

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127021.2 - phase: 0
(377 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
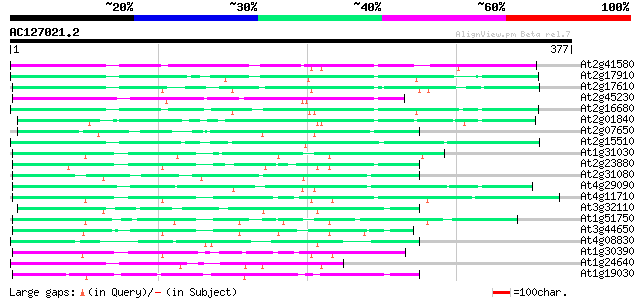
Score E
Sequences producing significant alignments: (bits) Value
At2g41580 putative non-LTR retroelement reverse transcriptase 94 9e-20
At2g17910 putative non-LTR retroelement reverse transcriptase 90 2e-18
At2g17610 putative non-LTR retroelement reverse transcriptase 90 2e-18
At2g45230 putative non-LTR retroelement reverse transcriptase 86 4e-17
At2g16680 putative non-LTR retroelement reverse transcriptase 82 4e-16
At2g01840 putative non-LTR retroelement reverse transcriptase 79 3e-15
At2g07650 putative non-LTR retrolelement reverse transcriptase 77 1e-14
At2g15510 putative non-LTR retroelement reverse transcriptase 76 3e-14
At1g31030 F17F8.5 75 8e-14
At2g23880 putative non-LTR retroelement reverse transcriptase 72 4e-13
At2g31080 putative non-LTR retroelement reverse transcriptase 72 6e-13
At4g29090 putative protein 71 8e-13
At4g11710 putative protein 70 1e-12
At3g32110 non-LTR reverse transcriptase, putative 70 1e-12
At1g51750 hypothetical protein 69 3e-12
At3g44650 putative protein 67 2e-11
At4g08830 putative protein 64 2e-10
At1g30390 hypothetical protein 63 2e-10
At1g24640 hypothetical protein 62 4e-10
At1g19030 hypothetical protein 59 6e-09
>At2g41580 putative non-LTR retroelement reverse transcriptase
Length = 1094
Score = 94.4 bits (233), Expect = 9e-20
Identities = 93/375 (24%), Positives = 155/375 (40%), Gaps = 53/375 (14%)
Query: 1 MSKGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRL-REGGVTGS 59
+ K +GG G + LQ FN ALL K R D + L ++L SRY L G S
Sbjct: 575 LPKFLGGFGFKDLQCFNQALLAKQASRLHTDSDSLLSQILKSRYYMNSDFLSATKGTRPS 634
Query: 60 AWWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLSI 119
W+ I + G + K +GNG NT+ W D W+ F ++ RR L I
Sbjct: 635 YAWQSI--------LYGRELLVSGLKKIIGNGENTYVWMDNWI----FDDKPRRPESLQI 682
Query: 120 HKDLSVGEMHALGWGEDGEAWRWRRRLLAWEE-ELVVEIRNLLTNVTLQDTQPDVWLWQP 178
D+ + + R L W+E +++ + R + ++ D + W
Sbjct: 683 MVDIQLKVSQLIDPFSRNWNLNMLRDLFPWKEIQIICQQRPMA-------SRQDSFCWFG 735
Query: 179 NIGDGYTVRGVYQMLMRQEVHNH--DVVSDAP---------WHKSVPLKVSICAWRLLRN 227
YTV+ Y + RQ VH + P W+ + K+ + W++L+
Sbjct: 736 TNHGLYTVKSEYDLCSRQ-VHKQMFKEAEEQPSLNPLFGKIWNLNSAPKIKVFLWKVLKG 794
Query: 228 RWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQ 287
+D L RGV+ D C KNET++H++ C + +W ++ P
Sbjct: 795 AVAVEDRLRTRGVLIEDG--CSMCPEKNETLNHILFQCPLARQVW----------ALTPM 842
Query: 288 QVLDH-YYQFVYSS------GCYNPRRS-FLQLIWLCGIWVLSNERNQRLFTNTARTTVQ 339
Q +H + ++++ C+N S L+ + IW+L RN+RLF ++
Sbjct: 843 QSPNHGFGDSIFTNVNHVIGNCHNTELSPHLRYVSPWIIWILWKNRNKRLFEGIGSVSLS 902
Query: 340 LLENVKITSLKWLKA 354
++ +WLKA
Sbjct: 903 IVGKALEDCKEWLKA 917
>At2g17910 putative non-LTR retroelement reverse transcriptase
Length = 1344
Score = 89.7 bits (221), Expect = 2e-18
Identities = 93/385 (24%), Positives = 153/385 (39%), Gaps = 71/385 (18%)
Query: 1 MSKGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRL-REGGVTGS 59
+ K GG G + LQ FN ALL K WR L ++ L+ +V SRY L G S
Sbjct: 824 LPKDQGGFGFKDLQCFNQALLAKQAWRVLQEKGSLFSRVFQSRYFSNSDFLSATRGSRPS 883
Query: 60 AWWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLSI 119
WR I + G + + +GNG TF W+D W+ S R +++
Sbjct: 884 YAWRSI--------LFGRELLMQGLRTVIGNGQKTFVWTDKWLHDGSNRRPLNRRRFINV 935
