

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC129091.14 + phase: 0
(439 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
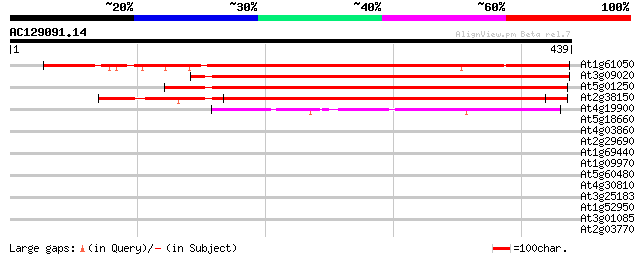
Score E
Sequences producing significant alignments: (bits) Value
At1g61050 hypothetical protein 396 e-110
At3g09020 unknown protein 339 2e-93
At5g01250 unknown protein 332 2e-91
At2g38150 hypothetical protein 315 4e-86
At4g19900 putative protein 131 6e-31
At5g18660 unknown protein 33 0.24
At4g03860 putative transposon protein 32 0.69
At2g29690 anthranilate synthase, alpha subunit 30 2.6
At1g69440 hypothetical protein 29 4.5
At1g09970 leucine-rich repeat receptor-like kinase At1g09970 29 4.5
At5g60480 putative protein 29 5.9
At4g30810 serine carboxypeptidase II like protein 29 5.9
At3g25183 hypothetical protein 29 5.9
At1g52950 replication protein A1, putative 29 5.9
At3g01085 putative protein 28 7.7
At2g03770 putative steroid sulfotransferase 28 7.7
>At1g61050 hypothetical protein
Length = 435
Score = 396 bits (1018), Expect = e-110
Identities = 210/431 (48%), Positives = 283/431 (64%), Gaps = 33/431 (7%)
Query: 27 LHNFKKKILSILFKLPTSFFALLVLFLLAYNAFTVFCIHIPHHYSPHKPL--LLPPPS-- 82
+ K+ I+S +F LP S LL++ LL YN+F+VF +H+ P++P+ L P
Sbjct: 13 VQRLKRLIVSFVFCLPMSLLGLLLMLLLIYNSFSVFSLHLV----PNQPIQSTLSPTHLQ 68
Query: 83 --HTKTKLSSSVFFAHKEDNSP--IINKTHFPLLHKISIHKF---KRVKKHKRGLKTLHS 135
H +T SSSV D+S ++ +T + K ++ K+ ++ KR + +
Sbjct: 69 ILHHQTSTSSSV-----SDSSLLLVVKETSLGFIQKQNVSSTRIEKKTRRFKRSTELTPA 123
Query: 136 DTKF------PLFQKRLGAFFNGNSSSCSCKLRFFMTWISPLKAFGDRELLSVESLFKSH 189
T+ FQ R+ + S SC+ FFMTWIS +++FGDRE ++ESLFK H
Sbjct: 124 ITQRLQVKSRQRFQTRVKSLL----SKSSCESLFFMTWISSIESFGDRERFTIESLFKFH 179
Query: 190 PKACLVIVSKSMDSDKGTQILRPFVKNGFRVIAIEPDFNYIFKNTHAESWFNRLIQGNVN 249
P CL++VS S D D+GT IL+PF G +V+ I+PDF YIFK+T AE WF RL +G ++
Sbjct: 180 PNGCLILVSNSFDCDRGTLILKPFTDKGLKVLPIKPDFAYIFKDTSAEKWFERLKKGTLS 239
Query: 250 PGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGAQNIDVKTKKWSRLN 309
PG I L QNLSNLLRL LLYK+GGIY+D D+II+KS S N IGAQ +D TKKWSRLN
Sbjct: 240 PGVIPLEQNLSNLLRLVLLYKYGGIYLDTDVIILKSLSNLHNVIGAQTVDPVTKKWSRLN 299
Query: 310 NAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVVSR--VSGREGYNFSVVPPS 367
NAVLIFDK HPLL +FI+EF+ TF+GNKWGHNGPYL+SRV++R +S FSV+PPS
Sbjct: 300 NAVLIFDKNHPLLKRFIDEFSRTFNGNKWGHNGPYLVSRVITRIKISSSSDLGFSVLPPS 359
Query: 368 AFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKESYAVHLWNRQSGKLEVVKGSIIDS 427
