANIVSLLGFCYEGRKKAIIYELMPNGSLDKFISKNMSAKM 429
Query: 120 GYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNREN 179
++ L+ IA+G + G+ YLH C RI+H+DIKP NIL+D + PK++DFGLAK C
Sbjct: 430 EWKTLYNIAVGVSHGLEYLHSHCVSRIVHFDIKPQNILIDGDLCPKISDFGLAKLCKNNE 489
Query: 180 THITMTGGRGTPGYAAPELWMP--FPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQE- 236
+ I+M RGT GY APE++ ++HK DVYS+GM++ E+IG R +N S
Sbjct: 490 SIISMLHARGTIGYIAPEVFSQNFGGVSHKSDVYSYGMVVLEMIGARNIGRAQNAGSSNT 549
Query: 237 --WFPIWVWKKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVK 294
+FP W++K + G + + EE++++I ++M+ V LWC+Q P RP MS VV+
Sbjct: 550 SMYFPDWIYKDLEKGEIMSFLADQITEEEDEKIVKKMVLVGLWCIQTNPYDRPPMSKVVE 609
Query: 295 MLEGSLE 301
MLEGSLE
Sbjct: 610 MLEGSLE 616
>At1g67000 hypothetical protein
Length = 717
Score = 234 bits (598), Expect = 4e-62
Identities = 124/300 (41%), Positives = 190/300 (63%), Gaps = 15/300 (5%)
Query: 16 FTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTI 75
+T +++ T +++ ++G GGFG VY G S+ +MVAVKVL+ S E F+ EV ++
Sbjct: 371 YTYAEVKKMTKSFTEVVGRGGFGIVYSGTLSDSSMVAVKVLKDSKGTD-GEDFINEVASM 429
Query: 76 GRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV-LGYEKLHEIAIGTARG 134
+ H N+V L GFC E + A++YE++GNGSLD+++ ++ V L + L+ IA+G ARG
Sbjct: 430 SQTSHVNIVSLLGFCCEGSRRAIIYEFLGNGSLDKFISDKSSVNLDLKTLYGIALGVARG 489
Query: 135 IAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYA 194
+ YLH C+ RI+H+DIKP N+LLD N PKV+DFGLAK C ++ + +++ RGT GY
Sbjct: 490 LEYLHYGCKTRIVHFDIKPQNVLLDDNLCPKVSDFGLAKLCEKKESILSLLDTRGTIGYI 549
Query: 195 APELWMPF--PITHKCDVYSFGMLLFEIIGRRRNLAI----KNTESQEWFPIWVWKKKDA 248
APE+ ++HK DVYS+GML+ E+IG R+ ++ S +FP W++K +
Sbjct: 550 APEMISRLYGSVSHKSDVYSYGMLVLEMIGARKKERFDQNSRSDGSSIYFPEWIYKDLEK 609
Query: 249 GLLGE-------AMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLE 301
+ + +I GI + +EIA +M V LWC+Q P RP M+ VV+M+EGSL+
Sbjct: 610 ANIKDIEKTENGGLIENGISSEEEEIARKMTLVGLWCIQSSPSDRPPMNKVVEMMEGSLD 669
>At4g18250 receptor serine/threonine kinase-like protein
Length = 687
Score = 233 bits (595), Expect = 9e-62
Identities = 132/348 (37%), Positives = 215/348 (60%), Gaps = 24/348 (6%)
Query: 5 LNDMEREKPI---RFTGQQLRIATDNYSNLLGSGGFGTVYKGIF--SNGTMVAVKVLRGS 59
LND E + R++ ++++ T+++ +++G GGFGTVYKG ++G +A+K+L+ S
Sbjct: 329 LNDENIEAVVMLKRYSFEKVKKMTNSFDHVIGKGGFGTVYKGKLPDASGRDIALKILKES 388
Query: 60 SNKKIDEQFMAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV- 118
E+F+ E+ ++ R H N+V L+GFC+E + A++YE+M NGSLD+++
Sbjct: 389 KGN--GEEFINELVSMSRASHVNIVSLFGFCYEGSQRAIIYEFMPNGSLDKFISENMSTK 446
Query: 119 LGYEKLHEIAIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRE 178
+ ++ L+ IA+G ARG+ YLH C +I+H+DIKP NIL+D++ PK++DFGLAK C ++
Sbjct: 447 IEWKTLYNIAVGVARGLEYLHNSCVSKIVHFDIKPQNILIDEDLCPKISDFGLAKLCKKK 506
Query: 179 NTHITMTGGRGTPGYAAPELWMP--FPITHKCDVYSFGMLLFEIIGRRRNLAIKNT---E 233
+ I+M RGT GY APE++ ++HK DVYS+GM++ E+IG + ++ + +
Sbjct: 507 ESIISMLDARGTVGYIAPEMFSKNYGGVSHKSDVYSYGMVVLEMIGATKREEVETSATDK 566
