

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC127019.10 + phase: 0
(113 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
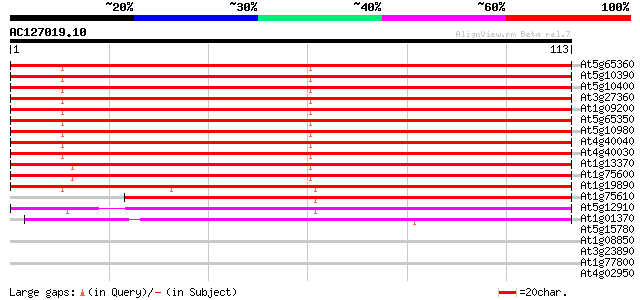
Sequences producing significant alignments: (bits) Value
At5g65360 histone H3 (sp|P05203) 135 4e-33
At5g10390 histone H3 - like protein 135 4e-33
At5g10400 histone H3 - like protein 135 4e-33
At3g27360 histone H3 like protein 135 4e-33
At1g09200 histone H3 like protein 135 4e-33
At5g65350 histone H3 130 1e-31
At5g10980 histon H3 protein 129 2e-31
At4g40040 Histon H3 129 2e-31
At4g40030 histone H3.3 129 2e-31
At1g13370 unknown protein 128 5e-31
At1g75600 histone H3, putative 121 6e-29
At1g19890 hypothetical protein 113 1e-26
At1g75610 histone H3, putative 94 8e-21
At5g12910 histone H3 -like protein 86 4e-18
At1g01370 centromeric histone H3 HTR12 51 8e-08
At5g15780 proline-rich protein 26 2.8
At1g08850 hypothetical protein, 3' partial 26 2.8
At3g23890 topoisomerase II 25 4.8
At1g77800 putative phorbol ester / diacylglycerol binding protein 25 4.8
At4g02950 putative protein 25 6.3
>At5g65360 histone H3 (sp|P05203)
Length = 136
Score = 135 bits (339), Expect = 4e-33
Identities = 77/116 (66%), Positives = 84/116 (72%), Gaps = 3/116 (2%)
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA A G VKKPH F GTVALREIR YQ+ST
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQSSAV+A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAK 116
>At5g10390 histone H3 - like protein
Length = 136
Score = 135 bits (339), Expect = 4e-33
Identities = 77/116 (66%), Positives = 84/116 (72%), Gaps = 3/116 (2%)
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA A G VKKPH F GTVALREIR YQ+ST
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQSSAV+A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAK 116
>At5g10400 histone H3 - like protein
Length = 136
Score = 135 bits (339), Expect = 4e-33
Identities = 77/116 (66%), Positives = 84/116 (72%), Gaps = 3/116 (2%)
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA A G VKKPH F GTVALREIR YQ+ST
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQSSAV+A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAK 116
>At3g27360 histone H3 like protein
Length = 136
Score = 135 bits (339), Expect = 4e-33
Identities = 77/116 (66%), Positives = 84/116 (72%), Gaps = 3/116 (2%)
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA A G VKKPH F GTVALREIR YQ+ST
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQSSAV+A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAK 116
>At1g09200 histone H3 like protein
Length = 136
Score = 135 bits (339), Expect = 4e-33
Identities = 77/116 (66%), Positives = 84/116 (72%), Gaps = 3/116 (2%)
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA A G VKKPH F GTVALREIR YQ+ST
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPATGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQSSAV+A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAK 116
>At5g65350 histone H3
Length = 139
Score = 130 bits (326), Expect = 1e-31
Identities = 74/116 (63%), Positives = 83/116 (70%), Gaps = 3/116 (2%)
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLA KAAR+SA A G VKKPH F GTVALR+IR YQ+ST
Sbjct: 1 MARTKQTARISTGGKAPRKQLAPKAARQSAPATGGVKKPHRFRPGTVALRDIRKYQKSTE 60
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQSSAV+A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 ILIRKLPFQRLVREIAQDFKTDLRFQSSAVAALQEAAEAYLVGLFEDTNLCAIHAK 116
>At5g10980 histon H3 protein
Length = 136
Score = 129 bits (324), Expect = 2e-31
Identities = 74/116 (63%), Positives = 81/116 (69%), Gaps = 3/116 (2%)
