

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127021.6 + phase: 0 /pseudo
(408 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
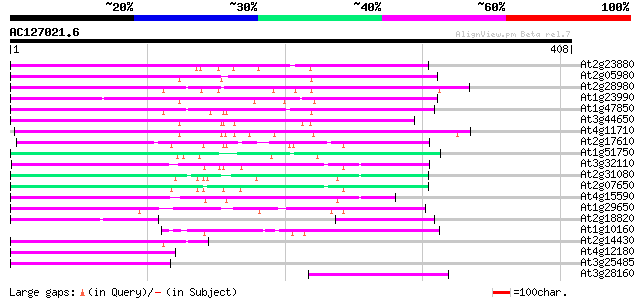
Score E
Sequences producing significant alignments: (bits) Value
At2g23880 putative non-LTR retroelement reverse transcriptase 119 2e-27
At2g05980 putative non-LTR retroelement reverse transcriptase 119 2e-27
At2g28980 putative non-LTR retroelement reverse transcriptase 116 2e-26
At1g23990 reverse transcriptase, putative 108 4e-24
At1g47850 hypothetical protein 107 9e-24
At3g44650 putative protein 106 2e-23
At4g11710 putative protein 106 3e-23
At2g17610 putative non-LTR retroelement reverse transcriptase 104 1e-22
At1g51750 hypothetical protein 102 3e-22
At3g32110 non-LTR reverse transcriptase, putative 96 3e-20
At2g31080 putative non-LTR retroelement reverse transcriptase 91 9e-19
At2g07650 putative non-LTR retrolelement reverse transcriptase 90 2e-18
At4g15590 reverse transcriptase like protein 82 7e-16
At1g29650 reverse transcriptase, putative 79 3e-15
At2g18820 putative non-LTR retroelement reverse transcriptase 77 2e-14
At1g10160 putative reverse transcriptase 77 2e-14
At2g14430 putative non-LTR retroelement reverse transcriptase 75 6e-14
At4g12180 putative reverse transcriptase 74 1e-13
At3g25485 unknown protein 74 1e-13
At3g28160 non-LTR reverse transcriptase, putative 71 9e-13
>At2g23880 putative non-LTR retroelement reverse transcriptase
Length = 1216
Score = 119 bits (299), Expect = 2e-27
Identities = 89/336 (26%), Positives = 147/336 (43%), Gaps = 35/336 (10%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
D+I+ ++ W R LS GR L+N V+ + + + R P+ +NEI +I +WSG
Sbjct: 512 DQIRRRIGMWTSRYLSFAGRLSLINSVLWSITNFWMNAFRLPRECINEINRISSALLWSG 571
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQ-WAQLCRHIL 119
K V+W+ +C+PK EGGL ++ N S L L + L+ QD W + R L
Sbjct: 572 PELNPKKAKVSWDEICKPKKEGGLGLQSLREANKVSSLKLIWRLLSCQDSLWVKWTRMNL 631
Query: 120 LSNAKPKNHYIHSSI--WIG---------LKSWVDVVIQHSN-----LEHW*WKICFI-- 161
L + HS++ WI KS+ + + + ++W K I
Sbjct: 632 LKKESFWSIGTHSTLGSWIWRRLLKHREVAKSFCKIEVNNGVNTSFWFDNWSEKGPLINL 691
Query: 162 --LHSALSMSVSDYIVDGE-W-------NLPTYFHVKDDVLTNKILQTTLPIKPVPDALQ 211
A+ M +S ++ E W + + +++L K + ++ DA+
Sbjct: 692 TGARGAIDMGISRHMTLAEAWSRRRRKRHRVEILNEFEEILLQKYQHRNIELE---DAIL 748
Query: 212 WLSTTD---GHLTNKLTYLFVRGTCIKVPWHKLVWNTYIPPTRVFICWRFIHNRLPTDEN 268
W D + K T+ +R + + WHK VW + P F W I NRL T +
Sbjct: 749 WRGKEDVFKARFSTKDTWNHIRTSSNQRAWHKGVWFAHATPKFSFCAWLAIRNRLSTGDR 808
Query: 269 LRKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIW 304
+ + C FC +ET DH+F +C +S+IW
Sbjct: 809 MMTWNNGTPTTCVFCSSPMETRDHLFFQCCYSSEIW 844
>At2g05980 putative non-LTR retroelement reverse transcriptase
Length = 1352
Score = 119 bits (299), Expect = 2e-27
Identities = 89/343 (25%), Positives = 151/343 (43%), Gaps = 36/343 (10%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
++I++++++W R LS GR QL+ V+S + + + R PK L EI K+ F+WSG
Sbjct: 938 EKIRARITSWTNRFLSFAGRLQLIKSVLSSITNFWLSVFRLPKACLQEIEKMFSAFLWSG 997
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQ-WAQ-LCRHI 118
