

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
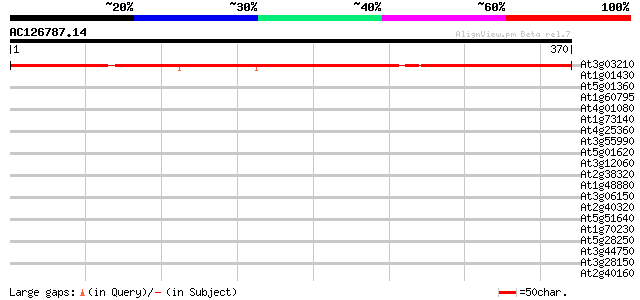
Query= AC126787.14 - phase: 0
(370 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At3g03210 unknown protein 422 e-118
At1g01430 unknown protein 39 0.005
At5g01360 unknown protein 36 0.039
At1g60795 unknown protein 36 0.039
At4g01080 unknown protein (At4g01080) 33 0.19
At1g73140 hypothetical protein 33 0.25
At4g25360 unknown protein (At4g25360) 33 0.33
At3g55990 unknown protein 33 0.33
At5g01620 unknown protein 32 0.43
At3g12060 hypothetical protein 32 0.43
At2g38320 unknown protein 32 0.56
At1g48880 hypothetical protein 32 0.56
At3g06150 unknown protein 32 0.73
At2g40320 unknown protein 32 0.73
At5g51640 unknown protein (At5g51640) 31 0.95
At1g70230 unknown protein 31 0.95
At5g28250 putative protein 30 1.6
At3g44750 histone deacetylase like protein 30 2.8
At3g28150 unknown protein (At3g28150) 30 2.8
At2g40160 unknown protein 30 2.8
>At3g03210 unknown protein
Length = 368
Score = 422 bits (1084), Expect = e-118
Identities = 217/376 (57%), Positives = 274/376 (72%), Gaps = 14/376 (3%)
Query: 1 MFGAVQLGLFAACVVLFVPMGMAGWHLSRNKVLFFSGALFITLAVGVHLTPYFPSVSDFM 60
M GA+ LG+ AAC VLFVPM MAGWHLSRNK+LFFSGALFI+LAV VHLTPYFPSVSD +
Sbjct: 1 MLGAIHLGVLAACFVLFVPMAMAGWHLSRNKMLFFSGALFISLAVCVHLTPYFPSVSDIV 60
Query: 61 SSVKSSNDDVVFVEDRGLCVSLLHEIAWEVMPSKVFDPLLRNN-SLNYDKF---WSWSRL 116
+SV S VV + R C++ +++I W+V P + + RNN S D F W W +
Sbjct: 61 ASVSS----VVVYDHRISCINEVNQIVWDVKPVPNPESVRRNNGSTKLDYFVKNWDWMKS 116
Query: 117 NSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVL--DSERVESVRSFLF 174
V CEFQ+L + DV DLLNGSWV++ GDSQAR LSLL+LVL DS+ ++SVR LF
Sbjct: 117 RKVLSCEFQKLDKFDVSDLLNGSWVVVAGDSQARFVALSLLNLVLGSDSKAMDSVRGDLF 176
Query: 175 KRHSDYHIDIDEMGLKLDFIWAPYPNNLTDLVMRFKKKHLYPDVLVMGSGLWHMLHITNA 234
+RHSDY I + E+G+KLDF+WAPY +L DLV+ +KK YPDV++MG+GLWHMLH+ NA
Sbjct: 177 RRHSDYSIVVKEIGMKLDFVWAPYEKDLDDLVVSYKKMKKYPDVVIMGTGLWHMLHVNNA 236
Query: 235 SDYGGLLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLINSMLNTQEK 294
SD+G L L + V S++P++PK ++ GSVS RS HLFW+GMP LIN MLNT EK
Sbjct: 237 SDFGFRLRQLSSHVESLVPLTPK---EQEGGGSVSG-RSVHLFWIGMPVLINGMLNTDEK 292
Query: 295 RVKMNDVMQGEYEREVRKSGILREFGGPFQLLDIGSLSLNCGIKCTDDGMHYDGAVYEAE 354
+ KM+D + EY+R + +S ILR+ GGP LLDI S + NCG +CT DGMHYD AVY+A
Sbjct: 293 KEKMSDTVWHEYDRSLGESKILRQMGGPLILLDIQSFTWNCGPQCTLDGMHYDSAVYDAA 352
Query: 355 LHIMFNALLIESHQKL 370
+H+M NALLIESHQ L
Sbjct: 353 VHVMLNALLIESHQSL 368
>At1g01430 unknown protein
Length = 456