AFYPVDW IK +R P +E WL K++ +RK ++AVHLWNR+S KL + +GSII
Sbjct: 360 AFYPVDWTRIKGFYRAPTNE-SDAWLRKRLTHLRKNTFAVHLWNRESKKLRIEEGSIIHQ 418
Query: 428 IISSCCIFCNT 438
++S CIFCN+
Sbjct: 419 LMSHSCIFCNS 429
>At3g09020 unknown protein
Length = 411
Score = 339 bits (870), Expect = 2e-93
Identities = 156/297 (52%), Positives = 219/297 (73%), Gaps = 5/297 (1%)
Query: 142 FQKRLGAFFNGNSSSCSCKLRFFMTWISPLKAFGDRELLSVESLFKSHPKACLVIVSKSM 201
FQ+R F + C+++F MTWISP + FG RE+LSVES+FKSH + CL+I+S +M
Sbjct: 115 FQQRATEFLRDD-----CEVKFMMTWISPAELFGKREILSVESVFKSHARGCLMILSSTM 169
Query: 202 DSDKGTQILRPFVKNGFRVIAIEPDFNYIFKNTHAESWFNRLIQGNVNPGEISLGQNLSN 261
DS +G +IL+PF+ G+RV+A+ PD ++ K+T ESW + G +PG+ISL QNLSN
Sbjct: 170 DSLQGFRILKPFLDRGYRVMAVTPDLPFLLKDTAGESWLEEIQTGKRDPGKISLAQNLSN 229
Query: 262 LLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGAQNIDVKTKKWSRLNNAVLIFDKKHPL 321
L+RL+ L+KFGG+Y+D D+I++KSF RN IGAQ ++ ++ W+RLNNAVLIFDK HP
Sbjct: 230 LMRLAYLFKFGGVYLDTDMIVLKSFKTLRNVIGAQTLEPVSRNWTRLNNAVLIFDKNHPF 289
Query: 322 LLKFIEEFALTFDGNKWGHNGPYLISRVVSRVSGREGYNFSVVPPSAFYPVDWRGIKSLF 381
LLK IEEFALTF+GN WGHNGPYL+SRV V G +GYNF+++ P AFYPV+W I+ LF
Sbjct: 290 LLKSIEEFALTFNGNVWGHNGPYLVSRVARAVEGTDGYNFTILTPPAFYPVNWVEIEKLF 349
Query: 382 RGPGDEIHSKWLVKKMVQIRKESYAVHLWNRQSGKLEVVKGSIIDSIISSCCIFCNT 438
+ P E SK + K+++++K SY +HLWN+ S K E+ +GS +D ++S+ CI C++
Sbjct: 350 KVPRTEKDSKRVQVKVLEMQKRSYGLHLWNKFSRKFEIEQGSAMDKLVSNQCIICDS 406
>At5g01250 unknown protein
Length = 407
Score = 332 bits (851), Expect = 2e-91
Identities = 154/315 (48%), Positives = 215/315 (67%), Gaps = 5/315 (1%)
Query: 122 RVKKHKRGLKTLHSDTKFPLFQKRLGAFFNGNSSSCSCKLRFFMTWISPLKAFGDRELLS 181
+V + + ++ D FQKR+ F C++ F MTWISP FG+RE+L+
Sbjct: 91 QVNEKLQVIEVFSGDNLSDKFQKRVNEFVGDG-----CEVNFVMTWISPADFFGNREVLA 145
Query: 182 VESLFKSHPKACLVIVSKSMDSDKGTQILRPFVKNGFRVIAIEPDFNYIFKNTHAESWFN 241
+ES+FKSHP CL+I+S +MDS +G L+PF+ G++V+A+ PD ++ K T E W +
Sbjct: 146 IESVFKSHPYGCLMILSATMDSPQGYATLKPFIDRGYKVLAVTPDLPFLLKGTAGELWLD 205
Query: 242 RLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGAQNIDVK 301
+ G +PG+ISL QNLSNL+RL+ LYK+GG+Y+D D+I++KSF RN IGAQ +D
Sbjct: 206 EIKSGKRDPGKISLAQNLSNLMRLAYLYKYGGVYLDTDMIVLKSFKGLRNVIGAQTLDPS 265
Query: 302 TKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVVSRVSGREGYNF 361
+ W+RLNNAVLIFDK HPLLLKF+EEFA TF+GN WG+NGPYL+SRV V G GYNF
Sbjct: 266 STNWTRLNNAVLIFDKNHPLLLKFMEEFAKTFNGNIWGYNGPYLVSRVARAVEGSSGYNF 325
Query: 362 SVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKESYAVHLWNRQSGKLEVVK 421
