

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130200.9 - phase: 0
(393 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
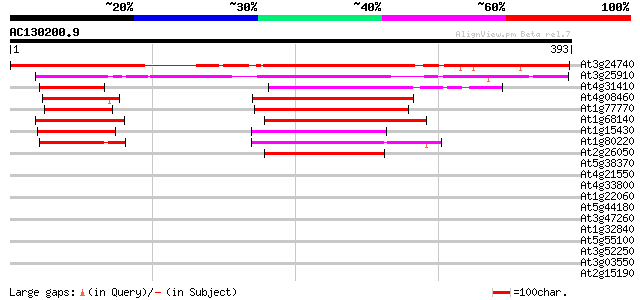
Score E
Sequences producing significant alignments: (bits) Value
At3g24740 unknown protein 381 e-106
At3g25910 unknown protein 198 4e-51
At4g31410 unknown protein 139 2e-33
At4g08460 unknown protein 124 1e-28
At1g77770 unknown protein 119 4e-27
At1g68140 unknown protein 119 4e-27
At1g15430 unknown protein 83 2e-16
At1g80220 hypothetical protein 80 3e-15
At2g26050 hypothetical protein 79 6e-15
At5g38370 similar to unknown protein (pir||T02618) 40 0.003
At4g21550 VP1 like protein 34 0.12
At4g33800 unknown protein 33 0.21
At1g22060 hypothetical protein 33 0.27
At5g44180 unknown protein 32 0.79
At3g47260 putative protein 32 0.79
At1g32840 putative protein FARE2.4 (cds3) 32 0.79
At5g55100 unknown protein 31 1.3
At3g52250 putative protein 31 1.3
At3g03550 unknown protein 31 1.3
At2g15190 putative protein on transposon FARE2.7 (cds3) 30 2.3
>At3g24740 unknown protein
Length = 354
Score = 381 bits (979), Expect = e-106
Identities = 209/402 (51%), Positives = 261/402 (63%), Gaps = 60/402 (14%)
Query: 1 MAGFKRRLCNDSDMHALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHS 60
MAG KR+L +SD+HALH+ELDEVSCP+CMDHPHNAVLLLCSSHDKGCRSYICDTSYRHS
Sbjct: 1 MAGVKRKLSTESDVHALHKELDEVSCPVCMDHPHNAVLLLCSSHDKGCRSYICDTSYRHS 60
Query: 61 NCLDRFKKLRDNSKENPNLQSSLINTNNSSGNSFDINFTMQSDMHDVDELLLYQNEINTL 120
NCLDRFKKL S +P +++L + +++ + ++
Sbjct: 61 NCLDRFKKLHSESANDPTPEANLASREHNNESLYE------------------------- 95
Query: 121 LSVGIPQGSRQGDAQDPSRHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKSSL 180
G A S H + + G + DSE+L+ R +EEE++ SED ++L
Sbjct: 96 ----------HGTASRSSFHRESGNRG--SSWDSESLRRRRRVEEEVE----SEDI-TNL 138
Query: 181 HCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVD 240
CPLCRGTVLGW+VVEE R YL++K RSCSR+SCSF G+Y +LRRHARR HPT+RPSD D
Sbjct: 139 KCPLCRGTVLGWKVVEEVRTYLDHKNRSCSRESCSFTGNYQDLRRHARRTHPTTRPSDTD 198
Query: 241 PTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGNG 300
P+RE+AW++ E QREYGDIVSAI+SA+PGAVVVGDYV+ENGD G RE NG
Sbjct: 199 PSRERAWRRLENQREYGDIVSAIRSAMPGAVVVGDYVIENGDRF-----AGERETGNGGS 253
Query: 301 NVPWLTTTTILFQM---MDNTIEIVR----EPRARSSNGWSRHRRSSDRRRYLWGENLLG 353
+ L TT +LFQM +DN R+ S W HRRSS R YLWGENLLG
Sbjct: 254 D---LWTTLLLFQMIGSLDNGGSSASGSGGGSRSHRSRAWRNHRRSSSDRPYLWGENLLG 310