K +AW+ +C+ K+EGGL ++ N SLL L + L+++D W + + +H+
Sbjct: 998 PDLNTKKAKIAWSEVCKLKEEGGLGLKPLKEANEVSLLKLIWRILSARDSLWVKWVNKHL 1057
Query: 119 LLSN---AKPKNHYIHSSIWIGLKSWVDVVIQHSNLE------------HW*WKICFILH 163
+ + +N + S +W + D +E HW C +
Sbjct: 1058 IRKETFWSVKENTGLGSWLWRKILKQRDKARLFHRMEVRSGTFTSFWHDHW----CPLGR 1113
Query: 164 SALSMSVSDYIVDGEWNLPTYFHVKDDVLTNKILQTTL-PIKPVPDALQWLSTTDG---- 218
M I G N T V + + L IK + + +TDG
Sbjct: 1114 LHQHMGSRGTIDLGIPNNATVAEVMNTHRRKRHRADFLNQIKSQIELARQDRSTDGDRSL 1173
Query: 219 ----------HLTNKLTYLFVRGTCIKVPWHKLVWNTYIPPTRVFICWRFIHNRLPTDEN 268
++ T+ +R ++ W++ VW + P F+ W HNRL T +
Sbjct: 1174 WKQKEDTFKSSFSSSKTWQQIRSISLRCDWYRGVWFSASTPKYSFVTWLAFHNRLTTSDK 1233
Query: 269 LRKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIWDWLNQGI 311
+ K C FC ++LET DH+F CP +S +W L +G+
Sbjct: 1234 ICKWNSGARYDCVFCGEELETRDHLFFSCPYSSHVWFSLTKGL 1276
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 116 bits (291), Expect = 2e-26
Identities = 97/368 (26%), Positives = 155/368 (41%), Gaps = 36/368 (9%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
+ +K+K+S+W R LS GR L+N VI + + +R P + EI K+ F+WSG
Sbjct: 1088 EAVKTKISSWTARSLSYAGRLALLNSVIVSIANFWMSAYRLPAGCIREIEKLCSAFLWSG 1147
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQ------WAQL 114
K +AW+++C+PK EGGL I+ N S L L + L++Q W +
Sbjct: 1148 PVLNPKKAKIAWSSICQPKKEGGLGIKSLAEANKVSCLKLIWRLLSTQPSLWVTWIWTFI 1207
Query: 115 CRHILLSNAKPKNHYIHSSIWIGL-------KSWVDVVIQHSN-----LEHW*WKICFIL 162
R +A ++ + S +W L KS V +++ + +HW + +L
Sbjct: 1208 IRKGTFWSANERSS-LGSWMWKKLLKYRELAKSMHKVEVRNGSSTSFWYDHW-SHLGRLL 1265
Query: 163 HSALSMSVSDYIVDGEWNLPTYFHVKDD-----VLTNKILQTTLPIKPV-----PDALQW 212
+ V D + E NL T + N+I ++ PD W
Sbjct: 1266 DITGTRRVIDLGIPLETNLETVLRTHQHRQHRAAIYNRINAEIQRLQQQEREAGPDISLW 1325
Query: 213 LSTTDG---HLTNKLTYLFVRGTCIKVPWHKLVWNTYIPPTRVFICWRFIHNRLPTDENL 269
S + K+T+ VR + W+K VW Y P F+ W + NRL T + +
Sbjct: 1326 RSLKNDFNKRFITKVTWNNVRTHQPQQNWYKGVWFPYSTPKYSFLLWLTVQNRLSTGDRI 1385
Query: 270 RKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIWDWLNQGI---NQQLDLSSCQNLFVK 326
+ + C C ET DH+F C TS +W+ L Q + N D + L
Sbjct: 1386 KAWNSGQLVTCTLCNNAEETRDHLFFSCQYTSYVWEALTQRLLSTNYSRDWNRLFTLLCT 1445
Query: 327 QNWRRQYV 334
N R ++
Sbjct: 1446 SNLPRDHL 1453
>At1g23990 reverse transcriptase, putative
Length = 657
Score = 108 bits (271), Expect = 4e-24
Identities = 85/326 (26%), Positives = 146/326 (44%), Gaps = 17/326 (5%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
+ IK K+ TW R LS GR L+ V+ + + R P+ + EI KI F+WSG
Sbjct: 240 EHIKKKIGTWTTRYLSYAGRLNLITSVLWSICNFWLAAFRLPRECIREIDKICSAFLWSG 299
Query: 61 -DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLT-SQDQWAQLCRHI 118
D N +K V W +C+PK EGGL +R +N S L L + ++ + W +
Sbjct: 300 PDLNPRKT-RVCWGDVCKPKQEGGLGLRSLKEMNEVSCLKLIWRIVSHTNSLWVRWIEQY 358
Query: 119 LLSN----AKPKNHYIHSSIWIGLKSWVDVVIQHSNLEHW*WK-ICFILHSALSMSVSDY 173
LL + + + S +W L + D+ Q +E K + F + ++ V +
Sbjct: 359 LLKHDTFWSVQTTTNMGSWMWRKLLKYRDIAKQFCKVEVKNGKRVAFWYENWSNLGVLND 418
Query: 174 IVD--GEWNLPTYFHVK-DDVLTNKILQ--TTLPIKPVPDALQWLSTTDGHL---TNKLT 225