Query: 120 HKDLSVGEMHALGWGEDGEAWRWR----RRLLAWEE-ELVVEIRNLLTNVTLQDTQPDVW 174
DL V ++ D + W R L W++ E++++ R L + D +
Sbjct: 936 --DLKVSQL------IDPTSRNWNLNMLRDLFPWKDVEIILKQRPLF-------FKEDSF 980
Query: 175 LWQPNIGDGYTVRGVYQMLMRQEVH----------NHDVVSDAPWHKSVPLKVSICAWRL 224
W + Y+V+ Y+ L +Q H + + + D W+ K+ I W+
Sbjct: 981 CWLHSHNGLYSVKTGYEFLSKQVHHRLYQEAKVKPSVNSLFDKIWNLHTAPKIRIFLWKA 1040
Query: 225 LRNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLW------------ 272
L P +D L RG+ ++D C+ +NETI+H++ C + +W
Sbjct: 1041 LHGAIPVEDRLRTRGIRSDDG--CLMCDTENETINHILFECPLARQVWAITHLSSAGSEF 1098
Query: 273 -QRIKTWIG-VFSVDPQQVLDHYYQFVYSSGCYNPRRSFLQLIWLCGIWVLSNERNQRLF 330
+ T + + + Q L H+ +FV W+ +W L RN LF
Sbjct: 1099 SNSVYTNMSRLIDLTQQNDLPHHLRFVSP--------------WI--LWFLWKNRNALLF 1142
Query: 331 TNTARTTVQLLENVKITSLKWLKAK 355
T L++ +W A+
Sbjct: 1143 EGKGSITTTLVDKAYEAYHEWFSAQ 1167
>At2g17610 putative non-LTR retroelement reverse transcriptase
Length = 773
Score = 89.7 bits (221), Expect = 2e-18
Identities = 97/382 (25%), Positives = 152/382 (39%), Gaps = 68/382 (17%)
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLREGGVTGSAW- 61
K +GGL +R L++FNIALL K WR L L +V ++Y + L + S++
Sbjct: 303 KKMGGLAIRDLKDFNIALLAKQSWRILQQPFSLMARVFKAKYFPKERLLDAKATSQSSYA 362
Query: 62 WREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCW---------VGTVSFMERFR 112
W+ I + G + GNG N W D W VGT
Sbjct: 363 WKSI--------LHGTKLISRGLKYIAGNGNNIQLWKDNWLPLNPPRPPVGTCD------ 408
Query: 113 RLYDLSIHKDLSVGEMHALGWGEDGEAWRWRRRLLA--WEEELVVEIRNLLTNVTLQDTQ 170
SI+ L V ++ G RW LL + + IR + ++T +
Sbjct: 409 -----SIYSQLKVSDLLIEG--------RWNEDLLCKLIHQNDIPHIRAIRPSITGAN-- 453
Query: 171 PDVWLWQPNIGDGYTVRGVYQMLMRQEVHNH-----------DVVSDAPWHKSVPLKVSI 219
D W Y+V+ Y +L + H V W ++ P K+
Sbjct: 454 -DAITWIYTHDGNYSVKSGYHLLRKLSQQQHASLPSPNEVSAQTVFTNIWKQNAPPKIKH 512
Query: 220 CAWRLLRNRWPTKDNLVRRGVITNDTQLCVRGCGK-NETIDHLIIHCLIFGDLWQR--IK 276
WR N PT NL RR +IT+DT C R CG+ +E ++HL+ C + ++W++ IK
Sbjct: 513 FWWRSAHNALPTAGNLKRRRLITDDT--CQR-CGEASEDVNHLLFQCRVSKEIWEQAHIK 569
Query: 277 TWIG--VFSVDPQQVLDHYYQFVYSSGCYNPRRSFLQLIWLCGIWVLSNERNQRLFTNTA 334
G + S Q L+ + S+ R + L G W + RN +F N
Sbjct: 570 LCPGDSLMSNSFNQNLESIQKLNQSA------RKDVSLFPFIG-WRIWKMRNDLIFNNKR 622
Query: 335 RTTVQLLENVKITSLKWLKAKN 356
+ ++ I +W ++ N
Sbjct: 623 WSIPDSIQKALIDQQQWKESLN 644
>At2g45230 putative non-LTR retroelement reverse transcriptase
Length = 1374
Score = 85.5 bits (210), Expect = 4e-17
Identities = 75/276 (27%), Positives = 116/276 (41%), Gaps = 27/276 (9%)
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLREG-GVTGSAW 61
K VGGLG + ++ FNIALLGK WR + +++ L KV SRY + L G S
Sbjct: 848 KAVGGLGFKEIEAFNIALLGKQLWRMITEKDSLMAKVFKSRYFSKSDPLNAPLGSRPSFA 907
Query: 62 WREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGT--VSFMERFRRLYDLSI 119
W+ I + Q I ++ I +GNG W+D W+G + +R + +S
Sbjct: 908 WKSIYEAQVLI--------KQGIRAVIGNGETINVWTDPWIGAKPAKAAQAVKRSHLVSQ 959
Query: 120 HKDLSVGEMHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDTQPDVWLWQPN 179
+ S+ + L DG W W L + + N+L D + W+ +
Sbjct: 960 YAANSIHVVKDL-LLPDGRDWNWNLVSLLFPDNTQ---ENILALRPGGKETRDRFTWEYS 1015
Query: 180 IGDGYTVRGVYQMLMR--------QEV--HNHDVVSDAPWHKSVPLKVSICAWRLLRNRW 229
Y+V+ Y ++ QEV + D + W VP K+ WR + N
Sbjct: 1016 RSGHYSVKSGYWVMTEIINQRNNPQEVLQPSLDPIFQQIWKLDVPPKIHHFLWRCVNNCL 1075
Query: 230 PTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHC 265
NL R + + CVR ET++HL+ C
