

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130200.5 - phase: 0
(312 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
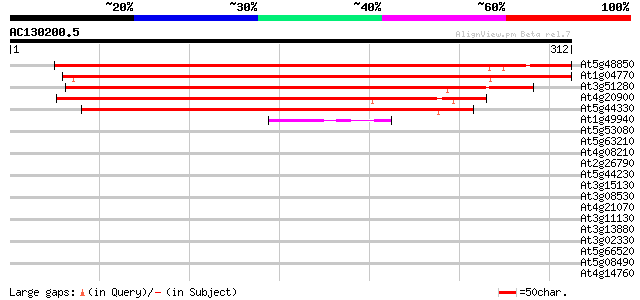
Score E
Sequences producing significant alignments: (bits) Value
At5g48850 unknown protein 385 e-107
At1g04770 unknown protein 325 2e-89
At3g51280 MS5 like protein 275 2e-74
At4g20900 putative protein 243 1e-64
At5g44330 male sterility MS5; pollenless3 213 1e-55
At1g49940 unknown protein 41 7e-04
At5g53080 unknown protein 37 0.014
At5g63210 putative protein 36 0.023
At4g08210 putative protein 35 0.068
At2g26790 putative salt-inducible protein 34 0.089
At5g44230 selenium-binding protein-like 33 0.26
At3g15130 hypothetical protein 32 0.34
At3g08530 hypothetical protein 32 0.34
At4g21070 putative protein (fragment) 32 0.44
At3g11130 hypothetical protein 31 0.76
At3g13880 hypothetical protein 31 0.99
At3g02330 hypothetical protein 31 0.99
At5g66520 selenium-binding protein-like 30 1.3
At5g08490 selenium-binding protein-like 30 1.3
At4g14760 centromere protein homolog 30 1.3
>At5g48850 unknown protein
Length = 306
Score = 385 bits (989), Expect = e-107
Identities = 190/292 (65%), Positives = 247/292 (84%), Gaps = 7/292 (2%)
Query: 26 KSSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKD 85
KS+ K ++L+HVIHKVP GDTPYV+AKHAQL++K+PE+AIV+FWKAIN GD+VDSALKD
Sbjct: 17 KSNLMKDDELFHVIHKVPCGDTPYVRAKHAQLIEKNPEMAIVWFWKAINTGDRVDSALKD 76
Query: 86 MAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRL 145
MAVVMKQLDR+EEAIEAIKSFR C+K+SQ+SLDNVL+DLYKKCGR+EEQ+ELLKRKLR
Sbjct: 77 MAVVMKQLDRSEEAIEAIKSFRPRCSKNSQDSLDNVLIDLYKKCGRMEEQVELLKRKLRQ 136
Query: 146 IYQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMI 205
IYQGEAFNG+ TKTARSHGKKFQV+++QE +RLLGNLGWAYMQ+ Y+ AE V++KAQM+
Sbjct: 137 IYQGEAFNGKPTKTARSHGKKFQVTVQQEISRLLGNLGWAYMQQAKYLSAEAVYRKAQMV 196
Query: 206 DADANKALNLALCLMRQSRYEEAYLILEQVLQGKLPGSDEIKSRNRAEELLVELSANLPQ 265
+ DANK+ NLA+CL++Q R+EE L+L+ VL+ ++ G+D+ ++R RAEELL EL ++LP+
Sbjct: 197 EPDANKSCNLAMCLIKQGRFEEGRLVLDDVLEYRVLGADDCRTRQRAEELLSELESSLPR 256
Query: 266 ---PKFMDDLG--LDDDLLKGIDGLLNVWSPTRSRRLPIFEEISSFRDQLAC 312
+ D LG LDDD + G++ + + + +S+RLPIFE+ISSFR+ L C
Sbjct: 257 MRDAEMEDVLGNILDDDFVLGLEEMTS--TSFKSKRLPIFEQISSFRNTLVC 306
>At1g04770 unknown protein
Length = 303
Score = 325 bits (833), Expect = 2e-89
Identities = 168/294 (57%), Positives = 223/294 (75%), Gaps = 11/294 (3%)
Query: 30 GKKED----LYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKD 85
G+++D Y+V+HK+P+GD+PYV+AKH QLV+KD E AI FW AI A D+VDSALKD
Sbjct: 10 GERQDSSAAAYNVVHKLPHGDSPYVRAKHVQLVEKDAEAAIELFWIAIKARDRVDSALKD 69
Query: 86 MAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRL 145
MA++MKQ +RAEEAI+AI+SFR LC++ +QESLDNVL+DLYKKCGR+EEQ+ELLK+KL +
