

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126786.5 - phase: 0
(487 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
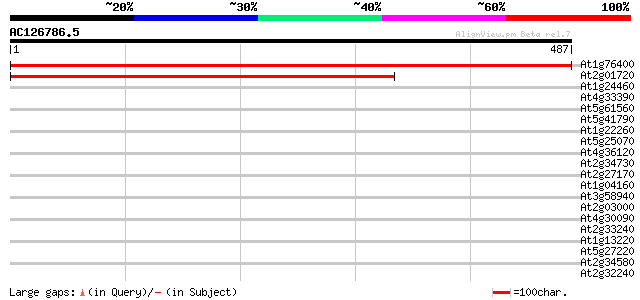
Score E
Sequences producing significant alignments: (bits) Value
At1g76400 putative ribophorin I (dolichyl-diphosphooligosacchari... 719 0.0
At2g01720 putative ribophorin I 406 e-113
At1g24460 unknown protein 36 0.042
At4g33390 putative protein 35 0.12
At5g61560 putative protein 32 0.60
At5g41790 myosin heavy chain-like protein 32 0.60
At1g22260 hypothetical protein 32 0.60
At5g25070 unknown protein 32 0.79
At4g36120 myosin-like protein 32 0.79
At2g34730 putative myosin heavy chain 31 1.3
At2g27170 putative chromosome associated protein 31 1.3
At1g04160 myosin heavy chain MYA2 31 1.3
At3g58940 putative protein 31 1.8
At2g03000 hypothetical protein 31 1.8
At4g30090 putative protein 30 2.3
At2g33240 putative myosin heavy chain 30 2.3
At1g13220 putative nuclear matrix constituent protein 30 2.3
At5g27220 putative protein 30 3.0
At2g34580 hypothetical protein 30 3.0
At2g32240 putative myosin heavy chain 30 3.0
>At1g76400 putative ribophorin I
(dolichyl-diphosphooligosaccharide-protein
glycosyltransferase)
Length = 614
Score = 719 bits (1855), Expect = 0.0
Identities = 350/487 (71%), Positives = 416/487 (84%)
Query: 1 MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKL 60
++++ FT++LQPFPEKITQ +I L++ QESAQYLSPY V+ QSL++KLP ARIESYTK
Sbjct: 128 LEVVAAFTNVLQPFPEKITQGEIHLVMLQESAQYLSPYAVESQSLSIKLPNARIESYTKF 187
Query: 61 ENTKLQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHY 120
ENTKLQGSELKYGPY+NL +SY PIV+HFE+ FAVA++LVREIE+SHWGNVQ+TE+Y
Sbjct: 188 ENTKLQGSELKYGPYKNLQSYSYSPIVVHFESKAAFAVAEKLVREIEVSHWGNVQVTENY 247
Query: 121 NLVHGGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWG 180
N+VH GAQ KGEFSRLD+QARP RGASAFR L A+LPPRAHS+YYRD+IGNISTS +
Sbjct: 248 NVVHRGAQLKGEFSRLDFQARPNPRGASAFRHLLARLPPRAHSIYYRDDIGNISTSEMKS 307
Query: 181 DSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQDFLFGVDGKRFLNISFGSPINELVIDTL 240
DSKKTEL IEPR+PLFGGWKT FTIGYGLPL DFLF +GKRFLNISFGSPI +LV + L
Sbjct: 308 DSKKTELLIEPRFPLFGGWKTFFTIGYGLPLTDFLFASEGKRFLNISFGSPILDLVTEKL 367
Query: 241 VVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYYK 300
+V+VVLPEGSKDIS + PF VK+ E K+SHLDIAGRPVVVLEKNN VP+HN+H QVYYK
Sbjct: 368 IVQVVLPEGSKDISVTTPFAVKQSQEIKYSHLDIAGRPVVVLEKNNVVPDHNQHIQVYYK 427
Query: 301 FNRLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSI 360
F+ +++L EPLMLISGFF LF+ I+Y AD+SISK+S SYLAK+QW+EV AT+ +V SI
Sbjct: 428 FSNINLLSEPLMLISGFFILFITCIIYTRADISISKSSPSYLAKLQWDEVLATLQEVQSI 487
Query: 361 VSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQ 420