+V+ PS FY V+W IK LF+ P E SKW+ K++ +++ Y +HLWN+ S K E+ +
Sbjct: 326 TVMRPSVFYSVNWLEIKKLFKVPKTEKDSKWVKTKLLHMQRNGYGLHLWNKFSRKYEIEQ 385
Query: 422 GSIIDSIISSCCIFC 436
GS + ++S CI C
Sbjct: 386 GSAMWKLVSEHCIIC 400
>At2g38150 hypothetical protein
Length = 736
Score = 315 bits (806), Expect = 4e-86
Identities = 158/372 (42%), Positives = 234/372 (62%), Gaps = 17/372 (4%)
Query: 70 YSPHKPLLLP-PPSHTKTKLSSSVFFAHKEDNSPIINKTHFPLLHKISIHKFKRVKKHKR 128
+ P+ PL P P + ++S V HKE K PLL K +R+ +R
Sbjct: 369 FKPNVPLPRPYGPMSSSCNINSVVDSEHKE-------KELDPLLPPRKASKNQRIDWFRR 421
Query: 129 GL---KTLHSDTKFPLFQKRLGAFFNGNSSSCSCKLRFFMTWISPLKAFGDRELLSVESL 185
L + L S TK F R+ +N N C +FFM W+SP +FG RE+L++++L
Sbjct: 422 KLPELEILKSTTKSKSFHTRVLDLYNKN-----CSAQFFMIWLSPANSFGPREMLAIDTL 476
Query: 186 FKSHPKACLVIVSKSMDSDKGTQILRPFVKNGFRVIAIEPDFNYIFKNTHAESWFNRLIQ 245
F ++P ACL I+S S+DS G IL+P GF +IA+ D ++ KNT AE+W RL
Sbjct: 477 FTTNPGACLAILSNSLDSPNGYTILKPLFDQGFNLIAVTIDIPFLVKNTPAEAWLKRLKS 536
Query: 246 GNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGAQNIDVKTKKW 305
GN++PG I L NLS+L RL++LYK+GG+Y+D DII + + RN IGAQ+ D TK+W
Sbjct: 537 GNMDPGSIPLFMNLSDLTRLAVLYKYGGVYLDTDIIFLNDMTGLRNAIGAQSSDPATKRW 596
Query: 306 SRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLISRVVSRVSGREGY-NFSVV 364
+RLNNAV++FD HPL+ +F++E+A TFDGNKWG+N PYL+SRV+ R+ + GY N ++
Sbjct: 597 TRLNNAVMVFDIYHPLMREFLQEYATTFDGNKWGYNSPYLVSRVIKRLGNKPGYNNLTIF 656
Query: 365 PPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKESYAVHLWNRQSGKLEVVKGSI 424
P AFYPV+W I+ LF+ P +KW+ K + + K SY +HLWN+ + K+++ +GS+
Sbjct: 657 SPDAFYPVNWIKIQKLFKKPATTREAKWVEKTVQDMNKGSYMIHLWNKVTRKIKIEEGSV 716
Query: 425 IDSIISSCCIFC 436
+ +++S+ C C
Sbjct: 717 MHTLVSTHCTVC 728
Score = 270 bits (691), Expect = 9e-73
Identities = 129/253 (50%), Positives = 177/253 (68%), Gaps = 1/253 (0%)
Query: 168 ISPLKAFGDRELLSVESLFKSHPKACLVIVSKSMDSDKGTQILRPFVKNGFRVIAIEPDF 227
+S + FG RE+L+VES+FK+HP+ CL+IVS S+DS +G IL+P G++V A PD
Sbjct: 103 VSSDEYFGKREMLAVESVFKAHPQGCLMIVSGSLDSLQGDSILKPLNDRGYKVFAATPDM 162
Query: 228 NYIFKNTHAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFS 287
+ + +NT A+SWF + +PG I L QNLSNL RL+ LYK+GG+Y+D D I+ +SF
Sbjct: 163 SLLLENTPAKSWFQEMKSCKRDPGRIPLHQNLSNLARLAFLYKYGGVYLDTDFIVTRSFK 222
Query: 288 KFRNTIGAQNI-DVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFDGNKWGHNGPYLI 346
+N+IGAQ + + +K W+RLNNAVLIF+K HPL+ FIEEFA TFDGNKWGHNGPYL+
Sbjct: 223 GLKNSIGAQTVVEGDSKNWTRLNNAVLIFEKDHPLVYSFIEEFASTFDGNKWGHNGPYLV 282