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA G VKKPH + GTVALREIR YQ+ST
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 60
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQS AV A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAK 116
>At4g40040 Histon H3
Length = 136
Score = 129 bits (324), Expect = 2e-31
Identities = 74/116 (63%), Positives = 81/116 (69%), Gaps = 3/116 (2%)
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA G VKKPH + GTVALREIR YQ+ST
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 60
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQS AV A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAK 116
>At4g40030 histone H3.3
Length = 136
Score = 129 bits (324), Expect = 2e-31
Identities = 74/116 (63%), Positives = 81/116 (69%), Gaps = 3/116 (2%)
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR +TGGKAPRKQLATKAARKSA G VKKPH + GTVALREIR YQ+ST
Sbjct: 1 MARTKQTARKSTGGKAPRKQLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 60
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FKT LRFQS AV A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 LLIRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAK 116
>At1g13370 unknown protein
Length = 136
Score = 128 bits (321), Expect = 5e-31
Identities = 73/116 (62%), Positives = 80/116 (68%), Gaps = 3/116 (2%)
Query: 1 MAGTKQTARNT-GGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQ+AR + GGKAP KQLATKAARKSA G VKKPH F GTVALREIR YQ+ST
Sbjct: 1 MARTKQSARKSHGGKAPTKQLATKAARKSAPTTGGVKKPHRFRPGTVALREIRKYQKSTE 60
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
N++ FKT LRFQS AV A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 LLNRKLPFQRLVREIAQDFKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAK 116
>At1g75600 histone H3, putative
Length = 136
Score = 121 bits (303), Expect = 6e-29
Identities = 70/116 (60%), Positives = 78/116 (66%), Gaps = 3/116 (2%)
Query: 1 MAGTKQTARNT-GGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA TKQTAR + GGKAPR LATKAARKSA G VKKPH + GTVALREIR YQ+ST
Sbjct: 1 MARTKQTARKSHGGKAPRTLLATKAARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTE 60
Query: 60 --NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ +KT LRFQS AV A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 LLIRKLPFQRLVREIAQDYKTDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAK 116
>At1g19890 hypothetical protein
Length = 137
Score = 113 bits (283), Expect = 1e-26
Identities = 68/117 (58%), Positives = 77/117 (65%), Gaps = 4/117 (3%)
Query: 1 MAGTKQTAR-NTGGKAPRKQLATKAARKSALA-AGTVKKPHHFCHGTVALREIRNYQRST 58
MA TKQTAR +TGGK PRK+LATKAARK+ G VK+ H F GTVALREIR YQ+ST
Sbjct: 1 MARTKQTARKSTGGKGPRKELATKAARKTRRPYRGGVKRAHRFRPGTVALREIRKYQKST 60
Query: 59 SN--KEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ FK LRFQS AV A+QE EA V LFEDTN+CAIHAK
Sbjct: 61 DLLIRKLPFQRLVREIAQDFKVDLRFQSHAVLALQEAAEAYLVGLFEDTNLCAIHAK 117
>At1g75610 histone H3, putative
Length = 115
Score = 94.4 bits (233), Expect = 8e-21
Identities = 53/92 (57%), Positives = 59/92 (63%), Gaps = 2/92 (2%)
Query: 24 AARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTSN--KEASVSETCEGNCSGFKTGLR 81
AARKSA G VKKPH + GTVALREIR YQ+ST ++ FKT LR
Sbjct: 4 AARKSAPTTGGVKKPHRYRPGTVALREIRKYQKSTELLIRKLPFQRLVREIAQDFKTDLR 63
Query: 82 FQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
FQS AV A+QE EA V LFEDTN+CAIHAK
Sbjct: 64 FQSHAVLALQEAAEAYLVGLFEDTNLCAIHAK 95
>At5g12910 histone H3 -like protein
Length = 131
Score = 85.5 bits (210), Expect = 4e-18
Identities = 53/116 (45%), Positives = 68/116 (57%), Gaps = 8/116 (6%)