V +CL THDKLEASLRDLSRTGDIQ CKA RKS DS LK+L+KELK + FLQS P A+
Sbjct: 488 VQKCLATHDKLEASLRDLSRTGDIQTCKAARKSTDSLLKDLSKELKPLLGFLQSFPSASH 547
Query: 421 ILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDL 480
I PKVEE++ KE++LQEKLMAKH+TVV+ YEKK GR+IENRIAS QQKI AL+QEI+DL
Sbjct: 548 ISPKVEELVVKEKELQEKLMAKHTTVVEGYEKKSSGRDIENRIASQQQKIIALRQEIEDL 607
Query: 481 MDLIDEI 487
++ IDEI
Sbjct: 608 LEFIDEI 614
>At2g01720 putative ribophorin I
Length = 464
Score = 406 bits (1043), Expect = e-113
Identities = 193/335 (57%), Positives = 250/335 (74%), Gaps = 1/335 (0%)
Query: 1 MDILTVFTHILQPFPEKITQADIQLLLFQESAQYLSPYPVKVQSLNVKLPEARIESYTKL 60
+++L + TH L+PFP +ITQ++ QL+ + +SA LSPY VK Q+ +K P R+ES+T +
Sbjct: 127 LEVLYILTHSLEPFPVEITQSESQLVYYHDSAVILSPYHVKQQTTFIKTPSTRVESFTSI 186
Query: 61 ENTKLQGSELKYGPYENLPPFSYLPIVIHFENNQPFAVAKELVREIEISHWGNVQITEHY 120
E G E+KYGPYEN +SY P++IHFENN PFAV +ELVREIEISHWG++QITE+Y
Sbjct: 187 EPANRAGKEIKYGPYENRASYSYTPVIIHFENNSPFAVVEELVREIEISHWGSLQITENY 246
Query: 121 NLVHGGAQSKGEFSRLDYQARPYVRGASAFRRLTAKLPPRAHSVYYRDEIGNISTSSLWG 180
L HGGA+ KG FSR+DYQ++ V GAS+F L A LPPR +SVYYRDEIGNISTS L
Sbjct: 247 RLTHGGARHKGVFSRVDYQSKRSVSGASSFNALLAVLPPRVNSVYYRDEIGNISTSHLRT 306
Query: 181 DSKKTELEIEPRYPLFGGWKTAFTIGYGLPLQDFLF-GVDGKRFLNISFGSPINELVIDT 239
+K+ELE EPRYPLFGGW F IGY +PL+D+LF DG+R+LN +FG P+ E +++
Sbjct: 307 GFRKSELEFEPRYPLFGGWSATFIIGYRVPLEDYLFEASDGRRYLNFTFGCPLVETIVNK 366
Query: 240 LVVKVVLPEGSKDISPSVPFPVKERHETKFSHLDIAGRPVVVLEKNNAVPEHNEHFQVYY 299
L +KVVLPEGSKD S +PF V + + K+S+LDI GR VVVL+K+N VP HN FQVYY
Sbjct: 367 LTIKVVLPEGSKDPSAVLPFTVNQELQVKYSYLDIVGRTVVVLQKDNVVPTHNVPFQVYY 426
Query: 300 KFNRLSMLREPLMLISGFFFLFLASIVYMHADLSI 334
F + ML EP ML+S FF +F+AS+ Y+H DL+I
Sbjct: 427 TFKPIYMLAEPFMLVSAFFLVFVASLAYVHIDLNI 461
>At1g24460 unknown protein
Length = 1791
Score = 36.2 bits (82), Expect = 0.042
Identities = 44/178 (24%), Positives = 78/178 (43%), Gaps = 40/178 (22%)
Query: 339 ASYLAKIQWEEVQATIHQV-HSIVSRCLTTH-DKLEASLRDLS-RTGDIQGCKATRKSVD 395
A+ L ++Q +E+ +V + + LT D+LE +L+ R+ ++ + T+K ++
Sbjct: 354 ANRLTELQEKEIALESSEVMKGQLEQSLTEKTDELEKCYAELNDRSVSLEAYELTKKELE 413
Query: 396 SSLKELTKELKSPVTFLQSSPQAAQILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLG 455
SL E TKEL+ +T LQ A + +L + +AK +V Y++ L
Sbjct: 414 QSLAEKTKELEECLTKLQEMSTALD-----------QSELDKGELAKSDAMVASYQEMLS 462
Query: 456 GRE--IEN------------------------RIASHQQKITALKQEIDDLMDLIDEI 487
R IEN +A ++++T + QE + L DLI I
Sbjct: 463 VRNSIIENIETILSNIYTPEEGHSFDIVEKVRSLAEERKELTNVSQEYNRLKDLIVSI 520
>At4g33390 putative protein
Length = 779
Score = 34.7 bits (78), Expect = 0.12
Identities = 35/146 (23%), Positives = 69/146 (46%), Gaps = 9/146 (6%)
Query: 340 SYLAKIQWEEVQATIHQVHSIVSRCLTTHDKLE-ASLRDLSRTGDIQGCKATRKSVDSSL 398
