

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126792.7 - phase: 0
(253 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
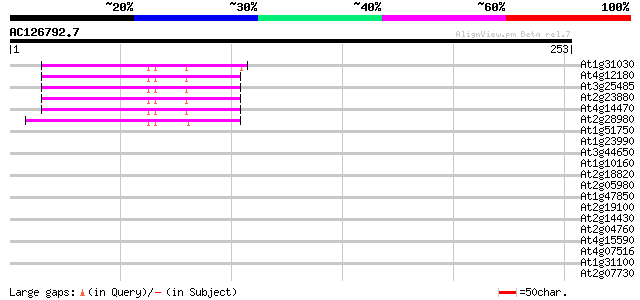
Score E
Sequences producing significant alignments: (bits) Value
At1g31030 F17F8.5 46 2e-05
At4g12180 putative reverse transcriptase 45 4e-05
At3g25485 unknown protein 45 4e-05
At2g23880 putative non-LTR retroelement reverse transcriptase 44 8e-05
At4g14470 reverse transcriptase like protein 44 1e-04
At2g28980 putative non-LTR retroelement reverse transcriptase 41 7e-04
At1g51750 hypothetical protein 40 0.001
At1g23990 reverse transcriptase, putative 40 0.001
At3g44650 putative protein 39 0.002
At1g10160 putative reverse transcriptase 39 0.002
At2g18820 putative non-LTR retroelement reverse transcriptase 39 0.003
At2g05980 putative non-LTR retroelement reverse transcriptase 38 0.005
At1g47850 hypothetical protein 37 0.008
At2g19100 putative non-LTR retroelement reverse transcriptase 37 0.010
At2g14430 putative non-LTR retroelement reverse transcriptase 37 0.010
At2g04760 putative non-LTR retroelement reverse transcriptase 37 0.010
At4g15590 reverse transcriptase like protein 35 0.039
At4g07516 putative protein 35 0.039
At1g31100 hypothetical protein 35 0.039
At2g07730 putative non-LTR retroelement reverse transcriptase 35 0.050
>At1g31030 F17F8.5
Length = 872
Score = 46.2 bits (108), Expect = 2e-05
Identities = 34/110 (30%), Positives = 55/110 (49%), Gaps = 17/110 (15%)
Query: 15 ASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA--------PRC-- 64
AS SV+ F GL QG +LSPY+F + +DVL++ + + A P+C
Sbjct: 283 ASFSVQVNGDLVGYFQSKRGLRQGCSLSPYLFVICMDVLSKMLDKAAGVRKFGFHPKCQR 342
Query: 65 -----MLFADDVVLVGDSR-EEVNGRLETWRQALEAYGFCLSRSK-TEYM 107
+ FADD++++ D + + G LE + + + G +S K T YM
Sbjct: 343 LGLTHLSFADDLMVLSDGKTRSIEGILEVFDEFCKRSGLRISLEKSTLYM 392
>At4g12180 putative reverse transcriptase
Length = 662
Score = 45.1 bits (105), Expect = 4e-05
Identities = 31/106 (29%), Positives = 53/106 (49%), Gaps = 16/106 (15%)
Query: 15 ASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA--------PRC-- 64
AS SV+ F GL QG +L+PY+F +V+DVL++ + A P+C
Sbjct: 154 ASFSVQVNRELAGYFNSLRGLRQGCSLTPYLFVIVMDVLSKKLDRAAGLRKFGYHPKCKN 213
Query: 65 -----MLFADDVVLVGDSR-EEVNGRLETWRQALEAYGFCLSRSKT 104
+ FADD++++ D + + G +E + + G +S +KT
Sbjct: 214 LGLTHLSFADDIMVLTDGKLRSLEGIVEVFDSFAKQSGLKISMAKT 259
>At3g25485 unknown protein
Length = 979
Score = 45.1 bits (105), Expect = 4e-05
Identities = 31/106 (29%), Positives = 52/106 (48%), Gaps = 16/106 (15%)
Query: 15 ASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA--------PRC-- 64
AS SV+ F + GL QG +LSPY+F + +DVL++ + A PRC
Sbjct: 552 ASFSVQVNGELVGFFNSSRGLRQGCSLSPYLFVIAMDVLSKMLDRAAGFKKFGYHPRCQT 611
Query: 65 -----MLFADDVVLVGDSR-EEVNGRLETWRQALEAYGFCLSRSKT 104
+ FADD++++ D + V G + + + + G +S K+
Sbjct: 612 IGLTHLTFADDLMVLSDGKVRSVEGIVSVFDEFAKKSGLKISMEKS 657
>At2g23880 putative non-LTR retroelement reverse transcriptase
Length = 1216
Score = 43.9 bits (102), Expect = 8e-05
Identities = 31/106 (29%), Positives = 50/106 (46%), Gaps = 16/106 (15%)