Score = 38.9 bits (89), Expect = 0.005
Identities = 22/88 (25%), Positives = 36/88 (40%), Gaps = 6/88 (6%)
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVR 170
W W +C+ R + +D + W+ +GDS +R SLL ++ E VE +
Sbjct: 142 WRWQP----RDCDLPRFNPEQFLDNMRNKWLAFIGDSISRNHVQSLLCILSQVEEVEDI- 196
Query: 171 SFLFKRHSDYHIDIDEMGLKLDFIWAPY 198
F K + L IW+P+
Sbjct: 197 -FHDKEYKSRIWRFPSYNFTLSVIWSPF 223
>At5g01360 unknown protein
Length = 434
Score = 35.8 bits (81), Expect = 0.039
Identities = 46/180 (25%), Positives = 70/180 (38%), Gaps = 19/180 (10%)
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVR 170
W W +C R K M+ L G ++ VGDS R S + LV +S E +
Sbjct: 136 WEWQP----DDCTIPRFSPKLAMNKLRGKRLLFVGDSLQRSQWESFVCLV-ESIIPEGEK 190
Query: 171 SFLFKRHSDYHI-DIDEMGLKLDFIWAPY---PNNLTDLVMRFKKKHLYPDVLVMGSGLW 226
S KR Y + E ++F WAPY N ++ KK+ + D + + W
Sbjct: 191 S--MKRSQKYFVFKAKEYNATIEFYWAPYIVESNTDIPVISDPKKRIVKVDSVKDRAKFW 248
Query: 227 HMLHITNASDYGGLLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLFWLGMPTLIN 286
I + Y + LR M + FGN E ++ + + LG+ T N
Sbjct: 249 EGADILVFNTYVWWMSGLR-----MKALWGSFGNGE---SGAEALDTQVAYRLGLKTWAN 300
>At1g60795 unknown protein
Length = 541
Score = 35.8 bits (81), Expect = 0.039
Identities = 31/136 (22%), Positives = 56/136 (40%), Gaps = 24/136 (17%)
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSL----VLDSERV 166
W W + C+ RL D ++ L G ++ VGDS R SL+ + + D +RV
Sbjct: 235 WRWQP----NGCDIPRLNGTDFLEKLRGKKLVFVGDSINRNMWESLICILRHSLKDKKRV 290
Query: 167 ESVRSFL-FKRHSDYHIDIDEMGLKLDFIWAPY---------PNNLT------DLVMRFK 210
+ FK+ Y ++ +DF+ +P+ N T D++ +
Sbjct: 291 YEISGRREFKKKGFYAFRFEDYNCTVDFVGSPFFVRESSFKGVNGTTLETLRLDMMDKTT 350
Query: 211 KKHLYPDVLVMGSGLW 226
+ D+L+ +G W
Sbjct: 351 SMYRDADILIFNTGHW 366
>At4g01080 unknown protein (At4g01080)
Length = 442
Score = 33.5 bits (75), Expect = 0.19
Identities = 21/89 (23%), Positives = 33/89 (36%), Gaps = 6/89 (6%)
Query: 110 FWSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESV 169
FW W +C+ R + + W +GDS AR SL+ ++ E VE +
Sbjct: 133 FWRWKP----RDCDLPRFSPSQFLASVKNKWWAFIGDSIARNHVQSLICILSQVEEVEEI 188
Query: 170 RSFLFKRHSDYHIDIDEMGLKLDFIWAPY 198
+ K L IW+P+
Sbjct: 189 --YHDKEFRSKIWRFPSHNFTLSVIWSPF 215
>At1g73140 hypothetical protein
Length = 403
Score = 33.1 bits (74), Expect = 0.25
Identities = 21/100 (21%), Positives = 42/100 (42%), Gaps = 20/100 (20%)
Query: 106 NYDKFWSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSER 165
+Y + W W S C+ R ++D+L +M +GDS R S++ +