Query: 347 SRVVSRVSGREGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIHSKWLVKKMVQIRKESYA 406
+RV R G NF+V+PP AFYP +W I LF+ P S L +V++ +ESY
Sbjct: 283 TRVAQRARETIGDNFTVLPPVAFYPFNWLDIPRLFQTPRGSNDSTLLKTDLVKLNRESYG 342
Query: 407 VHLWNRQSGKLEV 419
+HLWN+ + KL++
Sbjct: 343 LHLWNKITRKLKI 355
>At4g19900 putative protein
Length = 1302
Score = 131 bits (330), Expect = 6e-31
Identities = 88/282 (31%), Positives = 142/282 (50%), Gaps = 23/282 (8%)
Query: 159 CKLRFFMTWISPLKAFGDRELLSVESLFKSHPKACLVIVSKSMDSDKGTQILRPFVKNGF 218
C +R FM W SP F R +ESL H AC+V+ S++++ D FVK+ +
Sbjct: 368 CSMRVFMVWNSPGWMFSVRHQRGLESLLSQHRDACVVVFSETVELDF---FRNSFVKDSY 424
Query: 219 RVIAIEPDFNYIFKNT----HAESWFNRLIQGNVNPGEISLGQNLSNLLRLSLLYKFGGI 274
+V P+ + + ++T A WF+ + P + S L+RL+ LYK+GG+
Sbjct: 425 KVAVAMPNLDELLQDTPTHVFASVWFDWR-KTKFYP------THYSELVRLAALYKYGGV 477
Query: 275 YIDADIIIMKSFSKFRNTIGAQNIDVKTKKWSRLNNAVLIFDKKHPLLLKFIEEFALTFD 334
Y+D+D+I++ S S RNTIG ++ LN AV+ F+KK P LL+ + E+ LT+D
Sbjct: 478 YLDSDVIVLGSLSSLRNTIGMED----QVAGESLNGAVMSFEKKSPFLLECLNEYYLTYD 533
Query: 335 GNKWGHNGPYLISRVVSR-VSGR----EGYNFSVVPPSAFYPVDWRGIKSLFRGPGDEIH 389
NG L++RV R ++G+ ++ P S F+P++ + I + F P E
Sbjct: 534 DKCLRCNGADLLTRVAKRFLNGKNRRMNQQELNIRPSSVFFPINSQQITNYFAYPAIEDE 593
Query: 390 SKWLVKKMVQIRKESYAVHLWNRQSGKLEVVKGSIIDSIISS 431
+ +I ES H WN + L S++ +ISS
Sbjct: 594 RSQQDESFKKILNESLTFHFWNSVTSSLIPEPESLVAKLISS 635
>At5g18660 unknown protein
Length = 417
Score = 33.5 bits (75), Expect = 0.24
Identities = 62/249 (24%), Positives = 104/249 (40%), Gaps = 27/249 (10%)
Query: 131 KTLHSDTKF-PLFQKR---LGAFFNGNSSSCSCKLRFFMTWISPLKAF--GDRELLSVES 184
KT+ D+KF FQ + L + F+ N SS S L++ + P+ + G E+ + S
Sbjct: 18 KTIFKDSKFISQFQVKSSPLASTFHTNESSTS--LKYKRARLKPISSLDSGISEIATSPS 75
Query: 185 LFKSHPKACLVIVSKSMDSDKGTQILRPFVKNGFRVIAIEPDFNYIFKNTHAESWFNRLI 244
PK V+V S G +++ +K GF VIA+ + + I E +L
Sbjct: 76 FRNKSPKDINVLVVGSTGYI-GRFVVKEMIKRGFNVIAVAREKSGIRGKNDKEETLKQLQ 134
Query: 245 QGNVNPGEISLGQNLSNLLRLSLLYK-FGGIYIDADIIIMKSFSKFRNTIGAQNIDVKTK 303
NV S++ L +L K + D+++ S+ + ID +
Sbjct: 135 GANV---------CFSDVTELDVLEKSIENLGFGVDVVVSCLASRNGGIKDSWKIDYEAT 185
Query: 304 KWSRLNNAVLIFDKKHPLLLKFI--EEFALTFDGNKWGHNGPYLISRVVSRVSGREGYNF 361
K S + A F KH +LL I ++ L F K L+ + S + +
Sbjct: 186 KNSLV--AGKKFGAKHFVLLSAICVQKPLLEFQRAKLKFEAE-LMDLAEQQDS---SFTY 239
Query: 362 SVVPPSAFY 370
S+V P+AF+
Sbjct: 240 SIVRPTAFF 248
>At4g03860 putative transposon protein
Length = 745
Score = 32.0 bits (71), Expect = 0.69