Query: 354 LQD---NEVEEDLRIFNELVEDASHVPRRRRRLNRTRSNEDH 392
LQD N +E+ R+ N+ ++ VPRRRRR R RS+ +H
Sbjct: 311 LQDERNNNDDEEFRLQNDAGGASTPVPRRRRRFGRPRSSGNH 352
>At3g25910 unknown protein
Length = 372
Score = 198 bits (504), Expect = 4e-51
Identities = 124/382 (32%), Positives = 192/382 (49%), Gaps = 60/382 (15%)
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSKENPN 78
+E +E CP+CM+HPHN +LL+CSS++ GCR Y+CDTS+RHSNC D+F+K SKE P+
Sbjct: 36 KEWEEARCPVCMEHPHNGILLICSSYENGCRPYMCDTSHRHSNCFDQFRKA---SKEKPS 92
Query: 79 LQSSLINTNNSSGNSFDINFTMQSDMHDVDELLLYQNEINTLLSVGIPQGSRQGDAQDPS 138
L SL+ S ++ + SD V+ L +EI + +G + + ++
Sbjct: 93 L--SLLREEEESNEPTEME-DVDSDSTAVNLLGEAASEITVVDLSDGERGEEEVEEEEEE 149
Query: 139 RHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKSSLHCPLCRGTVLGWEVVEEA 198
+++ +EGI+ T + + ++ L CPLCRG + W VV+ A
Sbjct: 150 VVVEEEEEGIVTTEEDQE-----------------KNKPQKLTCPLCRGHIKEWVVVKAA 192
Query: 199 RNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPTREQAWQQFERQREYGD 258
R ++N+K RSCS ++C F+G Y +LR+HAR +HP RPS+ DP R+++W++ ERQ + GD
Sbjct: 193 RCFMNSKHRSCSCETCDFSGSYSDLRKHARLLHPGVRPSEADPERQRSWRRLERQSDLGD 252
Query: 259 IVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGNGNVPWLTTTTILFQMMDNT 318
++S +QS+ GG E SN +G +L T L +
Sbjct: 253 LLSTLQSSF-----------------------GGDEISNDDG---FLFADTRLLTVYFLI 286
Query: 319 IEIVREPRARSSNGWS---------RHRRSSDRRRYLWGENLLGLQDNEVEEDLRIFNEL 369
E S+ WS RR S R LWGE+ G ++ N+
Sbjct: 287 RVFRPESSGSRSSSWSGTSRARTHTSGRRRSSRPASLWGESYEGNTGTSPRDEEN--NQS 344
Query: 370 VEDASHVPRRRRRLNRTRSNED 391
++ RRRR RT ++D
Sbjct: 345 SDEQVSGTRRRRSRRRTVIDDD 366
>At4g31410 unknown protein
Length = 308
Score = 139 bits (351), Expect = 2e-33
Identities = 76/164 (46%), Positives = 95/164 (57%), Gaps = 10/164 (6%)
Query: 182 CPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDP 241
CPLCRG V GW VVEEAR L+ KKR C + C F G YLELR+HA+ HP SRPS++DP
Sbjct: 102 CPLCRGEVTGWLVVEEARLRLDEKKRCCEEERCRFMGTYLELRKHAQSEHPDSRPSEIDP 161
Query: 242 TREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGGREGSNGNGN 301
R+ W+ F++ E D++S I S +P VV+GDYV+E GD G E N
Sbjct: 162 ARKLDWENFQQSSEIIDVLSTIHSEVPRGVVLGDYVIEYGDD----DTGDEFEDVPNNEG 217
Query: 302 VPWLTTTTILFQMMDNTIEIVREPRARSSNGWSRHRRSSDRRRY 345
W T+ IL+QM DN +R R R + S RR S R Y
Sbjct: 218 NWW--TSCILYQMFDN----IRNARNRRRSRMSESRRGSRRSSY 255
Score = 75.9 bits (185), Expect = 4e-14
Identities = 28/45 (62%), Positives = 38/45 (84%)
Query: 22 DEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRF 66
