

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.3 + phase: 0
(170 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
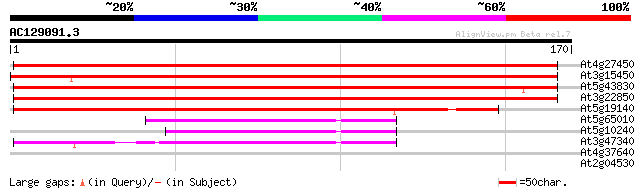
Score E
Sequences producing significant alignments: (bits) Value
At4g27450 unknown protein 257 2e-69
At3g15450 unknown protein 243 5e-65
At5g43830 aluminum-induced protein-like 191 2e-49
At3g22850 unknown protein 188 1e-48
At5g19140 aluminium-induced protein - like 147 4e-36
At5g65010 asparagine synthetase (gb|AAC72837.1) 46 8e-06
At5g10240 asparagine synthetase ASN3 45 2e-05
At3g47340 glutamine-dependent asparagine synthetase 44 4e-05
At4g37640 plasma membrane-type calcium ATPase (ACA2) 27 4.1
At2g04530 unknown protein 26 9.1
>At4g27450 unknown protein
Length = 250
Score = 257 bits (657), Expect = 2e-69
Identities = 119/165 (72%), Positives = 143/165 (86%)
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G+L+NL QLNKQYGL+K NEAMF+IEAYRTLRDRGPYPADQV+K L+GSF+FV+YD K
Sbjct: 85 GSLNNLCQLNKQYGLTKTTNEAMFVIEAYRTLRDRGPYPADQVVKDLDGSFSFVVYDSKA 144
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNF 121
G+VF A GSDG + LYWGIAADGSVVIS++L+++K CAKSFAPFPTGC+FHSE GL++F
Sbjct: 145 GSVFTALGSDGGVKLYWGIAADGSVVISDDLDVIKEGCAKSFAPFPTGCMFHSEGGLMSF 204
Query: 122 EHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWA 166
EHP K+KAMPR+DSEGV+CGANF VD +R +PR GSEANW+
Sbjct: 205 EHPMNKIKAMPRVDSEGVLCGANFKVDVYNRVNSIPRRGSEANWS 249
>At3g15450 unknown protein
Length = 253
Score = 243 bits (619), Expect = 5e-65
Identities = 114/167 (68%), Positives = 139/167 (82%), Gaps = 1/167 (0%)
Query: 1 MGNLHNLSQLNKQYGLS-KGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDH 59
+G L+NL LN+QYGLS K NEAMF+IEAYRTLRDRGPYPADQVL+GLEGSFAFV+YD
Sbjct: 82 LGRLNNLCTLNRQYGLSGKNSNEAMFVIEAYRTLRDRGPYPADQVLRGLEGSFAFVVYDT 141
Query: 60 KDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLL 119
+ +VF+A SDG LYWGI+ DGSVV+S++++++K CAKSFAPFP GC+FHSE GL
Sbjct: 142 QTSSVFSALSSDGGESLYWGISGDGSVVMSDDIQIIKQGCAKSFAPFPNGCMFHSETGLK 201
Query: 120 NFEHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWA 166
+F+HPT MKAMPRIDSEGV+CGA+F VD+ S+ +PR GSEANWA
Sbjct: 202 SFDHPTNMMKAMPRIDSEGVLCGASFKVDACSKINSIPRRGSEANWA 248
>At5g43830 aluminum-induced protein-like
Length = 251
Score = 191 bits (485), Expect = 2e-49
Identities = 87/167 (52%), Positives = 119/167 (71%), Gaps = 2/167 (1%)
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G++ NL L +QYGL+K NEA+ +IEAYRTLRDRGPYP D+V++ G FAF+++D
Sbjct: 82 GHIENLPFLKQQYGLNKITNEAIIVIEAYRTLRDRGPYPVDKVVRDFHGKFAFILFDSVK 141
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNF 121
TVFAA+ +DG + +WG A+G +V S+N E+VK CAKS+ PFP GC F S GL +F
Sbjct: 142 KTVFAAADADGSVPFFWGTDAEGHLVFSDNTEMVKKGCAKSYGPFPKGCFFTSSGGLRSF 201
Query: 122 EHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQ--MMPRVGSEANWA 166
EHP ++K +PR+DS G +CGA F VD++++ + MPRV S NWA
Sbjct: 202 EHPKNELKPVPRVDSSGDVCGATFKVDAETKREGTKMPRVDSSQNWA 248
>At3g22850 unknown protein
Length = 248
Score = 188 bits (478), Expect = 1e-48
Identities = 83/165 (50%), Positives = 115/165 (69%)
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G++ N+ L +QYGL+K E +IEAYRTLRDRGPY A+QV++ +G F F++YD
Sbjct: 81 GHIENVPILKQQYGLTKTATEVTIVIEAYRTLRDRGPYSAEQVVRDFQGKFGFMLYDCST 140
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHGLLNF 121
VF A DG + LYWG A+G +V+S+++E VK C KSFAPFP GC F S GL ++
Sbjct: 141 QNVFLAGDVDGSVPLYWGTDAEGHLVVSDDVETVKKGCGKSFAPFPKGCFFTSSGGLRSY 200
Query: 122 EHPTKKMKAMPRIDSEGVMCGANFNVDSQSRNQMMPRVGSEANWA 166
EHP+ ++K +PR+DS G +CG F VDS+++ + MPRVGS NW+
Sbjct: 201 EHPSNELKPVPRVDSSGEVCGVTFKVDSEAKKEAMPRVGSVQNWS 245
