

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
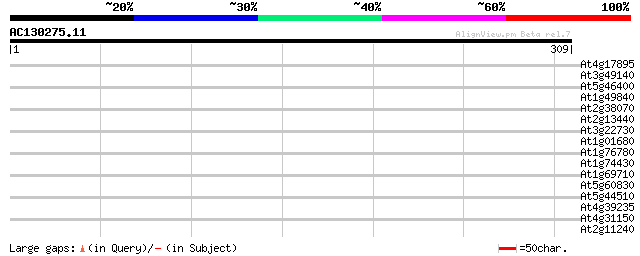
Query= AC130275.11 - phase: 0
(309 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g17895 ubiquitin-specific protease 20 (UBP20) 37 0.010
At3g49140 putative protein 36 0.030
At5g46400 putative protein 32 0.44
At1g49840 unknown protein 29 2.8
At2g38070 unknown protein 29 3.7
At2g13440 similar to glucose inhibited division protein A from p... 29 3.7
At3g22730 hypothetical protein 28 4.8
At1g01680 hypothetical protein 28 4.8
At1g76780 putative heat shock protein 28 6.3
At1g74430 transcription like factor (MYB95) 28 6.3
At1g69710 putative regulator of chromosome condensation 28 6.3
At5g60830 bZip transcription factor AtbZip70 28 8.2
At5g44510 disease resistance protein-like 28 8.2
At4g39235 unknown protein 28 8.2
At4g31150 unknown protein 28 8.2
At2g11240 pseudogene 28 8.2
>At4g17895 ubiquitin-specific protease 20 (UBP20)
Length = 695
Score = 37.4 bits (85), Expect = 0.010
Identities = 25/95 (26%), Positives = 41/95 (42%)
Query: 73 NFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINEYQEYSK 132
N SP V + + + I G A P + + + K K+ L N+ S
Sbjct: 498 NSSPKSVLDSSTNGECLSEISYENGDKASKPCDSAGVCNQHVKTKEDFVSLSNDDVFLSA 557
Query: 133 DSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCI 167
+SSS +E +++D +D + YS C E SC+
Sbjct: 558 ESSSGEESPMGELLDPLDPDDSYSPCTEKESDSCL 592
>At3g49140 putative protein
Length = 1229
Score = 35.8 bits (81), Expect = 0.030
Identities = 27/88 (30%), Positives = 43/88 (48%), Gaps = 8/88 (9%)
Query: 202 ANKDIINW----VDYKFYNQTVSSADELVNLYN--KLLNEYGTDVKLLPGVSTDPDSNTN 255
+ +D+++W V Y NQ A E+ K+ ++ GT LLP VS N
Sbjct: 202 SRRDVVSWNSLVVGYA-QNQRFDDALEVCREMESVKISHDAGTMASLLPAVSNTTTENVM 260
Query: 256 MTRDVFIK-GCKSLLESESLPGIFVWNA 282
+D+F K G KSL+ + G+++ NA
Sbjct: 261 YVKDMFFKMGKKSLVSWNVMIGVYMKNA 288
>At5g46400 putative protein
Length = 1022
Score = 32.0 bits (71), Expect = 0.44
Identities = 14/57 (24%), Positives = 28/57 (48%)
Query: 177 SSKSIKLVSIAPTELLKPHYHKLYWANKDIINWVDYKFYNQTVSSADELVNLYNKLL 233
S + ++ +S T++ +P++H + NW Y + +T D +NLY + L
Sbjct: 269 SRQLMEKISCFETQIRRPYFHVKPLDTNQLDNWHAYLSFGETYGDFDWAINLYERCL 325
>At1g49840 unknown protein
Length = 494
Score = 29.3 bits (64), Expect = 2.8
Identities = 15/45 (33%), Positives = 21/45 (46%)
Query: 163 FSSCIGELIRKLKKSSKSIKLVSIAPTELLKPHYHKLYWANKDII 207
FSS E++R L S + S +P+ PH L W K +I
Sbjct: 8 FSSLRDEVVRGLSPSRSRPRSRSASPSRSTTPHMKALLWGRKKLI 52
>At2g38070 unknown protein
Length = 619
Score = 28.9 bits (63), Expect = 3.7