D+++CPIC+D PHN VLL CSS+ GCR+++C+T + HSNCLDRF
Sbjct: 30 DDLTCPICLDFPHNGVLLQCSSYGNGCRAFVCNTDHLHSNCLDRF 74
>At4g08460 unknown protein
Length = 274
Score = 124 bits (310), Expect = 1e-28
Identities = 53/113 (46%), Positives = 73/113 (63%)
Query: 171 NSSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRV 230
+ D L CPLCRG V GW VVE+ R YLN+KKRSC D C F G Y +L++H +
Sbjct: 96 DEKSDKPPELLCPLCRGQVKGWTVVEKERKYLNSKKRSCMNDECLFYGSYRQLKKHVKEN 155
Query: 231 HPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDG 283
HP ++P +DP E W++ E +RE D++S + S+ PGA+V GDYV+E +G
Sbjct: 156 HPRAKPRAIDPVLEAKWKKLEVERERSDVISTVMSSTPGAMVFGDYVIEPYNG 208
Score = 72.0 bits (175), Expect = 5e-13
Identities = 29/56 (51%), Positives = 42/56 (74%), Gaps = 2/56 (3%)
Query: 24 VSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKK--LRDNSKENP 77
V+CP+C++ PHN+V+LLCSS+ KGCR Y+C T R SNCL+++KK +D + P
Sbjct: 47 VTCPVCLEVPHNSVVLLCSSYHKGCRPYMCATGNRFSNCLEQYKKAYAKDEKSDKP 102
>At1g77770 unknown protein
Length = 265
Score = 119 bits (297), Expect = 4e-27
Identities = 49/108 (45%), Positives = 76/108 (70%)
Query: 172 SSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVH 231
+ + L CPLCRG V GW VV++AR + N+K+R+C +D+CSF G++ +L++H + H
Sbjct: 77 NENSGQPELLCPLCRGQVKGWTVVKDARMHFNSKRRTCMQDNCSFLGNFRKLKKHMKEKH 136
Query: 232 PTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLE 279
P + P +DP E W++ ER+R+ D++S I S+ PGAVV+GDYV+E
Sbjct: 137 PHACPRAIDPALETKWKRLERERDRRDVISTIMSSTPGAVVLGDYVIE 184
Score = 73.6 bits (179), Expect = 2e-13
Identities = 29/48 (60%), Positives = 39/48 (80%)
Query: 25 SCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDN 72
+CP+C++ PHNAVLLLCSS+ KGCR Y+C TS R +NCLD+++K N
Sbjct: 30 TCPVCLESPHNAVLLLCSSYHKGCRPYMCATSSRFANCLDQYRKSYGN 77
>At1g68140 unknown protein
Length = 334
Score = 119 bits (297), Expect = 4e-27
Identities = 52/114 (45%), Positives = 78/114 (67%)
Query: 179 SLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSD 238
+L CPLCRG V GW +V+ AR++LN KKR C +++C +AG + ELR+H + HP+++P +
Sbjct: 117 NLTCPLCRGQVKGWTIVQPARDFLNLKKRICMQENCVYAGTFKELRKHMKVDHPSAKPRE 176
Query: 239 VDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLPPGGG 292
VDP EQ W++ E + + D++S I+S +PG VV GDYV+E + G GG
Sbjct: 177 VDPDVEQNWRRLEIEHDRDDVMSTIRSTMPGTVVYGDYVIERNNANGSDSDEGG 230
Score = 84.7 bits (208), Expect = 8e-17
Identities = 36/62 (58%), Positives = 47/62 (75%)
Query: 19 RELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSKENPN 78