Sbjct: 96 SYYQNWRWKP----SSCDLPRFNALKLLDVLRNKRLMFIGDSVQRSTFESMVCM------ 145
Query: 166 VESVRSFLFKRHSDYH-------IDIDEMGLKLDFIWAPY 198
V+S + ++ +H +E +++ WAP+
Sbjct: 146 ---VQSVIPEKKKSFHRIPPMKIFKAEEYNASIEYYWAPF 182
>At4g25360 unknown protein (At4g25360)
Length = 533
Score = 32.7 bits (73), Expect = 0.33
Identities = 32/132 (24%), Positives = 53/132 (39%), Gaps = 24/132 (18%)
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSER----- 165
W W S+CE R + ++L+ G + +GDS AR S+L L+ E
Sbjct: 218 WRWKP----SQCELPRFDARKFLELMKGKTLAFIGDSVARNQMESMLCLLWQVETPVNRG 273
Query: 166 VESVRSFLFKRHSDYHIDI------DEMGLKLDFIWAPYPNNLTDLVMRFKKKHLYP--- 216
++ + FK+ S I + K D+ P +T L + + +
Sbjct: 274 SRKMQRWYFKQSSVMIARIWSSWLVHQFNEKFDYA----PEGVTKLKLDLPDERIMEAIP 329
Query: 217 --DVLVMGSGLW 226
DV+V+ SG W
Sbjct: 330 KFDVVVLSSGHW 341
>At3g55990 unknown protein
Length = 487
Score = 32.7 bits (73), Expect = 0.33
Identities = 24/88 (27%), Positives = 38/88 (42%), Gaps = 6/88 (6%)
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVR 170
W W +C + K K +++ L +M VGDS R S++ LV V R
Sbjct: 184 WRWQP----RDCSLPKFKAKLLLEKLRNKRMMFVGDSLNRNQWESMVCLV--QSVVPPGR 237
Query: 171 SFLFKRHSDYHIDIDEMGLKLDFIWAPY 198
L K S +++ ++F WAP+
Sbjct: 238 KSLNKTGSLSVFRVEDYNATVEFYWAPF 265
>At5g01620 unknown protein
Length = 449
Score = 32.3 bits (72), Expect = 0.43
Identities = 31/137 (22%), Positives = 52/137 (37%), Gaps = 32/137 (23%)
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVL-----DSER 165
W W C +R ++ + L G +M VGDS R +S++ L+ D +
Sbjct: 155 WRWQP----HACNLKRWNAIEMWEKLRGKRLMFVGDSLNRGQWISMVCLLQSVIPRDKQS 210
Query: 166 VESVRSFLFKRHSDYHIDIDEMGLKLDFIWAPY--------PNN--------LTDLVMRF 209
+ R DY+ ++ F+WAP P N D V++
Sbjct: 211 MSPNAHLTIFRAEDYNATVE-------FLWAPLLVESNSDDPVNHRLSERIIRPDSVLKH 263
Query: 210 KKKHLYPDVLVMGSGLW 226
K + D+L+ + LW
Sbjct: 264 ASKWQHADILIFNTYLW 280
>At3g12060 hypothetical protein
Length = 556
Score = 32.3 bits (72), Expect = 0.43
Identities = 29/126 (23%), Positives = 54/126 (42%), Gaps = 20/126 (15%)
Query: 121 ECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRH--- 177
+C RL +++++ G ++ VGDS R SL+ ++ S + ES RH
Sbjct: 245 QCSLPRLNGGKLLEMIRGRRLVFVGDSLNRNMWESLVCILKGSVKDESQVFEAHGRHQFR 304
Query: 178 --SDYHIDIDEMGLKLDFIWAPY--------PNNLT-------DLVMRFKKKHLYPDVLV 220
++Y + ++F +P+ N T DLV + +++ D+LV
Sbjct: 305 WEAEYSFVFKDYNCTVEFFASPFLVQEWEVTEKNGTKKETLRLDLVGKSSEQYKGADILV 364
Query: 221 MGSGLW 226
+G W
Sbjct: 365 FNTGHW 370
>At2g38320 unknown protein
Length = 410
Score = 32.0 bits (71), Expect = 0.56