Query: 15 ASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA--------PRC-- 64
AS S++ F GL QG +LSPY+F + +DVL+ + + A PRC
Sbjct: 357 ASFSIQVNGELAGYFRSARGLRQGCSLSPYLFVISMDVLSRMLDKAAGAREFGYHPRCKT 416
Query: 65 -----MLFADDVVLVGDSR-EEVNGRLETWRQALEAYGFCLSRSKT 104
+ FADD++++ D + V+G ++ Q G + KT
Sbjct: 417 LGLTHLCFADDLMILTDGKIRSVDGIVKVLNQFAAKLGLKICMEKT 462
>At4g14470 reverse transcriptase like protein
Length = 318
Score = 43.5 bits (101), Expect = 1e-04
Identities = 31/106 (29%), Positives = 52/106 (48%), Gaps = 16/106 (15%)
Query: 15 ASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA--------PRC-- 64
AS SV+ F + GL QG +LSPY+F +V+DVL++ + A P+C
Sbjct: 72 ASFSVQVNGELAGYFNSSRGLRQGCSLSPYLFVIVMDVLSKKLDRAAGLRKFGYHPKCKN 131
Query: 65 -----MLFADDVVLVGDSR-EEVNGRLETWRQALEAYGFCLSRSKT 104
+ FADD++++ D + + G +E + + +S KT
Sbjct: 132 LGLTHLSFADDIMVLTDGKLRSLEGIVEVFDSFAKQSCLKISMEKT 177
>At2g28980 putative non-LTR retroelement reverse transcriptase
Length = 1529
Score = 40.8 bits (94), Expect = 7e-04
Identities = 31/113 (27%), Positives = 52/113 (45%), Gaps = 16/113 (14%)
Query: 8 IKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA------ 61
IK A+ SV+ F + GL QG LSPY+F + ++VL+ I E A
Sbjct: 926 IKLCISTATFSVQVNGELAGFFGSSRGLRQGCALSPYLFVICMNVLSHMIDEAAVHRNIG 985
Query: 62 --PRC-------MLFADDVVLVGDSRE-EVNGRLETWRQALEAYGFCLSRSKT 104
P+C + FADD+++ D + + G + +++ G +S K+
Sbjct: 986 YHPKCEKIGLTHLCFADDLMVFVDGHQWSIEGVINVFKEFAGRSGLQISLEKS 1038
>At1g51750 hypothetical protein
Length = 629
Score = 40.4 bits (93), Expect = 0.001
Identities = 30/117 (25%), Positives = 53/117 (44%), Gaps = 16/117 (13%)
Query: 4 YIRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA-- 61
+I I S SV+ F G+ QG LSPY+F + ++VL++ + + A
Sbjct: 30 FIHWISRCITTTSFSVQVNGELAGYFRSARGIRQGCALSPYLFVISMEVLSKMLDQAAGG 89
Query: 62 ------PRC-------MLFADDVVLVGDSR-EEVNGRLETWRQALEAYGFCLSRSKT 104
P+C + FADD++++ D + V+G +E + G ++ KT
Sbjct: 90 KRFGFHPKCKNLGLTHLCFADDLMILTDGKVRSVDGIVEVMNLFAKRSGLQINMEKT 146
>At1g23990 reverse transcriptase, putative
Length = 657
Score = 40.4 bits (93), Expect = 0.001
Identities = 30/106 (28%), Positives = 52/106 (48%), Gaps = 16/106 (15%)
Query: 15 ASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA--------PRC-- 64
AS SV+ F GL QG +LSPY+F + +DVL++ + + A RC
Sbjct: 85 ASFSVQVNGELVGFFQSKRGLRQGCSLSPYLFVMSMDVLSKLLDQAASAKKFGYHSRCKE 144
Query: 65 -----MLFADDVVLVGDSR-EEVNGRLETWRQALEAYGFCLSRSKT 104
+ FADD++++ D + ++G +E + + G +S K+
Sbjct: 145 LSLTHLSFADDLMVLSDGKVRSIDGIVEVFDIFAKFSGLKISMEKS 190
>At3g44650 putative protein
Length = 762
Score = 39.3 bits (90), Expect = 0.002
Identities = 29/113 (25%), Positives = 51/113 (44%), Gaps = 16/113 (14%)
Query: 16 STSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA--------PRC--- 64
S SV+ F + GL QG LSPY+F + +DVL++ + ++ P C
Sbjct: 190 SFSVQVNGELAGFFQSSRGLRQGCALSPYLFVICMDVLSKLLDKVVGIGRIGYHPHCKRM 249
Query: 65 ----MLFADDVVLVGDSR-EEVNGRLETWRQALEAYGFCLSRSKTEYMECNFS 112
+ FADD++++ D + + G +E + + G +S K+ S
Sbjct: 250 GLTHLSFADDLMILTDGQCRSIEGIIEVFDLFSKWSGLKISMEKSTIFSAGLS 302