R+ + V C +CM+ PHNAVLLLCSSHDKGCR Y+C TS+R+SNCLD++KK K + +
Sbjct: 48 RDWENVICSVCMECPHNAVLLLCSSHDKGCRPYMCGTSFRYSNCLDQYKKASAKLKTSGH 107
Query: 79 LQ 80
Q
Sbjct: 108 QQ 109
>At1g15430 unknown protein
Length = 259
Score = 83.2 bits (204), Expect = 2e-16
Identities = 37/95 (38%), Positives = 56/95 (58%)
Query: 170 DNSSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARR 229
++SS S +LHCP CRG V G AR ++N + R C+ D C F+G Y +L+ H +
Sbjct: 97 NSSSRCSGKTLHCPYCRGEVQGTMKSTCARRFMNARPRCCTVDKCDFSGTYAQLKNHLKT 156
Query: 230 VHPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQ 264
HP P +DP + W+Q ER+ EY ++++A Q
Sbjct: 157 EHPGFTPPKLDPWEQHMWEQLEREAEYIEMLNARQ 191
Score = 76.3 bits (186), Expect = 3e-14
Identities = 31/55 (56%), Positives = 42/55 (76%)
Query: 20 ELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSK 74
E ++V C ICM+ PHNAVLL CSS KGCR+Y+CDTS RHSNC ++++ +S+
Sbjct: 47 EWEDVRCVICMEPPHNAVLLQCSSFSKGCRAYMCDTSARHSNCFKQYRRSNSSSR 101
>At1g80220 hypothetical protein
Length = 255
Score = 79.7 bits (195), Expect = 3e-15
Identities = 46/136 (33%), Positives = 67/136 (48%), Gaps = 5/136 (3%)
Query: 170 DNSSEDSKSSLHCPLCRGTVLGWEVVEE-ARNYLNNKKRSCSRDSCSFAGDYLELRRHAR 228
+ + +LHCPLCRG VL + + AR ++N K RSC D C F+G Y L +H +
Sbjct: 91 NRKKRSNTKTLHCPLCRGEVLETKKASKTARRFMNAKPRSCPVDGCEFSGTYAHLNKHLK 150
Query: 229 RVHPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQSAIPGAVVVGDYVLENGDGIGRLP 288
H P+ VDP R+ W+ R EY +++SA + IP V + L N +
Sbjct: 151 TEHQGLVPAKVDPQRQSRWEMLVRHAEYVNLMSA--AGIPHMSGVVHHQLPNNHHLPMFQ 208
Query: 289 PG--GGREGSNGNGNV 302
G G + +G NV
Sbjct: 209 VGFSGTMQNLDGPSNV 224
Score = 69.3 bits (168), Expect = 3e-12
Identities = 31/60 (51%), Positives = 41/60 (67%), Gaps = 2/60 (3%)
Query: 22 DEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCLDRFKKLRDNSKENPNLQS 81
++V C ICM+ PH AVLL CSS GCR Y+C TS+RHSNC +F R+N K+ N ++
Sbjct: 43 EDVRCMICMEPPHEAVLLTCSSSLNGCRPYMCGTSFRHSNCFKQF--CRNNRKKRSNTKT 100
>At2g26050 hypothetical protein
Length = 221
Score = 78.6 bits (192), Expect = 6e-15
Identities = 36/85 (42%), Positives = 52/85 (60%), Gaps = 1/85 (1%)
Query: 179 SLHCPLCRGTVLGW-EVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPS 237
+LHCPLCRG V +V AR ++N K RSCS + C F+G + +L +H + H P
Sbjct: 71 TLHCPLCRGEVSETTKVTSTARRFMNAKPRSCSVEDCKFSGTFSQLTKHLKTEHRGIVPP 130
Query: 238 DVDPTREQAWQQFERQREYGDIVSA 262
VDP R+Q W+ ER EY ++++A
Sbjct: 131 KVDPLRQQRWEMMERHSEYVELMTA 155