Identities = 35/142 (24%), Positives = 58/142 (40%), Gaps = 25/142 (17%)
Query: 109 KFWSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVES 168
KFW W C+ R +++ L ++ VGDS R +S++ +V S + +
Sbjct: 101 KFWRWQP----HTCDLPRFNGTKLLERLRNKRMVYVGDSLNRGQWVSMVCMV--SSVITN 154
Query: 169 VRSFLFKRHSDYHIDID--EMGLKLDFIWAPY--------PNN--LTDLVMRFK--KKH- 213
++ + I E +D+ WAP P N D ++R + +KH
Sbjct: 155 PKAMYMHNNGSNLITFKALEYNATIDYYWAPLLVESNSDDPTNHRFPDRIVRIQSIEKHA 214
Query: 214 ---LYPDVLVMGSGL-WHMLHI 231
D++V S L W M HI
Sbjct: 215 RHWTNSDIIVFNSYLWWRMPHI 236
>At1g48880 hypothetical protein
Length = 485
Score = 32.0 bits (71), Expect = 0.56
Identities = 25/97 (25%), Positives = 41/97 (41%), Gaps = 18/97 (18%)
Query: 103 NSLNYDKF--WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLV 160
N +D++ W W C R + +DV+ L G ++ VGDS +R SL+ ++
Sbjct: 148 NGRGHDEYLKWRWKP----KHCTVPRFEVRDVLKRLRGKRIVFVGDSMSRTQWESLICML 203
Query: 161 L----DSERVESVRS--------FLFKRHSDYHIDID 185
+ D V V FL R S Y+ ++
Sbjct: 204 MTGLEDKRSVYEVNGNNITKRIRFLGVRFSSYNFTVE 240
>At3g06150 unknown protein
Length = 594
Score = 31.6 bits (70), Expect = 0.73
Identities = 28/128 (21%), Positives = 53/128 (40%), Gaps = 14/128 (10%)
Query: 120 SECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVRSFLFKRHSD 179
S C F+ + D L G W+ GDS +LL+ VL + +V + S+
Sbjct: 336 SHCSFKLFSAEKAWDCLKGKWIFFWGDSNHVDSIRNLLNFVLGHPEIPAVPRRFDMKFSN 395
Query: 180 YHIDIDEMGLKLDF--IWAPYPN----------NLTDLVMRF--KKKHLYPDVLVMGSGL 225
+ + + F W N + +L+ ++ ++ + PDV+++ SGL
Sbjct: 396 PKNPSETVRITSIFNGHWNETKNYQGLDSLKDRDFRELLKKYFNEETNRVPDVMIVNSGL 455
Query: 226 WHMLHITN 233
+H T+
Sbjct: 456 HDGIHWTS 463
>At2g40320 unknown protein
Length = 425
Score = 31.6 bits (70), Expect = 0.73
Identities = 23/92 (25%), Positives = 41/92 (44%), Gaps = 10/92 (10%)
Query: 109 KFWSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLV--LDSERV 166
+FW W + C+ +++ L G +M VGDS R +S++ L+ L E
Sbjct: 123 QFWRWQP----NHCDLPSFNASLMLETLRGKRMMYVGDSLNRGMFVSMICLLHRLIPEDQ 178
Query: 167 ESVRSFLFKRHSDYHIDIDEMGLKLDFIWAPY 198
+S+++ S E ++F WAP+
Sbjct: 179 KSIKT----NGSLTVFTAKEYNATIEFYWAPF 206
>At5g51640 unknown protein (At5g51640)
Length = 501
Score = 31.2 bits (69), Expect = 0.95
Identities = 16/54 (29%), Positives = 26/54 (47%), Gaps = 4/54 (7%)
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSE 164
W W S+C+ R K ++L+ G + +GDS AR S++ L+ E
Sbjct: 181 WRWKP----SQCDLPRFDAKKFLELMRGKTLAFIGDSVARNQMESMMCLLWQVE 230
>At1g70230 unknown protein
Length = 416
Score = 31.2 bits (69), Expect = 0.95