>At5g19140 aluminium-induced protein - like
Length = 234
Score = 147 bits (370), Expect = 4e-36
Identities = 71/148 (47%), Positives = 99/148 (65%), Gaps = 3/148 (2%)
Query: 2 GNLHNLSQLNKQYGLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G+L NL L +QYGL+K NE + +IEAY+TLRDR PYPA+ V+ L G FAFV++D
Sbjct: 82 GSLDNLGSLKQQYGLAKNANEVLLVIEAYKTLRDRAPYPANHVVAHLSGDFAFVVFDKST 141
Query: 62 GTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSE-HGLLN 120
T+F AS G + LYWGI ADG V +++++L+K +C KS A FP GC + + GL +
Sbjct: 142 STLFVASDQVGKVPLYWGITADGYVAFADDVDLLKGACGKSLASFPQGCYYSTALGGLRS 201
Query: 121 FEHPTKKMKAMPRIDSEGVMCGANFNVD 148
FE+P K+ A+P +EG + GA F V+
Sbjct: 202 FENPKNKITAVPA--NEGEIWGATFKVE 227
>At5g65010 asparagine synthetase (gb|AAC72837.1)
Length = 578
Score = 46.2 bits (108), Expect = 8e-06
Identities = 25/76 (32%), Positives = 41/76 (53%), Gaps = 1/76 (1%)
Query: 42 DQVLKGLEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAK 101
++ + L+G FAFV+ D +D + AA + G LY G DGSV + ++ + C +
Sbjct: 111 EEFIDMLDGMFAFVLLDTRDKSFIAARDAIGITPLYIGWGLDGSVWFASEMKALSDDC-E 169
Query: 102 SFAPFPTGCLFHSEHG 117
F FP G ++ S+ G
Sbjct: 170 QFMSFPPGHIYSSKQG 185
>At5g10240 asparagine synthetase ASN3
Length = 578
Score = 45.1 bits (105), Expect = 2e-05
Identities = 25/70 (35%), Positives = 38/70 (53%), Gaps = 1/70 (1%)
Query: 48 LEGSFAFVIYDHKDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFP 107
L+G FAFV+ D +D + AA + G LY G DGSV + ++ + C + F FP
Sbjct: 117 LDGMFAFVLLDTRDKSFIAARDAIGITPLYIGWGLDGSVWFASEMKALSDDC-EQFMCFP 175
Query: 108 TGCLFHSEHG 117
G ++ S+ G
Sbjct: 176 PGHIYSSKQG 185
>At3g47340 glutamine-dependent asparagine synthetase
Length = 584
Score = 43.9 bits (102), Expect = 4e-05
Identities = 36/118 (30%), Positives = 54/118 (45%), Gaps = 10/118 (8%)
Query: 2 GNLHNLSQLNKQYGLSK--GGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDH 59
G ++N +L K+ K G++ I Y Y D V L+G F+FV+ D
Sbjct: 76 GEIYNHEELRKRLKNHKFRTGSDCEVIAHLYEE------YGVDFV-DMLDGIFSFVLLDT 128
Query: 60 KDGTVFAASGSDGHIGLYWGIAADGSVVISENLELVKASCAKSFAPFPTGCLFHSEHG 117
+D + A + G LY G DGSV IS ++ + C + F FP G + S+ G
Sbjct: 129 RDNSFMVARDAIGVTSLYIGWGLDGSVWISSEMKGLNDDC-EHFETFPPGHFYSSKLG 185
>At4g37640 plasma membrane-type calcium ATPase (ACA2)
Length = 1014
Score = 27.3 bits (59), Expect = 4.1
Identities = 21/78 (26%), Positives = 36/78 (45%), Gaps = 2/78 (2%)
Query: 18 KGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKDGTVFAASGSDGHIGLY 77
K E + +I + + P ++K L +F V+ DGT A + + IGL
Sbjct: 712 KNQEELLELIPKIQVMARSSPMDKHTLVKQLRTTFDEVVAVTGDGTNDAPALHEADIGLA 771
Query: 78 WGIAADGSVVISENLELV 95
GIA G+ V E+ +++
Sbjct: 772 MGIA--GTEVAKESADVI 787
>At2g04530 unknown protein
Length = 354
Score = 26.2 bits (56), Expect = 9.1
Identities = 13/47 (27%), Positives = 24/47 (50%), Gaps = 4/47 (8%)
Query: 15 GLSKGGNEAMFIIEAYRTLRDRGPYPADQVLKGLEGSFAFVIYDHKD 61
G+S GG+E I+ + + D G P+ + ++ F F+ + H D
Sbjct: 85 GISVGGHETCVIVPELKCVFDIGRCPS----RAIQQKFLFITHAHLD 127
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.135 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,009,670
Number of Sequences: 26719
Number of extensions: 162183
Number of successful extensions: 283
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 272
Number of HSP's gapped (non-prelim): 10
length of query: 170
length of database: 11,318,596
effective HSP length: 92
effective length of query: 78
effective length of database: 8,860,448
effective search space: 691114944
effective search space used: 691114944
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC129091.3