S LAK++ +E++ I S+ S+ +LE A R S +++ K +++ +
Sbjct: 252 SELAKLRVQEMEQGIADEASVASKA-----QLEVAQARHTSAISELESVKEELQTLQNEY 306
Query: 399 KELTKELKSPVTFLQSSPQAA-QILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGR 457
L KE V + + A+ ++ KVEE+ + +E L HS+ ++ E ++G
Sbjct: 307 DALVKEKDLAVKEAEEAVIASKEVERKVEELTIELIATKESLECAHSSHLEAEEHRIGAA 366
Query: 458 EIENRIASHQQKITALKQEIDDLMDL 483
+ ++ +K LKQ ++L L
Sbjct: 367 MLRDQETHRWEK--ELKQAEEELQRL 390
>At5g61560 putative protein
Length = 815
Score = 32.3 bits (72), Expect = 0.60
Identities = 47/207 (22%), Positives = 88/207 (41%), Gaps = 31/207 (14%)
Query: 284 KNNAVPEHNEHFQVYYKFNRLSMLREPLMLISGFFFLFLASIVYMHADLSISKTSASYLA 343
K N P H+ H + L + L+ GF+ + I Y +D+S ++S A
Sbjct: 241 KQNNEPHHHHHNRA----GSLDVDESKLLNQKGFYRTSSSGIGYGGSDISSWRSSQMEEA 296
Query: 344 KIQ--WEEVQATIHQVHSIVSRCLTTHDKLEASLRDLSRTGDIQGCKATRKS--VDSSLK 399
+ + ++ Q+H +KL+ LR I+G A +S +D+S K
Sbjct: 297 SSSSTYSDPTSSSSQIHKDFEL-----EKLKIELRH------IKGMYAVAQSEVIDASKK 345
Query: 400 --ELTKELKSPVTFLQSSPQAAQILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGR 457
+L + T L++ + + +EV+ ER+ QE + V +C E R
Sbjct: 346 MQDLNQRRSEEATRLKNLTIREE---EADEVVEMERERQEDAENEAELVRECIE-----R 397
Query: 458 EIENRI--ASHQQKITALKQEIDDLMD 482
E E R+ + +++ KQ ++D ++
Sbjct: 398 ETEERLEAEARAEEVRKEKQRLEDALE 424
>At5g41790 myosin heavy chain-like protein
Length = 1305
Score = 32.3 bits (72), Expect = 0.60
Identities = 32/138 (23%), Positives = 62/138 (44%), Gaps = 28/138 (20%)
Query: 354 IHQVHSIVSRCLTTHDKLEASLRDLSR-----TGDIQGCKATRKSVDSSLKELTKELKSP 408
IH+ H S T +LE L+ L + + + + +KS+ S + E+T ELK
Sbjct: 475 IHETHQRESS--TRLSELETQLKLLEQRVVDLSASLNAAEEEKKSLSSMILEITDELK-- 530
Query: 409 VTFLQSSPQAAQILPKVEEVIAKERDLQEKLMAKH---STVVDCYE-----KKLGGREIE 460
Q KV+E++ + + ++ L K S+ V+ +E +E+E
Sbjct: 531 -----------QAQSKVQELVTELAESKDTLTQKENELSSFVEVHEAHKRDSSSQVKELE 579
Query: 461 NRIASHQQKITALKQEID 478
R+ S ++++ L Q ++
Sbjct: 580 ARVESAEEQVKELNQNLN 597
>At1g22260 hypothetical protein
Length = 851
Score = 32.3 bits (72), Expect = 0.60
Identities = 30/127 (23%), Positives = 60/127 (46%), Gaps = 18/127 (14%)
Query: 363 RCLT--THDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQ 420
RC T T DKLE+ + +G + +S++ +L +E+++ + +++S Q
Sbjct: 386 RCSTSQTIDKLES---------EAKGLVSKHADAESAISQLKEEMETLLESVKTSEDKKQ 436
Query: 421 ILP-KVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDD 479
L K+ + + ++ EKL A V+ E ++ SHQ + L +E++
Sbjct: 437 ELSLKLSSLEMESKEKCEKLQADAQRQVEELET------LQKESESHQLQADLLAKEVNQ 490
Query: 480 LMDLIDE 486
L +I+E
Sbjct: 491 LQTVIEE 497
Score = 28.5 bits (62), Expect = 8.7
Identities = 27/120 (22%), Positives = 49/120 (40%), Gaps = 16/120 (13%)