Sbjct: 70 MALLMKQQNRAEEAIDAIQSFRDLCSRQAQESLDNVLIDLYKKCGRIEEQVELLKQKLWM 129
Query: 146 IYQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMI 205
IYQGEAFNG+ TKTARSHGKKFQV++++ET+R+LGNLGWAYMQ +Y AE V++KAQ+I
Sbjct: 130 IYQGEAFNGKPTKTARSHGKKFQVTVEKETSRILGNLGWAYMQLMDYTAAEAVYRKAQLI 189
Query: 206 DADANKALNLALCLMRQSRYEEAYLIL-EQVLQGKLPGSDEIKSRNRAEELLVELSANLP 264
+ DANKA NL CL++Q +++EA IL VL GS + + R +ELL EL
Sbjct: 190 EPDANKACNLCTCLIKQGKHDEARSILFRDVLMENKEGSGDPRLMARVQELLSELKPQEE 249
Query: 265 QP----KFMDDLGLDD-DLLKGIDGLLNVW-SPTRSRRLPIFEEISSFRDQLAC 312
+ ++G+D+ +++G+D + W P R+RRLPIFEEI RDQLAC
Sbjct: 250 EAAASVSVECEVGIDEIAVVEGLDEFVKEWRRPYRTRRLPIFEEILPLRDQLAC 303
>At3g51280 MS5 like protein
Length = 430
Score = 275 bits (704), Expect = 2e-74
Identities = 144/264 (54%), Positives = 188/264 (70%), Gaps = 5/264 (1%)
Query: 32 KEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMK 91
+ + +H IHKVP GD+PYV+AK+ QLV+KDPE AI FWKAINAGD+VDSALKDMA+VMK
Sbjct: 26 QSESFHAIHKVPVGDSPYVRAKNVQLVEKDPERAIPLFWKAINAGDRVDSALKDMAIVMK 85
Query: 92 QLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGEA 151
Q +RAEEAIEAIKS R C+ +QESLDN+LLDLYK+CGR+++QI LLK KL LI +G A
Sbjct: 86 QQNRAEEAIEAIKSLRVRCSDQAQESLDNILLDLYKRCGRLDDQIGLLKHKLFLIQKGLA 145
Query: 152 FNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADANK 211
FNG+ TKTARS GKKFQVS++QE RLLGNLGWA MQ+ N++ AE +++A I D NK
Sbjct: 146 FNGKRTKTARSQGKKFQVSVEQEATRLLGNLGWALMQRDNFVEAEDAYRRALSIAPDNNK 205
Query: 212 ALNLALCLMRQSRYEEAYLILEQVLQGKLPG----SDEIKSRNRAEELLVELSANLPQPK 267
NL +CLM+Q R +EA L +V + G +K+ RA+++L +L + + + +
Sbjct: 206 MCNLGICLMKQGRIDEAKETLRRVKPAVVDGPRGVDSHLKAYERAQQMLNDLGSEMMR-R 264
Query: 268 FMDDLGLDDDLLKGIDGLLNVWSP 291
DD L I G ++W P
Sbjct: 265 GGDDKVEQRRLFDAIFGSSSIWQP 288
>At4g20900 putative protein
Length = 450
Score = 243 bits (620), Expect = 1e-64
Identities = 130/261 (49%), Positives = 173/261 (65%), Gaps = 24/261 (9%)
Query: 27 SSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDM 86
SS ++ D +H++HKVP GD+PYV+AKHAQL+DKDP AI FW AINAGD+VDSALKDM
Sbjct: 42 SSSSERRDPFHIVHKVPSGDSPYVRAKHAQLIDKDPNRAISLFWTAINAGDRVDSALKDM 101
Query: 87 AVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLI 146
AVVMKQL R++E IEAIKSFR LC+ SQ+S+DN+LL+LYKK GR+EE+ LL+ KL+ +
Sbjct: 102 AVVMKQLGRSDEGIEAIKSFRYLCSFESQDSIDNLLLELYKKSGRIEEEAVLLEHKLQTL 161
Query: 147 YQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFK------ 200
QG F GR ++ R GK ++I+QE AR+LGNLGW ++Q NY +AE ++
Sbjct: 162 EQGMGFGGRVSRAKRVQGKHVIMTIEQEKARILGNLGWVHLQLHNYGIAEQHYRFGFVTK 221
Query: 201 ----------KAQMIDADANKALNLALCLMRQSRYEEAYLILEQVLQGKLPGSDE----- 245
+A ++ D NK NLA+CLMR SR EA +L+ V P E
Sbjct: 222 IPNIDYCLVMRALGLERDKNKLCNLAICLMRMSRIPEAKSLLDDVRDS--PAESECGDEP 279
Query: 246 -IKSRNRAEELLVELSANLPQ 265