+V G +L H+ D ++ + + + + +A+ L D ++ + + T
Sbjct: 419 LVGDRGRIDLGITSHMTLSDAWASRQQRRHRSATLNQIEEAIM-LGRNDEYMPKFSTRDT 477
Query: 226 YLFVRGTCIKVPWHKLVWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTIVSICCFCKK 285
+ R T V WH +W + P F W + NRL T + + + + C C
Sbjct: 478 WNQTRNTSTPVTWHMGIWFAHATPKFSFCAWLAVQNRLSTGDKMLQWNRRLSPTCVLCNN 537
Query: 286 QLETSDHIFLRCPVTSQIWDWLNQGI 311
+ET +H+F C T++IW+ L + I
Sbjct: 538 NIETRNHLFFSCCYTAEIWENLAKNI 563
>At1g47850 hypothetical protein
Length = 799
Score = 107 bits (268), Expect = 9e-24
Identities = 88/341 (25%), Positives = 142/341 (40%), Gaps = 35/341 (10%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
D+++SK+S+W R LS GR L+N VI + + +R P + EI K+ F+WSG
Sbjct: 364 DKVRSKISSWTARSLSYAGRLALINSVIVSLSNFWMSAYRLPAGCIKEIEKLCSAFLWSG 423
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQ------WAQL 114
K + W ++C+ K EGGL I+ N S L L + ++ Q W +
Sbjct: 424 PELNPKKAKITWTSLCKLKQEGGLGIKSLLEANKVSCLKLIWRLVSRQSSLWVNWVWTYI 483
Query: 115 CRHILLSNAKPKNHYIHSSIWIGLKSWVDV--------VIQHSNLEHW--*W-------- 156
R +A ++ + S +W L + DV + S+ W W
Sbjct: 484 IRKGSFWSANDRSS-LGSWMWKKLLKYRDVAKSMCKVEIKSGSSTSFWYDNWSQLGQLVD 542
Query: 157 ----KICFILHSALSMSVSDYIVDGEW-NLPTYFHVKDDVLTNKILQTTLPIKPVPDALQ 211
+ + L+ +V+ + + T + K + ILQ PD
Sbjct: 543 VTNARRTIDMGIPLAATVATVLASHRTKHHRTAIYNKIEAEIQSILQRER--SGAPDIFL 600
Query: 212 WLSTTDG---HLTNKLTYLFVRGTCIKVPWHKLVWNTYIPPTRVFICWRFIHNRLPTDEN 268
W S+ D K+T+ +R W+K VW +Y P F+ W IH+RL T +
Sbjct: 601 WRSSGDNFRQSFITKVTWHNIRVIHTHRQWYKGVWFSYNTPKYSFLLWLAIHDRLSTGDR 660
Query: 269 LRKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIWDWLNQ 309
++K C C E+ +H+F C T+ IWD L +
Sbjct: 661 IKKWNSGQQVNCVLCGNVEESRNHLFFSCHYTTAIWDNLTR 701
>At3g44650 putative protein
Length = 762
Score = 106 bits (265), Expect = 2e-23
Identities = 83/324 (25%), Positives = 136/324 (41%), Gaps = 30/324 (9%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
++I+ ++ +W R LS GR L++ +I + + P+ + EI K+ +F+WSG
Sbjct: 344 EQIRKRIGSWSSRFLSFAGRFNLISSIIWSSCNFWLSAFQLPRACIQEIEKLCSSFLWSG 403
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQ-WAQLCRHIL 119
K ++WN +C+PK EGGL +R N+ L L + ++ D W + H L
Sbjct: 404 TNLNSKKAKISWNQVCKPKSEGGLGLRSLKEANDVCCLKLVWRIISHGDSLWVKWVEHNL 463
Query: 120 LSN----AKPKNHYIHSSIWIGLKSWVDVVIQHSNLE--------HW--*WKICFIL--- 162
L +N + S IW + + V + E W W + L
Sbjct: 464 LKREIFWIVKENANLGSWIWKKILKYRGVAKRFCKAEVGNGESTSFWFDDWSLLGRLIDV 523
Query: 163 ---HSALSMSVSDYI-VDGEWNLPTYFHVKDDVL-TNKILQTTLPIKPVPDALQ----WL 213
+ M +S + V W H + ++L T + + +T K Q W
Sbjct: 524 AGIRGTIDMGISRTMSVADAWTSRRRRHHRQEILNTIEEVLSTQHQKRTQQQQQGRVLWK 583
Query: 214 STTD---GHLTNKLTYLFVRGTCIKVPWHKLVWNTYIPPTRVFICWRFIHNRLPTDENLR 270
D + K T+ ++R T +V WHK VW + P F W H+RL T +
Sbjct: 584 GKNDIYKDKFSTKNTWNYLRTTSNEVAWHKGVWFPHATPKYSFCLWLAAHDRLATGARMI 643
Query: 271 KRGCTIVSICCFCKKQLETSDHIF 294
K C FC++ +ET DH+F
Sbjct: 644 KWNRGETGDCTFCRQGIETRDHLF 667
>At4g11710 putative protein
Length = 473
Score = 106 bits (264), Expect = 3e-23
Identities = 84/362 (23%), Positives = 149/362 (40%), Gaps = 30/362 (8%)
Query: 4 KSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSGDCN 63