Sbjct: 1076 SVASNLAYRHLAREKS--CVRCPSHGETVNHLLFKC 1109
>At2g16680 putative non-LTR retroelement reverse transcriptase
Length = 1319
Score = 82.4 bits (202), Expect = 4e-16
Identities = 90/373 (24%), Positives = 144/373 (38%), Gaps = 47/373 (12%)
Query: 1 MSKGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLR-EGGVTGS 59
+ K +GG G + LQ FN ALL K WR D + + ++ SRY L G S
Sbjct: 798 LPKSLGGFGFKDLQCFNQALLAKQAWRLFSDSKSIVSQIFKSRYFMNTDFLNARQGTRPS 857
Query: 60 AWWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLSI 119
WR I + G + + +GNG T W D W+ F RR +L
Sbjct: 858 YTWRSI--------LYGRELLNGGLKRLIGNGEQTNVWIDKWL----FDGHSRRPMNLHS 905
Query: 120 HKDLSVGEMHALGWGEDGEAWRWRRRLLA-----WEEELVVEIRNLLTNVTLQDTQPDVW 174
++ + H + D W + L + +L++ R L+++ D +
Sbjct: 906 LMNIHMKVSHLI----DPLTRNWNLKKLTELFHEKDVQLIMHQRPLISS-------EDSY 954
Query: 175 LWQPNIGDGYTVRGVYQMLMRQEVHN----HDV------VSDAPWHKSVPLKVSICAWRL 224
W YTV+ Y+ R+ N DV + D W K+ + W+
Sbjct: 955 CWAGTNNGLYTVKSGYERSSRETFKNLFKEADVYPSVNPLFDKVWSLETVPKIKVFMWKA 1014
Query: 225 LRNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLW--QRIKTWIGVF 282
L+ +D L RG+ T D C+ + ETI+HL+ C +W I+ F
Sbjct: 1015 LKGALAVEDRLRSRGIRTADG--CLFCKEEIETINHLLFQCPFARQVWALSLIQAPATGF 1072
Query: 283 SVDPQQVLDHYYQFVYSSGCYNPRRSFLQLIWLCGIWVLSNERNQRLFTNTARTTVQLLE 342
++H Q + G PR WL +W + RN+ LF T T+ +++
Sbjct: 1073 GTSIFSNINHVIQNSQNFGI--PRHMRTVSPWL--LWEIWKNRNKTLFQGTGLTSSEIVA 1128
Query: 343 NVKITSLKWLKAK 355
W+ A+
Sbjct: 1129 KAYEECNLWINAQ 1141
>At2g01840 putative non-LTR retroelement reverse transcriptase
Length = 1715
Score = 79.3 bits (194), Expect = 3e-15
Identities = 88/363 (24%), Positives = 145/363 (39%), Gaps = 42/363 (11%)
Query: 6 GGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLR--EGGVTGSAWWR 63
G LG + L +FN ALL K WR L + + L ++ Y LR +GG W
Sbjct: 1195 GDLGFKDLHQFNRALLAKQAWRILTNPQSLLARLYKGLYYPNTTYLRANKGGHASYGW-- 1252
Query: 64 EIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLSIHKDL 123
N I EG ++ + RLG+G T W D W+ T+ + D +D+
Sbjct: 1253 ------NSI-QEGKLLLQQGLRVRLGDGQTTKIWEDPWLPTLPPRPARGPILD----EDM 1301
Query: 124 SVGEMHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQD-TQPDVWLWQPNIGD 182
V ++ W E+ W + +E L E + L ++ L + D + W
Sbjct: 1302 KVADL----WRENKREW----DPVIFEGVLNPEDQQLAKSLYLSNYAARDSYKWAYTRNT 1353
Query: 183 GYTVRGVYQMLMRQEVHNHDVVS----DAP-----WHKSVPLKVSICAWRLLRNRWPTKD 233
YTVR Y + + ++++ D P W + K+ WR L T
Sbjct: 1354 QYTVRSGYWVATHVNLTEEEIINPLEGDVPLKQEIWRLKITPKIKHFIWRCLSGALSTTT 1413
Query: 234 NLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQVLDHY 293
L R + + T C R C +ETI+H+I C +W R + G + L+
Sbjct: 1414 QLRNRNIPADPT--CQRCCNADETINHIIFTCSYAQVVW-RSANFSGSNRLCFTDNLEEN 1470
Query: 294 YQFVYSSGCYN---PRRSFLQLIWLCGIWVLSNERNQRLFTNTARTTVQLLENVKITSLK 350
+ + G N P + L W+ +W L RN+ LF R ++ + + + +
Sbjct: 1471 IRLIL-QGKKNQNLPILNGLMPFWI--MWRLWKSRNEYLFQQLDRFPWKVAQKAEQEATE 1527
Query: 351 WLK 353
W++
Sbjct: 1528 WVE 1530
>At2g07650 putative non-LTR retrolelement reverse transcriptase
Length = 732
Score = 77.4 bits (189), Expect = 1e-14
Identities = 72/284 (25%), Positives = 110/284 (38%), Gaps = 40/284 (14%)
Query: 6 GGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLREGGVTG-----SA 60
GGLG+R+ Q+ N ALL K WR + D LW +++ Y Q +R+G T S+
Sbjct: 376 GGLGIRKAQDMNKALLSKVGWRLIQDYHSLWARIMRCNYRVQ--DVRDGAWTKVRSVCSS 433
Query: 61 WWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLSIH 120