Query: 369 DKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSP-QAAQILPKVEE 427
D L +RD+S D K S D L+EL E + F Q+ A ++ K +
Sbjct: 154 DSLNQQMRDMSLRLD--AAKEEITSRDKELEELKLEKQQKEMFYQTERCGTASLIEKKDA 211
Query: 428 VIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDLMDLIDEI 487
VI K E+KL + +++ ++T + E+ DL+ + +++
Sbjct: 212 VITKLE-------------ASAAERKLNIENLNSQLEKVHLELTTKEDEVKDLVSIQEKL 258
>At5g25070 unknown protein
Length = 736
Score = 32.0 bits (71), Expect = 0.79
Identities = 27/121 (22%), Positives = 57/121 (46%), Gaps = 13/121 (10%)
Query: 361 VSRCLTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQ 420
V +T+ + A LRDL+R + C+ +E+ K K ++++ + +
Sbjct: 463 VDEFMTSEKERGAKLRDLARVSADEACE---------YEEVIKLRKGLMSYVSKTREERA 513
Query: 421 ILPKVEEVIAKE-RDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDD 479
L +EE +++E + LQE++ + + + KK I+ I S KI +++ + +
Sbjct: 514 KLVNIEEKLSEEVQKLQEEVSSTRELLKERSSKK---SIIQQNITSFMDKIMFIEKRMPE 570
Query: 480 L 480
L
Sbjct: 571 L 571
>At4g36120 myosin-like protein
Length = 981
Score = 32.0 bits (71), Expect = 0.79
Identities = 34/118 (28%), Positives = 53/118 (44%), Gaps = 27/118 (22%)
Query: 361 VSRCLTTHDKLEASLRD-----------LSRTGDIQGCKATR-KSVDSSLKEL---TKEL 405
+SRCL + +A L + L+ + D+Q T+ K V S K L KEL
Sbjct: 761 LSRCLQNLESTKAWLEEKEQLISKLKSQLTSSEDLQSLAETQLKCVTESYKSLDLHAKEL 820
Query: 406 KSPVTFLQSSPQAAQILPKVE-----EVIAKERDLQEKL-------MAKHSTVVDCYE 451
++ V L+ + ++ E E +AK RDLQEK+ + + ST+ C E
Sbjct: 821 EAKVKSLEEETKRLEMAFTTEKHGHEETLAKCRDLQEKMQRYKNHNLLRSSTMHTCQE 878
>At2g34730 putative myosin heavy chain
Length = 829
Score = 31.2 bits (69), Expect = 1.3
Identities = 36/170 (21%), Positives = 72/170 (42%), Gaps = 15/170 (8%)
Query: 328 MHADLSISKTSASYLAKIQWEEVQATIHQVHSIVSRCLTTHDKL----EASLRDLSRTGD 383
+ D+ S+ AS + V + Q+ H L E L DL+R
Sbjct: 457 LETDVEDSRNEASIYEDVYGCFVTEFVGQIKCTKQETDLEHSMLREAYELLLEDLARK-- 514
Query: 384 IQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILPKVE-EVIAKERDLQEKLMAK 442
+ K+ DS +K + E V + ++ +A + + ++ V KE L+ +++ K
Sbjct: 515 -EARKSKEDFEDSCVKSVMMEECCSVIYKEAVKEAHKKIVELNLHVTEKEGTLRSEMVDK 573
Query: 443 HSTVVDCY-------EKKLGGREIENRIASHQQKITALKQEIDDLMDLID 485
+ + EK+ + EN +A+ ++KI + Q+I+DL ++
Sbjct: 574 ERLKEEIHRLGCLVKEKENLVQTAENNLATERKKIEVVSQQINDLQSQVE 623
>At2g27170 putative chromosome associated protein
Length = 1163
Score = 31.2 bits (69), Expect = 1.3
Identities = 23/92 (25%), Positives = 45/92 (48%), Gaps = 4/92 (4%)
Query: 392 KSVDSSLKELTKELKSPVTFLQS-SPQAAQILPKVEEVIAKERDLQEKLMAKHSTVVDCY 450
KS+D SLKELTKEL++ ++ Q + L K ++ +D Q+++ + D
Sbjct: 268 KSLDESLKELTKELQTLYKEKETVEAQQTKALKKKTKLELDVKDFQDRITGNIQSKNDAL 327
Query: 451 EKKLGGREIENRIASHQQKITALKQEIDDLMD 482
E+ +E + +++ A+K + +D
Sbjct: 328 EQL---NTVEREMQDSLRELEAIKPLYESQVD 356