+ K+ +W R LS GR L++ V+ + + R P+ + EI K+ ++WSG
Sbjct: 43 RQKICSWSARFLSYAGRLNLISSVLWSICNFWMGAFRLPRDCIREIDKMCSAYLWSGGEL 102
Query: 64 QQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQ-WAQLCRHILLSN 122
+ W +C+PK+EGGL +R N+ L L + ++ D W + + LL
Sbjct: 103 NTSKAKITWAFVCKPKEEGGLGLRSLKEANDVCCLKLIWRIISHADSLWVKWIQSSLLKK 162
Query: 123 ----AKPKNHYIHSSIWIGLKSWVDVVIQHSNLE--------HW*--WKICFIL------ 162
A +N + S +W + + D+ +E W W L
Sbjct: 163 VSFWAVRENTSLGSWMWRKILKFRDIARTLCKVEINNGARTSFWYDDWSDLGRLIDSAGD 222
Query: 163 HSALSMSVSDY--IVDGEWNLPTYFHVKDDV--LTNKILQTTLPIKPVPDALQWLSTTDG 218
A+ + ++ + +V+ N H + + + +++ + D W +
Sbjct: 223 RGAIDLGINKHATVVEAWGNRRRRRHRTNFLNRVEERLILSWNSRNQAEDRALWKGKENR 282
Query: 219 H---LTNKLTYLFVRGTCIKVPWHKLVWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCT 275
+ K T+ +R KV W+K VW P F W +HNRL T + +
Sbjct: 283 FRSIFSTKDTWNHIRTVSNKVAWYKGVWFAQAIPKHAFCMWLAVHNRLSTGDRMTLWNMG 342
Query: 276 IVSICCFCKKQLETSDHIFLRCPVTSQIWDWLNQGINQQLDLSSCQNLF--VKQNWRRQY 333
+ + C C K LE+ DH+F CP ++IW+ L + I + Q + V +NW +
Sbjct: 343 VDATCILCNKALESRDHLFFSCPFATEIWEPLAKTIYNTCFYTDWQTIINNVSRNWPDRI 402
Query: 334 VG 335
G
Sbjct: 403 AG 404
>At2g17610 putative non-LTR retroelement reverse transcriptase
Length = 773
Score = 104 bits (259), Expect = 1e-22
Identities = 92/349 (26%), Positives = 148/349 (42%), Gaps = 63/349 (18%)
Query: 6 KLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSGDCNQQ 65
K+ W+ +LLS G+ L+ +++ + Y+ P ++++IT +R F WS +
Sbjct: 230 KMDNWYNKLLSPAGKEVLIKSIVTAIPTYSMSCFLLPMRLIHQITSAMRWFWWSNTKVKH 289
Query: 66 KICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQWAQLCR--------- 116
KI VAW+ + PK GGLAIRD N A L + + L Q ++ + R
Sbjct: 290 KIPWVAWSKLNDPKKMGGLAIRDLKDFNIALLAKQSWRIL--QQPFSLMARVFKAKYFPK 347
Query: 117 -HILLSNAKPKNHYIHSSIWIGLK----SWVDVVIQHSNLEHW--*W----------KIC 159
+L + A ++ Y SI G K + +N++ W W C
Sbjct: 348 ERLLDAKATSQSSYAWKSILHGTKLISRGLKYIAGNGNNIQLWKDNWLPLNPPRPPVGTC 407
Query: 160 FILHSALSMSVSDYIVDGEWNLPTYFHVKDDVLTNKILQTTLP----IKP----VPDALQ 211
++S L VSD +++G WN +D+L I Q +P I+P DA+
Sbjct: 408 DSIYSQL--KVSDLLIEGRWN--------EDLLCKLIHQNDIPHIRAIRPSITGANDAIT 457
Query: 212 WLSTTDGHLTNKLTYLFVRGTCIKVPWHKL---------------VWNTYIPPTRVFICW 256
W+ T DG+ + K Y +R + H +W PP W
Sbjct: 458 WIYTHDGNYSVKSGYHLLRK--LSQQQHASLPSPNEVSAQTVFTNIWKQNAPPKIKHFWW 515
Query: 257 RFIHNRLPTDENLRKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIWD 305
R HN LPT NL++R C C + E +H+ +C V+ +IW+
Sbjct: 516 RSAHNALPTAGNLKRRRLITDDTCQRCGEASEDVNHLLFQCRVSKEIWE 564
>At1g51750 hypothetical protein
Length = 629
Score = 102 bits (255), Expect = 3e-22
Identities = 89/357 (24%), Positives = 143/357 (39%), Gaps = 59/357 (16%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
++I++++ TW R LS GR L++ V+ + + R P L EI I F+WSG
Sbjct: 196 EQIRNRIGTWTSRYLSFAGRLNLISSVLWSTMNFWMSAFRLPSACLKEINSICSAFLWSG 255
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQ-WAQLCRHIL 119
++ V+W+ +C+PK EGGL +R N S+L L + ++ D W + + L
Sbjct: 256 PELHRRKAKVSWDDICKPKQEGGLGLRSLTEANVVSVLKLIWRVTSNDDSLWVKWSKMNL 315
Query: 120 L--------------------------SNAKP-----KNHYIHSSIWI----GLKSWVDV 144