Score = 38.9 bits (89), Expect = 0.005
Identities = 13/23 (56%), Positives = 18/23 (77%)
Query: 46 KGCRSYICDTSYRHSNCLDRFKK 68
+GC Y+CDTS RHSNC +F++
Sbjct: 38 RGCHPYMCDTSVRHSNCFKQFRR 60
>At5g38370 similar to unknown protein (pir||T02618)
Length = 100
Score = 39.7 bits (91), Expect = 0.003
Identities = 20/60 (33%), Positives = 31/60 (51%)
Query: 205 KKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPTREQAWQQFERQREYGDIVSAIQ 264
K RSC D C+ + Y +L +H + H VDP + +Q +R EY D++SA +
Sbjct: 16 KLRSCPLDDCNSSETYFQLDKHLKNEHDGLVLLMVDPQSQWRCEQMKRHVEYVDLMSATE 75
>At4g21550 VP1 like protein
Length = 739
Score = 34.3 bits (77), Expect = 0.12
Identities = 27/107 (25%), Positives = 41/107 (38%), Gaps = 24/107 (22%)
Query: 15 HALHRELDEVSCPICMDHPHNAVLLLCSSHDKGCRSYICDTSYRHSNCL----------- 63
H HR D +C IC+ P + HD+ C +CDT+ R L
Sbjct: 578 HPRHR--DGCTCIICIQSPSG----IGPKHDRCCSCAVCDTNKRRRRSLLLRREKKQMEK 631
Query: 64 -DRFKKLRDNSKENPNLQSSLINTNNSSGNS------FDINFTMQSD 103
D +KL + + L S N+ N ++ D+NF + D
Sbjct: 632 EDNARKLLEQLNSDNGLHQSANNSENHERHASPLKVQLDLNFKPEKD 678
>At4g33800 unknown protein
Length = 170
Score = 33.5 bits (75), Expect = 0.21
Identities = 19/51 (37%), Positives = 27/51 (52%), Gaps = 8/51 (15%)
Query: 328 RSSNGWSRHRRSSDRRRYLWGENLLGLQDNEVEEDLRIFNELVEDASHVPR 378
R SNG+ RRS D W +N + ++ E E+DL ++ DAS PR
Sbjct: 29 RDSNGFDSRRRSKDS----WDQNYVHQEEEEEEDDL----SMISDASSGPR 71
>At1g22060 hypothetical protein
Length = 1999
Score = 33.1 bits (74), Expect = 0.27
Identities = 44/156 (28%), Positives = 65/156 (41%), Gaps = 32/156 (20%)
Query: 48 CRSYICDTSYRH---SNCLDRFK-KLRDNSKENPNLQSSLINTNN-SSGNSFDINF---- 98
CR D + H +N FK K RDNS L + N+ SG FD++
Sbjct: 175 CRISPSDETLSHVDKTNIRGSFKEKFRDNS-----LVEETVGLNDLDSGLGFDVSSNTSG 229
Query: 99 TMQSDMHDVDELLLYQNEINTLLSVGIPQGSRQGDAQDPSRHLD----QHDEGI------ 148
++ ++ HD+ + NE+++L SV G G AQ P + D QH G
Sbjct: 230 SLNAEKHDISSI----NEVDSLKSV--VSGDLSGLAQSPQKEKDSLGWQHGWGSDYLGKN 283
Query: 149 --LETADSENLQDRAVLEEELDVDNSSEDSKSSLHC 182
L A +N + + LE+ N + SSL C
Sbjct: 284 SDLGNAIEDNNKLKGFLEDMESSINEIKIEVSSLQC 319
>At5g44180 unknown protein
Length = 1694
Score = 31.6 bits (70), Expect = 0.79
Identities = 31/101 (30%), Positives = 43/101 (41%), Gaps = 3/101 (2%)
Query: 277 VLENGDGIGRLPPGGGREGSNGNGNVPWLTTTTILFQMMDNTIEIVREPRARSSNGWSRH 336
V +G G GR P G GR + GNG P ++ N E + PRA+ G
Sbjct: 1463 VSSSGRG-GRPPRGRGRPRARGNGKKPAVSVKPPRGAANSNG-ETMLRPRAQPRGGRKNG 1520
Query: 337 RRSSDRRRYLWGENLLGLQDNEVEEDLRIFNELVEDASHVP 377