>At1g10160 putative reverse transcriptase
Length = 1118
Score = 39.3 bits (90), Expect = 0.002
Identities = 30/101 (29%), Positives = 48/101 (46%), Gaps = 18/101 (17%)
Query: 34 GLHQGSTLSPYIFTLVLDVLTEHIQ--------ELAPRCML-------FADDVVLVGD-S 77
G+ QG +S ++F LV+DVL++ + L P C+ FADDV++ D +
Sbjct: 649 GIRQGDPMSSHLFVLVMDVLSKSLDLGALNGLFNLHPNCLAPIITHLSFADDVLVFSDGA 708
Query: 78 REEVNGRLETWRQALEAYGFCLSRSKTEYM--ECNFSGRRS 116
+ G L + G ++R KTE + NF+ RS
Sbjct: 709 ASSIAGILTILDDFRQGSGLGINREKTELLLDGGNFARNRS 749
>At2g18820 putative non-LTR retroelement reverse transcriptase
Length = 1412
Score = 38.5 bits (88), Expect = 0.003
Identities = 31/113 (27%), Positives = 50/113 (43%), Gaps = 17/113 (15%)
Query: 25 TTEDFPITI-GLHQGSTLSPYIFTLVLDVLTEHIQELA--------PRC-------MLFA 68
+T F + + GL QG +LSPY+F + ++VL+ + + A PRC + FA
Sbjct: 900 STASFSVQVNGLRQGCSLSPYLFVICMNVLSAMLDKGAVEKRFGYHPRCRNMGLTHLCFA 959
Query: 69 DDV-VLVGDSREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGRRSKSTL 120
DD+ V S + G L ++ G +S K+ + S S L
Sbjct: 960 DDIMVFSAGSAHSLEGVLAIFKDFAAFSGLNISLEKSTLFMASISSETCASIL 1012
>At2g05980 putative non-LTR retroelement reverse transcriptase
Length = 1352
Score = 38.1 bits (87), Expect = 0.005
Identities = 31/123 (25%), Positives = 54/123 (43%), Gaps = 16/123 (13%)
Query: 15 ASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA--------PRC-- 64
AS SV+ + F GL QG +LSPY++ + ++VL+ + + A PRC
Sbjct: 783 ASFSVQVNGELSGFFRSERGLRQGCSLSPYLYVICMNVLSCMLDKAAVEKKISYHPRCRN 842
Query: 65 -----MLFADDVVLVGD-SREEVNGRLETWRQALEAYGFCLSRSKTEYMECNFSGRRSKS 118
+ FADD+++ D + + + G L + + +S K+ S S
Sbjct: 843 MNLTHLCFADDIMVFSDGTSKSIQGTLAIFEKFAAMSWLKISLEKSTIFMAGISPNAKTS 902
Query: 119 TLE 121
L+
Sbjct: 903 ILQ 905
>At1g47850 hypothetical protein
Length = 799
Score = 37.4 bits (85), Expect = 0.008
Identities = 31/113 (27%), Positives = 49/113 (42%), Gaps = 16/113 (14%)
Query: 8 IKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA------ 61
IK A+ SV+ F GL QG LSPY+F + ++VL+ I A
Sbjct: 202 IKLCISTATFSVQVNGELAGFFGSKRGLRQGCALSPYLFVICMNVLSHMIDVAAVHRNIG 261
Query: 62 --PRC-------MLFADD-VVLVGDSREEVNGRLETWRQALEAYGFCLSRSKT 104
P+C + FADD +V + + V G + +++ G +S K+
Sbjct: 262 YHPKCKKLSLTHLCFADDLMVFIDGQQRSVEGVINIFKEFAGKSGLHISLEKS 314
>At2g19100 putative non-LTR retroelement reverse transcriptase
Length = 1447
Score = 37.0 bits (84), Expect = 0.010
Identities = 25/94 (26%), Positives = 43/94 (45%), Gaps = 17/94 (18%)
Query: 15 ASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQE-------------LA 61
AS SV+ F GL QG +LSPY+F + +D+L++ + + L
Sbjct: 990 ASFSVQVNGELAGFFQSARGLRQGCSLSPYLFVISMDILSKMLDKAAGQLPVRYLGLPLV 1049
Query: 62 PRCMLFADDVVLVGDSREEVNGRLETWRQALEAY 95
+ + AD + L+ E++ R+ TW +Y
Sbjct: 1050 TKRLSSADSLPLI----EQIRHRIGTWTSRFLSY 1079
>At2g14430 putative non-LTR retroelement reverse transcriptase
Length = 1277
Score = 37.0 bits (84), Expect = 0.010
Identities = 31/113 (27%), Positives = 48/113 (42%), Gaps = 16/113 (14%)
Query: 8 IKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA------ 61
IK A+ SV+ F GL QG LSPY+F + ++VL+ I A