L AKP N+ +S W G+ +DV
Sbjct: 316 LKQESFWSLTPNSSLGSWMWKKMLKYRETAKPFSRVEVNNGARTSFWFDNWSGMGHLMDV 375
Query: 145 VIQHSNLEHW*WKICFILHSALSMSVSDYIVDGEWNLPTYFHVKD-----DVLTNKILQT 199
Q ++ L +S + + + N H + + N+ QT
Sbjct: 376 TGQRGQID-------------LGISRNKTVAEAWSNRRRRKHRTEQLNDIEAALNQKYQT 422
Query: 200 TLPIKPVPDALQWLSTTDGHLTN---KLTYLFVRGTCIKVPWHKLVWNTYIPPTRVFICW 256
++ DA W D T+ K T+ VR +V W+K VW ++ P F W
Sbjct: 423 RNLLR--EDATLWRGKGDVFKTSFSTKDTWNQVRKKSNEVAWYKGVWFSHSTPKYQFCTW 480
Query: 257 RFIHNRLPTDENLRKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIWDWLNQGINQ 313
+ NRL T ++ C FC +ET DH+F C S IW + + + Q
Sbjct: 481 LALRNRLSTGYRMQLWNNGSDVKCTFCSTSIETRDHLFFSCSYASAIWTAIAKNVLQ 537
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 96.3 bits (238), Expect = 3e-20
Identities = 86/345 (24%), Positives = 142/345 (40%), Gaps = 50/345 (14%)
Query: 2 RIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSGD 61
R S+L+ W GR+LS GR L V++ +LV+T + P+ L+ + K+ R F+W
Sbjct: 1310 RFSSRLAGWKGRMLSFAGRLTLTKAVLTSILVHTMSTIKLPQSTLDGLDKVSRAFLWGST 1369
Query: 62 CNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLT-SQDQWAQLCRHILL 120
++K VAW +C P+ EGGL IR +N A + + + L WAQ+ R
Sbjct: 1370 LEKKKQHLVAWTRVCLPRREGGLGIRSATAMNKALIAKVGWRVLNDGSSLWAQVVR---- 1425
Query: 121 SNAKPKNHYIHSSIWIGLKS-----WVDVVIQHSNL----EHW---------*WKICFIL 162
+K K +H W KS W V + ++ +HW W ++
Sbjct: 1426 --SKYKVGDVHDRNWTVAKSNWSSTWRSVGVGLRDVIWREQHWVIGDGRQIRFWTDRWLS 1483
Query: 163 HSALS-------------MSVSDYIVDGE-WNLPTYF-HVKDDVLTNKILQTTLPIKPVP 207
+ ++ + D DG W++ V D+ + + +
Sbjct: 1484 ETPIADDSIVPLSLAQMLCTARDLWRDGTGWDMSQIAPFVTDNKRLDLLAVIVDSVTGAH 1543
Query: 208 DALQWLSTTDGHLTNKLTYLFVRGTCIKVPWHKL------VWNTYIPP-TRVFICWRFIH 260
D L W T+DG T K + + T P + VW P RVF+ W ++
Sbjct: 1544 DRLAWGMTSDGRFTVKSAFAML--TNDDSPRQDMSSLYGRVWKVQAPERVRVFL-WLVVN 1600
Query: 261 NRLPTDENLRKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIWD 305
+ T+ ++R +C C+ +E+ H+ CP S IWD
Sbjct: 1601 QAIMTNSERKRRHLCDSDVCQVCRGGIESILHVLRDCPAMSGIWD 1645
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 91.3 bits (225), Expect = 9e-19
Identities = 85/345 (24%), Positives = 140/345 (39%), Gaps = 50/345 (14%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
+R+ ++L+ W GR LS+ GR L V+S + V+ P L+ + + R F+W
Sbjct: 633 ERVSARLAGWKGRSLSLAGRITLTKAVLSSIPVHVMSAILLPVSTLDTLDRYSRTFLWGS 692
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQ-WAQLCRHIL 119
++K ++W +C+PK EGG+ +R +N A + + + L ++ WA++ R
Sbjct: 693 TMEKKKQHLLSWRKICKPKAEGGIGLRSARDMNKALVAKVGWRLLQDKESLWARVVRKKY 752
Query: 120 -------LSNAKPKNHYIHSSIW----IGLK-------SWV--DVVIQHSNLEHW*WKIC 159
S KP+ + SS W +GL+ WV D L+ W
Sbjct: 753 KVGGVQDTSWLKPQPRW--SSTWRSVAVGLREVVVKGVGWVPGDGCTIRFWLDRW----- 805
Query: 160 FILHSALSMSVSDYIVDGE--------------WNLPTYFHVKDDVLTNKILQTTLPI-K 204
+L L +D I +GE WNL + + ++L + +
Sbjct: 806 -LLQEPLVELGTDMIPEGERIKVAADYWLPGSGWNLEILGLYLPETVKRRLLSVVVQVFL 864
Query: 205 PVPDALQWLSTTDGHLTNKLTYLFVRGTCIKVP----WHKLVWNTYIPP-TRVFICWRFI 259
D + W T DG T + Y ++G P + +W P RVFI W
Sbjct: 865 GNGDEISWKGTQDGAFTVRSAYSLLQGDVGDRPNMGSFFNRIWKLITPERVRVFI-WLVS 923