Query: 1 MAGTKQTARN-TGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTS 59
MA + QTAR TGGKAP A R + +KKP+ + GTVALREIR YQ++T
Sbjct: 1 MARSNQTARKATGGKAPHF-----AMRVWQHSTPPLKKPYRYKPGTVALREIRKYQKTTD 55
Query: 60 N--KEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
++ + K LRFQ+ AVSA+QE EA V +FEDTN+CA+HAK
Sbjct: 56 LVIRKLPFQRLVKEIAQSLKADLRFQTGAVSALQEAAEAFMVGMFEDTNLCAMHAK 111
>At1g01370 centromeric histone H3 HTR12
Length = 178
Score = 51.2 bits (121), Expect = 8e-08
Identities = 36/114 (31%), Positives = 59/114 (51%), Gaps = 6/114 (5%)
Query: 4 TKQTARNTGGKAPRKQLATKAARKSALAAGTVKKPHHFCHGTVALREIRNYQRSTSNKEA 63
T+QT T ++ A ++ + A+ G+ KK + + GTVAL+EIR++Q+ T+
Sbjct: 45 TQQTNPTTSPATGTRRGAKRS--RQAMPRGSQKKSYRYRPGTVALKEIRHFQKQTNLLIP 102
Query: 64 SVSETCEGNCSGFKTGL----RFQSSAVSAVQEVVEACFVSLFEDTNICAIHAK 113
+ S E R+ + A+ A+QE E V LF D+ +CAIHA+
Sbjct: 103 AASFIREVRSITHMLAPPQINRWTAEALVALQEAAEDYLVGLFSDSMLCAIHAR 156
>At5g15780 proline-rich protein
Length = 401
Score = 26.2 bits (56), Expect = 2.8
Identities = 30/112 (26%), Positives = 45/112 (39%), Gaps = 21/112 (18%)
Query: 12 GGKAPRKQLATKAARKSALAAGTV-----------KKPHHFCHGT-VALREIRNYQRSTS 59
GG + +Q K R SA+ GTV K P+H G VA+ I + +
Sbjct: 23 GGLSQGQQHVMKKTRSSAVVVGTVYCDTCFNGAFSKSPNHLISGALVAVECIDENSKPSF 82
Query: 60 NKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTN-ICAI 110
+E + E FK L F +VS + ++ C V L + C+I
Sbjct: 83 RQEVKTDKRGE-----FKVKLPF---SVSKHVKKIKRCSVKLLSSSQPYCSI 126
>At1g08850 hypothetical protein, 3' partial
Length = 835
Score = 26.2 bits (56), Expect = 2.8
Identities = 15/57 (26%), Positives = 24/57 (41%)
Query: 55 QRSTSNKEASVSETCEGNCSGFKTGLRFQSSAVSAVQEVVEACFVSLFEDTNICAIH 111
+R SN+ A+V+ F+ GL +S +V +QE E C +H
Sbjct: 418 ERLRSNEHATVALLGTLQHQVFQAGLSQESPSVDGLQEYASTVIEKSIESLYACGVH 474
>At3g23890 topoisomerase II
Length = 1473
Score = 25.4 bits (54), Expect = 4.8
Identities = 21/67 (31%), Positives = 24/67 (35%), Gaps = 2/67 (2%)
Query: 15 APRKQLATKAARKSALAAGTV--KKPHHFCHGTVALREIRNYQRSTSNKEASVSETCEGN 72
APRK+ A LA G KK + R NKE SE GN
Sbjct: 1363 APRKRGKQTVASTEVLAIGVSPEKKVRKMRSSPFNKKSSSVMSRLADNKEEESSENVAGN 1422
Query: 73 CSGFKTG 79
S K+G
Sbjct: 1423 SSSEKSG 1429
>At1g77800 putative phorbol ester / diacylglycerol binding protein
Length = 1506
Score = 25.4 bits (54), Expect = 4.8
Identities = 18/68 (26%), Positives = 30/68 (43%), Gaps = 8/68 (11%)
Query: 13 GKAPRKQLATKAARKSALAA-GTVKKPHHFCHGTVA-------LREIRNYQRSTSNKEAS 64
GK +++ TK+ R++A+ G V VA EI +T+N +
Sbjct: 634 GKNLKRKTTTKSERRAAICTEGIVMLDSDILDPAVAKAFSIERTHEINICNNTTNNTRCT 693
Query: 65 VSETCEGN 72
++E C GN
Sbjct: 694 LTENCTGN 701
>At4g02950 putative protein
Length = 318
Score = 25.0 bits (53), Expect = 6.3
Identities = 11/26 (42%), Positives = 17/26 (65%)
Query: 45 TVALREIRNYQRSTSNKEASVSETCE 70
TVA+ IRN + N+E+S S++ E
Sbjct: 194 TVAVGSIRNRDQEVKNRESSPSDSVE 219
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.315 0.125 0.355
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,131,038
Number of Sequences: 26719
Number of extensions: 64426
Number of successful extensions: 196
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 18
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 152
Number of HSP's gapped (non-prelim): 23
length of query: 113
length of database: 11,318,596
effective HSP length: 89
effective length of query: 24
effective length of database: 8,940,605
effective search space: 214574520
effective search space used: 214574520
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 52 (24.6 bits)
Medicago: description of AC127019.10