Identities = 43/158 (27%), Positives = 64/158 (40%), Gaps = 33/158 (20%)
Query: 7 HHHNPFLYFPLLLLHKL----HHFLHNFKKKILSILFKLPTSFFALLVLFLLAYNAFTVF 62
HHH LL L K HHF+H+ + + K P+ LLA N T+F
Sbjct: 47 HHHG------LLPLSKETTRPHHFIHS------TSITKSPSP--------LLALNHSTIF 86
Query: 63 CIHIPHHYSPHKPLLLPPPSH--TKTKLSSSVFFAHKEDNSPIINKTHFPLLHKISIHKF 120
I+ + + H PP H T TK +S F + + S N T+ PL ++ +
Sbjct: 87 TINQSNRFLLHSSSKTGPPLHSTTTTKPASLDFTSSLDRVSFFTNHTNSPL----TLSSY 142
Query: 121 KRVKKHKRGLKTLHSDTKFPLFQKR---LGAFFNGNSS 155
K K ++ T F R + + +NG SS
Sbjct: 143 STACKKKLTKQSDEPSTTLLAFAPRSSFIMSNYNGESS 180
>At2g29690 anthranilate synthase, alpha subunit
Length = 621
Score = 30.0 bits (66), Expect = 2.6
Identities = 23/75 (30%), Positives = 31/75 (40%), Gaps = 9/75 (12%)
Query: 59 FTVFCIHIPHHYSPHK--------PLLLPPPSHTKTKLSSSVFFAHKEDNSPIINKTHFP 110
FTV I + HH +PH PL L P T L + H P I ++ P
Sbjct: 14 FTVEAIAVTHHRTPHPPHFPSLRFPLSLKSPPATSLNLVAGSKLLHFSRRLPSIKCSYTP 73
Query: 111 LLHKISIHKFKRVKK 125
L +S +F + KK
Sbjct: 74 SL-DLSEEQFTKFKK 87
>At1g69440 hypothetical protein
Length = 990
Score = 29.3 bits (64), Expect = 4.5
Identities = 31/112 (27%), Positives = 48/112 (42%), Gaps = 20/112 (17%)
Query: 1 MEDHTKHHHN------PFLYFPLLLLHKLHH----------FLHNFKKKILSILFK-LPT 43
ME+ T HHH+ P LLHK +H LH + L+++ LP+
Sbjct: 1 MEEKTHHHHHSTNKHIPSSKSRTPLLHKPYHHHVQTNPPPFLLHPSSHQNLNLVASNLPS 60
Query: 44 SFFALLVLFLLAYNAFTVFCIHIPH--HYSPHKPLLLP-PPSHTKTKLSSSV 92
S++ + + ++ PH SP P LLP PP H+ T+ S+
Sbjct: 61 SYYYYYYCYFYSQFHNSLPPPPPPHLLPLSPPLPPLLPLPPPHSMTRFHKSL 112
>At1g09970 leucine-rich repeat receptor-like kinase At1g09970
Length = 976
Score = 29.3 bits (64), Expect = 4.5
Identities = 13/39 (33%), Positives = 19/39 (48%)
Query: 138 KFPLFQKRLGAFFNGNSSSCSCKLRFFMTWISPLKAFGD 176
+ PL FNGN CS ++ F I+P ++ GD
Sbjct: 568 RIPLSLSSYNGSFNGNPGLCSTTIKSFNRCINPSRSHGD 606
>At5g60480 putative protein
Length = 191
Score = 28.9 bits (63), Expect = 5.9
Identities = 27/86 (31%), Positives = 34/86 (39%), Gaps = 8/86 (9%)
Query: 69 HYSPHKPLLLPPPSHTKTKLSS--SVFFAHKEDNSPIINKTHFPLLHKISIHKFKRVKKH 126
H SP PL L P + L S S FF + K F + + + K +KKH
Sbjct: 64 HRSPPSPLQLQPLAPVPNLLLSLSSGFFGPSDQEV----KNKFTV--ERDVRKTAMIKKH 117
Query: 127 KRGLKTLHSDTKFPLFQKRLGAFFNG 152
KR T K F +R G NG
Sbjct: 118 KRTKFTAEQKVKMRGFAERAGWKING 143
>At4g30810 serine carboxypeptidase II like protein
Length = 479
Score = 28.9 bits (63), Expect = 5.9
Identities = 18/70 (25%), Positives = 31/70 (43%), Gaps = 3/70 (4%)