RRS + R + LG+ NEV R+ V + +P
Sbjct: 1521 RRSGTKGRKRPTQGTLGI-CNEVGGGRRVKEVAVTAKTSLP 1560
>At3g47260 putative protein
Length = 820
Score = 31.6 bits (70), Expect = 0.79
Identities = 42/192 (21%), Positives = 80/192 (40%), Gaps = 24/192 (12%)
Query: 84 INTNNSSGNSFDINFTMQSDMHDVDELLLYQNEINTLLSVGIPQGSRQGDAQDPSRHLDQ 143
IN S D T+ D++ EL I +L + + + + Q +D R Q
Sbjct: 370 INNKISDSEKLDRILTILEDLNKRVEL------IERILDIRMEEKNNQRSEEDEERK--Q 421
Query: 144 HDEGILETADSEN---LQDRAVLEEELDVDNSSEDSKSSLHCPLCRGTV---LGWEVVEE 197
DEG+ ++E L+ +A + E D+ E + P + +G E+VEE
Sbjct: 422 EDEGVERQPEAEEEGGLERKAENDNESFEDSIREPNTQYGSYPGDDENIQRDVGDELVEE 481
Query: 198 ARNYLNNKKRSCSRDSCSFAGDYLEL----------RRHARRVHPTSRPSDVDPTREQAW 247
+ ++ RS + + + + L++ R +RV P+ D TR+Q
Sbjct: 482 SSKDMSPTPRSSTLNFNILSEESLDVQKDKKRVSRGRNENKRVKPSVHAEDNFKTRKQMP 541
Query: 248 QQFERQREYGDI 259
++ ++Q + D+
Sbjct: 542 RKRQKQVDSADV 553
>At1g32840 putative protein FARE2.4 (cds3)
Length = 611
Score = 31.6 bits (70), Expect = 0.79
Identities = 43/195 (22%), Positives = 76/195 (38%), Gaps = 41/195 (21%)
Query: 84 INTNNSSGNSFDINFTMQSDMHDVDELLLYQNEINTLLSVGIPQGSRQGDAQDPSR---- 139
IN S D T+ D++ EL I +L + + + + Q +D R
Sbjct: 136 INNKISDSEKLDRILTILEDLNKRVEL------IERILDIRMEEKNNQRSEEDEERKQEH 189
Query: 140 -----HLDQHDEGILETA---DSENLQDRAVLEEEL-------DVDNSSEDSKSSLHCPL 184
L+ +EG LE D+E+ +D ++ E D +N+ D KS P
Sbjct: 190 EEVERQLEAEEEGGLERKAENDNESFED-SIREPNTQYGSYPGDDENTQRDDKS----PT 244
Query: 185 CRGTVLGWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPTRE 244
R + + ++ E + K+ SR R +RV P+ D TR+
Sbjct: 245 PRSSTPSFNILSEESLDVQKDKKRVSRG-----------RNENKRVKPSVHAEDNLKTRK 293
Query: 245 QAWQQFERQREYGDI 259
Q ++ ++Q + D+
Sbjct: 294 QVPRKRQKQVDSADV 308
>At5g55100 unknown protein
Length = 844
Score = 30.8 bits (68), Expect = 1.3
Identities = 25/94 (26%), Positives = 36/94 (37%), Gaps = 15/94 (15%)
Query: 145 DEGILETADSENLQDRAVLEEELDVDNSSEDSKSSLHCPLCRGTVLGWEVVEEARNYLNN 204
D+G E N + R+ +E+ ++ E+ S EV EE + +
Sbjct: 607 DDGNSERRSKRNYRSRSQRDEDGKMEQGEEEESSMD------------EVTEETKT---D 651
Query: 205 KKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSD 238
KK SCSR Y RH+R H SD
Sbjct: 652 KKHSCSRKRHKHKTRYSSKDRHSRDKHKHESSSD 685
>At3g52250 putative protein
Length = 1677
Score = 30.8 bits (68), Expect = 1.3
Identities = 35/119 (29%), Positives = 49/119 (40%), Gaps = 21/119 (17%)
Query: 138 SRHLDQHDEGILETADSENLQDRAVLEEELDVDNSSEDSKSSLHCPLCRGTVLGW----- 192