WR + + + G SW +G+G FW D W+ + E
Sbjct: 434 TWRSVALGMREVVIPGLSWV-------IGDGREILFWMDKWLTNIPLAE----------- 475
Query: 121 KDLSVGEMHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQD-------TQPDV 173
L V E A GW + A RR + W+ + + + LQ D
Sbjct: 476 --LLVQEPPA-GW-KGMRARDLRRNGIGWDMATIAPYISSYNRLQLQSFVLDDITGARDR 531
Query: 174 WLWQPNIGDGYTVRGVYQMLMRQEVHNHDV--VSDAPWHKSVPLKVSICAWRLLRNRWPT 231
W + + V Y L R E D+ D W + P +V + W + + T
Sbjct: 532 ISWGESQDGRFKVSTTYSFLTRDEAPRQDMKRFFDRVWSVTAPERVRLFLWLVAQQAIMT 591
Query: 232 KDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRI 275
RR + T T +C G E+I H++ C +W R+
Sbjct: 592 NQERHRRHLST--TTICQVCKGAKESIIHVLRDCPAMEGIWLRL 633
>At2g15510 putative non-LTR retroelement reverse transcriptase
Length = 1138
Score = 75.9 bits (185), Expect = 3e-14
Identities = 81/368 (22%), Positives = 147/368 (39%), Gaps = 34/368 (9%)
Query: 1 MSKGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLR-EGGVTGS 59
++K GGLG R ++ FN ALL K WR L L+ + SRY + L + GV S
Sbjct: 622 LAKESGGLGFRDIESFNQALLAKQSWRILQYPSFLFARFFKSRYFDDEEFLEADLGVRPS 681
Query: 60 AWWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLSI 119
I + G + + K +GNG + W D W+ ++ +R + S+
Sbjct: 682 YACCSI--------LFGRELLAKGLRKEVGNGKSLNVWMDPWIFDIAPRLPLQRHF--SV 731
Query: 120 HKDLSVGEMHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDTQPDVWLWQPN 179
+ DL V ++ + E W+R LL EE L+T D W+W
Sbjct: 732 NLDLKVNDL------INFEDRCWKRDLL--EELFYPTDVELITKRNPVVNMDDFWVWLHA 783
Query: 180 IGDGYTVRGVYQMLMRQE----------VHNHDVVSDAPWHKSVPLKVSICAWRLLRNRW 229
Y+V+ Y + + + + + + + W K+ + WR+L
Sbjct: 784 KTGEYSVKSGYWLAFQSNKLDLIQQANLLPSTNGLKEQVWSTKSSPKIKMFLWRILSAAL 843
Query: 230 PTKDNLVRRGV-ITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDPQQ 288
P D ++RRG+ I Q+C + E+ +H++ C + +W F
Sbjct: 844 PVADQIIRRGMSIDPRCQIC---GDEGESTNHVLFTCSMARQVWALSGVPTPEFGFQNAS 900
Query: 289 VLDHYYQFVYSSGCYNPRRSFLQLIWLCGIWVLSNERNQRLFTNTARTTVQLLENVKITS 348
+ + QF++ ++ W +W L RN+ F + + ++ +
Sbjct: 901 IFAN-IQFLFELKKMILVPDLVKRSWPWVLWRLWKNRNKLFFDGITFCPLNSIVKIQEDT 959
Query: 349 LKWLKAKN 356
L+W +A++
Sbjct: 960 LEWFQAQS 967
>At1g31030 F17F8.5
Length = 872
Score = 74.7 bits (182), Expect = 8e-14
Identities = 72/306 (23%), Positives = 123/306 (39%), Gaps = 44/306 (14%)
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRG--RLREGGVTGSA 60
K GGLG+R L+E N K WR + + LW K ++ ++ L++ GS
Sbjct: 516 KAEGGLGLRNLKEANDVSCLKLVWRIISNSNSLWTKWVAEYLIRKKSIWSLKQSTSMGSW 575
Query: 61 WWREIVKIQNGIGVEGGSWFEESISK-RLGNGFNTFFWSDCWVGTVSFMERF--RRLYDL 117
WR+I+KI++ +S S+ +GNG + FW D W ++ + DL
Sbjct: 576 IWRKILKIRD---------VAKSFSRVEVGNGESASFWYDHWSAHGRLIDTVGDKGTIDL 626
Query: 118 SIHKDLSVGEMHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDTQ-PDVWLW 176
I ++ SV +AW RR L+ EI ++ + + D LW
Sbjct: 627 GIPREASV-----------ADAWT-RRSRRRHRTSLLNEIEEMMAYQRIHHSDAEDTVLW 674
Query: 177 QPN---IGDGYTVRGVYQMLMRQEVHNHDVVSDAPWHKSV-----PLKVSICAWRLLRNR 228
+ ++ R + ++ S WHK V K ++C W + NR
Sbjct: 675 RGKNDVFKPHFSTRDTWHLIKATS-------STVSWHKGVWFRHATPKYALCTWLAIHNR 727
Query: 229 WPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRIK--TWIGVFSVDP 286
PT D +++ + + CV ++T++HL C +W + W +S
Sbjct: 728 LPTGDRMLKWNSSGSVSGNCVLCTNNSKTLEHLFFSCSYASTVWAALAKGIWKTRYSTRW 787
Query: 287 QQVLDH 292
+L H
Sbjct: 788 SHLLTH 793
>At2g23880 putative non-LTR retroelement reverse transcriptase
Length = 1216
Score = 72.4 bits (176), Expect = 4e-13