Query: 234 SQEWFPIWVW----KKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIM 289
S +FP WV+ +K+ LL + +I EE+ ++I +RM V LWC+Q P RP M
Sbjct: 567 SSMYFPDWVYEDLERKETMRLLEDHIIE---EEEEEKIVKRMTLVGLWCIQTNPSDRPPM 623
Query: 290 SVVVKMLEGS----LEIPKTFNPFQHLIDGTEFTTHSVQESNTYTTSV 333
VV+MLEGS L++P H++ E + S Q S T S+
Sbjct: 624 RKVVEMLEGSRLEALQVPPKPLLNLHVVTDWETSEDSQQTSRLSTQSL 671
>At5g38260 receptor serine/threonine protein kinase - like
Length = 638
Score = 232 bits (592), Expect = 2e-61
Identities = 128/299 (42%), Positives = 190/299 (62%), Gaps = 19/299 (6%)
Query: 15 RFTGQQLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGT 74
+++ ++R T +S+ LG GGFGTVY G +G VAVK+L+ K E F+ EV +
Sbjct: 310 QYSYAEVRKITKLFSHTLGKGGFGTVYGGNLCDGRKVAVKILKDF--KSNGEDFINEVAS 367
Query: 75 IGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV-LGYEKLHEIAIGTAR 133
+ + H N+V L GFC+E + A+VYE++ NGSLD++L + + L L+ IA+G AR
Sbjct: 368 MSQTSHVNIVSLLGFCYEGSKRAIVYEFLENGSLDQFLSEKKSLNLDVSTLYRIALGVAR 427
Query: 134 GIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGY 193
G+ YLH C+ RI+H+DIKP NILLD F PKV+DFGLAK C + + +++ RGT GY
Sbjct: 428 GLDYLHHGCKTRIVHFDIKPQNILLDDTFCPKVSDFGLAKLCEKRESILSLLDARGTIGY 487
Query: 194 AAPELW--MPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNT----ESQEWFPIWVWKK-- 245
APE++ M ++HK DVYS+GML+ E+IG +N I+ T S +FP W++K
Sbjct: 488 IAPEVFSGMYGRVSHKSDVYSYGMLVLEMIG-AKNKEIEETAASNSSSAYFPDWIYKNLE 546
Query: 246 --KDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEI 302
+D G+ I ++KE+A++M V LWC+Q P RP M+ +V+M+EGSL++
Sbjct: 547 NGEDTWKFGDE-----ISREDKEVAKKMTLVGLWCIQPSPLNRPPMNRIVEMMEGSLDV 600
>At5g39020 receptor protein kinase - like protein
Length = 813
Score = 227 bits (578), Expect = 9e-60
Identities = 135/335 (40%), Positives = 203/335 (60%), Gaps = 27/335 (8%)
Query: 20 QLRIATDNYSNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFMAEVGTIGRIH 79
+L+ T ++S+ +G GGFGTVY+G SNG VAVKVL+ D F+ EV ++ +
Sbjct: 490 ELKKITKSFSHTVGKGGFGTVYRGNLSNGRTVAVKVLKDLKGNGDD--FINEVTSMSQTS 547
Query: 80 HFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKVL-GYEKLHEIAIGTARGIAYL 138
H N+V L GFC+E + A++ E++ +GSLD+++ + L+ IA+G ARG+ YL
Sbjct: 548 HVNIVSLLGFCYEGSKRAIISEFLEHGSLDQFISRNKSLTPNVTTLYGIALGIARGLEYL 607
Query: 139 HEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGRGTPGYAAPEL 198
H C+ RI+H+DIKP NILLD NF PKVADFGLAK C + + +++ RGT GY APE+
Sbjct: 608 HYGCKTRIVHFDIKPQNILLDDNFCPKVADFGLAKLCEKRESILSLIDTRGTIGYIAPEV 667
Query: 199 --WMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTE-SQEWFPIWVWKKKDAG----LL 251
M I+HK DVYS+GML+ ++IG R + S +FP W++K + G ++
Sbjct: 668 VSRMYGGISHKSDVYSYGMLVLDMIGARNKVETTTCNGSTAYFPDWIYKDLENGDQTWII 727
Query: 252 GEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGS---LEIPKTFNP 308
G+ I E++ +I ++MI V+LWC++ P RP M+ VV+M+EGS LE+P P
Sbjct: 728 GDE-----INEEDNKIVKKMILVSLWCIRPCPSDRPPMNKVVEMIEGSLDALELPP--KP 780
Query: 309 FQHLIDGTEFTTHSV-----QESNTYTTSVSSVMV 338
+H+ TE S QE+ T ++ S ++