Query: 260 HNRLPTDENLRKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIW 304
N + T+ +R + +IC C ET H+ CP IW
Sbjct: 924 QNVIMTNVERVRRHLSENAICSVCNGAEETILHVLRDCPAMEPIW 968
>At2g07650 putative non-LTR retrolelement reverse transcriptase
Length = 732
Score = 90.1 bits (222), Expect = 2e-18
Identities = 82/340 (24%), Positives = 135/340 (39%), Gaps = 40/340 (11%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
+++ ++L+ W GR LS+ GR L V+S + V+T PK L+ + K+ R+F+W
Sbjct: 295 EKLTTRLAGWKGRFLSLAGRVTLTKAVLSSIPVHTMSTIALPKSTLDGLDKVSRSFLWGS 354
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTS-QDQWAQLCR--- 116
Q+K ++W +C+P+ EGGL IR +N A L + + + WA++ R
Sbjct: 355 SVTQRKQHLISWKRVCKPRSEGGLGIRKAQDMNKALLSKVGWRLIQDYHSLWARIMRCNY 414
Query: 117 --HILLSNAKPKNHYIHSSIW----IGLK-------SWVDVVIQHSNLEHW--*WKICFI 161
+ A K + SS W +G++ SW V+ + W W
Sbjct: 415 RVQDVRDGAWTKVRSVCSSTWRSVALGMREVVIPGLSW--VIGDGREILFWMDKWLTNIP 472
Query: 162 LHSAL---------SMSVSDYIVDG-EWNLPTYFHVKDDVLTNKILQTTL-PIKPVPDAL 210
L L M D +G W++ T ++ L I D +
Sbjct: 473 LAELLVQEPPAGWKGMRARDLRRNGIGWDMATIAPYISSYNRLQLQSFVLDDITGARDRI 532
Query: 211 QWLSTTDGHLTNKLTYLFVRGTCIKVPWHKL------VWNTYIPPTRVFICWRFIHNRLP 264
W + DG TY F+ T + P + VW+ P W +
Sbjct: 533 SWGESQDGRFKVSTTYSFL--TRDEAPRQDMKRFFDRVWSVTAPERVRLFLWLVAQQAIM 590
Query: 265 TDENLRKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIW 304
T++ +R + +IC CK E+ H+ CP IW
Sbjct: 591 TNQERHRRHLSTTTICQVCKGAKESIIHVLRDCPAMEGIW 630
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 81.6 bits (200), Expect = 7e-16
Identities = 70/302 (23%), Positives = 128/302 (42%), Gaps = 29/302 (9%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
+R+ S+LS W R LS+ GR L V+ + ++T P +L ++ K+ RNF+W
Sbjct: 568 ERVSSRLSGWKSRSLSLAGRITLTKAVLMSIPIHTMSSILLPASLLEQLDKVSRNFLWGS 627
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQ-DQWAQLCRHIL 119
++K ++W +CRPK GGL +R +N A L + + L + WA++ R
Sbjct: 628 TVEKRKQHLLSWKKVCRPKAAGGLGLRASKDMNRALLAKVGWRLLNDKVSLWARVLRR-- 685
Query: 120 LSNAKPKNHYIHSSIWIGLK-SW------VDVVIQHSNLEHW*W-------KICFILHSA 165
K K +H S W+ K +W + V ++ + W C +
Sbjct: 686 ----KYKVTDVHDSSWLVPKATWSSTWRSIGVGLREGVAKGWILHEPLCTRATCLLSPEE 741
Query: 166 LSMSVSDYIVDG-EWNLPTYFHVKDDVLTNKILQTTLP-IKPVPDALQWLSTTDGHLTNK 223
L+ V ++ +G W++ +T+++ + + + D + W T+DG T
Sbjct: 742 LNARVEEFWTEGVGWDMVKLGQCLPRSVTDRLHAVVIKGVLGLRDRISWQGTSDGDFTVG 801
Query: 224 LTYLFVRGTCIKVP----WHKLVWNTYIPP-TRVFICWRFIHNRLPTDENLRKRGCTIVS 278
Y+ + P + K +W P RVF+ W + T+ +R +
Sbjct: 802 SAYVLLTQEEESKPCMESFFKRIWGVIAPERVRVFL-WLVGQQVIMTNVERVRRHIGDIE 860
Query: 279 IC 280
+C
Sbjct: 861 VC 862
>At1g29650 reverse transcriptase, putative
Length = 1557
Score = 79.3 bits (194), Expect = 3e-15
Identities = 78/324 (24%), Positives = 133/324 (40%), Gaps = 44/324 (13%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
DR+KS+LS W R+LS+ G+ L+ V M VY + K ++ + +F W+
Sbjct: 1079 DRLKSRLSGWFARILSLGGKQILLKAVAMAMPVYAISCFKLTKTTCENLSSAMADFWWNA 1138
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVN---NASLLHLTRKFLTSQDQWAQLCRH 117
+++K V+W MC K+ GG D + A + R L +D