Identities = 82/291 (28%), Positives = 113/291 (38%), Gaps = 51/291 (17%)
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFK--VLSSRYGEQRGRLREGGVTGSA 60
K GGLG++ L+E N K WR L ++ LW K ++ E + GS
Sbjct: 590 KKEGGLGLQSLREANKVSSLKLIWRLLSCQDSLWVKWTRMNLLKKESFWSIGTHSTLGSW 649
Query: 61 WWREIVKIQNGIGVEGGSWFEESISK-RLGNGFNTFFWSDCW--VGTVSFMERFRRLYDL 117
WR ++K + +S K + NG NT FW D W G + + R D+
Sbjct: 650 IWRRLLKHRE---------VAKSFCKIEVNNGVNTSFWFDNWSEKGPLINLTGARGAIDM 700
Query: 118 SIHKDLSVGEMHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDTQ------P 171
I + +++ EAW RRR + VEI N + LQ Q
Sbjct: 701 GISRHMTL-----------AEAWSRRRR-----KRHRVEILNEFEEILLQKYQHRNIELE 744
Query: 172 DVWLWQPNIGDGYTVRGVYQMLMRQEVHNHDVVS--DAPWHKSV-----PLKVSICAWRL 224
D LW+ D + R ++ NH S WHK V K S CAW
Sbjct: 745 DAILWRGK-EDVFKAR-----FSTKDTWNHIRTSSNQRAWHKGVWFAHATPKFSFCAWLA 798
Query: 225 LRNRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRI 275
+RNR T D ++ T T CV ET DHL C ++W I
Sbjct: 799 IRNRLSTGDRMMTWNNGTPTT--CVFCSSPMETRDHLFFQCCYSSEIWTSI 847
>At2g31080 putative non-LTR retroelement reverse transcriptase
Length = 1231
Score = 71.6 bits (174), Expect = 6e-13
Identities = 69/287 (24%), Positives = 114/287 (39%), Gaps = 40/287 (13%)
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLREGGVTGSAW- 61
K GG+G+R ++ N AL+ K WR L D+E LW +V+ +Y + GGV ++W
Sbjct: 711 KAEGGIGLRSARDMNKALVAKVGWRLLQDKESLWARVVRKKY-------KVGGVQDTSWL 763
Query: 62 ---------WREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFR 112
WR + + V+G W G+G FW D W+ +E
Sbjct: 764 KPQPRWSSTWRSVAVGLREVVVKGVGWVP-------GDGCTIRFWLDRWLLQEPLVELGT 816
Query: 113 RLYDLSIHKDLSVGE--MHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDTQ 170
+ + GE A + G W L E + + +++ V L +
Sbjct: 817 DM--------IPEGERIKVAADYWLPGSGWNLEILGLYLPETVKRRLLSVVVQVFLGN-- 866
Query: 171 PDVWLWQPNIGDGYTVRGVYQMLMRQ--EVHNHDVVSDAPWHKSVPLKVSICAWRLLRNR 228
D W+ +TVR Y +L + N + W P +V + W + +N
Sbjct: 867 GDEISWKGTQDGAFTVRSAYSLLQGDVGDRPNMGSFFNRIWKLITPERVRVFIWLVSQNV 926
Query: 229 WPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRI 275
T VRR + ++ +C G ETI H++ C +W+R+
Sbjct: 927 IMTNVERVRRHL--SENAICSVCNGAEETILHVLRDCPAMEPIWRRL 971
>At4g29090 putative protein
Length = 575
Score = 71.2 bits (173), Expect = 8e-13
Identities = 85/367 (23%), Positives = 139/367 (37%), Gaps = 38/367 (10%)
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLREG-GVTGSAW 61
K GG+G + ++ FN+ALLGK WR L E L KV SRY + L G S
Sbjct: 51 KAEGGIGFKDIEAFNLALLGKQMWRMLSRPESLMAKVFKSRYFHKSDPLNAPLGSRPSFV 110
Query: 62 WREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLSIHK 121
W+ I Q + + +GNG + W W+ + R+ + +
Sbjct: 111 WKSIHASQEIL--------RQGARAVVGNGEDIIIWRHKWLDSKPASAAL-RMQRVPPQE 161
Query: 122 DLSVGEMHALGWGEDGEAWRWRRRLLAW-----EEELVVEIRNLLTNVTLQDTQPDVWLW 176
SV + + D WR+ ++ E +L+ E+R + D + W
Sbjct: 162 YASVSSILKVSDLIDESGREWRKDVIEMLFPEVERKLIGELRPGGRRIL------DSYTW 215
Query: 177 QPNIGDGYTVRGVYQMLMR--------QEVHNHDV--VSDAPWHKSVPLKVSICAWRLLR 226
YTV+ Y +L + QEV + + W K+ W+ L
Sbjct: 216 DYTSSGDYTVKSGYWVLTQIINKRSSPQEVSEPSLNPIYQKIWKSQTSPKIQHFLWKCLS 275
Query: 227 NRWPTKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRIKTWIGVFSVDP 286
N P L R + + C+R ET++HL+ C W I +
Sbjct: 276 NSLPVAGALAYRHL--SKESACIRCPSCKETVNHLLFKCTFARLTWAISSIPIPLGGEWA 333
Query: 287 QQVLDHYYQFVYSSGCYNPR-RSFLQLI-WLCGIWVLSNERNQRLFTNTARTTVQLLENV 344