Sbjct: 781 SRHI--STELVLESSSLSDGQEAEKQTQTLDSTII 813
>At1g56120 receptor-like protein kinase, putative
Length = 1045
Score = 225 bits (574), Expect = 3e-59
Identities = 145/331 (43%), Positives = 188/331 (55%), Gaps = 16/331 (4%)
Query: 12 KPIRFTGQQLRIATDNY--SNLLGSGGFGTVYKGIFSNGTMVAVKVLR-GSSNKKIDEQF 68
KP FT +L+ AT ++ SN LG GGFG VYKG ++G VAVK L GS K QF
Sbjct: 692 KPYTFTYSELKNATQDFDLSNKLGEGGFGAVYKGNLNDGREVAVKQLSIGSRQGK--GQF 749
Query: 69 MAEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV-LGYEKLHEI 127
+AE+ I + H NLV+LYG CFE + LVYEY+ NGSLD+ LF + + L + +EI
Sbjct: 750 VAEIIAISSVLHRNLVKLYGCCFEGDHRLLVYEYLPNGSLDQALFGDKSLHLDWSTRYEI 809
Query: 128 AIGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGG 187
+G ARG+ YLHEE RIIH D+K NILLD PKV+DFGLAK + + THI+ T
Sbjct: 810 CLGVARGLVYLHEEASVRIIHRDVKASNILLDSELVPKVSDFGLAKLYDDKKTHIS-TRV 868
Query: 188 RGTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVW---- 243
GT GY APE M +T K DVY+FG++ E++ R+N E +++ W W
Sbjct: 869 AGTIGYLAPEYAMRGHLTEKTDVYAFGVVALELVSGRKNSDENLEEGKKYLLEWAWNLHE 928
Query: 244 KKKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIP 303
K +D L+ + + E N E +RMI +AL C Q LRP MS VV ML G E+
Sbjct: 929 KNRDVELIDDE-----LSEYNMEEVKRMIGIALLCTQSSYALRPPMSRVVAMLSGDAEVN 983
Query: 304 KTFNPFQHLIDGTEFTTHSVQESNTYTTSVS 334
+ +L D T T S SN T S
Sbjct: 984 DATSKPGYLTDCTFDDTTSSSFSNFQTKDTS 1014
>At1g56140 receptor-like protein kinase, putative
Length = 2083
Score = 222 bits (565), Expect = 3e-58
Identities = 140/336 (41%), Positives = 190/336 (55%), Gaps = 16/336 (4%)
Query: 12 KPIRFTGQQLRIATDNY--SNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFM 69
KP FT +L+ AT ++ SN LG GGFG VYKG ++G VAVK+L S + QF+
Sbjct: 1727 KPYTFTYSELKSATQDFDPSNKLGEGGFGPVYKGKLNDGREVAVKLLSVGSRQG-KGQFV 1785
Query: 70 AEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHE-TKVLGYEKLHEIA 128
AE+ I + H NLV+LYG C+E LVYEY+ NGSLD+ LF E T L + +EI
Sbjct: 1786 AEIVAISAVQHRNLVKLYGCCYEGEHRLLVYEYLPNGSLDQALFGEKTLHLDWSTRYEIC 1845
Query: 129 IGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGR 188
+G ARG+ YLHEE + RI+H D+K NILLD PKV+DFGLAK + + THI+ T
Sbjct: 1846 LGVARGLVYLHEEARLRIVHRDVKASNILLDSKLVPKVSDFGLAKLYDDKKTHIS-TRVA 1904
Query: 189 GTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDA 248
GT GY APE M +T K DVY+FG++ E++ R N + + + W W +
Sbjct: 1905 GTIGYLAPEYAMRGHLTEKTDVYAFGVVALELVSGRPNSDENLEDEKRYLLEWAWNLHEK 1964
Query: 249 GLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIPKTFNP 308
G E +I + E N E +RMI +AL C Q LRP MS VV ML G +E+ +
Sbjct: 1965 GREVE-LIDHQLTEFNMEEGKRMIGIALLCTQTSHALRPPMSRVVAMLSGDVEVSDVTSK 2023
Query: 309 FQHLIDG----------TEFTTHSVQESNTYTTSVS 334
+L D + F + Q S ++T+ V+
Sbjct: 2024 PGYLTDWRFDDTTASSISGFPLRNTQASESFTSFVA 2059
Score = 213 bits (543), Expect = 1e-55
Identities = 131/334 (39%), Positives = 192/334 (57%), Gaps = 6/334 (1%)
Query: 12 KPIRFTGQQLRIATDNY--SNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFM 69