Identities = 16/51 (31%), Positives = 24/51 (46%), Gaps = 4/51 (7%)
Query: 110 FWSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLV 160
+W W +EC+ R +DL+ + +GDS AR SLL L+
Sbjct: 121 YWKWKP----NECDIPRFDSNRFLDLMRDKHLAFIGDSMARNQLESLLCLL 167
>At5g28250 putative protein
Length = 939
Score = 30.4 bits (67), Expect = 1.6
Identities = 29/110 (26%), Positives = 54/110 (48%), Gaps = 13/110 (11%)
Query: 139 SWVMIVGD---SQARIFTLSLLSLVLDSERVESVRSFLFKRHSDYHIDIDEMGLKLDFIW 195
S++ IVG+ S +IF + S+ ++++ F RH DID L++D +
Sbjct: 701 SFISIVGEVSLSAKKIFDIIERKKQFTSKVMDAL--IKFSRHLLRTDDIDGEKLRVDVLG 758
Query: 196 APYPNNLTDLVMRFKK-----KHLYPDVLV---MGSGLWHMLHITNASDY 237
+ + + LT L ++F K ++P VLV MG G + + + +D+
Sbjct: 759 SKFVSQLTRLFLKFSKTSPPEDFIFPAVLVELLMGVGEADRVRLFDEADF 808
>At3g44750 histone deacetylase like protein
Length = 245
Score = 29.6 bits (65), Expect = 2.8
Identities = 15/50 (30%), Positives = 26/50 (52%), Gaps = 5/50 (10%)
Query: 228 MLHITNASDYGGLLGLLRNSVTSMLPVSPKFGNDEPAMGSVSSVRSPHLF 277
++H++ AS LG +N +P+ K GN +G++S+ P LF
Sbjct: 23 LIHVSQAS-----LGECKNKKGEFVPLHVKVGNQNLVLGTLSTENIPQLF 67
>At3g28150 unknown protein (At3g28150)
Length = 414
Score = 29.6 bits (65), Expect = 2.8
Identities = 17/60 (28%), Positives = 26/60 (43%), Gaps = 4/60 (6%)
Query: 110 FWSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESV 169
FW W C+ R K + ++ G + +GDS AR SLL L+ E + +
Sbjct: 112 FWRWKP----DGCDLPRFNPKAFLSMVRGKKMNFIGDSVARNHMESLLCLLSMEETPKDI 167
>At2g40160 unknown protein
Length = 427
Score = 29.6 bits (65), Expect = 2.8
Identities = 21/88 (23%), Positives = 36/88 (40%), Gaps = 6/88 (6%)
Query: 111 WSWSRLNSVSECEFQRLKRKDVMDLLNGSWVMIVGDSQARIFTLSLLSLVLDSERVESVR 170
W W +C R K +++ L G +M +GDS S++ +V + S +
Sbjct: 121 WRWQP----QDCSLPRFDSKLLLEKLRGKKLMFIGDSIHYNQWQSMVCMV--QSVIPSGK 174
Query: 171 SFLFKRHSDYHIDIDEMGLKLDFIWAPY 198
L +I+E + F WAP+
Sbjct: 175 KTLKHTAQMSIFNIEEYNATISFYWAPF 202
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.139 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,492,417
Number of Sequences: 26719
Number of extensions: 368874
Number of successful extensions: 827
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 810
Number of HSP's gapped (non-prelim): 31
length of query: 370
length of database: 11,318,596
effective HSP length: 101
effective length of query: 269
effective length of database: 8,619,977
effective search space: 2318773813
effective search space used: 2318773813
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC126787.14