KS +RA E+L E+ + P+
Sbjct: 280 FAKSYDRAVEMLAEIESKKPE 300
>At5g44330 male sterility MS5; pollenless3
Length = 469
Score = 213 bits (542), Expect = 1e-55
Identities = 115/221 (52%), Positives = 148/221 (66%), Gaps = 3/221 (1%)
Query: 41 KVPYGDTPYVKAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAI 100
+V GD+PYV+AKHAQLV KDP AI FW AINAGD+VDSALKDM VV+KQL+R +E I
Sbjct: 49 RVRTGDSPYVRAKHAQLVSKDPNRAISLFWAAINAGDRVDSALKDMVVVLKQLNRFDEGI 108
Query: 101 EAIKSFRGLCNKHSQESLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGEAFNGRTTKTA 160
EAIKSFR LC SQ+S+DN+LL+LY K GR+ E ELL+ KLR + Q + + GR
Sbjct: 109 EAIKSFRYLCPFESQDSIDNLLLELYMKSGRITEVAELLEHKLRTLEQDKHYGGRIKIAK 168
Query: 161 RSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQMIDADANKALNLALCLM 220
RSH ++ +I+QE AR+LGNL W ++Q NY +AE ++ A ++ D NK NLA+CL+
Sbjct: 169 RSHEEQNNKTIEQEKARILGNLAWVHLQLHNYGIAEQYYRNALSLEPDNNKLCNLAICLI 228
Query: 221 RQSRYEEAYLILEQVLQ---GKLPGSDEIKSRNRAEELLVE 258
R R EA +LE V Q + KS RA E+L E
Sbjct: 229 RMERTHEAKSLLEDVKQSLGNQWKNEPFCKSFERATEMLAE 269
>At1g49940 unknown protein
Length = 230
Score = 41.2 bits (95), Expect = 7e-04
Identities = 25/68 (36%), Positives = 37/68 (53%), Gaps = 18/68 (26%)
Query: 145 LIYQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKAQM 204
+IYQ EAFNG+ K ARSHG+ F+ +++ LG+ + +T Q+
Sbjct: 1 MIYQEEAFNGKPAKIARSHGRNFRSRSRRKP------LGYCRVSET------------QV 42
Query: 205 IDADANKA 212
I+ DANKA
Sbjct: 43 IEPDANKA 50
>At5g53080 unknown protein
Length = 564
Score = 37.0 bits (84), Expect = 0.014
Identities = 62/265 (23%), Positives = 108/265 (40%), Gaps = 46/265 (17%)
Query: 21 ISVLKKSSKGKKEDLYHVIHKVPYGDTPYVKAKHAQLVDKD-PEVAIVYFWKAI--NAGD 77
+++L+++ + EDL VP + K + + + P +IV +K I
Sbjct: 255 LTILERNRGSESEDLV-----VPLFSLGKLLLKEGKAAEAEIPFTSIVNIYKKIYGERDG 309
Query: 78 KVDSALKDMAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLD--------LYKKC 129
+V A+ +A A EA++ ++ + + ++DN +L+ L
Sbjct: 310 RVGMAMCSLANAKCSKGDANEAVDIYRNALRIIKDSNYMTIDNSILENMRIDLAELLHFV 369
Query: 130 GRVEEQIELLKRKLRLIYQGEAFNGRTTKTARSHGKKFQVSIKQETARLLGNLGWAYMQK 189
GR +E ELL+ L + E F G+ + +H L NL +Y +
Sbjct: 370 GRGDEGRELLEECLLI---NERFKGKNHPSMATH---------------LINLAASYSRS 411
Query: 190 TNYMMAEVVFKKAQMI---------DADANKALNLALCLMRQSRYEEAYLILEQVL--QG 238
NY+ AE + + I + LNLA+ L + +R EEA I +VL +
Sbjct: 412 KNYVEAERLLRTCLNIMEVSVGSEGQSITFPMLNLAVTLSQLNRDEEAEQIALKVLRIRE 471
Query: 239 KLPGSDEIKSRNRAEELLVELSANL 263
K G D + A + LV + A L
Sbjct: 472 KAFGEDSLPV-GEALDCLVSIQARL 495
>At5g63210 putative protein
Length = 262
Score = 36.2 bits (82), Expect = 0.023
Identities = 23/70 (32%), Positives = 37/70 (52%), Gaps = 1/70 (1%)
Query: 169 VSIKQETARLLGNLGWAYMQKTNYMMAEVVFKKA-QMIDADANKALNLALCLMRQSRYEE 227