+P F+ +LR AT ++ SN LG GGFG V+KG ++G +AVK L +S + QF+
Sbjct: 657 RPYTFSYSELRTATQDFDPSNKLGEGGFGPVFKGKLNDGREIAVKQLSVASRQG-KGQFV 715
Query: 70 AEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHETKV-LGYEKLHEIA 128
AE+ TI + H NLV+LYG C E N LVYEY+ N SLD+ LF E + LG+ + EI
Sbjct: 716 AEIATISAVQHRNLVKLYGCCIEGNQRMLVYEYLSNKSLDQALFEEKSLQLGWSQRFEIC 775
Query: 129 IGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGR 188
+G A+G+AY+HEE RI+H D+K NILLD + PK++DFGLAK + + THI+ T
Sbjct: 776 LGVAKGLAYMHEESNPRIVHRDVKASNILLDSDLVPKLSDFGLAKLYDDKKTHIS-TRVA 834
Query: 189 GTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVWKKKDA 248
GT GY +PE M +T K DV++FG++ EI+ R N + + + +++ W W
Sbjct: 835 GTIGYLSPEYVMLGHLTEKTDVFAFGIVALEIVSGRPNSSPELDDDKQYLLEWAWSLHQE 894
Query: 249 GLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEIPKTFNP 308
E ++ + E +KE +R+I VA C Q +RP MS VV ML G +EI +
Sbjct: 895 QRDME-VVDPDLTEFDKEEVKRVIGVAFLCTQTDHAIRPTMSRVVGMLTGDVEITEANAK 953
Query: 309 FQHLIDGTEFTTHSVQESNTYTTSVSSVMVSDSS 342
++ + T S +T ++ + DSS
Sbjct: 954 PGYVSERTFENAMSFMSGSTSSSWILPETPKDSS 987
>At1g56130 receptor-like protein kinase, putative
Length = 1032
Score = 219 bits (558), Expect = 2e-57
Identities = 133/298 (44%), Positives = 178/298 (59%), Gaps = 14/298 (4%)
Query: 12 KPIRFTGQQLRIATDNY--SNLLGSGGFGTVYKGIFSNGTMVAVKVLRGSSNKKIDEQFM 69
KP FT +L+ AT ++ SN LG GGFG VYKG ++G +VAVK+L S + QF+
Sbjct: 678 KPYIFTYSELKSATQDFDPSNKLGEGGFGPVYKGNLNDGRVVAVKLLSVGSRQG-KGQFV 736
Query: 70 AEVGTIGRIHHFNLVRLYGFCFERNLIALVYEYMGNGSLDRYLFHE-TKVLGYEKLHEIA 128
AE+ I + H NLV+LYG CFE LVYEY+ NGSLD+ LF + T L + +EI
Sbjct: 737 AEIVAISSVLHRNLVKLYGCCFEGEHRMLVYEYLPNGSLDQALFGDKTLHLDWSTRYEIC 796
Query: 129 IGTARGIAYLHEECQHRIIHYDIKPGNILLDKNFYPKVADFGLAKNCNRENTHITMTGGR 188
+G ARG+ YLHEE RI+H D+K NILLD P+++DFGLAK + + THI+ T
Sbjct: 797 LGVARGLVYLHEEASVRIVHRDVKASNILLDSRLVPQISDFGLAKLYDDKKTHIS-TRVA 855
Query: 189 GTPGYAAPELWMPFPITHKCDVYSFGMLLFEIIGRRRNLAIKNTESQEWFPIWVW----K 244
GT GY APE M +T K DVY+FG++ E++ R N E +++ W W K
Sbjct: 856 GTIGYLAPEYAMRGHLTEKTDVYAFGVVALELVSGRPNSDENLEEEKKYLLEWAWNLHEK 915
Query: 245 KKDAGLLGEAMIVCGIEEKNKEIAERMIKVALWCVQYRPELRPIMSVVVKMLEGSLEI 302
+D L+ + + + N E A+RMI +AL C Q LRP MS VV ML G +EI
Sbjct: 916 SRDIELIDDK-----LTDFNMEEAKRMIGIALLCTQTSHALRPPMSRVVAMLSGDVEI 968
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,377,090
Number of Sequences: 26719
Number of extensions: 372936
Number of successful extensions: 4173
Number of sequences better than 10.0: 984
Number of HSP's better than 10.0 without gapping: 843
Number of HSP's successfully gapped in prelim test: 141
Number of HSP's that attempted gapping in prelim test: 869
Number of HSP's gapped (non-prelim): 1093
length of query: 362
length of database: 11,318,596
effective HSP length: 101
effective length of query: 261
effective length of database: 8,619,977
effective search space: 2249813997
effective search space used: 2249813997
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC130275.4