Identities = 30/117 (25%), Positives = 51/117 (42%), Gaps = 14/117 (11%)
Query: 6 QINAVVKPIIFREYIGVKDIPKNLKDFPAEMINDDIEEFHFILGTLREVYSGDGKGKGEF 65
+IN++VK +F E ++ N KD E +E ++ + E + K E
Sbjct: 165 RINSIVKSPVFEEETEIESEQDNEKDIKFE----TFKEPRSVIDEIVEEEEEEETKKVED 220
Query: 66 YRTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKK 122
+ T FN + AK K +D K + G F + K+++ W + KQ +KK
Sbjct: 221 F-TMEFNPQTTAK--KTNRDFKEI-----AGSFWSAASVFSKKLQKW--RQKQKLKK 267
>At2g13440 similar to glucose inhibited division protein A from
prokaryotes
Length = 723
Score = 28.9 bits (63), Expect = 3.7
Identities = 23/97 (23%), Positives = 40/97 (40%), Gaps = 17/97 (17%)
Query: 67 RTWNFNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLIN- 125
R W A++++ KK K VK+ +++G AE + S K +++ L+
Sbjct: 528 RRWKLYQEKQARISEEKKRLKTVKISVAVGDLAAE----VSSVSSQPVKESATLESLLKK 583
Query: 126 --------EYQEYSKDSSSTDECHCDDIIDGIDINYE 154
E + ++ S E C + IDI YE
Sbjct: 584 PHIHYKLLEKHGFGNETLSRMEKDCVE----IDIKYE 616
>At3g22730 hypothetical protein
Length = 372
Score = 28.5 bits (62), Expect = 4.8
Identities = 13/46 (28%), Positives = 22/46 (47%), Gaps = 5/46 (10%)
Query: 124 INEYQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGE 169
+ Y YS++ + CHCD ++ ++Y NP C+GE
Sbjct: 86 LKNYPPYSEEVDIHEACHCDGLLLCTTMDYRIVVWNP-----CLGE 126
>At1g01680 hypothetical protein
Length = 282
Score = 28.5 bits (62), Expect = 4.8
Identities = 15/44 (34%), Positives = 23/44 (52%)
Query: 101 ENPFNPKEIESWSTKAKQSIKKLINEYQEYSKDSSSTDECHCDD 144
E P +PKE W + ++S KK E E SK + S ++ +D
Sbjct: 162 EEPESPKEHPGWILEPEESPKKGRKETIEKSKSNESDEDPRLED 205
>At1g76780 putative heat shock protein
Length = 1871
Score = 28.1 bits (61), Expect = 6.3
Identities = 12/37 (32%), Positives = 19/37 (50%)
Query: 107 KEIESWSTKAKQSIKKLINEYQEYSKDSSSTDECHCD 143
K E W+ + ++ K L+ E + Y KD + E H D
Sbjct: 1037 KSYEDWTHEKREKRKVLVEEEETYPKDKHTGGEDHND 1073
>At1g74430 transcription like factor (MYB95)
Length = 271
Score = 28.1 bits (61), Expect = 6.3
Identities = 26/90 (28%), Positives = 39/90 (42%), Gaps = 7/90 (7%)
Query: 71 FNNFSPAKVAKLKKDHKNVKVIISIGGFGAENPFNPKEIESWSTKAKQSIKKLINEYQEY 130
F N A ++ K +++K +I AE P + E T K+LI+ Y E
Sbjct: 167 FLNKLAAGISSRKHGLESIKTVIL-----AEQPREAVDEEKMMT-INMKEKELISCYMEI 220
Query: 131 SKDSSSTDECHCDDIIDGIDINYEYSNCNP 160
++ S DE CDD G +YS +P
Sbjct: 221 D-ETMSIDELPCDDSTSGFVAFDDYSLIDP 249
>At1g69710 putative regulator of chromosome condensation
Length = 1028
Score = 28.1 bits (61), Expect = 6.3
Identities = 20/64 (31%), Positives = 34/64 (52%), Gaps = 7/64 (10%)