VS + + +L NLG Y QK Y +++ +F KA ++ A NL L + + R+EE
Sbjct: 159 VSKEDQAHAILSNLGNLYRQKKQYEVSKAMFSKALELKPGYAPAYNNLGLVFVAERRWEE 218
Query: 228 AYLILEQVLQ 237
A E+ L+
Sbjct: 219 AKSCFEKSLE 228
>At4g08210 putative protein
Length = 686
Score = 34.7 bits (78), Expect = 0.068
Identities = 26/89 (29%), Positives = 34/89 (37%), Gaps = 10/89 (11%)
Query: 104 KSFRGLCNKHSQES---LDNVLLDLYKKCGRVEEQIELLKRKLRL-------IYQGEAFN 153
K GLC K ES L+D+Y KCG ++ + L L I G N
Sbjct: 463 KQIHGLCIKKGYESEPVTATALVDMYVKCGEIDNGVVLFDGMLERDVVSWTGIIVGFGQN 522
Query: 154 GRTTKTARSHGKKFQVSIKQETARLLGNL 182
GR + R K + I+ LG L
Sbjct: 523 GRVEEAFRYFHKMINIGIEPNKVTFLGLL 551
>At2g26790 putative salt-inducible protein
Length = 799
Score = 34.3 bits (77), Expect = 0.089
Identities = 25/77 (32%), Positives = 37/77 (47%), Gaps = 2/77 (2%)
Query: 65 AIVYFWKAINAGDKVDSALKDMAVVMK-QLDRAEEAIEAIKSFRGLCNKHSQESLDNVLL 123
A+ + K + G KV+ + + + ++D EA+E K FR + N NV
Sbjct: 337 ALGFLDKMLGKGLKVNCVIVSLILQCYCKMDMCLEALEKFKEFRDM-NIFLDRVCYNVAF 395
Query: 124 DLYKKCGRVEEQIELLK 140
D K GRVEE ELL+
Sbjct: 396 DALSKLGRVEEAFELLQ 412
>At5g44230 selenium-binding protein-like
Length = 657
Score = 32.7 bits (73), Expect = 0.26
Identities = 35/135 (25%), Positives = 55/135 (39%), Gaps = 18/135 (13%)
Query: 62 PEVAIVYFWKAINAGDKVDSA-LKDMAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDN 120
P+ A+ YF + +G + D + QL ++ A A++ + K D+
Sbjct: 262 PQEALEYFDRMEKSGIRADEVTVAGYISACAQLGASKYADRAVQ----IAQKSGYSPSDH 317
Query: 121 V-----LLDLYKKCGRVEEQIEL---LKRKLRLIYQ----GEAFNGRTTKTAR-SHGKKF 167
V L+D+Y KCG VEE + + + K Y G A +GR + H
Sbjct: 318 VVIGSALIDMYSKCGNVEEAVNVFMSMNNKNVFTYSSMILGLATHGRAQEALHLFHYMVT 377
Query: 168 QVSIKQETARLLGNL 182
Q IK T +G L
Sbjct: 378 QTEIKPNTVTFVGAL 392
>At3g15130 hypothetical protein
Length = 689
Score = 32.3 bits (72), Expect = 0.34
Identities = 26/107 (24%), Positives = 44/107 (40%), Gaps = 22/107 (20%)
Query: 56 QLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAE------------------ 97
++ D PE +V W A+ +G ++ LK + ++ R
Sbjct: 62 KVFDSMPERNVVS-WSALMSGHVLNGDLKGSLSLFSEMGRQGIYPNEFTFSTNLKACGLL 120
Query: 98 EAIEAIKSFRGLCNKHSQESL---DNVLLDLYKKCGRVEEQIELLKR 141
A+E G C K E + N L+D+Y KCGR+ E ++ +R
Sbjct: 121 NALEKGLQIHGFCLKIGFEMMVEVGNSLVDMYSKCGRINEAEKVFRR 167
>At3g08530 hypothetical protein
Length = 1516
Score = 32.3 bits (72), Expect = 0.34
Identities = 21/68 (30%), Positives = 33/68 (47%), Gaps = 5/68 (7%)
Query: 79 VDSALKDMAVVMKQLDRAEEAIEAIKSFRGL-----CNKHSQESLDNVLLDLYKKCGRVE 133
V+ AL ++ V + DR E+I+ SF + KH + V +YKK GR +
Sbjct: 1290 VNEALNEIYVEEEDYDRLRESIDLHDSFDQIGLAQKIEKHELVEMRRVAAYIYKKAGRWK 1349
Query: 134 EQIELLKR 141
+ I L K+
Sbjct: 1350 QSIALSKK 1357
>At4g21070 putative protein (fragment)
Length = 1495
Score = 32.0 bits (71), Expect = 0.44