Sbjct: 771 IKLCISTATFSVQVNSEQAGFFGSKRGLRQGCALSPYLFVICMNVLSHMIDVAAVHRNIG 830
Query: 62 --PRC-------MLFADD-VVLVGDSREEVNGRLETWRQALEAYGFCLSRSKT 104
P+C + FADD +V + + V G + ++ G +S K+
Sbjct: 831 YHPKCKKLSLTHLCFADDLMVFIDGQQRSVEGVINIFKDFAGKSGLHISLEKS 883
>At2g04760 putative non-LTR retroelement reverse transcriptase
Length = 631
Score = 37.0 bits (84), Expect = 0.010
Identities = 30/113 (26%), Positives = 48/113 (41%), Gaps = 16/113 (14%)
Query: 8 IKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELA------ 61
IK A+ SV+ F GL QG LSPY+F + ++VL+ I E A
Sbjct: 199 IKLCISTATFSVQVNGELAGFFGNKRGLRQGCALSPYLFVICMNVLSHMIDEAAVRRNIG 258
Query: 62 --PRC-------MLFADD-VVLVGDSREEVNGRLETWRQALEAYGFCLSRSKT 104
P+C + F DD +V + + + G + + + G +S K+
Sbjct: 259 YHPKCKKLSLTHLCFVDDLMVFIDGQQRSIEGVINIFHEFAGKSGLHISLEKS 311
>At4g15590 reverse transcriptase like protein
Length = 929
Score = 35.0 bits (79), Expect = 0.039
Identities = 23/90 (25%), Positives = 38/90 (41%), Gaps = 16/90 (17%)
Query: 4 YIRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQELAPR 63
+I+ I + G S+ T+ F GL QG +SPY+F L ++ L I+ R
Sbjct: 417 WIKRIMECVAGPEMSLLWNGEKTDSFTPERGLRQGDPISPYLFVLCIERLCHQIETAVGR 476
Query: 64 ----------------CMLFADDVVLVGDS 77
+ FADD++L ++
Sbjct: 477 GDWKSISISQGGPKVSHVCFADDLILFAEA 506
>At4g07516 putative protein
Length = 740
Score = 35.0 bits (79), Expect = 0.039
Identities = 20/56 (35%), Positives = 28/56 (49%)
Query: 3 AYIRAIKDMYEGASTSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQ 58
++I I G S S+ TE F + GL QG LSPY+F L L+ L ++
Sbjct: 253 SWIWWIMQCVTGPSMSLMWNGEKTEPFRPSRGLRQGDPLSPYLFVLCLERLCHQVE 308
>At1g31100 hypothetical protein
Length = 1090
Score = 35.0 bits (79), Expect = 0.039
Identities = 30/106 (28%), Positives = 47/106 (44%), Gaps = 22/106 (20%)
Query: 18 SVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQE---------------LAP 62
SV T F T GL QG+ LSP++F L ++V + + L+
Sbjct: 548 SVCVNGNTGGFFKSTRGLRQGNPLSPFLFVLAMEVFSSLLNSRFQAGYIHYHPKTSPLSI 607
Query: 63 RCMLFADDVVLVGDSREEVNGRLETWRQALEAY----GFCLSRSKT 104
++FADD+++ D + L +ALE + G L+R KT
Sbjct: 608 SHLMFADDIMVFFDGG---SSSLHGISEALEDFAFWSGLVLNREKT 650
>At2g07730 putative non-LTR retroelement reverse transcriptase
Length = 970
Score = 34.7 bits (78), Expect = 0.050
Identities = 19/43 (44%), Positives = 23/43 (53%)
Query: 16 STSVRTQDGTTEDFPITIGLHQGSTLSPYIFTLVLDVLTEHIQ 58
S SV G T+ F GL QG LSPY+F L L+ L I+
Sbjct: 327 SMSVLWNGGKTDSFVPARGLRQGDPLSPYLFVLCLERLCHLIE 369
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.135 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,498,845
Number of Sequences: 26719
Number of extensions: 219388
Number of successful extensions: 598
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 35
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 561
Number of HSP's gapped (non-prelim): 52
length of query: 253
length of database: 11,318,596
effective HSP length: 97
effective length of query: 156
effective length of database: 8,726,853
effective search space: 1361389068
effective search space used: 1361389068
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC126792.7