>At1g04160 myosin heavy chain MYA2
Length = 1519
Score = 31.2 bits (69), Expect = 1.3
Identities = 29/133 (21%), Positives = 57/133 (42%), Gaps = 14/133 (10%)
Query: 320 LFLASIVYMHADLSISKTSASYLAKIQWEEVQATIHQVHSIVSRCLTTHDKLEASLRDLS 379
++LA + Y + T ++ K+ +E++ + +KLE + +L+
Sbjct: 844 VYLARLHYRKLKKAAITTQCAWRGKVARKELK-NLKMAARETGALQEAKNKLEKQVEELT 902
Query: 380 ---------RTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAA----QILPKVE 426
RT + K +SSL+E+ + K L +AA ++LP ++
Sbjct: 903 WRLQLEKRMRTDLEEAKKQENAKYESSLEEIQNKFKETEALLIKEREAAKTVSEVLPIIK 962
Query: 427 EVIAKERDLQEKL 439
EV +++L EKL
Sbjct: 963 EVPVVDQELMEKL 975
>At3g58940 putative protein
Length = 618
Score = 30.8 bits (68), Expect = 1.8
Identities = 25/82 (30%), Positives = 39/82 (47%), Gaps = 6/82 (7%)
Query: 411 FLQSSPQAAQILPKVEEVIAKERDLQEKLMAKHSTVVD---CYEKKLGGREI---ENRIA 464
+LQ + A + + + AK DL+E+L A +VD L RE E A
Sbjct: 492 YLQMQLEVADLKERQRDQEAKLADLKERLRAMLDMIVDQNPIIASALRAREATNSEREKA 551
Query: 465 SHQQKITALKQEIDDLMDLIDE 486
S Q++T +++ D L D+I E
Sbjct: 552 SSGQEMTEKERDYDALFDIIVE 573
>At2g03000 hypothetical protein
Length = 535
Score = 30.8 bits (68), Expect = 1.8
Identities = 28/94 (29%), Positives = 46/94 (48%), Gaps = 6/94 (6%)
Query: 366 TTHDKLEASLRDLSRTGDIQ-GCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILPK 424
+T + +SL L+R D Q +AT S ++ + LT E + + + P+ A + K
Sbjct: 420 STSNMRTSSLMTLTRFIDEQISLRATVSSTRTTARLLTNESPATIRAVAMLPRVAMV-EK 478
Query: 425 VEEVIAKER----DLQEKLMAKHSTVVDCYEKKL 454
E VI E D++ +L KH ++C EK L
Sbjct: 479 GECVICFEEWSKSDMETELPCKHKYHLECVEKWL 512
>At4g30090 putative protein
Length = 312
Score = 30.4 bits (67), Expect = 2.3
Identities = 24/101 (23%), Positives = 47/101 (45%), Gaps = 4/101 (3%)
Query: 382 GDIQGCKATRKSVDSSLKELTKELKSPVTFLQS--SPQAAQILPKVEEVIAKERDLQEKL 439
G+ + + L ELKS V+ LQS + ++L K E++ E ++EK
Sbjct: 24 GEASSSSSENNGCNGDQNHLLNELKSTVSALQSIIKEKNQELLSKEEKIRGLELYIREKP 83
Query: 440 MAKHSTV-VDCYEKKLG-GREIENRIASHQQKITALKQEID 478
S + +E + E+E ++ Q+++ LK+E++
Sbjct: 84 YLFESEIDFSQFENPVKHASEVEEKVYELQKQVFGLKREVE 124
>At2g33240 putative myosin heavy chain
Length = 1611
Score = 30.4 bits (67), Expect = 2.3
Identities = 24/126 (19%), Positives = 64/126 (50%), Gaps = 10/126 (7%)
Query: 370 KLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILP----KV 425
KL+ ++++ +I ++ + + +EL KEL+ + Q ++ K+
Sbjct: 963 KLQLGETQVTKSEEILKLQSALQDMQLEFEELAKELEMTNDLAAENEQLKDLVSSLQRKI 1022
Query: 426 EEVIAKERD---LQEKLMAKHSTVVD---CYEKKLGGREIENRIASHQQKITALKQEIDD 479
+E +K + L E+ + + V+D + + ++++ +++ ++KI +L ++ DD
Sbjct: 1023 DESDSKYEETSKLSEERVKQEVPVIDQGVIIKLEAENQKLKALVSTLEKKIDSLDRKHDD 1082
Query: 480 LMDLID 485
L+DL++
Sbjct: 1083 LVDLLE 1088