Sbjct: 1139 LEHKRKTHWVSWEKMCLSKENGGRYFPDEDFLEADLGARPSYAWRSVLHGRD-------- 1190
Query: 118 ILLSNAKPKNHYIHSSIWIGLKSWV-DVVIQHSNLEHW*WKICFILHSALSMSVSDYI-V 175
LL+ K +SI + + +W+ D + +H+ +L++ V D I +
Sbjct: 1191 -LLTKGLRKEIGNGNSISVWMDNWIYDNGPRIPLQKHF--------SVSLNLRVKDLINL 1241
Query: 176 DGE-WN----LPTYFHVKDDVLTNKILQTTLPIKPVPDALQWLSTTDGHLTNKLTYLFVR 230
DG WN +F V D++ K P+ + D WL + G + K Y V
Sbjct: 1242 DGRCWNRDLLQDLFFQVDIDLICKK-----KPVVDMDDFWVWLHSKSGEYSVKSGYWLVF 1296
Query: 231 GT---------CIKVPWHKL---VWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTIVS 278
+ C + + L +W+T P WR + LP + + +RG +
Sbjct: 1297 QSNKPELLFKACRQPSTNGLKEKIWSTSTFPKIKLFLWRILSAALPVADQILRRGMNVDP 1356
Query: 279 ICCFCKKQLETSDHIFLRCPVTSQ 302
C C ++ E+ +H+ C V Q
Sbjct: 1357 CCQICGQEGESINHVLFTCSVARQ 1380
>At2g18820 putative non-LTR retroelement reverse transcriptase
Length = 1412
Score = 76.6 bits (187), Expect = 2e-14
Identities = 40/109 (36%), Positives = 62/109 (56%), Gaps = 2/109 (1%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
++I+S++S+W R LS GR QL+N VIS + + R P+ + EI +I F+WSG
Sbjct: 1046 EKIRSRISSWKNRFLSYAGRLQLLNSVISSLTKFWISAFRLPRACIREIEQISAAFLWSG 1105
Query: 61 -DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQ 108
D N K VAW+ +C+PK EGGL +R N L + ++++
Sbjct: 1106 TDLNPHK-AKVAWHDVCKPKSEGGLGLRSLVDANKICCFKLIWRLVSAK 1153
Score = 60.8 bits (146), Expect = 1e-09
Identities = 26/72 (36%), Positives = 36/72 (49%)
Query: 238 WHKLVWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTIVSICCFCKKQLETSDHIFLRC 297
WHK +W + P FI W H+RL T + + I S+C C E+ DH+F C
Sbjct: 1239 WHKAIWFSGATPKFTFISWLAAHDRLTTGDKMASWNRGISSVCVLCNISAESRDHLFFSC 1298
Query: 298 PVTSQIWDWLNQ 309
+S IWD L +
Sbjct: 1299 NFSSHIWDRLTR 1310
>At1g10160 putative reverse transcriptase
Length = 1118
Score = 76.6 bits (187), Expect = 2e-14
Identities = 60/214 (28%), Positives = 94/214 (43%), Gaps = 20/214 (9%)
Query: 111 WAQLCRHILLSNAKPKNHYIHSSIWIGLKS--WVDVVIQHSNLEHW*WKICFILHSA-LS 167
W +LC+ L A+P +I + G+ + W D H L H +L L+
Sbjct: 812 WRKLCK--LRPFARP---FIICEVGSGVTASFWHDNWTDHGPLLHLTGPAGPLLAGLPLN 866
Query: 168 MSVSDYIVDGEWNLPTYFHVKDDVLTNKILQTTLPIK------PVPDALQWL---STTDG 218
V D + D W + + ++ V+T +LQ LP P D W
Sbjct: 867 SVVRDALRDDTWRISSS-RSRNPVIT--LLQRVLPSAASLIDCPHDDTYLWKIGHHAPSN 923
Query: 219 HLTNKLTYLFVRGTCIKVPWHKLVWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTIVS 278
+ T+ +++ + V WHK VW P + FICW HNRL T + LR+ G +I
Sbjct: 924 RFSTADTWSYLQPSSTSVLWHKAVWFKDHVPKQAFICWVVAHNRLHTRDRLRRWGFSIPP 983
Query: 279 ICCFCKKQLETSDHIFLRCPVTSQIWDWLNQGIN 312
C C E+ +H+F RC +S+IW + + +N
Sbjct: 984 TCVLCNDLDESREHLFFRCQFSSEIWSFFMRALN 1017
Score = 32.3 bits (72), Expect = 0.48
Identities = 13/27 (48%), Positives = 19/27 (70%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYV 27
DRI S+ ++W R LS GR QL+N++
Sbjct: 785 DRINSRFTSWTARHLSFAGRLQLLNWI 811
>At2g14430 putative non-LTR retroelement reverse transcriptase
Length = 1277
Score = 75.1 bits (183), Expect = 6e-14
Identities = 45/150 (30%), Positives = 73/150 (48%), Gaps = 7/150 (4%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
D+++SK+S+W R LS GR L+N VI + + +R P + EI K+ F+WSG