Query: 151 NGNSSSCSCKLRFFMTWISPLKAFGDRELLSVESLFKSH--PKACLVIVSKSMDSDKGTQ 208
NG+ + L+F + W+ + R+ V + H P+ IV + SDK +
Sbjct: 152 NGDKRTAEDSLKFLLKWVERFPEYKGRDFYIVGESYAGHYIPQLSEAIVKHNQGSDKNSI 211
Query: 209 ILRPF-VKNG 217
L+ + V NG
Sbjct: 212 NLKGYMVGNG 221
>At3g25183 hypothetical protein
Length = 362
Score = 28.9 bits (63), Expect = 5.9
Identities = 20/59 (33%), Positives = 26/59 (43%), Gaps = 5/59 (8%)
Query: 247 NVNPGEISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGAQNIDVKTKKW 305
+VNP + L +NLLR L ID DI K +GA +D + KKW
Sbjct: 225 DVNPTQNRLSMPFNNLLRNDFLTSVESRIIDKDIKNDKKIG-----VGAILVDQRCKKW 278
>At1g52950 replication protein A1, putative
Length = 566
Score = 28.9 bits (63), Expect = 5.9
Identities = 22/86 (25%), Positives = 41/86 (47%), Gaps = 2/86 (2%)
Query: 252 EISLGQNLSNLLRLSLLYKFGGIYIDADIIIMKSFSKFRNTIGAQNIDVKTKKWSRLNNA 311
+I G +LL+L K G I + + K+ +R+T I++++ +R +N+
Sbjct: 52 KIQAGIKKEHLLKLQRYVKIGHWTIIEEFSVTKASGLYRSTTHPYRINIQSS--TRFSNS 109
Query: 312 VLIFDKKHPLLLKFIEEFALTFDGNK 337
I D+ L+ F + + T D NK
Sbjct: 110 PTISDEIWLDLVNFNDVLSGTLDQNK 135
>At3g01085 putative protein
Length = 536
Score = 28.5 bits (62), Expect = 7.7
Identities = 14/40 (35%), Positives = 21/40 (52%), Gaps = 1/40 (2%)
Query: 64 IHIPHHYSPHKPLLLP-PPSHTKTKLSSSVFFAHKEDNSP 102
+H H PH +LP PPSH + SS + +D++P
Sbjct: 26 VHRSHTQDPHNRHVLPGPPSHRRVVNSSPKKHRNNDDDAP 65
>At2g03770 putative steroid sulfotransferase
Length = 324
Score = 28.5 bits (62), Expect = 7.7
Identities = 29/108 (26%), Positives = 41/108 (37%), Gaps = 13/108 (12%)
Query: 7 HHHNPFLYFPLLL---LHKLHHFLHNFKKKILSILFKLPTSFFALLVLFLLAYNAFTVFC 63
+ + F Y P LL L+ HF +L+ + K T++ LV L+ F
Sbjct: 40 YQYQGFWYPPNLLEGVLYSQKHFQARDSDIVLASIPKSGTTWLKSLVFALIHRQEFQTPL 99
Query: 64 IHIPHHYSPHKPLLLPPPSHTKTKLSSSVFFAHKEDNSPIINKTHFPL 111
+ P LL HT F H +D SP I TH P+
Sbjct: 100 VSHP---------LLDNNPHTLVTFIEG-FHLHTQDTSPRIFSTHIPV 137
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.326 0.141 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,657,954
Number of Sequences: 26719
Number of extensions: 470123
Number of successful extensions: 1573
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 1555
Number of HSP's gapped (non-prelim): 18
length of query: 439
length of database: 11,318,596
effective HSP length: 102
effective length of query: 337
effective length of database: 8,593,258
effective search space: 2895927946
effective search space used: 2895927946
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC129091.14