+ + Y +V++ G NP+ QL+ WL +W L RN+ +F ++L
Sbjct: 334 DSIYVNLY-WVFNLGNGNPQWEKASQLVPWL--LWRLWKNRNELVFRGREFNAQEVLRRA 390
Query: 345 KITSLKW 351
+ +W
Sbjct: 391 EDDLEEW 397
>At4g11710 putative protein
Length = 473
Score = 70.5 bits (171), Expect = 1e-12
Identities = 92/382 (24%), Positives = 151/382 (39%), Gaps = 44/382 (11%)
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRG--RLREGGVTGSA 60
K GGLG+R L+E N K WR + + LW K + S ++ +RE GS
Sbjct: 118 KEEGGLGLRSLKEANDVCCLKLIWRIISHADSLWVKWIQSSLLKKVSFWAVRENTSLGSW 177
Query: 61 WWREIVKIQNGIGVEGGSWFEESISK-RLGNGFNTFFWSDCW--VGTVSFMERFRRLYDL 117
WR+I+K ++ ++ K + NG T FW D W +G + R DL
Sbjct: 178 MWRKILKFRD---------IARTLCKVEINNGARTSFWYDDWSDLGRLIDSAGDRGAIDL 228
Query: 118 SIHKDLSVGEMHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDTQPDVWLWQ 177
I+K +V EAW RRR L L+ + ++ D LW+
Sbjct: 229 GINKHATV-----------VEAWGNRRRRRHRTNFLNRVEERLILSWNSRNQAEDRALWK 277
Query: 178 PNIGDGYTVRGVYQMLMRQEVHNH--DVVSDAPWHKSVPL-----KVSICAWRLLRNRWP 230
G R ++ ++ NH V + W+K V K + C W + NR
Sbjct: 278 ---GKENRFRSIFS---TKDTWNHIRTVSNKVAWYKGVWFAQAIPKHAFCMWLAVHNRLS 331
Query: 231 TKDNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRIKTWI--GVFSVDPQQ 288
T D + + + T C+ E+ DHL C ++W+ + I F D Q
Sbjct: 332 TGDRMTLWNMGVDAT--CILCNKALESRDHLFFSCPFATEIWEPLAKTIYNTCFYTDWQT 389
Query: 289 VLDHYYQFVYSSGCYNPRRSFLQLIWLCGIWVLSNERNQRLFTNTARTTVQLLENVKITS 348
++++ + R LQ + + +W NER N++ + ++
Sbjct: 390 IINNVSRNWPDRIAGFLARCILQ-VTIYTLWRERNERKHGASPNSSSRLISWIDKHIRNH 448
Query: 349 LKWLK-AKNVCFPFGYHMWWQR 369
L +K + + F G+ +W QR
Sbjct: 449 LMAIKQSGDRRFDRGFQVWLQR 470
>At3g32110 non-LTR reverse transcriptase, putative
Length = 1911
Score = 70.5 bits (171), Expect = 1e-12
Identities = 69/279 (24%), Positives = 107/279 (37%), Gaps = 30/279 (10%)
Query: 6 GGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLREGGVTGSAW---W 62
GGLG+R N AL+ K WR L D LW +V+ S+Y R V S W W
Sbjct: 1390 GGLGIRSATAMNKALIAKVGWRVLNDGSSLWAQVVRSKYKVGDVHDRNWTVAKSNWSSTW 1449
Query: 63 REIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLSIHKD 122
R + G+G+ W E+ +G+G FW+D W+ + D +
Sbjct: 1450 RSV-----GVGLRDVIWREQHWV--IGDGRQIRFWTDRWLSETPIAD------DSIVPLS 1496
Query: 123 LSVGEMHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDT---QPDVWLWQPN 179
L+ A DG W ++ V + + L + D+ D W
Sbjct: 1497 LAQMLCTARDLWRDGTGWD-----MSQIAPFVTDNKRLDLLAVIVDSVTGAHDRLAWGMT 1551
Query: 180 IGDGYTVRGVYQMLMRQEVHNHDVVS--DAPWHKSVPLKVSICAWRLLRNRWPTKDNLVR 237
+TV+ + ML + D+ S W P +V + W ++ T R
Sbjct: 1552 SDGRFTVKSAFAMLTNDDSPRQDMSSLYGRVWKVQAPERVRVFLWLVVNQAIMTNSERKR 1611
Query: 238 RGVITNDT-QLCVRGCGKNETIDHLIIHCLIFGDLWQRI 275
R + +D Q+C G E+I H++ C +W RI
Sbjct: 1612 RHLCDSDVCQVCRGGI---ESILHVLRDCPAMSGIWDRI 1647
>At1g51750 hypothetical protein
Length = 629
Score = 69.3 bits (168), Expect = 3e-12
Identities = 95/366 (25%), Positives = 140/366 (37%), Gaps = 71/366 (19%)
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRG--RLREGGVTGSA 60
K GGLG+R L E N+ + K WR + + LW K +Q L GS
Sbjct: 274 KQEGGLGLRSLTEANVVSVLKLIWRVTSNDDSLWVKWSKMNLLKQESFWSLTPNSSLGSW 333
Query: 61 WWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFME--RFRRLYDLS 118
W++++K + E F + NG T FW D W G M+ R DL
Sbjct: 334 MWKKMLKYR-----ETAKPFSR---VEVNNGARTSFWFDNWSGMGHLMDVTGQRGQIDLG 385
Query: 119 IHKDLSVGEMHALGWGEDGEAWRWRRRLLAWEEEL---------VVEIRNLLTNVTLQDT 169
I ++ +V EAW RRR E+L + RNLL
Sbjct: 386 ISRNKTV-----------AEAWSNRRRRKHRTEQLNDIEAALNQKYQTRNLL-------- 426