>At1g13220 putative nuclear matrix constituent protein
Length = 1128
Score = 30.4 bits (67), Expect = 2.3
Identities = 32/142 (22%), Positives = 61/142 (42%), Gaps = 11/142 (7%)
Query: 348 EEVQATIHQVHSIVSRC-LTTHDKL-EASLRDLSRTGDIQGCKATRKSVDSSLKELTKEL 405
+E++ + ++ S+ L++ KL EA+ S G + S +S L E T++
Sbjct: 164 QELEKALREIQEENSKIRLSSEAKLVEANALVASVNGRSSDVENKIYSAESKLAEATRKS 223
Query: 406 KSPVTFLQSSPQAAQILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIAS 465
L+ +L + KER+ E K ++ +EKKL G+E
Sbjct: 224 SELKLRLKEVETRESVLQQERLSFTKERESYEGTFQKQREYLNEWEKKLQGKE------- 276
Query: 466 HQQKITALKQEIDDLMDLIDEI 487
+ IT K+ ++ + ++EI
Sbjct: 277 --ESITEQKRNLNQREEKVNEI 296
>At5g27220 putative protein
Length = 1181
Score = 30.0 bits (66), Expect = 3.0
Identities = 23/96 (23%), Positives = 46/96 (46%), Gaps = 12/96 (12%)
Query: 399 KELTKELKSPVTFLQSSPQAAQILPKVEEVIAKERDLQEKLMAKHSTVVDCYEKKLGG-- 456
+E+ ++ K L + + + + VEE +A LQ+KL+ S+ + +K+L G
Sbjct: 340 EEIERKRKELTAVLDKTAEYGKTIELVEEELA----LQQKLLDIRSSELVSKKKELDGLS 395
Query: 457 ------REIENRIASHQQKITALKQEIDDLMDLIDE 486
+ N + Q+I + +E++D+ LI E
Sbjct: 396 LDLELVNSLNNELKETVQRIESKGKELEDMERLIQE 431
>At2g34580 hypothetical protein
Length = 312
Score = 30.0 bits (66), Expect = 3.0
Identities = 19/57 (33%), Positives = 32/57 (55%), Gaps = 7/57 (12%)
Query: 428 VIAKERDLQ----EKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDDL 480
++ E++LQ + L +KH+T+ + L +E +IA+ KIT L+ IDDL
Sbjct: 113 LLQAEKELQVNKAQILASKHATIQSIERRCL---MLEQKIAAQNLKITILRSNIDDL 166
>At2g32240 putative myosin heavy chain
Length = 1333
Score = 30.0 bits (66), Expect = 3.0
Identities = 28/115 (24%), Positives = 56/115 (48%), Gaps = 11/115 (9%)
Query: 365 LTTHDKLEASLRDLSRTGDIQGCKATRKSVDSSLKELTKELKSPVTFLQSSPQAAQILPK 424
L +HD + L ++ D G + ++SS K+L EL+ L+ S + AQ +
Sbjct: 169 LQSHDAKDKELTEVKEAFDALGIE-----LESSRKKLI-ELEEG---LKRSAEEAQKFEE 219
Query: 425 VEEVIAKERDLQEKLMAKHSTVVDCYEKKLGGREIENRIASHQQKITALKQEIDD 479
+ + A D + + + S ++ K +E+E ++AS QQ+I L +++ +
Sbjct: 220 LHKQSASHADSESQKALEFSELLK--STKESAKEMEEKMASLQQEIKELNEKMSE 272
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.318 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,659,882
Number of Sequences: 26719
Number of extensions: 459477
Number of successful extensions: 1347
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 37
Number of HSP's that attempted gapping in prelim test: 1331
Number of HSP's gapped (non-prelim): 52
length of query: 487
length of database: 11,318,596
effective HSP length: 103
effective length of query: 384
effective length of database: 8,566,539
effective search space: 3289550976
effective search space used: 3289550976
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC126786.5