Query: 81 KLKKDHKNVKVIISIGGFGAENPFNPKE-IESWSTKAKQSIKKLINEYQEYSKDSSSTDE 139
+L+K + +KV+ ++ AE + KE I S +T+ K+ +K + KDS ST+
Sbjct: 866 ELEKTKRQLKVVTAMAADEAEENRSAKEVIRSLTTQLKEMAEK------QSQKDSISTNS 919
Query: 140 CHCD 143
H D
Sbjct: 920 KHTD 923
>At5g60830 bZip transcription factor AtbZip70
Length = 188
Score = 27.7 bits (60), Expect = 8.2
Identities = 10/25 (40%), Positives = 17/25 (68%)
Query: 133 DSSSTDECHCDDIIDGIDINYEYSN 157
+SSS HC DI+DG+ ++ ++ N
Sbjct: 10 ESSSVHRSHCFDILDGVPLHDDHFN 34
>At5g44510 disease resistance protein-like
Length = 1187
Score = 27.7 bits (60), Expect = 8.2
Identities = 23/62 (37%), Positives = 29/62 (46%), Gaps = 9/62 (14%)
Query: 220 SSADELVNLYNKLLNEYGTDVKLLPGVSTDPDSNTNMT--RDVFIKGCKSLLESESLPGI 277
SS L NL LN + VKL P S N+T +++ + GC SLLE S G
Sbjct: 722 SSIGNLTNLKKLFLNRCSSLVKL-------PSSFGNVTSLKELNLSGCSSLLEIPSSIGN 774
Query: 278 FV 279
V
Sbjct: 775 IV 776
>At4g39235 unknown protein
Length = 86
Score = 27.7 bits (60), Expect = 8.2
Identities = 19/68 (27%), Positives = 34/68 (49%), Gaps = 7/68 (10%)
Query: 114 TKAKQSIKKLINEYQEYSKDSSSTDECHCDDIIDGIDINYEYSNCNPDEFSSCIGELIRK 173
T+ QS K + ++ KD D +D+ D ID ++ + + DE SC + I+K
Sbjct: 3 TQTNQSPKPAMTSCRKKVKD----DATFFEDVKDHID---DFIHASMDEHKSCFQKTIKK 55
Query: 174 LKKSSKSI 181
+ SK++
Sbjct: 56 MFGLSKAV 63
>At4g31150 unknown protein
Length = 277
Score = 27.7 bits (60), Expect = 8.2
Identities = 20/74 (27%), Positives = 39/74 (52%), Gaps = 9/74 (12%)
Query: 97 GFGAENPFNPKEIESWSTKAKQSIKKLINEYQEYS-KDSSSTDECHCDDI---IDGIDIN 152
G +E+ + ++E W+ + + KKLI Y +++ K SSST+ +I + G+D++
Sbjct: 5 GASSESSSSGDQLEKWTEEQDELKKKLI-AYDDFTWKLSSSTELSQGSEILKYVGGVDMS 63
Query: 153 YEYSNCNPDEFSSC 166
+ C D +C
Sbjct: 64 F----CKEDSSVAC 73
>At2g11240 pseudogene
Length = 1044
Score = 27.7 bits (60), Expect = 8.2
Identities = 14/45 (31%), Positives = 27/45 (59%), Gaps = 1/45 (2%)
Query: 108 EIESWSTKAKQSIKKLINEYQEYSKDSSSTDECHCDDIIDGIDIN 152
EI WS +QS +KLI E ++ +++ S+ + H +++ I+ N
Sbjct: 115 EIVKWSRAKQQSSQKLIEENRQKLEEAMSSQD-HNQELLSTINTN 158
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,717,941
Number of Sequences: 26719
Number of extensions: 354077
Number of successful extensions: 723
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 714
Number of HSP's gapped (non-prelim): 18
length of query: 309
length of database: 11,318,596
effective HSP length: 99
effective length of query: 210
effective length of database: 8,673,415
effective search space: 1821417150
effective search space used: 1821417150
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC130275.11