Query: 170 QPDVWLWQPNIGD----GYTVRGVYQMLMRQEVHNHDVVSDAPWHKSV-----PLKVSIC 220
+ D LW+ GD ++ + + + ++ ++ W+K V K C
Sbjct: 427 REDATLWRGK-GDVFKTSFSTKDTWNQVRKKS-------NEVAWYKGVWFSHSTPKYQFC 478
Query: 221 AWRLLRNRWPTKDNLVRRGVITNDTQLCVRGCGKN-ETIDHLIIHCLIFGDLWQRIKTWI 279
W LRNR T R + N + + C + ET DHL C +W I +
Sbjct: 479 TWLALRNRLSTG---YRMQLWNNGSDVKCTFCSTSIETRDHLFFSCSYASAIWTAIAKNV 535
Query: 280 --GVFSVDPQQVLDHYYQFVYSSGCYNPR-RSFL-QLIWLCGIWVLSNERNQRLFTNTAR 335
FS D Q +++ Y S R RSFL + I+ + + ERN R R
Sbjct: 536 LQHRFSTDWQTIVN------YISETQTDRIRSFLSRYIFQLTVHTVWKERNDRRHGEEPR 589
Query: 336 TTVQLL 341
T+ L+
Sbjct: 590 TSANLI 595
>At3g44650 putative protein
Length = 762
Score = 66.6 bits (161), Expect = 2e-11
Identities = 74/285 (25%), Positives = 114/285 (39%), Gaps = 46/285 (16%)
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQR--GRLREGGVTGSA 60
K GGLG+R L+E N K WR + + LW K + ++ ++E GS
Sbjct: 422 KSEGGLGLRSLKEANDVCCLKLVWRIISHGDSLWVKWVEHNLLKREIFWIVKENANLGSW 481
Query: 61 WWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCW--VGTVSFMERFRRLYDLS 118
W++I+K + G+ + +GNG +T FW D W +G + + R D+
Sbjct: 482 IWKKILKYR-GVA-------KRFCKAEVGNGESTSFWFDDWSLLGRLIDVAGIRGTIDMG 533
Query: 119 IHKDLSVGEMHALGWGEDGEAWRWRRRLLAWEEEL--VVEIRNLLTNVTLQDTQPDVWLW 176
I + +SV +AW RRR +E L + E+ + Q Q LW
Sbjct: 534 ISRTMSV-----------ADAWTSRRRRHHRQEILNTIEEVLSTQHQKRTQQQQQGRVLW 582
Query: 177 QPN---IGDGYTVRGVYQMLMRQEVHNHDVVSDAPWHKSV-----PLKVSICAWRLLRNR 228
+ D ++ + + L ++ WHK V K S C W +R
Sbjct: 583 KGKNDIYKDKFSTKNTWNYL-------RTTSNEVAWHKGVWFPHATPKYSFCLWLAAHDR 635
Query: 229 WPTKDNLVR--RGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDL 271
T +++ RG T D C +G ET DHL L G L
Sbjct: 636 LATGARMIKWNRGE-TGDCTFCRQGI---ETRDHLFFMQLHIGSL 676
>At4g08830 putative protein
Length = 947
Score = 63.5 bits (153), Expect = 2e-10
Identities = 69/283 (24%), Positives = 109/283 (38%), Gaps = 68/283 (24%)
Query: 1 MSKGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLREGGVTGSA 60
+ K GGLG+R + N AL+ K WR + DR LW ++L S+Y R LRE GS
Sbjct: 523 LPKSEGGLGIRTSKCMNKALVSKVGWRLINDRYSLWARILRSKY---RVGLREVVSRGSR 579
Query: 61 WWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLSIH 120
W +GNG + FWSD W+ + + R + +
Sbjct: 580 W-------------------------VVGNGRDILFWSDNWLSHEALINR-AVIEIPNSE 613
Query: 121 KDLSVGEMHA--LGWG----EDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDTQPDVW 174
K+L V ++ A LGW E ++ R L A + V R+ L+
Sbjct: 614 KELRVKDLWANGLGWKLDKIEPYISYHTRLELAAVVVDSVTGARDRLS------------ 661
Query: 175 LWQPNIGDGYTVRGVYQMLMRQEVHNHDVVS--DAPWHKSVPLKVSICAWRLLRNRWPTK 232
W + +TV+ Y++L ++ + D W +V W +
Sbjct: 662 -WGYSADGVFTVKSAYRLLTEDHDPRPNMAAFFDRLWRVVALERVKTFLWHI-------- 712
Query: 233 DNLVRRGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRI 275
DT +C G +ETI H++ C +W+R+
Sbjct: 713 ----------GDTSVCQVCKGGDETILHVLKDCPSIAGIWRRL 745
>At1g30390 hypothetical protein
Length = 504
Score = 63.2 bits (152), Expect = 2e-10
Identities = 75/277 (27%), Positives = 112/277 (40%), Gaps = 39/277 (14%)
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQ--RGRLREGGVTGSA 60
K GGLG+R L+E N K WR + + LW K + S ++ +RE GS
Sbjct: 199 KEEGGLGLRSLKEANDVCCLKLIWRIISHADSLWVKWIQSSLLKKVFFWAVRENTSLGSW 258
Query: 61 WWREIVKIQNGIGVEGGSWFEESISK-RLGNGFNTFFWSDCW--VGTVSFMERFRRLYDL 117
WR+I+K ++ ++ K + NG T FW D W +G + R DL