Identities = 16/24 (66%), Positives = 17/24 (70%), Gaps = 4/24 (16%)
Query: 111 NKHSQESLDNVLLDLYKKCGRVEE 134
N HS NVLLDLY +CGRVEE
Sbjct: 256 NLHSS----NVLLDLYARCGRVEE 275
>At3g11130 hypothetical protein
Length = 1673
Score = 31.2 bits (69), Expect = 0.76
Identities = 28/96 (29%), Positives = 43/96 (44%), Gaps = 11/96 (11%)
Query: 51 KAKHAQLVDKDPEVAIVYFWKAINAGDKVDSALKDMAVVMKQLDRAEEAIEAIKSFRGL- 109
KA H +L+ K VA+ N V+ AL ++ + DR E+I+ SF +
Sbjct: 1423 KAGHLRLI-KPYMVAV-----QSNNVSAVNEALNEIYAEEEDYDRLRESIDLHDSFDQIG 1476
Query: 110 ----CNKHSQESLDNVLLDLYKKCGRVEEQIELLKR 141
KH + V +YKK GR ++ I L K+
Sbjct: 1477 LAQKIEKHELVEMRRVAAYIYKKAGRWKQSIALSKK 1512
>At3g13880 hypothetical protein
Length = 748
Score = 30.8 bits (68), Expect = 0.99
Identities = 12/27 (44%), Positives = 19/27 (69%)
Query: 115 QESLDNVLLDLYKKCGRVEEQIELLKR 141
Q L NVL+D+Y KCG++++ + L R
Sbjct: 182 QVFLINVLIDMYSKCGKLDQAMSLFDR 208
>At3g02330 hypothetical protein
Length = 861
Score = 30.8 bits (68), Expect = 0.99
Identities = 16/55 (29%), Positives = 27/55 (49%), Gaps = 4/55 (7%)
Query: 114 SQESLDNVLLDLYKKCGRVEEQIELLKRKLRLIYQGEAFNGRTTKTARSHGKKFQ 168
S S+ L+D+Y KCG +EE ++ R +Q +G + + H K+ Q
Sbjct: 473 SNSSVGCSLIDMYSKCGMIEEAEKIHSR----FFQRANVSGTMEELEKMHNKRLQ 523
>At5g66520 selenium-binding protein-like
Length = 620
Score = 30.4 bits (67), Expect = 1.3
Identities = 11/20 (55%), Positives = 16/20 (80%)
Query: 121 VLLDLYKKCGRVEEQIELLK 140
VL+D+Y KCG +EE +E+ K
Sbjct: 287 VLIDMYAKCGEMEEALEVFK 306
>At5g08490 selenium-binding protein-like
Length = 849
Score = 30.4 bits (67), Expect = 1.3
Identities = 13/26 (50%), Positives = 15/26 (57%)
Query: 108 GLCNKHSQESLDNVLLDLYKKCGRVE 133
GL + + L N LLD Y KCG VE
Sbjct: 462 GLLHDEEEPKLGNALLDAYAKCGNVE 487
>At4g14760 centromere protein homolog
Length = 1676
Score = 30.4 bits (67), Expect = 1.3
Identities = 20/69 (28%), Positives = 37/69 (52%), Gaps = 7/69 (10%)
Query: 85 DMAVVMKQLDRAEEAIEAIKSFRGLCNKHSQESLDNVLLDLYK-----KCGRVEEQIELL 139
D+ +V +QL EEA+ +++ + +K +E+ D D+Y+ K E+IE L
Sbjct: 1554 DLVIVKRQLKEMEEAVSQLENTNEILSKEIEETGD--ARDIYRKVVVEKSRSGSEKIEQL 1611
Query: 140 KRKLRLIYQ 148
+ K++ I Q
Sbjct: 1612 QNKMQNIEQ 1620
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.135 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,674,539
Number of Sequences: 26719
Number of extensions: 277680
Number of successful extensions: 1232
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 1181
Number of HSP's gapped (non-prelim): 76
length of query: 312
length of database: 11,318,596
effective HSP length: 99
effective length of query: 213
effective length of database: 8,673,415
effective search space: 1847437395
effective search space used: 1847437395
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC130200.5