Sbjct: 933 DKVRSKISSWTARSLSYAGRLALINSVIVSLSNFWMSAYRLPAGCIKEIEKLCSAFLWSG 992
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQ------WAQL 114
K + W ++C+ K EGGL I+ N S L L + ++ Q W +
Sbjct: 993 PELNPKKAKITWTSLCKLKQEGGLGIKSLLEANKVSCLKLIWRLVSRQSSLWVNWVWTYI 1052
Query: 115 CRHILLSNAKPKNHYIHSSIWIGLKSWVDV 144
R +A ++ + S +W L ++ DV
Sbjct: 1053 IRKGSFWSANDRSS-LGSWMWKKLLNYRDV 1081
>At4g12180 putative reverse transcriptase
Length = 662
Score = 74.3 bits (181), Expect = 1e-13
Identities = 41/121 (33%), Positives = 61/121 (49%), Gaps = 1/121 (0%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
++IK ++ TW R LS GR LV+ V+ + + R P+ + EI K+ F+WSG
Sbjct: 309 EQIKRRIGTWTARFLSYAGRLNLVSSVLWSICNFWLSAFRLPRECVREIDKLCSAFLWSG 368
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQ-WAQLCRHIL 119
+AW T+CRPK EGGL ++ N+ L L + ++ D W Q R L
Sbjct: 369 PELSTNKAKIAWETVCRPKREGGLGLQSIKEANDVCCLKLIWRIVSQGDSLWVQWIRTYL 428
Query: 120 L 120
L
Sbjct: 429 L 429
>At3g25485 unknown protein
Length = 979
Score = 74.3 bits (181), Expect = 1e-13
Identities = 40/118 (33%), Positives = 62/118 (51%), Gaps = 1/118 (0%)
Query: 1 DRIKSKLSTWHGRLLSIMGRAQLVNYVISGMLVYTFHIHRWPKIILNEITKIIRNFVWSG 60
++IK K+ TW R LS GR L++ VI + + R P+ + EI K+ F+WSG
Sbjct: 707 EQIKRKIGTWTARYLSYAGRLNLISSVIWSICNFWIAAFRLPRECIREIDKLCSAFLWSG 766
Query: 61 DCNQQKICTVAWNTMCRPKDEGGLAIRDPFMVNNASLLHLTRKFLTSQDQ-WAQLCRH 117
K V W +C+PK EGGL +R N+ S L L ++++ D W +L ++
Sbjct: 767 PELNSKRAKVNWEAICKPKREGGLGLRSLKEANDVSCLKLIWRWVSRGDSLWRKLLKY 824
Score = 38.9 bits (89), Expect = 0.005
Identities = 23/74 (31%), Positives = 33/74 (44%), Gaps = 2/74 (2%)
Query: 259 IHNRLPTDENLRKRGCTIVSICCFCKKQLETSDHIFLRCPVTSQIWDWLNQGINQQLDLS 318
+ NRL T + + + + C C LET DH+F C S+IW L I +
Sbjct: 902 LQNRLSTGDIMNQWNGGQSATCVLCNNALETRDHLFFSCDFASEIWSNLATEIYGDRYST 961
Query: 319 SCQNLF--VKQNWR 330
S Q+L + WR
Sbjct: 962 SWQDLIQAISGCWR 975
>At3g28160 non-LTR reverse transcriptase, putative
Length = 630
Score = 71.2 bits (173), Expect = 9e-13
Identities = 35/102 (34%), Positives = 53/102 (51%)
Query: 218 GHLTNKLTYLFVRGTCIKVPWHKLVWNTYIPPTRVFICWRFIHNRLPTDENLRKRGCTIV 277
G ++ T+ +R + W++ VW P F+ W HNRL T + L K
Sbjct: 259 GSFSSPKTWQQIRTISNECEWYRGVWFPSSTPKYSFVTWLAFHNRLATGDRLYKWNSEAR 318
Query: 278 SICCFCKKQLETSDHIFLRCPVTSQIWDWLNQGINQQLDLSS 319
+ C FC ++LET DH+F CP +SQIW L +G+ ++SS
Sbjct: 319 ATCVFCDEELETRDHLFFSCPYSSQIWIALAKGLLNGRNVSS 360
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.333 0.142 0.502
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,575,443
Number of Sequences: 26719
Number of extensions: 404430
Number of successful extensions: 1309
Number of sequences better than 10.0: 91
Number of HSP's better than 10.0 without gapping: 84
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 1155
Number of HSP's gapped (non-prelim): 136
length of query: 408
length of database: 11,318,596
effective HSP length: 102
effective length of query: 306
effective length of database: 8,593,258
effective search space: 2629536948
effective search space used: 2629536948
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC127021.6