Sbjct: 259 MWRKILKFRD---------IARTLCKVEINNGAQTSFWYDDWSDLGRLIESAGDRGAIDL 309
Query: 118 SIHKDLSVGEMHALGWGEDGEAWRWRRRLLAWEEELVVEIRNLLTNVTLQDTQPDVWLWQ 177
I+K +V E WG R RRR A V E L+ + ++ D LW+
Sbjct: 310 GINKHATVVE----AWGN-----RRRRRHRANFLNRVEE--RLVLSWNSRNQAEDCALWK 358
Query: 178 PNIGDGYTVRGVYQMLMRQEVHNH--DVVSDAPWHKSVPL-----KVSICAWRLLRNRWP 230
G R ++ ++ NH V + W+K V K + C W + NR
Sbjct: 359 ---GKENRFRSIFS---TKDTWNHIRTVSNKVAWYKGVWFAQAIPKHAFCMWLAVHNRLS 412
Query: 231 TKDNLVRRGVITNDT-QLCVRGCGKNETIDHLIIHCL 266
T D + + + T LC + + HL H L
Sbjct: 413 TGDRMTLWNMGVDATCILCNNALESRDHLQHLFFHGL 449
>At1g24640 hypothetical protein
Length = 1270
Score = 62.4 bits (150), Expect = 4e-10
Identities = 60/242 (24%), Positives = 104/242 (42%), Gaps = 43/242 (17%)
Query: 1 MSKGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGRLREG-GVTGS 59
+ K GGLG R +Q FN ALL K WR L + L ++L SRY + L S
Sbjct: 809 LPKDSGGLGFRDIQSFNQALLAKQAWRLLHFPDCLLSRLLKSRYFDATDFLDAALSQRPS 868
Query: 60 AWWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRR--LYDL 117
WR I + G + + KR+G+G + F W D W+ F +R+ +YD+
Sbjct: 869 FGWRSI--------LFGRELLSKGLQKRVGDGASLFVWIDPWIDDNGFRAPWRKNLIYDV 920
Query: 118 SIHKDLSVGEMHALGWGEDGEAWRWRRRLLAWEEELVVEI---RNLLTNVTLQD--TQPD 172
++ ++ AL R W+EE++ ++ ++L ++ +Q D
Sbjct: 921 TL-------KVKAL----------LNPRTGFWDEEVLHDLFLPEDILRIKAIKPVISQAD 963
Query: 173 VWLWQPNIGDGYTVRGVYQMLMRQEVHN-HDVVSDAP---------WHKSVPLKVSICAW 222
++W+ N ++V+ Y + + + N VS P W+ K+ I W
Sbjct: 964 FFVWKLNKSGDFSVKSAYWLAYQTKSQNLRSEVSMQPSTLGLKTQVWNLQTDPKIKIFLW 1023
Query: 223 RL 224
++
Sbjct: 1024 KV 1025
>At1g19030 hypothetical protein
Length = 398
Score = 58.5 bits (140), Expect = 6e-09
Identities = 70/280 (25%), Positives = 121/280 (43%), Gaps = 30/280 (10%)
Query: 3 KGVGGLGVRRLQEFNIALLGKWCWRCLVDREGLWFKVLSSRYGEQRGR---LREGGVTGS 59
K GGLG+R + E N + K WR L R LW + L +Y +RG ++E GS
Sbjct: 98 KQEGGLGLRSIAEANKVCVLKLIWRILSARRSLWVE-LVKKYLIRRGSFWLVKENTTCGS 156
Query: 60 AWWREIVKIQNGIGVEGGSWFEESISKRLGNGFNTFFWSDCWVGTVSFMERFRRLYDLSI 119
W++I+K ++ S++ + + F WS+ +G + R DL
Sbjct: 157 WIWKKILKYRD----TAKSFYRVEVRNGEASSFGFDRWSE--MGCLYDRLGARGCIDLGS 210
Query: 120 HKDLSVGEMHALGWGEDGEAWRWRRRLLAWEEELVVE---IRNLLTNVTLQDTQPDVWLW 176
+VGE+ A ++R +L E +V+ IR+ T+V L + + D +
Sbjct: 211 PMTATVGEVMA-----GARRRKYRVAILNQVENEIVKQKLIRSNETDVALWNGKHDHYRK 265
Query: 177 QPNIGDGYTVRGVYQMLMRQEVHNHDVVSDAPWHKSVPLKVSICAWRLLRNRWPTKDNLV 236
+ + + + ++LM N++ + W K K S+ W + +NR T D L+
Sbjct: 266 EFHTKETWLQIRTAKLLM----ENYNGI----WFKHSTPKYSLITWLVQKNRVATGDKLI 317
Query: 237 R-RGVITNDTQLCVRGCGKNETIDHLIIHCLIFGDLWQRI 275
+ +T + LC ET DHL C +W ++
Sbjct: 318 KWNPQVTPNCILCQHTL---ETWDHLFFMCPYSFQVWTKL 354
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.141 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,478,138
Number of Sequences: 26719
Number of extensions: 423413
Number of successful extensions: 1279
Number of sequences better than 10.0: 83
Number of HSP's better than 10.0 without gapping: 45
Number of HSP's successfully gapped in prelim test: 38
Number of HSP's that attempted gapping in prelim test: 1109
Number of HSP's gapped (non-prelim): 116
length of query: 377
length of database: 11,318,596
effective HSP length: 101
effective length of query: 276
effective length of database: 8,619,977
effective search space: 2379113652
effective search space used: 2379113652
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC127021.2