S HL++ I E + + +L E L D+S SS+ T+L W
Sbjct: 488 SIHLERFPINIEELDNISMERFGCLLNELLGTDDSGTGDSSSVQLT-SMNTLLAWKGEIL 546
Query: 193 ---EVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPTSRPSDVDPT--REQA 246
E+ E + L NK R+ LE RRH+R V P+S D D +EQA
Sbjct: 547 KAVEMTESEIDLLENKHRTLK----------LEGRRHSRVVGPSSYCCDGDANVPKEQA 595
>At3g03550 unknown protein
Length = 356
Score = 30.8 bits (68), Expect = 1.3
Identities = 33/138 (23%), Positives = 50/138 (35%), Gaps = 15/138 (10%)
Query: 177 KSSLHCPLCRGTVL---GWEVVEEARNYLNNKKRSCSRDSCSFAGDYLELRRHARRVHPT 233
KS +CPLCR ++ E+V+ + + S S S L+L R
Sbjct: 192 KSHSNCPLCRAFIVTSSAVEIVDLTNQQIVTENNSISTGDDSVVVVNLDLENSRSRNETV 251
Query: 234 SRPSDVDPTREQAWQQFERQR-----------EYGDIVSAIQSAIPGAVVVGDYVLENGD 282
+ S P Q + E +R DI+ I+ A V +E G+
Sbjct: 252 NEGSTPKPPEMQDSRDGEERRSASLNSGGGVVSIADILREIEDDEESAGVGTSRWVEEGE 311
Query: 283 GIGRLPPGGGREGSNGNG 300
G + PP G + NG
Sbjct: 312 G-EKTPPPSGSAANQTNG 328
>At2g15190 putative protein on transposon FARE2.7 (cds3)
Length = 775
Score = 30.0 bits (66), Expect = 2.3
Identities = 45/197 (22%), Positives = 81/197 (40%), Gaps = 34/197 (17%)
Query: 84 INTNNSSGNSFDINFTMQSDMHDVDELLLYQNEINTLLSVGIPQGSRQGDAQDPSRHLDQ 143
IN S+ D T+ D++ EL I +L + + + + Q +D R Q
Sbjct: 331 INNKISNSEKLDRILTILEDLNKRVEL------IERILDIRMEEKNNQRSEEDEERK--Q 382
Query: 144 HDEGILETADSENLQDRAVLEEELDVDNSS-EDSKSSLHCPLCRGTV----------LGW 192
DEG+ ++E + LE + + DN S EDS + G+ +G
Sbjct: 383 EDEGVERQPEAE---EEGGLERKAENDNESFEDSIREPNTQY--GSYPGDDENTQRDVGD 437
Query: 193 EVVEEARNYLNNKKRSCSRDSCSFAGDYLEL----------RRHARRVHPTSRPSDVDPT 242
EV EE+ + RS + + + + L++ R +RV P+ D T
Sbjct: 438 EVAEESSKDKSPTPRSSTPNFNILSEESLDVQKDKKRVSRGRNENKRVKPSVHAEDNLKT 497
Query: 243 REQAWQQFERQREYGDI 259
R+Q ++ ++Q + D+
Sbjct: 498 RKQVPRKRQKQVDSADV 514
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.133 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,833,748
Number of Sequences: 26719
Number of extensions: 460554
Number of successful extensions: 1671
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 33
Number of HSP's that attempted gapping in prelim test: 1637
Number of HSP's gapped (non-prelim): 57
length of query: 393
length of database: 11,318,596
effective HSP length: 101
effective length of query: 292
effective length of database: 8,619,977
effective search space: 2517033284
effective search space used: 2517033284
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC130200.9


